Abstract
Helicobacter pylori is a gram-negative bacteria which colonizes the gastric mucosa of humans and is implicated in a wide range of gastroduodenal diseases. This paper reviews the physiology of this bacterium as predicted from the sequenced genomes of two unrelated strains and reconciles these predictions with the literature. In general, the predicted capabilities are in good agreement with reported experimental observations. H. pylori is limited in carbohydrate utilization and will use amino acids, for which it has transporter systems, as sources of carbon. Energy can be generated by fermentation, and the bacterium possesses components necessary for both aerobic and anaerobic respiration. Sulfur metabolism is limited, whereas nitrogen metabolism is extensive. There is active uptake of DNA via transformation and ample restriction-modification activities. The cell contains numerous outer membrane proteins, some of which are porins or involved in iron uptake. Some of these outer membrane proteins and the lipopolysaccharide may be regulated by a slipped-strand repair mechanism which probably results in phase variation and plays a role in colonization. In contrast to a commonly held belief that H. pylori is a very diverse species, few differences were predicted in the physiology of these two unrelated strains, indicating that host and environmental factors probably play a significant role in the outocme of H. pylori-related disease.
Helicobacter pylori is a gram-negative bacterium which colonizes the gastric mucosa of humans, causes gastritis and peptic ulcer disease, and is associated with certain types of gastric cancer (27, 65, 87). Once colonized, the host can be chronically infected for life unless antimicrobial therapy is administered. The ability to colonize and persist in the human stomach for many years indicates that H. pylori is specifically adapted to occupy only this niche, and such adaptation should be reflected in a unique complement of physiological capabilities. Furthermore, physiological differences resulting from the apparent genomic variation among strains have been suggested to be responsible for the diversity of diseases associated with H. pylori infection (16, 95).
Bacterial genomics, the identification and annotation of the entire coding potential of a bacterium, allows a more complete understanding of bacterial physiology and pathogenesis. The recent analysis of the complete genomic sequence of two unrelated, pathogenic H. pylori strains (J99 and 26695) demonstrated that even though the chromosomes are organized differently in a limited number of discrete regions, the genome size, genetic content, and gene order of these two strains are remarkably similar (6). We have used the data resulting from this comparative sequence analysis as the starting point to examine, from a functional perspective, the genes that are common and unique to the two strains. The presence or absence of orthologous genes or metabolic pathways in both unrelated H. pylori strains implies that these genes or pathways are present or absent, respectively, in this species. Our comparison has defined the set of common H. pylori metabolic capabilities as well as a small number that are strain specific.
ANALYSES OF GENETIC AND FUNCTIONAL CONSERVATION
The predicted genes from both H. pylori genomes (6) were assigned a likely function, if their predicted amino acid sequence exhibited similarity to a protein of known function, and categorized as shown in Table 1. Function was annotated conservatively; e.g., proteins which showed sequence similarity to transporters, the majority of which did not have the same substrate specificity, were assigned to the general category of transporters pending experimental evidence for their specificity. The two H. pylori genomes are highly conserved with respect to gene content (1,495 and 1,552 open reading frames [ORFs] in J99 and 26695, respectively [6]), functional categorization (Table 2), and gene order (Fig. 1). In both strains, approximately 58% of the gene products were assigned a putative function based upon their having significant sequence similarity to a protein of known function; nearly 18% were conserved in other species but had no known function; and about 23% were specific to H. pylori (Table 2). Eighty-nine genes were specific to strain J99, and 117 were specific to strain 26695; 26 of these genes in each of the strains J99 and 26695 had an assigned function. Preliminary analysis of the Campylobacter jejuni genome (based on analysis of the recently completed genome by the Sanger Centre) indicated that approximately 90 of the H. pylori specific genes will have an orthologue in this closely related species. This will reduce the proportion of H. pylori-specific genes to approximately 17%.
