Abstract

Despite the abundance of phage-borne antibiotic resistance genes (ARGs) in the environment, the frequency of ARG propagation via phage-mediated transduction (relative to via conjugation) is poorly understood. We investigated the influence of bacterial concentration and water turbulence level [quantified as Reynold’s number (Re)] in suspended growth systems on the frequency of ARG transfer by two mechanisms: delivery by a lysogenic phage (phage λ carrying gentamycin-resistance gene, genR) and conjugation mediated by the self-transmissible plasmid RP4. Using Escherichia coli (E. coli) as the recipient, phage delivery had a comparable frequency (1.2 ± 0.9 × 10–6) to that of conjugation (1.1 ± 0.9 × 10–6) in suspensions with low cell concentration (104 CFU/mL) and moderate turbulence (Re = 5 × 104). Turbulence affected cell (or phage)-to-cell contact rates and detachment (due to shear force), and thus, it affected the relative importance of conjugation versus phage delivery. At 107 CFU/mL, no significant difference was observed between the frequencies of ARG transfer by the two mechanisms under quiescent water conditions (2.8 ± 0.3 × 10–5 for conjugation vs 2.2 ± 0.5 × 10–5 for phage delivery, p = 0.19) or when Re reached 5 × 105 (3.4 ± 1.5 × 10–5 for conjugation vs 2.9 ± 1.0 × 10–5 for phage delivery, p = 0.52). Transcriptomic analysis of genes related to conjugation and phage delivery and simulation of cell (or phage)-to-cell collisions at different Re values corroborate that the importance of phage delivery relative to conjugation increases under either quiescent or turbulent conditions. This finding challenges the prevailing view that conjugation is the dominant ARG transfer mechanism and underscores the need to consider and mitigate potential ARG dissemination via transduction.
Keywords: ARG propagation, conjugative plasmid, lysogenic phage, horizontal gene transfer
Introduction
The global spread of antibiotic resistance has resulted in significant societal and economic costs of dealing with resistant infections,1 which annually exceeds $4.6 billion in just the United States.2 Thus, elucidating the origins and system-specific dissemination pathways of the associated antibiotic resistance genes (ARGs) has become a research priority.3,4 Of the three main horizontal gene transfer (HGT) mechanisms (i.e., conjugation, transduction, and transformation), conjugation is often considered to have the greatest influences on ARG dissemination in the environment due to the high transfer efficiency5,6 and general abundance of plasmid-borne ARGs.6,7 Transformation generally has a minor influence on ARG dissemination due to limited species of naturally transformable bacteria in the environment (about 80 species8) and low probability of incorporating ARG-bearing extracellular DNA strands into the bacterial chromosome or plasmids.4 In contrast, little is known about the overall contribution of ARG delivery by phages to antibiotic resistance propagation despite increasing attention to this mechanism in recent years (Figure 1a).9
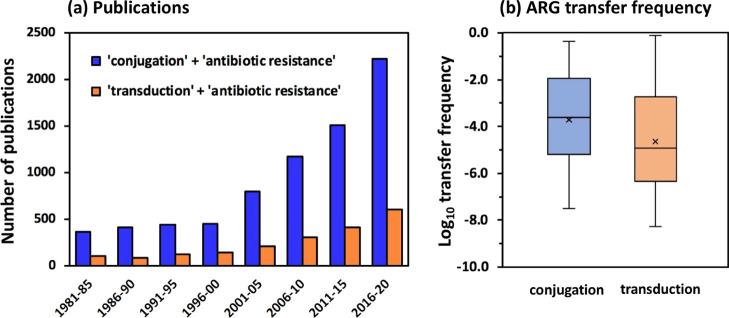
Figure 1.
Publication trends (a) and the reported ARG transfer frequencies (b) of plasmid-mediated conjugation and phage-mediated transduction. (a) Publications (https://pubmed.ncbi.nlm.nih.gov/) from 1981 to 2020 on conjugation and transduction-associated antibiotic resistance both show an exponential increase, with the number of papers related to conjugation 3–5 times higher than that of transduction. (b) Reported frequencies of ARG transfer via conjugation and transduction both showed high variability. Average frequencies are indicated by “×”.
Phages are the most abundant and diverse entities in the biosphere,10 and phage-mediated transduction can result in genetic exchange between distantly related bacterial species.9,11 Errors in phage replication may incidentally incorporate bacterial DNA (including ARGs) into the phage genome and transfer it to another bacterium upon the next infection.3 Recently, mounting evidence suggests that phages are a potentially significant ARG reservoir and serve as vectors in numerous environments including wastewater,12 sludge,13 rivers,14 sediment,15 soils,16 fresh vegetables,17 air,18 and animal and human feces.19,20 The abundance of phage-borne ARGs is sometimes comparable or even higher than that found inside bacteria, as determined by quantitative polymerase chain reaction (qPCR) or metagenomic analyses,15,16,21 although qPCR may sometimes detect unfunctional fragments of phage-borne ARGs. Therefore, it is important to discern conditions under which functional ARG delivery by phages is as significant as conjugation and assess the magnitude of their relative HGT contributions to prioritize associated mitigation and risk management efforts.
The contributions of conjugative plasmids and phages to ARG dissemination depend not only on the abundance of plasmid- or phage-borne ARGs but also on the ARG transfer frequency,5 which is highly variable (Figure 1b)6,22−45 and dependent on system-specific conditions.46−52 However, the frequency of ARG transfer via conjugation versus phage delivery has rarely been systematically compared under the same environmental conditions.53
In this study, we compare ARG dissemination in suspended growth systems that prevail in some wastewater treatment processes and natural water bodies, via conjugation versus phage delivery. We investigate how two system-specific factors known to influence HGT frequency51 (i.e., bacterial concentration and water turbulence) affect the relative importance of these two ARG delivery mechanisms. The conjugative plasmid RP4 and lysogenic phage λ harboring genR were selected for conjugation and phage delivery assays, respectively, using Escherichia coli as the recipient. Frequency of ARG dissemination was quantified as the percentage of recipient E. coli receiving genR, determined by plate counting methods and qPCR analysis. Transcriptomic analysis was performed to mechanistically explore the influence of bacterial concentration and water turbulence on the relative importance of conjugation versus phage delivery in ARG transfer. Collision theory to assess how turbulence affects donor-recipient contact frequency was also considered. Overall, this study advances the understanding of prevalent ARG dissemination pathways and rates as a function of system-specific conditions to inform potential mitigation and control strategies.
Materials and Methods
Experimental Approach
ARG transfer experiments were conducted with commonly used vectors to facilitate comparison with other studies. Conjugation tests were run with plasmid RP4, which is a model for plasmid biology (i.e., for plasmid maintenance and stability, synthesis of type IV secretion pili, and regulation of conjugation-related genes54,55) and is a representative plasmid of the incompatibility group P (IncP). Experiments involving ARG delivery by phages were run with phage λ, which belongs to the Siphoviridae family56 and is representative of long-tailed phages.57 Both conjugation and phage delivery experiments were conducted with genR, which is a well-studied ARG prevalent in both clinical and natural environments.58,59
Two system-specific factors, bacterial concentration and water turbulence (represented by Re), were selected as they exhibited significant influence on HGT frequency with great variability in the aquatic environments.51E. coli concentration was varied between 103 and 108 CFU/mL, which represents a wide range of conditions—from 0 in some drinking water systems60 to about 106 CFU/mL in sewage61 and 108 CFU/mL in activated sludge.62 The range of Re in this study was varied from 0 to 5 × 105 which are typical values in groundwater (1 to at least 105),63−65 lakes, and streams (0 to at least 5 × 106)66−68 and some wastewater transport and treatment processes (10 to 106).69−72Re was calculated in shaking flasks as73
| 1 |
where ρ is the fluid density (kg/m3), n is the shaking frequency (s–1), D is the maximum inner diameter of the flasks (m), and η is the dynamic viscosity (Pa·s). Parameter values are provided in Table S1.
When investigating the influence of bacterial concentration on conjugation and phage delivery frequencies, Re was fixed at 5 × 104. In the study of turbulence level, bacterial concentration was fixed at 107 CFU/mL which is representative of some systems where nutrients are abundant such as sewage and wastewater treatment plants.74
Bacterial Strains, Plasmids, and Phage Construction
Conjugative plasmid RP4 was obtained from Addgene (plasmid 79813). Plasmid RP4 contains a set of transfer genes enabling horizontal transmission75 and has been genetically modified to confer resistance to ampicillin, gentamycin, and tetracycline (AmpR, GmR, and TcR), as previously reported.76E. coli DH5α (Invitrogen, USA) harboring plasmid RP4 was selected as the donor strain, and E. coli MG1655 (a kind gift from Dr. Lauren Stadler, Rice University) carrying chloramphenicol resistance gene (cat) in the chromosome as the recipient (Table 1). The λ phage was obtained from Agilent Technologies, USA. GenR was amplified from plasmid RP4 by PCR with the EcoRI endonuclease restriction site added at both ends. After EcoRI digestion, genR was ligated into the genome of λ gt11 (Agilent Technologies, USA) according to the manufacturer’s instructions and electroporated into E. coli strain Y1088 (Agilent Technologies, USA) using Gene Pulser Xcell Electroporation Systems (Bio-Rad, USA). Phage plaques were screened to identify recombinant λ phage by PCR assays and gene sequencing. Further details of phage engineering and plaque screening are available in the Supporting Information (Text S1).
Table 1. Plasmid, Phage, and Bacterial Strains Used in This Study.
| plasmid or phage or strain | purpose | selective marker | ref. or source |
|---|---|---|---|
| plasmid RP4 | self-transmissible plasmid for conjugation | AmpR, GmR, and TcR | Addgene76 |
| λ phage | lysogenic phage for phage delivery assays | GmR | Agilent Technologies |
| E. coli DH5α | donor strain carrying plasmid RP4 for conjugation | Invitrogen | |
| E. coli MG1655 | recipient strain for both conjugation and phage delivery assays | ChlR | a gift from Dr. Lauren Stadler |
| E. coli Y1088 | propagation of λ phage | AmpR and TcR | Agilent Technologies |
Conjugation and Phage Delivery Assays
E. coli strains DH5α and MG1655 were grown to the stationary phase at 37 °C in Luria broth (LB) medium (BD Biosciences, USA) supplemented with gentamycin (10 mg/L) and chloramphenicol (25 mg/L), respectively. Bacteria were centrifuged at 14,000g for 2 min and washed three times with phosphate-buffered saline (PBS) to remove extracellular DNA and nutrients (thus obviating potential confounding effects by transformation) and then resuspended in the M9 mineral salt medium.77 Viable bacteria were counted by a plate assay using standard Difco plate count agar (BD Biosciences, USA) and expressed as colony-forming units (CFUs). Recombinant λ phage was propagated in E. coli Y1088 at 37 °C overnight in a LB medium. Bacterial cells were removed via centrifugation (4 °C, 8000g, 8 min), followed by filtration through 0.22 μm PES filter membranes (Millipore, USA). Phages were further concentrated and purified by PEG precipitation and suspended in PBS or SM buffer stored at 4 °C. Phage concentration was determined using the double-layer plaque assay and expressed as plaque-forming units (PFUs).78
The donor (E. coli DH5α harboring plasmid RP4 for conjugation assays or λ phage for assays using phages as gene delivery vectors) and recipient (E. coli MG1655) of genR transfer were mixed at a 1:1 initial ratio79−83 in all tests to facilitate fair comparisons. ARG donors and recipient cells were mixed in an M9 minimal medium spiked with 3 g/L glucose and shaken at targeted Re representative of a wide range of environmental conditions.38 Conjugation and phage delivery assays were performed under the same experimental conditions but in different flasks to separately quantify conjugation and phage delivery events. Assays were performed at room temperature (20 °C) for 1 h,79 without significant bacterial replication (i.e., no vertical genR transfer) (Figure S1), plasmid lost (Figure S2), or phage decay (Figure S3). Samples were taken and recovered afterward following previously reported procedures with minor adjustments.77,84 Briefly, samples were vigorously vortexed to stop ARG transfer events and washed with M9 medium three times. Recovered bacteria were well-dispersed using vortex and then plated onto LB agar. Plates containing both gentamycin (5 mg/L) and chloramphenicol (25 mg/L) were used to count transconjugants (i.e., chloramphenicol-resistant E. coli MG1655 that were recipients of plasmid-borne genR) or transductants (i.e., recipients of phage-borne genR) because only recipients of genR could offer resistance to gentamycin and grow on these plates. Colony PCR was used to verify that plasmid RP4 or phage λ carrying genR had been transferred into the recipients. Detailed procedures for colony PCR are included in the Supporting Information (Text S2).
To correct for conjugation and phage delivery events that could occur on the agar plates, donor and recipient were mixed in a 1:1 ratio at different concentrations and then immediately transferred on agar plates as a control. The number of transconjugants or transductants colonies were subtracted from the counting results mentioned above.
Quantification of Prophage Integration
After 1 h of phage delivery assays, bacteria were centrifuged at 8000g for 40 s, and microbial DNA was extracted from the pellet using a QIAamp DNA Micro Kit (Qiagen, USA) following the manufacturer’s instructions. The qPCR analysis was performed to quantify E. coli DNA and λ prophage integration using primers as previously reported85 (Table 2). The qPCR mixture contained 7.5 μL of PowerUp SYBR Green Master Mix (Applied Biosystems, USA), 1 μL of DNA template, 0.3 μM of each forward and reverse primers, and DNA-free water to a total volume of 15 μL. Triplicate qPCR reactions were conducted on a CFX96TM real-time system (Bio-Rad, USA) with the temperature setup provided in the Supporting Information (Text S3).
Table 2. Primers for qPCR Analysis.
| target genes | primer sequence | annealing temp. (°C) | purposes | refs |
|---|---|---|---|---|
| attB-upstream | 5′-GCCGACAACAAAGTCAGGTT-3′ | 59.5 | detecting E. coli DNA | (85) |
| 5′-AAAAGAAGCGCAGAATTTCG-3′ | ||||
| attB-attP | 5′-AGACGGGAAACTGAAAATGTG-3′ | 59.5 | detecting the integration of λ phage genome | (85) |
| 5′-CTGATAGTGACCTGTTCGTTGC-3′ | ||||
| traI | 5′-TTGAACTCTGCTGTGCCGTTGAC-3′ | 58.0 | transcriptomic analysis of conjugation | a |
| ATCACGAAGAAGGGAACCATCATC-3′ | ||||
| traJ | 5’-GACGTGCTCATAGTCCA-3′ | 58.0 | a | |
| 5’-TGTACTGCCTTCCAGAC-3′ | ||||
| trfAp | 5′-GAAGCCCATCGCCGTCGCCTGTAG-3′ | 58.0 | (84) | |
| 5′-GCCGACGATGACGAACTGGTGTGG-3′ | ||||
| lamB | 5′-GGTGGTTCTTCCTCTTTC-3′ | 59.0 | transcriptomic analysis of phage delivery assays | a |
| 5′-CGACACCCAGTTCTAATG-3′ | ||||
| malT | 5′-GCAGGCCGGACGTAAAAGTGAC-3′ | 59.0 | a | |
| 5′-ATGCTGTTCCAGTTCCGGCA-3′ |
Primers were custom designed and ordered from Integrated DNA Technology (IDT, USA).
Differential Gene Expression Analysis
Bacteria samples were collected by centrifugation after 1 h of conjugation and phage delivery assays. Total bacterial RNA was extracted from the pellets using a PureLink RNA mini kit (Invitrogen, USA) with on-column DNase treatment (PureLink DNase Set, Invitrogen, USA) to remove residual DNA. The quality and quantity of RNA were determined by using a NanoDrop 1000 spectrophotometer (Thermo Scientific, USA). RNA was transcribed to cDNA by reverse transcription PCR (RT-PCR) using a High-Capacity RNA-to-cDNA kit (Invitrogen, USA).
qPCR was performed to quantify the transcribed cDNA of conjugation-related genes (traI, traJ, and trbAp) and phage delivery-related genes (lamB and malT) (Table 2). 16S rRNA gene was included as a reference gene.86 Differential gene expression relative to the 16S rRNA gene was quantified by the 2–ΔΔCT method,87 and the cycle threshold (CT) values used were the means of independent triplicates. Heatmap presenting the overall RT-PCR and qPCR data of targeted genes was performed in Origin Pro 2021. Details of the RT-PCR and qPCR protocols are available in the Supporting Information (Text S3).
Simulation of Cell (or Phage)–Cell Collision Frequency
The frequency of collision Z (total collision event/s) between donor (E. coli cells for conjugation or phages) and recipient (E. coli cells) at different water turbulence levels was simulated according to the collision theory72,74 with several assumptions: (1) bacterial cells and phages all have a spherical shape with hydrodynamic diameters of 1.2 × 10–688 and 7.0 × 10–8 m;89 (2) Brownian motion prevails under static conditions (Re = 0) and neither cells nor phages move by themselves, although E. coli strains used in this study have flagella; (3) the concentrations of the donor and recipient cells are constant through the assays; and (4) under nonstatic conditions (Re > 0), the velocities (m/s) of cells and phages (u) are the same as water velocity, which is simplified as
| 2 |
where r is the distance from cells (or phages) to the center of the flask (m).