TABLE 1.
List of H. pylori J99 genes and corresponding 26695 orthologs with putative functional assignmentsa
| Gene no. in:
|
Gene name | Function | |
|---|---|---|---|
| J99 | 26695 | ||
| Amino acid biosynthesis | |||
| General | |||
| 206 | 0220 | Aminotransferase | |
| 568 | 0624 | Aminotransferase | |
| 632 | 0696 | Hydantoin utilization | |
| 633 | 0695 | Hydantoin utilization | |
| 673 | 0736 | Aminotransferase | |
| 976 | 0405 | Aminotransferase | |
| Aromatic amino acid family | |||
| 122 | 0134 | aroF | Phospho-2-dehydro-3-deoxyheptonate aldolase |
| 145 | 0157 | aroK | Shikamate kinase I |
| 268 | 0283 | aroB | 3-Dehydroquinate synthase |
| 386 | 1038 | aroD | 3-Dehydroquinate dehydratase |
| 608 | 0663 | aroC | Chorismate synthase |
| 980 | 0401 | aroA | 3-Phosphoshikimate 1-carboxyvinyl transferase |
| 1170 | 1249 | aroE | Shikimate 5-dehydrogenase |
| 1198 | 1277 | trpA | Tryptophan synthase alpha chain |
| 1199 | 1278 | trpB | Tryptophan synthase beta chain |
| 1200 | 1279 | trpC | Indole-3-glycerol phosphate synthase |
| 1201 | 1280 | trpD | Anthranilate phosphoribosyltransferase |
| 1202 | 1281 | trpG | Anthranilate synthase component II |
| 1203 | 1282 | trpE | Anthranilate synthase component I |
| 1294 | 1380 | tyrA | Prephenate dehydrogenase |
| Aspartate family | |||
| 90 | 0098 | thrC | Threonine synthase |
| 98 | 0106 | metB | Cystathionine gamma-synthase |
| 198 | 0212 | dapE | Succinyl-diaminopimelate desuccinylase |
| 275 | 0290 | lysA | Diaminopimelate decarboxylase |
| 375 | 1050 | thrB | Homoserine kinase |
| 410 | 1013 | dapA | Dihydrodipicolinate synthase |
| 460 | 0510 | dapB | Dihydrodipicolinate reductase |
| 513 | 0566 | dapF | Diaminopimelate epimerase |
| 570 | 0626 | dapD | 2,3,4,5-Tetrahydropyridine-2-carboxylate-N-succinyltransferase |
| 594 | 0649 | aspA | Aspartate ammonia-lyase |
| 615 | 0672 | aspB | Aspartate aminotransferase |
| 761 | 0822 | hom | Homoserine dehydrogenase |
| 1114 | 1189 | asd | Aspartate-semialdehyde dehydrogenase |
| 1150 | 1229 | lysC | Aspartokinase 2 alpha and beta subunits |
| Glutamate family | |||
| 461 | 0512 | glnA | Glutamine synthetase |
| 1001 | 0380 | gdhA | Glutamate dehydrogenase |
| 1085 | 1158 | proC | Pyrroline-5-carboxylate reductase |
| Pyruvate family | |||
| 313 | 0330 | ilvC | Ketol-acid reductoisomerase |
| 1361 | 1468 | ilvE | Branched-chain amino acid aminotransferase |
| Serine family | |||
| 99 | 0107 | cysK | Cysteine synthase |
| 171 | 0183 | glyA | Serine hydroxymethyltransferase |
| 597 | 0652 | serB | Phosphoserine phosphatase |
| 984 | 0397 | serA | Phosphoglycerate dehydrogenase |
| 1133 | 1210 | cysE | Serine acetyltransferase |
| Biosynthesis of cofactors, prosthetic groups, and carriers | |||
| Biotin | |||
| 25 | 0029 | bioD | Dethiobiotin synthetase |
| 545 | 0598 | bioF | 8-Amino-7-oxononanoate synthase |
| 910 | 0976 | bioA | Adenosylmethionine-8-amino-7-oxononanoate aminotransferase |
| 1068 | 1140 | Biotin activation protein | |
| 1298 | 1406 | bioB | Biotin synthetase |
| Folic acid | |||
| 278 | 0293 | pabB | p-Aminobenzoate synthetase |
| 388 | 1036 | folK | 7,8-Dihydro-6-hydroxymethylpterin-pyrophosphokinase |
| 524 | 0577 | folD | Methylenetetrahydrofolate dehydrogenase/methenyltetrahydrofolate cyclohydrolase |
| 862 | 0928 | folE | GTP cyclohydrolase I |
| 863 | 0928 | folE | GTP cyclohydrolase I |
| 1153 | 1232 | folP | Dihydropteroate synthase |
| 1403 | 1510 | folB | Dihydroneopterin aldolase |
| 1454 | 1545 | folC | Folylpolyglutamate synthase |
| Heme and porphyrin | |||
| 150 | 0163 | hemB | δ-Aminolevulinic acid dehydratase |
| 222 | 0237 | hemC | Porphobilinogen deaminase |
| 224 | 0239 | hemA | Glutamyl-tRNA reductase |
| 291 | 0306 | hemL | Glutamate-1-semialdehyde-2,1-aminomutase |
| 551 | 0604 | hemE | Uroporphyrinogen decarboxylase |
| 610 | 0665 | hemN | Oxygen-independent coproporphyrinogen III oxidase |
| 1000 | 0381 | hemG | Protoporphyrinogen oxidase |
| 1005 | 0376 | hemH | Ferrochelatase |
| 1145 | 1224 | hemD | Uroporphyrinogen III synthase |
| 1147 | 1226 | hemN | Oxygen-independent coproporphyrinogen III oxidase |
| Menaquinone and ubiquinone | |||
| 225 | 0240 | ispB | Octaprenyl-diphosphate synthase |
| 864 | 0929 | ispA | Geranyltransferase |
| 1278 | 1360 | ubiA | 4-Hydroxybenzoate octaprenyltransferase |
| 1369 | 1476 | ubiD | Octaprenyl-4-hydroxybenzoate carboxy-lyase |
| 1376 | 1483 | ubiE | Ubiquinone/menaquinone biosynthesis methyltransferase |
| Molybdopterin | |||
| 158 | 0172 | moeA | Molybdopterin biosynthesis protein |
| 705 | 0768 | moaA | Molybdopterin cofactor biosynthetic protein |
| 706 | 0769 | mobA | Molybdopterin-guanine dinucleotide biosynthesis protein A |
| 734 | 0798 | moaC | Molybdenum cofactor biosynthesis protein C |
| 735 | 0799 | mog | Molybdopterin biosynthesis protein |
| 736 | 0800 | moaE | Molybdopterin-converting factor, subunit 2 |
| 737 | 0801 | moaD | Molybdopterin-converting factor, subunit 1 |
| 750 | 0814 | moeB | Molybdopterin-synthase sulfurylase |
| Pantothenate | |||
| 6 | 0006 | panC | Pantoate-β-alanine ligase |
| 30 | 0034 | panD | Aspartate-1-decarboxylase |
| 367 | 1058 | panB | 3-Methyl-2-oxobutanoate hydroxymethyltransferase |
| 779 | 0841 | dfp | Pantothenate metabolism flavoprotein |
| Pyridoxine | |||
| 328 | 0354 | dxs | 1-Deoxyxylulose-5-phosphate synthase |
| 1489 | 1582 | pdxJ | Pyridoxal phosphate synthetase |
| 1490 | 1583 | pdxA | Pyridoxal phosphate biosynthetic protein A |
| Riboflavin | |||
| 2 | 0002 | ribE | Riboflavin synthase beta chain |
| 338 | 1087 | ribF | Riboflavin kinase |
| 738 | 0802 | ribA | GTP cyclohydrolase II |
| 740 | 0804 | ribBA | GTP cyclohydrolase II/3,4-dihydroxy-2-butanone 4-phosphate synthase |
| 1398 | 1505 | ribD | Riboflavin-specific deaminase |
| 1482 | 1574 | ribC | Riboflavin synthase alpha chain |
| Thioredoxin, glutaredoxin, and glutathione | |||
| 763 | 0824 | trxA | Thioredoxin |
| 764 | 0825 | trxB | Thioredoxin reductase |
| 1046 | 1118 | ggt | Gamma-glutamyl transpeptidase |
| 1091 | 1164 | trxB | Thioredoxin reductase |
| 1351 | 1458 | Thioredoxin | |
| Thiamine | |||
| 781 | 0843 | thiE | Thiamine phosphate pyrophosphorylase |
| 782 | 0844 | thiD | Phosphomethylpyrimidine kinase |
| 783 | 0845 | thiM | Hydroxyethylthiazole kinase |
| Pyridine nucleotides | |||
| 312 | 0329 | nadE | NH3-dependent NAD+ synthetase |
| 1273 | 1355 | nadC | Nicotinate-nucleotide pyrophosphorylase |
| 1274 | 1356 | nadA | Quinolinate synthetase |
| Cell envelope | |||
| Membranes and porins | |||
| 7 | 0009 | Outer membrane protein | |
| 21 | 0025 | Outer membrane protein | |
| 73 | 0078 | Outer membrane protein | |
| 0079 | |||
| 117 | 0127 | Outer membrane protein | |
| 195 | 0209 | Outer membrane protein | |
| 212 | 0227 | Outer membrane protein | |
| 214 | 0229 | hopA | Outer membrane protein—porin |
| 237 | 0252 | Outer membrane protein | |
| 238 | 0253, 0254 | Outer membrane protein | |
| 307 | 0324 | Outer membrane protein | |
| 342 | 1083 | Outer membrane protein | |
| 359 | 1066 | Outer membrane protein | |
| 424 | 0472 | Outer membrane protein | |
| 429 | 0477 | Outer membrane protein | |
| 438 | 0486 | Outer membrane protein | |
| 439 | 0487 | Outer membrane protein | |
| 456 | 0506 | Outer membrane protein | |
| 514 | 0567 | Inner membrane protein | |
| 581 | 0638 | Outer membrane protein | |
| 600 | 0655 | Outer membrane protein | |
| 614 | 0671 | Outer membrane protein | |
| 634 | 0694 | Outer membrane protein | |
| 645 | 0706 | hopE | Outer membrane protein—porin |
| 649 | 0710 | Outer membrane protein | |
| 659 | 0722 | Outer membrane protein | |
| 662 | 0725 | Outer membrane protein | |
| 663 | 0726 | Outer membrane protein | |
| 719 | 0782 | Outer membrane protein | |
| 725 | 0788 | Outer membrane protein | |
| 732 | 0796 | Outer membrane protein | |
| 777 | 0839 | Outer membrane protein | |
| 810 | 0876 | frpB | Iron-regulated outer membrane protein |
| 833 | 1243 | babB | Outer membrane protein—adhesin |
| 848 | 0912 | hopC | Outer membrane protein—porin |
| 849 | 0913 | hopB | Outer membrane protein—porin |
| 850 | 0914 | Outer membrane protein | |
| 851 | 0915, 0916 | frpB | Iron-regulated outer membrane protein |
| 857 | 0923 | Outer membrane protein | |
| 870 | Outer membrane protein | ||
| 1008 | 0373 | Outer membrane protein | |
| 1022 | 0358 | Outer membrane protein | |
| 1034 | 1107 | Outer membrane protein | |
| 1040 | 1113 | Outer membrane protein | |
| 1054 | 1125 | Outer membrane protein | |
| 1083 | 1156 | Outer membrane protein | |
| 1084 | 1157 | Outer membrane protein | |
| 1094 | 1167 | Outer membrane protein | |
| 1103 | 1177 | Outer membrane function | |
| 1138 | 1215, 1216 | imp | Role in outer membrane permeability |
| 1164 | 0896 | babA | Outer membrane protein—adhesin |
| 1261 | 1342 | Outer membrane protein | |
| 1343 | 1450 | Inner membrane protein | |
| 1346 | 1453 | Outer membrane protein | |
| 1349 | 1456 | lpp20 | Conserved lipoprotein |
| 1360 | 1467 | Outer membrane protein | |
| 1362 | 1469 | Outer membrane protein | |
| 1394 | 1501 | Outer membrane protein | |
| 1405 | 1512 | frpB | Iron-regulated outer membrane protein |
| 1432 | 1395 | Outer membrane protein | |
| 1472 | 1564 | Outer membrane protein | |
| 1479 | 1571 | Outer membrane protein | |
| Murein sacculus and peptidoglycan | |||
| 445 | 0493 | mraY | Phospho-N-acetylmuramoyl-pentapeptide-transferase |
| 446 | 0494 | murD | UDP-N-acetylmuramoylalanine-d-glutamate ligase |
| 496 | 0549 | murI | Glutamate racemase |
| 544 | 0597 | Penicillin-binding protein | |