Under static conditions (Re = 0), the collision between donors and recipients resulted from Brownian motions (ZB). Collision frequency Z (total collision event/s) was determined as90
| 3 |
where α is the collision efficiency (assumed as 1), k is the Boltzmann constant (J/K), T is the thermodynamic temperature (K), μ is the fluid viscosity (Pa·s), dd is the hydrodynamic diameter of donor (m), dr is the hydrodynamic diameter of recipient (m), Nd is the donor concentration [CFU (or PFU)/m3], Nr is the recipient concentration (CFU/m3), and V is the fluid volume (m3).
Under nonstatic conditions (Re > 0), the collision between ARG donor and recipient resulted from both Brownian motion (eq 3) and shear force. The simulation of shear force-related collision frequency,88ZS (total collision event/s), was determined as
| 4 |
where H is the height of the fluid (m) calculated from the fluid volume and flask diameter.
Details of formula derivation and the values of aforementioned parameters are provided in the Supporting Information (Text S4 and Table S1).
Statistical Analyses
ANOVA analysis was performed to compare the differences between conjugation and phage delivery frequencies under various experimental settings. Pearson correlation analysis was also conducted to characterize the relationship between the system-dependent factors and corresponding genR transfer frequency using SPSS 10.0 software. Differences were considered to be significant at the 95% confidence level (p < 0.05).
Results and Discussion
Phages Deliver ARGs at a Comparable Frequency to Conjugation at Low Bacterial Concentration
Both conjugation and phage delivery frequencies (determined by plate counting) were positively correlated with bacterial concentration, ranging from 104 to 108 CFU/mL (Pearson correlation coefficient r = 0.95, p = 0.013 for conjugation; r = 0.92, p = 0.027 for phage delivery) (Figure 2a), which facilitates contact between ARG donor and recipient, as previously reported.79,91 The relative importance of conjugation versus phage delivery was also affected by bacterial concentration. At high bacterial concentration, the frequency of conjugation was significantly higher than that of genR delivery by phages, exceeding it by 9.3-, 5.6-, and 5.9-fold at 108, 107, and 106 CFU/mL, respectively. There was no significant difference between conjugation and phage-mediated transfer frequencies when bacterial concentration was at 105 or 104 CFU/mL (8.8 ± 3.2 × 10–6 vs 5.9 ± 2.4 × 10–6 and 1.2 ± 0.9 × 10–6 vs 1.1 ± 0.9 × 10–6, respectively), indicating that the contribution of phages to ARG dissemination can rival that of conjugation at low bacterial concentration.
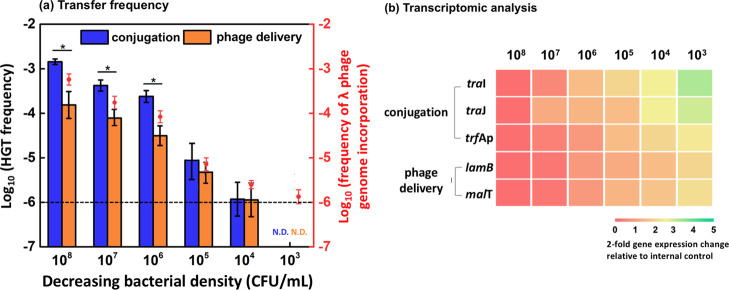
Figure 2.
Lower frequency but higher relative importance of genR transfer via phage delivery (relative to conjugation) occurs at lower bacterial concentration. (a) Transfer frequency was determined after 1 h of conjugation and phage delivery assays in minimal medium with 3.0 g/L glucose and shaken at an Re of 5 × 104. The genR donor (E. coli DH5α harboring plasmid RP4 for conjugation assays or λ phage for phage delivery assays) and recipient (E. coli MG1655) were mixed at a 1:1 ratio. Frequency determined by the plate counting method is presented as bars with the detection limit (10–6) indicated by the horizontal dashed line; orange dots represent the frequency determined by qPCR analysis. Asterisks (*) indicate significant differences (p < 0.05) between conjugation and phage delivery frequency based on Student’s t-test. “N.D.” refers to “not detected”. (b) Upregulation of conjugation-related genes (traI, traJ, and trbAp) and phage delivery-related genes (lamB and malT) at lower bacterial concentration (twofold gene expression change). The top X-axis depicts decreasing bacterial concentration (CFU/mL). The expression level at a bacterial concentration of 108 CFU/mL was used as the control. Error bars depict ± one standard deviation from the mean of at least three independent replicates.
No ARG recipients were detected by plate counting at a bacterial concentration of 103 CFU/mL, which contrasts previous reports that bacterial concentration thresholds for phage infection can be as low as 102 CFU/mL.82 Thus, further analysis using qPCR was needed to evaluate the phage delivery frequency at the genetic level. The frequency of ARG delivery by phages was determined as the percentage of recipient E. coli that was infected by λ phage and incorporated phage genome in the chromosome (Figure 2a) and was generally a little higher (0–0.5 log) than that determined by the plate counting method. The difference in frequency detected by these two methods might be due to insufficient accumulation of genR-coded gentamycin acetyltransferase92 in some of the phage-infected recipients. The λ phage genome incorporation was detected at a bacterial concentration of 103 CFU/mL, indicating that ARG delivery by phages might be more frequent than by conjugation at such low bacterial concentrations.
The expression of genes related to conjugation and phage delivery was upregulated as bacterial concentration decreased, possibly due to more abundant nutrients per cell at lower bacterial concentration to support energy-dependent gene expression93 (Figure 2b). The expression levels of conjugation-related genes traI (coding DNA relaxosome required for conjugation55), traJ (coding positive regulator of plasmid transfer operon94), and trfAp (coding positive regulator of plasmid transfer and replication system95) at a bacterial concentration of 103 CFU/mL were 9.21-, 7.82-, and 5.51-fold higher than that at 108 CFU/mL, respectively. Phage delivery-related gene lamB (coding the receptor of λ phage96) and malT (coding the positive regulator of lamB97) were overexpressed by 4.3- and 4.8-fold. The upregulation of conjugation-related genes was more significant than that of phage delivery, possibly due to quorum sensing regulation.98 The regulation of gene expression did not explain the influence of bacterial concentration on the relative importance of conjugation versus phage delivery. The frequency of ARG delivery by phages was less dependent on bacterial concentration than that of conjugation, which collaborates previous studies,38,82 although the reason for this observation remains unclear.
Water Turbulence Significantly Affects the Relative Frequency of ARG Conjugation Versus Phage Delivery
Water turbulence, quantified as Re, had a considerable effect on the frequency of ARG dissemination by conjugation versus phage delivery, and shaped their relative importance (Figure 3). The transfer frequency of genR peaked at Re of 5 × 104 for both conjugation and phage delivery, with a conjugation frequency 5.6-fold higher than that via phages. As Re decreased or increased from the peak value, the frequencies of ARG delivery by both conjugation and phages decreased due to lower collision frequency or higher detachment rate, respectively.88 However, the importance of phage delivery relative to conjugation increased under either quiescent or turbulent conditions. Specifically, the frequency of genR transfer mediated by phages was comparable to conjugation at Re of 0 (static) and 5 × 105 as no significant difference was observed. This implies that ARG transfer by phages should receive increasing attention in turbulent aquatic systems where Re can exceed 5 × 105 or in quiescent systems where a high level of phage-borne ARGs is detected.
Figure 3.
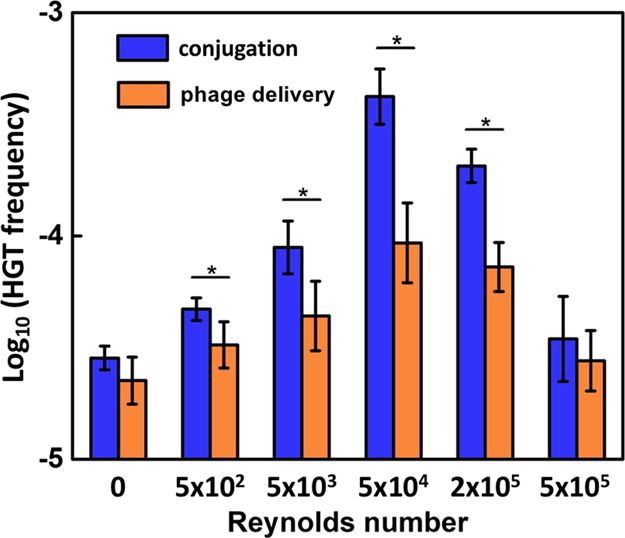
Turbulence (represented by Reynolds number) significantly impacts the relative frequency of conjugation vs phage delivery for the horizontal transfer of genR in a bell-shaped fashion. The transfer frequency was determined after 1 h of conjugation and phage delivery assays in minimal medium containing 3.0 g/L glucose with bacterial concentration at 107 CFU/mL. The genR donor (E. coli DH5α harboring plasmid RP4 for conjugation assays or λ phage for phage delivery assays) and recipient (E. coli MG1655) were mixed at a 1:1 ratio. Asterisks (*) indicate significant differences (p < 0.05) between conjugation and phage delivery frequency based on Student’s t-test. Error bars depict ± one standard deviation from the mean of at least three independent replicates.
Transcriptomic analysis indicated that genetic regulation was not as influential as turbulence in shaping the relative importance of conjugation versus phage delivery, as no significant difference was observed for the expression level of selected conjugation- and phage delivery-related genes among different turbulence levels (Figure S4).
Collision theory might help explain how hydrodynamic conditions influence ARG delivery because they greatly influence cell (or phage)-to-cell contact frequency (initiating gene transfer) and the stability of mating pairs (for the completion of ARG transfer).99 Under static conditions (Re = 0, no turbulence shear), the simulated frequency of collision between the conjugation donor (E. coli cells) and recipient (E. coli cells) was lower than that of phage delivery (phages as the donor and E. coli cells as the recipient), which helps explain the insignificant difference between conjugation and phage delivery frequencies (Figure 4a). Collision under static conditions was assumed to result only from the Brownian motion,100,101 according to which the smaller particles (i.e., phages) diffuse faster than the larger ones (i.e., E. coli cells) and therefore lead to higher collision frequency. The self-motion of E. coli cells powered by flagella100 was not included in this simple model.
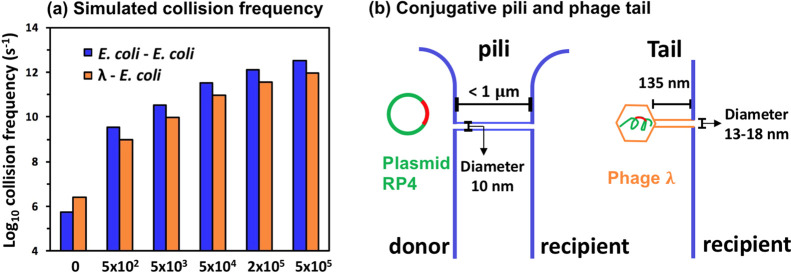
Figure 4.
Higher water turbulence increases the theoretical cell (or phage)-to-cell collision frequency and interferes with the stability of ARG transfer channels. (a) Collision frequency (total collision events/s) under static (Re = 0) and dynamic conditions (Re > 0) was simulated based on the Brownian motion (eq 3) and turbulence shear (eq 4), respectively. Brownian motion was not considered in the simulation under turbulent conditions because its induced collision has a frequency that is over 3 orders of magnitude lower than that of shear-induced collision (eq S7). (b) Shear-induced detachment frequency would be higher for conjugating cells than for attached phages due to postulated weaker links formed by thinner and longer multiprotein, hair-like links.57,102−105
Water turbulence leads to a trade-off between increased cell (or phage)-to-cell collision rate and higher propensity for cell (or phage)–cell detachment due to the shearing force,100 which explains the bell-shaped ARG transfer data (Figure 3). Elevated turbulence levels resulted in higher collision frequency and enhanced both conjugation and phage delivery at Re ranging from 0 to 5 × 104 (Figure 4a). The cell-to-cell collision frequency was higher than that of phage-to-cell collision because E. coli cells have a larger hydrodynamic radius,88,89 which contributed to higher peak conjugation frequency than phage delivery frequency (Figure 3). Decreased ARG transfer frequency at Re higher than 5 × 104 was probably due to an increase in the detachment rate. A lack of literature data on the stability of RP4 conjugation mating pairs and λ phages attached to E. coli under different Re values precludes accurate simulation of detachment. Nevertheless, the higher likelihood of detachment of conjugating cells than phages adsorbed onto cells at high Re values is supported by the fact that conjugative pili produced by the RP4 conjugal system would be a more fragile link than that formed by the phage tails that attach to bacteria (Figure 4b). Specifically, both links consist of multiprotein hair-like structures, and while RP4 pili typically measure 10 nm in diameter102 and under 1.0 μm in length (retractable),103,104 the λ phage tails are only about 135 nm long105 and have a thicker diameter of 13–18 nm.57,105
Further studies should corroborate the observed influence of hydrodynamic conditions on the relative importance of conjugation versus transduction using lower recipient bacterial concentrations representative of natural aquatic systems (e.g., 102–105 CFU/mL106,107), which would advance the understanding of ARG transfer from wastewater treatment plants or animal agriculture sources to natural environments (e.g., freshwater).
Environmental Implications
ARG dissemination mediated by phages is receiving increasing attention due to the abundance of phage-borne ARGs. This work demonstrates that both bacterial concentration and water turbulence can be influential in determining the relative importance of conjugation versus phage delivery of ARGs, with the latter reaching comparable transfer frequencies at relatively low cell concentrations under either quiescent or turbulent conditions that hinder conjugation. Accordingly, ARG transfer via phages may become relatively important in oligotrophic aquatic systems (Figure S5) not only due to low bacterial concentration but also because phage transfer is less energy intensive than conjugation93,108−112 and thus it is potentially less susceptible to lower energy source availability.
The fact that ARG transduction may be important in some systems underscores the need to control phage-borne ARGs, especially at hotspots that may serve as sources for environmental dissemination. For example, municipal wastewater treatment plants are widely acknowledged as potential reservoirs of ARGs,113 and high concentrations of phage-borne ARGs have been detected in the raw wastewater and activated sludge—sometimes comparable to intracellular ARG concentrations.13 Phages can persist and even propagate in activated sludge processes,114 sometimes leading to higher phage concentrations in the effluent (8.4 × 107 virion/mL) compared to raw wastewater (2.2 × 107 virion/mL).115 Effluent disinfection processes such as chlorination usually have a high efficacy of bacteria removal but could be less effective on phages.12,116,117 The phages discharged from ARG hotspots may migrate to new environments and might transfer antibiotic resistance to some indigenous bacteria, enhancing resistance propagation. Therefore, the efficacy of disinfection strategies on phage removal ought to be assessed.118
We recognize that the environmental factors considered by this study may affect ARG transfer differently in biofilms that prevail in many systems. Furthermore, other system-specific factors such as pH, ionic strength, chemical agents, and solar ultraviolet radiation might also influence the relative importance of conjugation versus phage delivery to ARG dissemination due to their differential effects on HGT vectors and recipients.51 Nevertheless, this study underscores the importance of transduction as an ARG propagation mechanism under hydrodynamic and cell concentration conditions that favor phage infection, which may include some groundwater systems,63−65,119 rivers and streams,66−68 and wastewater treatment processes.69−72 The data also suggest the importance to address hotspots with high concentrations of phage-borne ARGs, which may require additional attention to mitigate this relatively overlooked, potential antibiotic resistance dissemination pathway.
Acknowledgments
This work was supported by the U.S. National Science Foundation (NSF) ERC on Nanotechnology-Enabled Water Treatment (EEC-1449500). R.S. received financial support from China Scholarship of Council (CSC). We gratefully acknowledge Jacques Mathieu, Kaiqi Yang, and Xiaochuan Huang for their valuable insights and suggestions.
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acsenvironau.1c00027.
Phage manipulations, genetic analysis, collision simulation, and complementary results (PDF)
Author Present Address
† College of Environment and Resource Sciences, Zhejiang University, Hangzhou, Zhejiang, 310058, China
The authors declare no competing financial interest.