| 567 | 0623 | murC | UDP-N-acetylmuramate-alanine ligase |
| 590 | 0645 | Lytic murein transglycosylase | |
| 593 | 0648 | murA | UDP-N-acetylglucosamine enolpyruvyltransferase |
| 675 | 0738 | ddl | d-Alanine–d-alanine ligase |
| 677 | 0740 | murF | d-Alanyl–d-alanine-adding enzyme |
| 709 | 0772 | amiA | Probable N-acetylmuramoyl-l-alanine amidase |
| 876 | 0941 | alr | Alanine racemase, biosynthetic |
| 1082 | 1155 | murG | UDP-N-acetylglucosamine-N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenoln-acetylglucosaminetransferase |
| 1313 | 1418 | murB | UDP-N-acetylenolpyruvoyl glucosamine reductase |
| 1387 | 1494 | murE | UDP-N-acetylmuramyl-tripeptide synthetase |
| 1464 | 1556 | Penicillin-binding protein | |
| 1473 | 1565 | Penicillin-binding protein | |
| Surface polysaccharides, lipopolysaccharides, and antigens | |||
| 3 | 0003 | kdsA | 3-Deoxy-d-manno-octulosonic acid 8-phosphate synthase |
| 37 | 0043 | manA/manC | Phosphomannose isomerase/GDP-mannose pyrophosphorylase |
| 38 | 0044 | Gmd | GDP-d-mannose dehydratase |
| 39 | 0045 | Sugar nucleotide biosynthesis | |
| 70 | 0075 | glmM | Phosphoglucosamine mutase |
| 86 | 0093, 0094 | α-(1,2)-Fucosyltransferase | |
| 147 | 0159 | Lipopolysaccharide biosynthesis protein | |
| 166 | 0178 | neuB | Sialic acid synthase |
| 182 | 0196 | lpxD | UDP-3-O-[3-hydroxymyristoyl] glucosamine N-acetyltransferase |
| 194 | 0208 | Lipopolysaccharide biosynthesis protein | |
| 215 | 0230 | kdsB | 3-Deoxy-manno-octulosonate cytidylyltransferase |
| 264 | 0279 | waaC | Lipopolysaccharide heptosyltransferase-1 |
| 265 | 0280 | waaM | Lipid A biosynthesis acyltransferase |
| 309 | 0326 | neuA | Acylneuraminate cytidylyltransferase |
| 311 | 0328 | lpxK | Tetraacyldisaccharide-1-p 4′-kinase |
| 373 | 1052 | lpxC | UDP-3-O-[3-hydroxymyristoyl]-N-acetylglucosamine deacetylase |
| 562 | Lipopolysaccharide biosynthesis protein | ||
| 563 | 0619 | Lipopolysaccharide biosynthesis protein | |
| 596 | 0651 | fucT | α-(1,3)-Fucosyltransferase |
| 620 | 0679 | Lipopolysaccharide biosynthesis protein | |
| 741 | 0805 | Lipopolysaccharide biosynthesis protein | |
| 765 | 0826 | Lipopolysaccharide biosynthesis protein | |
| 778 | 0840 | Sugar nucleotide biosynthesis protein | |
| 791 | 0857 | gmhA | Phosphoheptose isomerase |
| 792 | 0858 | waaE | ADP-d-glycero-d-mannoheptose synthase |
| 793 | 0859 | gmhD | ADP-l-glycero-d-mannoheptose-6-epimerase |
| 801 | 0867 | lpxB | Lipid-A-disaccharide synthase |
| 820 | Lipopolysaccharide biosynthesis protein | ||
| 891 | 0957 | waaA | 3-Deoxy-d-manno-octulosonic-acid transferase |
| 963 | 0421 | Polysaccharide biosynthesis protein | |
| 1002 | 0379 | fucU | α-(1,3)-Fucosyltransferase |
| 1015 | 0366 | Sugar nucleotide biosynthesis | |
| 1020 | 0360 | galE | UDP-glucose 4-epimerase |
| 1031 | 1105 | Lipopolysaccharide biosynthesis protein | |
| 1032 | Lipopolysaccharide biosynthesis protein | ||
| 1116 | 1191 | waaF | ADP-heptose-lipopolysaccharide heptosyltransferase II |
| 1196 | 1275 | manB | Phosphomannomutase |
| 1289 | 1375 | lpxA | UDP-N-acetylglucosamine acyltransferase |
| 1311 | 1416 | Lipopolysaccharide biosynthesis protein | |
| 1368 | 1475 | kdtB | Lipopolysaccharide core biosynthesis protein |
| 1488 | 1581 | wecA | Undecaprenyl-phosphate-α-N-acetylglucosaminyltransferase |
| Surface structures | |||
| 107 | 0115 | flaB | Flagellin B |
| 159 | 0173 | fliR | Flagellar biosynthesis protein |
| 217 | 0232 | Motility protein | |
| 231 | 0246 | flgI | Flagellar P-ring protein |
| 280 | 0295 | flgL | Flagellar hook-associated protein 3 (hap3) |
| 308 | 0325 | flgH | Flagellar L-ring protein precursor (basal body L-ring protein) |
| 310 | 0327 | flaG | Flagellar biosynthesis protein |
| 325 | 0351 | fliF | Flagellar M-ring protein |
| 326 | 0352 | fliG | Flagellar motor switch protein |
| 327 | 0353 | fliH | Flagellar export apparatus |
| 333 | 1092 | flgG | Flagellar basal-body rod protein |
| 383 | 1041 | flhA | Flagellar biosynthesis protein |
| 389 | 1035 | flhF | Flagellar biosynthesis protein |
| 393 | 1031 | fliM | Flagellar motor switch protein |
| 394 | 1030 | Flagellar motor switch protein | |
| 444 | 0492 | Paralogue of hpaA | |
| 531 | 0584 | fliN | Flagellar motor switch protein |
| 548 | 0601 | flaA | Flagellin A |
| 625 | 0684, 0685 | fliP | Flagellar biosynthesis protein |
| 688 | 0751 | Flagellin protein | |
| 689 | 0752 | fliD | Flagellar hook-associated protein 2 (hap2) |
| 690 | 0753 | fliS | Flagellar protein |
| 707 | 0770 | flhB | Flagellar biosynthesis protein |
| 733 | 0797 | hpaA | Neuraminyllactose-binding hemagglutinin precursor |
| 745 | 0809 | fliL | Flagellar biosynthesis protein |
| 751 | 0815 | motA | Chemotaxis protein (motility protein a) |
| 752 | 0816 | motB | Flagellar motor protein |