Supplementary Material
References
- O’Neill J.Review on Antimicrobial Resistance: Tackling a Crisis for the Health and Wealth of Nations; HM Government: London, 2014. [Google Scholar]
- Nelson R. E.; Hatfield K. M.; Wolford H.; Samore M. H.; Scott R. D.; Reddy S. C.; Olubajo B.; Paul P.; Jernigan J. A.; Baggs J. National estimates of healthcare costs associated with multidrug-resistant bacterial infections among hospitalized patients in the United States. Clin. Infect. Dis. 2021, 72, S17–S26. 10.1093/cid/ciaa1581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vikesland P. J.; Pruden A.; Alvarez P. J. J.; Aga D.; Bürgmann H.; Li X.-d.; Manaia C. M.; Nambi I.; Wigginton K.; Zhang T.; Zhu Y.-G. Toward a comprehensive strategy to mitigate dissemination of environmental sources of antibiotic resistance. Environ. Sci. Technol. 2017, 51, 13061–13069. 10.1021/acs.est.7b03623. [DOI] [PubMed] [Google Scholar]
- Jiang X.; Ellabaan M. M. H.; Charusanti P.; Munck C.; Blin K.; Tong Y.; Weber T.; Sommer M. O. A.; Lee S. Y. Dissemination of antibiotic resistance genes from antibiotic producers to pathogens. Nat. Commun. 2017, 8, 15784. 10.1038/ncomms15784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- von Wintersdorff C. J.; Penders J.; van Niekerk J. M.; Mills N. D.; Majumder S.; van Alphen L. B.; Savelkoul P. H.; Wolffs P. F. Dissemination of antimicrobial resistance in microbial ecosystems through horizontal gene transfer. Front. Microbiol. 2016, 7, 173. 10.3389/fmicb.2016.00173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dang B.; Mao D.; Xu Y.; Luo Y. Conjugative multi-resistant plasmids in Haihe River and their impacts on the abundance and spatial distribution of antibiotic resistance genes. Water Res. 2017, 111, 81–91. 10.1016/j.watres.2016.12.046. [DOI] [PubMed] [Google Scholar]
- Che Y.; Xia Y.; Liu L.; Li A. D.; Yang Y.; Zhang T. Mobile antibiotic resistome in wastewater treatment plants revealed by Nanopore metagenomic sequencing. Microbiome 2019, 7, 44. 10.1186/s40168-019-0663-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston C.; Martin B.; Fichant G.; Polard P.; Claverys J.-P. Bacterial transformation: distribution, shared mechanisms and divergent control. Nat. Rev. Microbiol. 2014, 12, 181–196. 10.1038/nrmicro3199. [DOI] [PubMed] [Google Scholar]
- Lekunberri I.; Subirats J.; Borrego C. M.; Balcázar J. L. Exploring the contribution of bacteriophages to antibiotic resistance. Environ. Pollut. 2017, 220, 981–984. 10.1016/j.envpol.2016.11.059. [DOI] [PubMed] [Google Scholar]
- McGrath S.; Sinderen D. v.. Bacteriophage: Genetics and Molecular Biology; Caister Academic Press, 2007. [Google Scholar]
- Beumer A.; Robinson J. B. A broad-host-range, generalized transducing phage (SN-T) acquires 16S rRNA genes from different genera of bacteria. Appl. Environ. Microbiol. 2005, 71, 8301–8304. 10.1128/aem.71.12.8301-8304.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calero-Cáceres W.; Muniesa M. Persistence of naturally occurring antibiotic resistance genes in the bacteria and bacteriophage fractions of wastewater. Water Res. 2016, 95, 11–18. 10.1016/j.watres.2016.03.006. [DOI] [PubMed] [Google Scholar]
- Calero-Cáceres W.; Melgarejo A.; Colomer-Lluch M.; Stoll C.; Lucena F.; Jofre J.; Muniesa M. Sludge as a potential important source of antibiotic resistance genes in both the bacterial and bacteriophage fractions. Environ. Sci. Technol. 2014, 48, 7602–7611. 10.1021/es501851s. [DOI] [PubMed] [Google Scholar]
- Yang Y.; Shi W.; Lu S.-Y.; Liu J.; Liang H.; Yang Y.; Duan G.; Li Y.; Wang H.; Zhang A. Prevalence of antibiotic resistance genes in bacteriophage DNA fraction from Funan River water in Sichuan, China. Sci. Total Environ. 2018, 626, 835–841. 10.1016/j.scitotenv.2018.01.148. [DOI] [PubMed] [Google Scholar]
- Calero-Cáceres W.; Méndez J.; Martín-Díaz J.; Muniesa M. The occurrence of antibiotic resistance genes in a Mediterranean river and their persistence in the riverbed sediment. Environ. Pollut. 2017, 223, 384–394. 10.1016/j.envpol.2017.01.035. [DOI] [PubMed] [Google Scholar]
- Ross J.; Topp E. Abundance of antibiotic resistance genes in bacteriophage following soil fertilization with dairy manure or municipal biosolids, and evidence for potential transduction. Appl. Environ. Microbiol. 2015, 81, 7905–7913. 10.1128/aem.02363-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larrañaga O.; Brown-Jaque M.; Quirós P.; Gómez-Gómez C.; Blanch A. R.; Rodríguez-Rubio L.; Muniesa M. Phage particles harboring antibiotic resistance genes in fresh-cut vegetables and agricultural soil. Environ. Int. 2018, 115, 133–141. 10.1016/j.envint.2018.03.019. [DOI] [PubMed] [Google Scholar]
- He T.; Jin L.; Xie J.; Yue S.; Fu P.; Li X. Intracellular and extracellular antibiotic resistance genes in airborne PM2. 5 for respiratory exposure in urban areas. Environ. Sci. Technol. Lett. 2021, 8, 128–134. 10.1021/acs.estlett.0c00974. [DOI] [Google Scholar]
- Quirós P.; Colomer-Lluch M.; Martínez-Castillo A.; Miró E.; Argente M.; Jofre J.; Navarro F.; Muniesa M. Antibiotic resistance genes in the bacteriophage DNA fraction of human fecal samples. Antimicrob. Agents Chemother. 2014, 58, 606–609. 10.1128/aac.01684-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colomer-Lluch M.; Imamovic L.; Jofre J.; Muniesa M. Bacteriophages carrying antibiotic resistance genes in fecal waste from cattle, pigs, and poultry. Antimicrob. Agents Chemother. 2011, 55, 4908–4911. 10.1128/aac.00535-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subirats J.; Sànchez-Melsió A.; Borrego C. M.; Balcázar J. L.; Simonet P. Metagenomic analysis reveals that bacteriophages are reservoirs of antibiotic resistance genes. Int. J. Antimicrob. Agents 2016, 48, 163–167. 10.1016/j.ijantimicag.2016.04.028. [DOI] [PubMed] [Google Scholar]
- Yang D.; Wang J.; Qiu Z.; Jin M.; Shen Z.; Chen Z.; Wang X.; Zhang B.; Li J.-W. Horizontal transfer of antibiotic resistance genes in a membrane bioreactor. J. Biotechnol. 2013, 167, 441–447. 10.1016/j.jbiotec.2013.08.004. [DOI] [PubMed] [Google Scholar]
- Chandrasekaran S.; Venkatesh B.; Lalithakumari D. Transfer and expression of a multiple antibiotic resistance plasmid in marine bacteria. Curr. Microbiol. 1998, 37, 347–351. 10.1007/s002849900390. [DOI] [PubMed] [Google Scholar]
- Binh C. T. T.; Heuer H.; Kaupenjohann M.; Smalla K. Piggery manure used for soil fertilization is a reservoir for transferable antibiotic resistance plasmids. FEMS Microbiol. Ecol. 2008, 66, 25–37. 10.1111/j.1574-6941.2008.00526.x. [DOI] [PubMed] [Google Scholar]
- Li L.; Dechesne A.; He Z.; Madsen J. S.; Nesme J.; Sørensen S. J.; Smets B. F. Estimating the transfer range of plasmids encoding antimicrobial resistance in a wastewater treatment plant microbial community. Environ. Sci. Technol. Lett. 2018, 5, 260–265. 10.1021/acs.estlett.8b00105. [DOI] [Google Scholar]
- Inoue D.; Sei K.; Soda S.; Ike M.; Fujita M. Potential of predominant activated sludge bacteria as recipients in conjugative plasmid transfer. J. Biosci. Bioeng. 2005, 100, 600–605. 10.1263/jbb.100.600. [DOI] [PubMed] [Google Scholar]
- Soda S.; Otsuki H.; Inoue D.; Tsutsui H.; Sei K.; Ike M. Transfer of antibiotic multiresistant plasmid RP4 from Escherichia coli to activated sludge bacteria. J. Biosci. Bioeng. 2008, 106, 292–296. 10.1263/jbb.106.292. [DOI] [PubMed] [Google Scholar]
- Jacquiod S.; Brejnrod A.; Morberg S. M.; Abu Al-Soud W.; Sørensen S. J.; Riber L. Deciphering conjugative plasmid permissiveness in wastewater microbiomes. Mol. Ecol. 2017, 26, 3556–3571. 10.1111/mec.14138. [DOI] [PubMed] [Google Scholar]