| 804 | 0870 | flgE | Flagellar hook protein |
| 843 | 0907 | Flagellar biosynthesis protein | |
| 844 | 0908 | Flagellar basal-body/rod/hook protein | |
| 971 | 0410 | Paralogue of hpaA | |
| 1047 | 1119 | flgK | Flagellar hook-associated protein 1 (Hap1) |
| 1117 | 1192 | Motility protein | |
| 1195 | 1274 | pflA | Flagellar functional protein |
| 1314 | 1419 | fliQ | Flagellar biosynthesis protein |
| 1315 | 1420 | fliI | Flagellum-specific ATP synthase |
| 1355 | 1462 | Motility protein | |
| 1465 | 1557 | fliE | Flagellar hook–basal-body complex protein |
| 1466 | 1558 | flgC | Flagellar basal-body rod protein |
| 1467 | 1559 | flgB | Flagellar basal-body rod protein |
| 1483 | 1575 | flhB | Flagellar biosynthesis protein |
| 1492 | 1585 | flgG | Flagellar basal-body rod protein (distal rod protein) |
| Cellular processes | |||
| General | |||
| 4 | 0004 | icfA | Carbonic anhydrase |
| 161 | 0175 | Peptidyl-prolyl cis-trans isomerase | |
| 183 | 0197 | metK | S-Adenosylmethionine synthetase |
| 228 | 0243 | napA | Neutrophil-activating protein A |
| 466 | 0517 | era | GTP-binding protein |
| 678 | 0741 | HIT family protein | |
| 865 | 0930 | surE | Stationary-phase protein |
| 977 | 0404 | HIT family protein | |
| 1112 | 1186 | Carbonic anhydrase | |
| Cell division | |||
| 314 | 0331 | minD | Cell division inhibitor |
| 315 | 0332 | minE | Cell division topological specificity factor |
| 335 | 1090 | ftsK | Septum formation protein |
| 680 | 0743 | rodA | Rod shape-determining protein |
| 912 | 0978 | ftsA | Septum formation protein |
| 913 | 0979 | ftsZ | GTPase in circumferential ring formation |
| 1086 | 1159 | fic | cAMP-induced cell filamentation protein |
| 1287 | 1373 | mreB | Rod shape-determining protein |
| 1468 | 1560 | rodA | Rod shape-determining protein |
| Cell killing | |||
| 274 | 0289 | Vacuolating cytotoxin (VacA) paralogue | |
| 339 | 1086 | hlyA | Hemolysin |
| 556 | 0609, 0610 | Vacuolating cytotoxin (VacA) paralogue | |
| 819 | 0887 | vacA | Vacuolating cytotoxin |
| 856 | 0922 | Vacuolating cytotoxin (VacA) paralog | |
| Cag island proteins and transposable elements | |||
| 15 | 0017 | virB4a | DNA transfer protein |
| 36 | 0041, 0042 | virB10 | DNA transfer protein |
| 469 | 0520 | orf6 | Cag island protein |
| 470 | 0521 | orf7 | Cag island protein |
| 471 | 0522 | orf8 | Cag island protein |
| 472 | 0523 | orf9 | Cag island protein |
| 473 | 0524 | virD4 | Cag island protein, DNA transfer protein |
| 474 | 0525 | virB11a | Cag island protein, DNA transfer protein |
| 475 | 0526 | orf12 | Cag island protein |
| 476 | 0527 | orf13/14 | Cag island protein |
| 477 | 0528 | orf15 | Cag island protein |
| 478 | 0529 | orf16 | Cag island protein |
| 479 | 0530 | orf17 | Cag island protein |
| 480 | 0531 | orf18 | Cag island protein |
| 481 | 0532 | cagT | Cag island protein |
| 482 | 0534 | cagS | Cag island protein |
| 483 | 0535 | cagQ | Cag island protein |
| 484 | 0536 | cagP | Cag island protein |
| 485 | 0537 | cagM | Cag island protein |
| 486 | 0538 | cagN | Cag island protein |
| 487 | 0539 | cagL | Cag island protein |
| 488 | 0540 | cagI | Cag island protein |
| 489 | 0541 | cagH | Cag island protein |
| 490 | 0542 | cagG | Cag island protein |
| 491 | 0543 | cagF | Cag island protein |
| 492 | 0544 | cagE | DNA transfer protein (Agrobacterium VirB4 homologue) |
| 493 | 0545 | cagD | Cag island protein |
| 494 | 0546 | cagC | Cag island protein |
| 495 | 0547 | cagA | Cag island protein, cytotoxicity-associated immunodominant antigen |
| 826 | tnpB | IS606 transposase | |
| 827 | tnpA | IS606 transposase | |
| 917 | virB4b | DNA transfer protein | |
| 918 | virB4c | DNA transfer protein | |
| 1279 | 1361 | comEC | DNA transfer protein |
| 1316 | 1421 | virB11b | DNA transfer protein |
| Chaperones | |||
| 8 | 0010 | groEL | 60-kDa chaperone |
| 9 | 0011 | groES | 10-kDa chaperone |
| 101 | 0109 | dnaK | 70-kDa chaperone |
| 102 | 0110 | grpE | 24-kDa chaperone |
| 196 | 0210 | htpG | 90-kDa chaperone |
| 400 | 1024 | dnaJ2 | Cochaperone with DnaK |
| 861 | 0927 | htpX | Stress protein |
| 1252 | 1332 | dnaJ1 | Cochaperone with DnaK |
| Detoxification | |||
| 809 | 0875 | katA | Catalase |
| 991 | 0390 | tpx | Thiol peroxidase |
| 992 | 0389 | sodF | Iron-dependent superoxide dismutase |
| 1471 | 1563 | tsaA | Peroxidase |
| Protein and peptide secretion | |||
| 69 | 0074 | lspA | Lipoprotein signal peptidase |
| 168 | 0180 | lnt | Apolipoprotein N-acyltransferase |
| 329 | 0355 | lepA | GTP-binding protein |
| 523 | 0576 | lepB | Signal peptidase I |
| 700 | 0763 | ftsY | Functional homolog of srp receptor |
| 723 | 0786 | secA | Preprotein translocase subunit |
| 731 | 0795 | tig | Trigger factor |
| 889 | 0955 | lgt | Prolipoprotein diacylglyceryl transferase |
| 1079 | 1152 | ffh | Signal recognition particle protein |
| 1126A | 1203A | secE | Preprotein translocase subunit |
| 1176 | 1255 | secG | Protein export membrane protein |
| 1220 | 1300 | secY | Preprotein translocase subunit |
| 1449 | 1550 | secD | Protein export membrane protein |
| 1450 | 1549 | secF | Protein export membrane protein |
| Phosphorus compounds | |||
| 413 | 1010 | ppk | Polyphosphate kinase |
| 564 | 0620 | ppa | Inorganic pyrophosphatase |
| Polyamine biosynthesis | |||
| 18 | 0020 | nspC | Carboxynorspermidine decarboxylase |
| 771 | 0832 | speE | Spermidine synthase |
| 962 | 0422 | speA | Arginine decarboxylase |
| Urea | |||
| 62 | 0067 | ureH | Urease accessory protein |
| 63 | 0068 | ureG | Urease accessory protein |
| 64 | 0069 | ureF | Urease accessory protein |
| 65 | 0070 | ureE | Urease accessory protein |
| 67 | 0072 | ureB | Urease beta subunit |
| 68 | 0073 | ureA | Urease alpha subunit |
| DNA replication | |||
| 10 | 0012 | dnaG | DNA primase |
| 108 | 0116 | topA | DNA topoisomerase I |
| 199 | 0213 | gidA | Glucose-inhibited division protein A |
| 362 | 1063 | gidB | Glucose-inhibited division protein B |
| 452 | 0500 | dnaN | DNA polymerase III, beta chain |
| 453 | 0501 | gyrB | DNA gyrase subunit B |
| 558 | 0615 | lig | DNA ligase |
| 641 | 0701 | gyrA | DNA gyrase subunit A |
| 655 | 0717 | dnaX | DNA polymerase III subunits gamma and tau |
| 847 | 0911 | ATP-dependent helicase | |
| 919 | topA | Topoisomerase I | |
| 931 | topA | Topoisomerase I | |
| 994 | 0387 | priA | Primosomal protein n′ (replication factor y) |
| 1152 | 1231 | holB | DNA polymerase III subunit delta′ |
| 1166 | 1245 | ssb | Single-strand binding protein |
| 1280 | 1362 | dnaB | Replicative DNA helicase |
| 1353 | 1460 | dnaE | DNA polymerase III, alpha chain |
| 1363 | 1470 | polA | DNA polymerase I |
| 1371 | 1478 | rep | ATP-dependent DNA helicase |
| 1412 | 1523 | recG | ATP-dependent DNA helicase |
| 1417 | 1529 | dnaA | Chromosomal replication initiator protein |
| 1438 | 1387 | DNA polymerase III | |
| 1446 | 1553 | pcrA | ATP-dependent helicase |
| DNA restriction, modification, recombination, and repair | |||
| 43 | 0050 | Type II DNA modification enzyme (methyltransferase) | |
| 44 | Type II DNA modification enzyme (methyltransferase) | ||
| 45 | Type II DNA modification enzyme (methyltransferase) | ||
| 46 | Type II restriction enzyme | ||
| 84 | 0091 | Type II restriction enzyme | |
| 85 | 0092 | Type II DNA modification enzyme (methyltransferase) | |
| 130 | 0142 | mutY | A/G-specific adenine glycosylase |
| 141 | 0153 | recA | Recombination protein |
| 164 | Restriction enzyme | ||
| 209 | 0223 | radA | DNA repair protein |
| 243 | 0259 | xseA | Exodeoxyribonuclease large subunit |
| 244 | 0260 | Type II DNA modification enzyme (methyltransferase) | |
| 248 | 0263 | Type II DNA modification enzyme (methyltransferase) | |
| 306 | 0323 | Endonuclease | |
| 322 | 0348 | recJ | Single-stranded-DNA-specific exonuclease |
| 366 | 1059 | ruvB | Holliday junction DNA helicase |
| 414 | hsdS1 | Type I restriction enzyme (specificity subunit) | |
| 415 | 0463 | hsdM1 | Type I restriction enzyme (modification subunit) |
| 416 | 0464 | hsdR1 | Type I restriction enzyme (restriction subunit) |
| 430 | 0478 | Type II DNA modification enzyme (methyltransferase) | |
| 433 | 0481 | Type II DNA modification enzyme (methyltransferase) | |
| 435 | 0483 | Type II DNA modification enzyme (methyltransferase) | |
| 532 | 0585 | nth | Endonuclease III |
| 549 | 0602 | Endonuclease III | |
| 565 | 0621 | mutS | DNA mismatch repair protein |
| 606 | 0661 | rnhA | RNase HI |
| 617 | 0675 | Integrase-recombinase protein (xerCD family) | |
| 618 | 0676 | ogt | Methylated-DNA–protein-cysteine methyltransferase |
| 629 | Type II DNA modification enzyme (methyltransferase) | ||
| 630 | Type II restriction enzyme | ||
| 644 | 0705 | uvrA | Excinuclease ABC subunit A |
| 726 | hsdS | Type I restriction enzyme (specificity subunit) | |
| 756 | Type II DNA modification enzyme (methyltransferase) | ||
| 760 | 0821 | uvrC | Excinuclease ABC subunit C |
| 784 | 0846 | hsdR2 | Type I restriction enzyme (restriction subunit) |
| 785 | 0848, 0849 | hsdS2 | Type I restriction enzyme (specificity subunit) |
| 786 | 0850 | hsdM2 | Type I restriction enzyme (modification subunit) |
| 811 | 0877 | ruvC | Crossover junction endodeoxyribonuclease |
| 815 | 0883 | ruvA | Holliday junction DNA helicase |
| 846 | 0910 | Type II DNA modification enzyme (methyltransferase) | |
| 859 | 0925 | recR | Recombination protein |
| 941 | 0995 | Integrase/recombinase (xerCD family) | |
| 951 | Integrase/recombinase (xerCD family) | ||
| 1012 | 0369 | Type II DNA modification enzyme (methyltransferase) | |
| 1041 | 1114 | uvrB | Excinuclease ABC subunit B |