- Jutkina J.; Rutgersson C.; Flach C.-F.; Joakim Larsson D. G. An assay for determining minimal concentrations of antibiotics that drive horizontal transfer of resistance. Sci. Total Environ. 2016, 548–549, 131–138. 10.1016/j.scitotenv.2016.01.044. [DOI] [PubMed] [Google Scholar]
- Guo M.-T.; Yuan Q.-B.; Yang J. Distinguishing effects of ultraviolet exposure and chlorination on the horizontal transfer of antibiotic resistance genes in municipal wastewater. Environ. Sci. Technol. 2015, 49, 5771–5778. 10.1021/acs.est.5b00644. [DOI] [PubMed] [Google Scholar]
- Ubukata K.; Konno M.; Fujii R. Transduction of drug resistance to tetracycline, chloramphenicol, macrolides, lincomycin and clindamycin with phages induced from Streptococcus pyogenes. J. Antibiot. 1975, 28, 681–688. 10.7164/antibiotics.28.681. [DOI] [PubMed] [Google Scholar]
- Mazaheri Nezhad Fard R.; Barton M. D.; Heuzenroeder M. W. Bacteriophage-mediated transduction of antibiotic resistance in enterococci. Lett. Appl. Microbiol. 2011, 52, 559–564. 10.1111/j.1472-765x.2011.03043.x. [DOI] [PubMed] [Google Scholar]
- Morrison W. D.; Miller R. V.; Sayler G. S. Frequency of F116-mediated transduction of Pseudomonas aeruginosa in a freshwater environment. Appl. Environ. Microbiol. 1978, 36, 724–730. 10.1128/aem.36.5.724-730.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang S. C.; Paul J. H. Gene transfer by transduction in the marine environment. Appl. Environ. Microbiol. 1998, 64, 2780–2787. 10.1128/aem.64.8.2780-2787.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J.; Novick R. P. Phage-mediated intergeneric transfer of toxin genes. Science 2009, 323, 139–141. 10.1126/science.1164783. [DOI] [PubMed] [Google Scholar]
- Varga M.; Kuntová L.; Pantůček R.; Mašlaňová I.; Růžičková V.; Doškař J. Efficient transfer of antibiotic resistance plasmids by transduction within methicillin-resistant Staphylococcus aureus USA300 clone. FEMS Microbiol. Lett. 2012, 332, 146–152. 10.1111/j.1574-6968.2012.02589.x. [DOI] [PubMed] [Google Scholar]
- Saye D. J.; Ogunseitan O.; Sayler G. S.; Miller R. V. Potential for transduction of plasmids in a natural freshwater environment: Effect of plasmid donor concentration and a natural microbial community on transduction in Pseudomonas aeruginosa. Appl. Environ. Microbiol. 1987, 53, 987–995. 10.1128/aem.53.5.987-995.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kenzaka T.; Tani K.; Nasu M. High-frequency phage-mediated gene transfer in freshwater environments determined at single-cell level. ISME J. 2010, 4, 648–659. 10.1038/ismej.2009.145. [DOI] [PubMed] [Google Scholar]
- Zhang P.-Y.; Xu P.-P.; Xia Z.-J.; Wang J.; Xiong J.; Li Y.-Z. Combined treatment with the antibiotics kanamycin and streptomycin promotes the conjugation of Escherichia coli. FEMS Microbiol. Lett. 2013, 348, 149–156. 10.1111/1574-6968.12282. [DOI] [PubMed] [Google Scholar]
- Dahlberg C.; Bergström M.; Hermansson M. In situ detection of high levels of horizontal plasmid transfer in marine bacterial communities. Appl. Environ. Microbiol. 1998, 64, 2670–2675. 10.1128/aem.64.7.2670-2675.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Morchoe S. B.; Ogunseitan O.; Sayler G. S.; Miller R. V. Conjugal transfer of R68. 45 and FP5 between Pseudomonas aeruginosa strains in a freshwater environment. Appl. Environ. Microbiol. 1988, 54, 1923–1929. 10.1128/aem.54.8.1923-1929.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flett F.; Mersinias V.; Smith C. P. High efficiency intergeneric conjugal transfer of plasmid DNA from Escherichia coli to methyl DNA-restricting streptomycetes. FEMS Microbiol. Lett. 1997, 155, 223–229. 10.1111/j.1574-6968.1997.tb13882.x. [DOI] [PubMed] [Google Scholar]
- Mašlaňová I.; Stříbná S.; Doškař J.; Pantůček R. Efficient plasmid transduction to Staphylococcus aureus strains insensitive to the lytic action of transducing phage. FEMS Microbiol. Ecol. 2016, 363, fnw211. 10.1093/femsle/fnw211. [DOI] [PubMed] [Google Scholar]
- Raya R. R.; Klaenhammer T. R. High-frequency plasmid transduction by Lactobacillus gasseri bacteriophage φadh. Appl. Environ. Microbiol. 1992, 58, 187–193. 10.1128/aem.58.1.187-193.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kenzaka T.; Tani K.; Sakotani A.; Yamaguchi N.; Nasu M. High-frequency phage-mediated gene transfer among Escherichia coli cells, determined at the single-cell level. Appl. Environ. Microbiol. 2007, 73, 3291–3299. 10.1128/aem.02890-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alonso G.; Baptista K.; Ngo T.; Taylor D. E. Transcriptional organization of the temperature-sensitive transfer system from the IncHI1 plasmid R27. Microbiol 2005, 151, 3563–3573. 10.1099/mic.0.28256-0. [DOI] [PubMed] [Google Scholar]
- Stallions D. R.; Curtiss R. 3rd Bacterial conjugation under anaerobic conditions. J. Bacteriol. 1972, 111, 294–295. 10.1128/jb.111.1.294-295.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mills A. L.; Herman J. S.; Hornberger G. M.; DeJesús T. H. Effect of solution ionic strength and iron coatings on mineral grains on the sorption of bacterial cells to quartz sand. Appl. Environ. Microbiol. 1994, 60, 3300–3306. 10.1128/aem.60.9.3300-3306.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang H.; Chen Y.; Zheng X.; Su Y.; Wan R.; Yang S. Distribution of tetracycline resistance genes in anaerobic treatment of waste sludge: The role of pH in regulating tetracycline resistant bacteria and horizontal gene transfer. Bioresour. Technol. 2016, 218, 1284–1289. 10.1016/j.biortech.2016.07.097. [DOI] [PubMed] [Google Scholar]
- Jutkina J.; Marathe N. P.; Flach C.-F.; Larsson D. G. J. Antibiotics and common antibacterial biocides stimulate horizontal transfer of resistance at low concentrations. Sci. Total Environ. 2018, 616–617, 172–178. 10.1016/j.scitotenv.2017.10.312. [DOI] [PubMed] [Google Scholar]
- Drudge C. N.; Warren L. A. Prokaryotic horizontal gene transfer in freshwater lakes: Implications of dynamic biogeochemical zonation. J. Environ. Prot. 2012, 03, 1634–1654. 10.4236/jep.2012.312181. [DOI] [Google Scholar]
- Chiura H. X.; Kogure K.; Hagemann S.; Ellinger A.; Velimirov B. Evidence for particle-induced horizontal gene transfer and serial transduction between bacteria. FEMS Microbiol. Ecol. 2011, 76, 576–591. 10.1111/j.1574-6941.2011.01077.x. [DOI] [PubMed] [Google Scholar]
- Volkova V. V.; Lu Z.; Besser T.; Gröhn Y. T. Modeling the infection dynamics of bacteriophages in enteric Escherichia coli: Estimating the contribution of transduction to antimicrobial gene spread. Appl. Environ. Microbiol. 2014, 80, 4350–4362. 10.1128/aem.00446-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lanka E.; Barth P. T. Plasmid RP4 specifies a deoxyribonucleic-acid primase involved in its conjugal transfer and maintenance. J. Bacteriol. 1981, 148, 769–781. 10.1128/jb.148.3.769-781.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haase J.; Lurz R.; Grahn A. M.; Bamford D. H.; Lanka E. Bacterial conjugation mediated by plasmid RP4: RSF1010 mobilization, donor-specific phage propagation, and pilus production require the same Tra2 core components of a proposed DNA transport complex. J. Bacteriol. 1995, 177, 4779–4791. 10.1128/jb.177.16.4779-4791.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Virus Taxonomy: 2020 Release. International Committee on Taxonomy of Viruses (ICTV) March 2019.
- Pell L. G.; Kanelis V.; Donaldson L. W.; Lynne Howell P.; Davidson A. R. The phage λ major tail protein structure reveals a common evolution for long-tailed phages and the type VI bacterial secretion system. Proc. Natl. Acad. Sci. U. S. A. 2009, 106, 4160–4165. 10.1073/pnas.0900044106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heuer H.; Krogerrecklenfort E.; Wellington E. M. H.; Egan S.; Elsas J. D.; Overbeek L.; Collard J.-M.; Guillaume G.; Karagouni A. D.; Nikolakopoulou T. L.; Smalla K. Gentamicin resistance genes in environmental bacteria: Prevalence and transfer. FEMS Microbiol. Ecol. 2002, 42, 289–302. 10.1111/j.1574-6941.2002.tb01019.x. [DOI] [PubMed] [Google Scholar]
- Chen Y.; Mukherjee S.; Hoffmann M.; Kotewicz M. L.; Young S.; Abbott J.; Luo Y.; Davidson M. K.; Allard M.; McDermott P.; Zhao S. Whole-genome sequencing of gentamicin-resistant Campylobacter coli isolated from U.S. retail meats reveals novel plasmid-mediated aminoglycoside resistance genes. Antimicrob. Agents Chemother. 2013, 57, 5398–5405. 10.1128/aac.00669-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rufener S.; Mäusezahl D.; Mosler H.-J.; Weingartner R. Quality of drinking-water at source and point-of-consumption—drinking cup as a high potential recontamination risk: A field study in Bolivia. J. Health Popul. Nutr. 2010, 28, 34–41. 10.3329/jhpn.v28i1.4521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bréchet C.; Plantin J.; Sauget M.; Thouverez M.; Talon D.; Cholley P.; Guyeux C.; Hocquet D.; Bertrand X. Wastewater treatment plants release large amounts of extended-spectrum β-lactamase–producing Escherichia coli into the environment. Clin. Infect. Dis. 2014, 58, 1658–1665. 10.1093/cid/ciu190. [DOI] [PubMed] [Google Scholar]
- Mandilara G. D.; Smeti E. M.; Mavridou A. T.; Lambiri M. P.; Vatopoulos A. C.; Rigas F. P. Correlation between bacterial indicators and bacteriophages in sewage and sludge. FEMS Microbiol. Lett. 2006, 263, 119–126. 10.1111/j.1574-6968.2006.00414.x. [DOI] [PubMed] [Google Scholar]
- Wang W.; Zhang G. Numerical simulation of groundwater flowing to horizontal seepage wells under a river. Hydrogeol. J. 2007, 15, 1211–1220. 10.1007/s10040-007-0171-3. [DOI] [Google Scholar]
- Karay G.; Hajnal G. Modelling of groundwater flow in fractured rocks. Procedia Environ. Sci. 2015, 25, 142–149. 10.1016/j.proenv.2015.04.020. [DOI] [Google Scholar]
- Qian J.; Zhan H.; Zhao W.; Sun F. Experimental study of turbulent unconfined groundwater flow in a single fracture. J. Hydrol. 2005, 311, 134–142. 10.1016/j.jhydrol.2005.01.013. [DOI] [Google Scholar]
- Malverti L.; Lajeunesse E.; Métivier F. Small is beautiful: Upscaling from microscale laminar to natural turbulent rivers. J. Geophys. Res.: Earth Surf. 2008, 113, 1–14. 10.1029/2007jf000974. [DOI] [Google Scholar]
- Birjandi A. H.; Bibeau E.. Reynolds Effects on the Freewheeling Behavior of a High Solidity Vertical River Kinetic Turbine; ASME Interntional Mechanical Engineering Congress and Exposition, 2010; pp 1–6. [Google Scholar]
- Uijttewaal W. S. J.; Lehmann D.; Mazijk A. v. Exchange processes between a river and its groyne fields: Model experiments. J. Hydraul. Eng. 2001, 127, 928–936. 10.1061/(asce)0733-9429(2001)127:11(928). [DOI] [Google Scholar]
- Rosso D.; Stenstrom M. K. Alpha factors in full-scale wastewater aeration systems. Proc.-Water Environ. Fed. 2006, 2006, 4853–4863. 10.2175/193864706783762940. [DOI] [PubMed] [Google Scholar]
- Turano E.; Curcio S.; De Paola M. G.; Calabrò V.; Iorio G. An integrated centrifugation–ultrafiltration system in the treatment of olive mill wastewater. J. Membr. Sci. 2002, 209, 519–531. 10.1016/s0376-7388(02)00369-1. [DOI] [Google Scholar]
- Guo W. S.; Vigneswaran S.; Ngo H.; Xing W. Experimental investigation on acclimatized wastewater for membrane bioreactors. Desalination 2007, 207, 383–391. 10.1016/j.desal.2006.07.013. [DOI] [Google Scholar]
- Xu Y.; Wu Y.; Sun Q. Flow Characteristics of the raw sewage for the design of sewage-source heat pump systems. Sci. World J. 2014, 2014, 1–6. 10.1155/2014/503624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Büchs J.; Maier U.; Milbradt C.; Zoels B. Power consumption in shaking flasks on rotary shaking machines: I. Power consumption measurement in unbaffled flasks at low liquid viscosity. Biotechnol. Bioeng. 2000, 68, 589–593. . [DOI] [PubMed] [Google Scholar]
- Foladori P.; Bruni L.; Tamburini S.; Ziglio G. Direct quantification of bacterial biomass in influent, effluent and activated sludge of wastewater treatment plants by using flow cytometry. Water Res. 2010, 44, 3807–3818. 10.1016/j.watres.2010.04.027. [DOI] [PubMed] [Google Scholar]
- Wallden K.; Rivera-Calzada A.; Waksman G. Type IV secretion systems: Versatility and diversity in function. Cell. Microbiol. 2010, 12, 1203–1212. 10.1111/j.1462-5822.2010.01499.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quandt J.; Clark R. G.; Venter A. P.; Clark S. R. D.; Twelker S.; Hynes M. F. Modified RP4 and Tn5-Mob derivatives for facilitated manipulation of large plasmids in Gram-negative bacteria. Plasmid 2004, 52, 1–12. 10.1016/j.plasmid.2004.04.002. [DOI] [PubMed] [Google Scholar]
- Fox R. E.; Zhong X.; Krone S. M.; Top E. M. Spatial structure and nutrients promote invasion of IncP-1 plasmids in bacterial populations. ISME J. 2008, 2, 1024–1039. 10.1038/ismej.2008.53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu P.; Mathieu J.; Li M.; Dai Z.; Alvarez P. J. J. Isolation of polyvalent bacteriophages by sequential multiple-host approaches. Appl. Environ. Microbiol. 2016, 82, 808–815. 10.1128/aem.02382-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Handel N.; Otte S.; Jonker M.; Brul S.; ter Kuile B. H. Factors that affect transfer of the IncI1 beta-lactam resistance plasmid pESBL-283 between E. coli strains. PLoS One 2015, 10, e0123039 10.1371/journal.pone.0123039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beuls E.; Modrie P.; Deserranno C.; Mahillon J. High-salt stress conditions increase the pAW63 transfer frequency in Bacillus thuringiensis. Appl. Environ. Microbiol. 2012, 78, 7128–7131. 10.1128/aem.01105-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han X.; Lv P.; Wang L.-G.; Long F.; Ma X.-L.; Liu C.; Feng Y.-J.; Yang M.-F.; Xiao X. Impact of nano-TiO2 on horizontal transfer of resistance genes mediated by filamentous phage transduction. Environ. Sci.: Nano 2020, 7, 1214–1224. 10.1039/c9en01279f. [DOI] [Google Scholar]
- Kokjohn T. A.; Sayler G. S.; Miller R. V. Attachment and replication of Pseudomonas aeruginosa bacteriophages under conditions simulating aquatic environments. J. Gen. Microbiol. 1991, 137, 661–666. 10.1099/00221287-137-3-661. [DOI] [Google Scholar]
- Yu P.; Mathieu J.; Lu G. W.; Gabiatti N.; Alvarez P. J. Control of antibiotic-resistant bacteria in activated sludge using polyvalent phages in conjunction with a production host. Environ. Sci. Technol. Lett. 2017, 4, 137–142. 10.1021/acs.estlett.7b00045. [DOI] [Google Scholar]
- Wang Q.; Mao D.; Luo Y. Ionic liquid facilitates the conjugative transfer of antibiotic resistance genes mediated by plasmid RP4. Environ. Sci. Technol. 2015, 49, 8731–8740. 10.1021/acs.est.5b01129. [DOI] [PubMed] [Google Scholar]
- Shao Q.; Trinh J. T.; McIntosh C. S.; Christenson B.; Balazsi G.; Zeng L. Y. Lysis-lysogeny coexistence: Prophage integration during lytic development. Microbiologyopen 2017, 6, e00395 10.1002/mbo3.395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qiu Z.; Yu Y.; Chen Z.; Jin M.; Yang D.; Zhao Z.; Wang J.; Shen Z.; Wang X.; Qian D.; Huang A.; Zhang B.; Li J.-W. Nanoalumina promotes the horizontal transfer of multiresistance genes mediated by plasmids across genera. Proc. Natl. Acad. Sci. U.S.A. 2012, 109, 4944–4949. 10.1073/pnas.1107254109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Y.; Zhu H.; Colvin V. L.; Alvarez P. J. Cellular and transcriptional response of Pseudomonas stutzeri to quantum dots under aerobic and denitrifying conditions. Environ. Sci. Technol. 2011, 45, 4988–4994. 10.1021/es1042673. [DOI] [PubMed] [Google Scholar]