| 1050 | 1121 | Type II DNA modification enzyme (methyltransferase) | |
| 1131 | 1208 | M.HpyI | Type II DNA modification enzyme (methyltransferase) |
| 1149 | 1228 | mutT | dGTP pyrophosphohydrolase |
| 1243 | 1323 | rnhB | RNase HII |
| 1266 | 1347 | ung | Uracil-DNA glycosylase |
| 1271 | 1352 | Type II DNA modification enzyme (methyltransferase) | |
| 1284 | Type II DNA modification enzyme (methyltransferase) | ||
| 1295 | 1382 | Endonuclease | |
| 1296 | mod1 | Type III DNA modification enzyme (methyltransferase) | |
| 1297 | res1 | Type III restriction enzyme | |
| 1364 | 1471 | Type II restriction enzyme | |
| 1365 | 1472 | Type II DNA modification enzyme (methyltransferase) | |
| 1409 | Type II DNA modification enzyme (methyltransferase) | ||
| 1410 | 1521 | res2 | Type III restriction enzyme |
| 1411 | 1522 | mod2 | Type III DNA modification enzyme (methyltransferase) |
| 1415 | 1526 | exoA | Exodeoxyribonuclease |
| 1422 | hsdS3 | Type I restriction enzyme (specificity subunit) | |
| 1423 | 1403 | hsdM3 | Type I restriction enzyme (modification subunit) |
| 1424 | 1402 | hsdR3 | Type I restriction enzyme (restriction subunit) |
| 1434 | 1393 | recN | DNA repair protein |
| 1442 | 1366 | Type II restriction enzyme | |
| Energy metabolism | |||
| Amino acids and amines | |||
| 120 | 0132 | sdaB | l-Serine/l-Threonine deaminase |
| 279 | 0294 | aimE | Aliphatic amidase |
| 585 | 3-Hydroxyacid dehydrogenase | ||
| 661 | 0723 | ansB | l-Asparaginase II |
| 1159 | 1238 | Aliphatic amidase | |
| 1427 | 1399 | rocF | Arginase |
| 1428 | 1398 | ald | l-Alanine dehydrogenase |
| ATP-proton motive force interconversion | |||
| 767 | 0828 | atpB | ATP synthase F0, subunit a |
| 1059 | 1131 | atpC | ATP synthase F1, subunit epsilon |
| 1060 | 1132 | atpD | ATP synthase F1, subunit beta |
| 1061 | 1133 | atpG | ATP synthase F1, subunit gamma |
| 1062 | 1134 | atpA | ATP synthase F1, subunit alpha |
| 1063 | 1135 | atpH | ATP synthase F1, subunit delta |
| 1064 | 1136 | atpF | ATP synthase F0, subunit b |
| 1065 | 1137 | atpX | ATP synthase B′ |
| 1135 | 1212 | atpE | ATP synthase F0, subunit c |
| Electron transport | |||
| 40 | 0047 | hypE | Hydrogenase expression/formation protein |
| 48 | 0056 | putA | Proline/pyrroline-5-carboxylate dehydrogenase |
| 132 | 0144 | fixN | Cytochrome oxidase (CBB3-TYPE) |
| 133 | 0145 | fixO | Cytochrome oxidase (CBB3-TYPE) |
| 134 | 0146 | fixQ | Cytochrome oxidase (CBB3-TYPE) |
| 135 | 0147 | fixP | Cytochrome oxidase (CBB3-TYPE) |
| 177 | 0191 | frdB | Fumarate reductase |
| 178 | 0192 | frdA | Fumarate reductase |
| 179 | 0193 | frdC | Fumarate reductase |
| 250 | 0265 | ccdA | Cytochrome c biogenesis protein |
| 262 | 0277 | Ferredoxin | |
| 459 | 0509 | glcD | Glycolate oxidase |
| 574 | 0631 | hyaA | Hydrogenase, small subunit |
| 575 | 0632 | hyaB | Hydrogenase, large subunit |
| 576 | 0633 | hyaC | Hydrogenase, cytochrome subunit |
| 577 | 0634 | hyaD | Hydrogenase expression/formation protein |
| 611 | 0666 | glpC | Glycerol-3-phosphate dehydrogenase |
| 803 | 0869 | hypA | Hydrogenase expression/formation protein |
| 835 | 0898 | hypD | Hydrogenase expression/formation protein |
| 836 | 0899 | hypC | Hydrogenase expression/formation protein |
| 837 | 0900 | hypB | Hydrogenase expression/formation protein |
| 878 | 0943 | dadA | d-Amino acid dehydrogenase |
| 895 | 0961 | gpsA | Glycerol-3-phosphate dehydrogenase (NAD+) |
| 974 | 0407 | S/N-oxide reductase | |
| 1003 | 0378 | Cytochrome c biogenesis protein | |
| 1035 | 1108 | porG | Pyruvate ferrodoxin oxidoreductase |
| 1036 | 1109 | porD | Pyruvate ferrodoxin oxidoreductase |
| 1037 | 1110 | porA | Pyruvate ferrodoxin oxidoreductase |
| 1038 | 1111 | porB | Pyruvate ferrodoxin oxidoreductase |
| 1088 | 1161 | fldA | Flavodoxin |
| 1090 | 1163 | fixS | Component of cation transport for cbb3-type oxidase |
| 1143 | 1222 | dld | d-Lactate dehydrogenase |
| 1148 | 1227 | Periplasmic cytochrome c-553 | |
| 1181 | 1260 | nuoA | NADH oxidoreductase I |
| 1182 | 1261 | nuoB | NADH oxidoreductase I |
| 1183 | 1262 | nuoC | NADH oxidoreductase I |
| 1184 | 1263 | nuoD | NADH oxidoreductase I |
| 1185 | 1264 | nuoE | NADH oxidoreductase I |
| 1186 | 1265 | nuoF | NADH oxidoreductase I |
| 1187 | 1266 | nuoG | NADH oxidoreductase I |
| 1188 | 1267 | nuoH | NADH oxidoreductase I |
| 1189 | 1268 | nuoI | NADH oxidoreductase I |
| 1190 | 1269 | nuoJ | NADH oxidoreductase I |
| 1191 | 1270 | nuoK | NADH oxidoreductase I |
| 1192 | 1271 | nuoL | NADH oxidoreductase I |
| 1193 | 1272 | nuoM | NADH oxidoreductase I |
| 1194 | 1273 | nuoN | NADH oxidoreductase I |
| 1354 | 1461 | Cytochrome c peroxidase | |
| 1401 | 1508 | fixG | Component of cation transport for cbb3-type oxidase |