- Nakazawa S.; Haramiishi A.; Fukuda K.; Kanayama Y.; Watanabe T.; Yuki M.; Ohkuma M.; Takeda K.; Kimbara K.; Shintani M. Different transferability of incompatibility (Inc) P-7 plasmid pCAR1 and IncP-1 plasmid pBP136 in stirring liquid conditions. PLoS One 2017, 12, e0186248 10.1371/journal.pone.0186248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moldovan R.; Chapman-McQuiston E.; Wu X. L. On kinetics of phage adsorption. Biophys. J. 2007, 93, 303–315. 10.1529/biophysj.106.102962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benjamin M. M.; Lawler D. F.. Water Quality Engineering: Physical/chemical Treatment Processes; John Wiley & Sons, 2013. [Google Scholar]
- Replicon J.; Frankfater A.; Miller R. V. A continuous culture model to examine factors that affect transduction among Pseudomonas aeruginosa strains in freshwater environments. Appl. Environ. Microbiol. 1995, 61, 3359–3366. 10.1128/aem.61.9.3359-3366.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wohlleben W.; Arnold W.; Bissonnette L.; Pelletier A.; Tanguay A.; Roy P. H.; Gamboa G. C.; Barry G. F.; Aubert E.; Davies J. On the evolution of Tn 21-like multiresistance transposons: sequence analysis of the gene (aacC1) for gentamicin acetyltransferase-3-I (AAC (3)-I), another member of the Tn 21-based expression cassette. Mol. Gen. Genet. 1989, 217, 202–208. 10.1007/BF02464882. [DOI] [PubMed] [Google Scholar]
- Wagner A. Energy constraints on the evolution of gene expression. Mol. Biol. Evol. 2005, 22, 1365–1374. 10.1093/molbev/msi126. [DOI] [PubMed] [Google Scholar]
- Pansegrau W.; Ziegelin G.; Lanka E. Covalent association of the traI gene product of plasmid RP4 with the 5′-terminal nucleotide at the relaxation nick site. J. Biol. Chem. 1990, 265, 10637–10644. 10.1016/s0021-9258(18)86994-0. [DOI] [PubMed] [Google Scholar]
- Zatyka M.; Jagura-Burdzy G.; Thomas C. M. Transcriptional and translational control of the genes for the mating pair formation apparatus of promiscuous IncP plasmids. J. Bacteriol. 1997, 179, 7201–7209. 10.1128/jb.179.23.7201-7209.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emr S. D.; Schwartz M.; Silhavy T. J. Mutations altering the cellular localization of the phage lambda-receptor, an Escherichia coli outer membrane-protein. Proc. Natl. Acad. Sci. U. S. A. 1978, 75, 5802–5806. 10.1073/pnas.75.12.5802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marchal C.; Perrin D.; Hedgpeth J.; Hofnung M. Synthesis and maturation of lambda receptor in Escherichia coli K-12: in vivo and in vitro expression of gene lamB under lac promoter control. Proc. Natl. Acad. Sci. U. S. A. 1980, 77, 1491–1495. 10.1073/pnas.77.3.1491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu Y.; Zeng J.; Wu B.; Wang L.; Cai R.; Zhang N.; Li Y.; Huang X.; Huang B.; Chen C. Quorum sensing N-acyl homoserine lactones-SdiA suppresses Escherichia coli-Pseudomonas aeruginosa conjugation through inhibiting traI expression. Front. Cell. Infect. Microbiol. 2017, 7, 7. 10.3389/fcimb.2017.00007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhong X.; Kro J. E.; Top E. M.; Krone S. M. Accounting for mating pair formation in plasmid population dynamics. J. Theor. Biol. 2010, 262, 711–719. 10.1016/j.jtbi.2009.10.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murray A.; Jackson G. Viral dynamics - a model of the effects of size, shape, motion and abundance of single-celled planktonic organisms and other particles. Mar. Ecol.: Prog. Ser. 1992, 89, 103–116. 10.3354/meps089103. [DOI] [Google Scholar]
- Simmons M.; Drescher K.; Nadell C. D.; Bucci V. Phage mobility is a core determinant of phage-bacteria coexistence in biofilms. ISME J. 2018, 12, 531–543. 10.1038/ismej.2017.190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisenbrandt R.; Kalkum M.; Lai E.-M.; Lurz R.; Kado C. I.; Lanka E. Conjugative pili of IncP plasmids, and the Ti plasmid T pilus are composed of cyclic subunits. J. Biol. Chem. 1999, 274, 22548–22555. 10.1074/jbc.274.32.22548. [DOI] [PubMed] [Google Scholar]
- Lawley T. D.; Klimke W. A.; Gubbins M. J.; Frost L. S. F factor conjugation is a true type IV secretion system. FEMS Microbiol. Lett. 2003, 224, 1–15. 10.1016/s0378-1097(03)00430-0. [DOI] [PubMed] [Google Scholar]
- Bradley D. E. Morphological and serological relationships of conjugative pili. Plasmid 1980, 4, 155–169. 10.1016/0147-619x(80)90005-0. [DOI] [PubMed] [Google Scholar]
- Katsura I.Tail Assembly and Injection. Lambda II; Cold Spring Harbor Laboratory, Cold Spring Harbor: NY, 1983; pp 331–346. [Google Scholar]
- Bomberg M.; Claesson Liljedahl L.; Lamminmäki T.; Kontula A. Highly diverse aquatic microbial communities separated by permafrost in Greenland show distinct features according to environmental niches. Front. Microbiol. 2019, 10, 1583. 10.3389/fmicb.2019.01583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin D.; Kong X.; Cui B.; Jin S.; Xie Y.; Wang X.; Deng Y. Bacterial communities and potential waterborne pathogens within the typical urban surface waters. Sci. Rep. 2018, 8, 13368. 10.1038/s41598-018-31706-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bowater L.The Microbes Fight Back: Antibiotic Resistance; Royal Society of Chemistry, 2016. [Google Scholar]
- Lane A. N.; Fan T. W.-M. Regulation of mammalian nucleotide metabolism and biosynthesis. Nucleic Acids Res. 2015, 43, 2466–2485. 10.1093/nar/gkv047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adams D. W.; Pereira J. M.; Stoudmann C.; Stutzmann S.; Blokesch M. The type IV pilus protein PilU functions as a PilT-dependent retraction ATPase. PLoS Genet. 2019, 15, e1008393 10.1371/journal.pgen.1008393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molineux I. J.; Panja D. Popping the cork: Mechanisms of phage genome ejection. Nat. Rev. Microbiol. 2013, 11, 194–204. 10.1038/nrmicro2988. [DOI] [PubMed] [Google Scholar]
- Koch A. L. Shrinkage of growing Escherichia coli cells by osmotic challenge. J. Bacteriol. 1984, 159, 919–924. 10.1128/jb.159.3.919-924.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo J.; Li J.; Chen H.; Bond P. L.; Yuan Z. Metagenomic analysis reveals wastewater treatment plants as hotspots of antibiotic resistance genes and mobile genetic elements. Water Res. 2017, 123, 468–478. 10.1016/j.watres.2017.07.002. [DOI] [PubMed] [Google Scholar]
- Sinton L. W.; Hall C. H.; Lynch P. A.; Davies-Colley R. J. Sunlight inactivation of fecal indicator bacteria and bacteriophages from waste stabilization pond effluent in fresh and saline waters. Appl. Environ. Microbiol. 2002, 68, 1122–1131. 10.1128/aem.68.3.1122-1131.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ewert D. L.; Paynter M. J. B. Enumeration of bacteriophages and host bacteria in sewage and the activated-sludge treatment process. Appl. Environ. Microbiol. 1980, 39, 576–583. 10.1128/aem.39.3.576-583.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duran A. E.; Muniesa M.; Moce-Llivina L.; Campos C.; Jofre J.; Lucena F. Usefulness of different groups of bacteriophages as model micro-organisms for evaluating chlorination. J. Appl. Microbiol. 2003, 95, 29–37. 10.1046/j.1365-2672.2003.t01-1-01948.x. [DOI] [PubMed] [Google Scholar]
- Blatchley E. R.; Gong W.-L.; Alleman J. E.; Rose J. B.; Huffman D. E.; Otaki M.; Lisle J. T. Effects of wastewater disinfection on waterborne bacteria and viruses. Water Environ. Res. 2007, 79, 81–92. 10.2175/106143006x102024. [DOI] [PubMed] [Google Scholar]
- Maganha de Almeida Kumlien A. C.; Borrego C. M.; Balcázar J. L. Antimicrobial resistance and bacteriophages: An overlooked intersection in water disinfection. Trends Microbiol. 2021, 29, 517–527. 10.1016/j.tim.2020.12.011. [DOI] [PubMed] [Google Scholar]
- Griebler C.; Lueders T. Microbial biodiversity in groundwater ecosystems. Freshwater Biol. 2009, 54, 649–677. 10.1111/j.1365-2427.2008.02013.x. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.