| 1459 | 1540 | petA | Ubiquinol cytochrome c oxidoreductase, 2Fe-2S subunit |
| 1460 | 1539 | petB | Ubiquinol cytochrome c oxidoreductase, cytochrome b subunit |
| 1461 | 1538 | petC | Ubiquinol cytochrome c oxidoreductase, cytochrome c1 subunit |
| Entner-Doudoroff pathway | |||
| 1025 | 1099 | eda | 2-Keto-3-deoxy-6-phosphogluconate aldolase |
| 1026 | 1100 | edd | Phosphogluconate dehydratase |
| Fermentation | |||
| 840 | 0903 | ackA | Acetate kinase |
| 841 | 0904, 0905 | pta | Phosphotransacetylase |
| 1030 | 1104 | Zinc-dependent alcohol dehydrogenase | |
| 1429 | Zinc-dependent alcohol dehydrogenase | ||
| Gluconeogenesis | |||
| 111 | 0121 | ppsA | Phosphoenolpyruvate synthase |
| 142 | 0154 | eno | Enolase |
| 162 | 0176 | fba | Fructose-bisphosphate aldolase |
| 180 | 0194 | tpi | Triose-phosphate isomerase |
| 855 | 0921 | gap | Glyceraldehyde-3-phosphate dehydrogenase |
| 908 | 0974 | pgm | 2,3-Bisphosphoglycerate-independent phosphoglycerate mutase |
| 1093 | 1166 | pgi | Glucose-6-phosphate isomerase |
| 1264 | 1345 | pgk | Phosphoglycerate kinase |
| 1265 | 1346 | gap | Glyceraldehyde 3-phosphate dehydrogenase |
| 1440 | 1385 | fbp | Fructose-1,6-bisphosphatase |
| Phosphopentose pathway | |||
| 337 | 1088 | tktA | Transketolase |
| 521 | 0574 | rpi | Ribose 5-phosphate isomerase |
| 1027 | 1101 | zwf | Glucose-6-phosphate-1-dehydrogenase |
| 1388 | 1495 | tal | Transaldolase |
| 1439 | 1386 | rpe | Ribulose-phosphate-3-epimerase |
| Sugars | |||
| 1029 | 1103 | glk | Glucokinase |
| Tricarboxylic acid cycle | |||
| 22 | 0026 | gltA | Citrate synthase |
| 23 | 0027 | icd | Isocitrate dehydrogenase |
| 536 | 0588 | oorD | Subunit of 2-oxoglutarate oxidoreductase |
| 537 | 0589 | oorA | Subunit of 2-oxoglutarate oxidoreductase |
| 538 | 0590 | oorB | Subunit of 2-oxoglutarate oxidoreductase |
| 539 | 0591 | oorC | Subunit of 2-oxoglutarate oxidoreductase |
| 716 | 0779 | acnB | Aconitate hydratase |
| 1245 | 1325 | fumC | Fumarase |
| Other | |||
| 88 | 0096 | Keto-acid dehydrogenase | |
| 586 | 0642 | Oxidoreductase | |
| 888 | 0954 | Aldehyde dehydrogenase | |
| 1023 | 0357 | Oxidoreductase | |
| 1028 | 1102 | Dehydrogenase | |
| 1345 | 1452 | thdF | Thiophene/furan oxidation protein |
| Fatty acid and phospholipid metabolism | |||
| 83 | 0090 | fabD | Malonyl-CoA-ACP transacylase |
| 176 | 0190 | Cardiolipin synthase | |
| 181 | 0195 | fabI | Enoyl-ACP reductase |
| 187 | 0201 | plsX | Fatty acid/phospholipid synthesis protein |
| 188 | 0202 | fabH | β-Ketoacyl-ACP synthase III |
| 201 | 0215 | cdsA | CDP-diacylglycerol synthase |
| 354 | 1071 | pssA | Phosphatidylserine synthase |
| 407 | 1016 | pgsA | Phosphatidylglycerophosphate synthase |
| 409 | 1014 | Short-chain dehydrogenase | |
| 451 | 0499 | pldA | Phospholipase A1 |
| 504 | 0557 | accA | Acetyl-CoAa carboxylase subunit A |
| 505 | 0558 | fabB | β-Ketoacyl-ACP synthase I |
| 506 | 0559 | acpP | ACP |
| 508 | 0561 | fabG | Acetyl-CoA carboxylase subunit A |
| 636 | 0692 | scoB | 3-Oxoacid CoA-transferase, subunit B |
| 637 | 0691 | scoA | 3-Oxoacid CoA-transferase, subunit A |
| 638 | 0690 | thl | Acetyl-CoA acetyltransferase |
| 640 | 0700 | dgkA | Diacylglycerol kinase |
| 674 | 0737 | pgpA | Phosphatidylglycerophosphatase A |
| 744 | 0808 | acpS | Holo-ACP synthase |
| 805 | 0871 | cdh | CDP-diacylglycerol pyrophosphatase |
| 884 | 0950 | accD | Acetyl-CoA carboxylase subunit B |
| 968 | 0416 | cfa | Cyclopocyclopropane fatty acid synthase |
| 1010 | 0371 | accB | Biotin carboxyl carrier protein |
| 1011 | 0370 | accC | Biotin carboxylase |
| 1267 | 1348 | plsC | 1-Acyl-SN-glycerol-3-phosphate acyltransferase |
| 1275 | 1357 | psd | Phosphatidylserine decarboxylase |
| 1290 | 1376 | fabZ | Hydroxymyristoyl-ACP dehydratase |
| Purines, pyrimidines, nucleosides and nucleotides | |||
| 2′-Deoxyribonucleotide metabolism | |||
| 621 | 0680 | nrdA | Ribonucleoside-diphosphate reductase 1 alpha chain |
| 799 | 0865 | dut | Deoxyuridine 5′-triphosphate nucleotidohydrolase |
| 1009 | 0372 | dcd | Deoxycytidine triphosphate deaminase |
| 1016 | 0364 | nrdB | Ribonucleoside-diphosphate reductase 1 beta chain |
| Purine ribonucleotide biosynthesis | |||
| 1039 | 1112 | purB | Adenylosuccinate lyase |
| 1140 | 1218 | purD | Glycinamide ribonucleotide synthetase |
| 1327 | 1434 | purU | Formyltetrahydrofolate hydrolase |
| Pyrimidine ribonucleotide biosynthesis | |||
| 5 | 0005 | pyrF | Orotidine 5′-phosphate decarboxylase |
| 184 | 0198 | ndk | Nucleoside diphosphate kinase |
| 251 | 0266 | pyrC1 | Dihydroorotase |
| 323 | 0349 | pyrG | CTP synthase |
| 341 | 1084 | pyrB | Aspartate carbamoyltransferase catalytic chain |
| 412 | 1011 | pyrD | Dihydroorotate dehydrogenase |
| 528 | 0581 | pyrC2 | Dihydroorotase |
| 714 | 0777 | pyrH | Uridylate kinase |
| 853 | 0919 | pyrA1 | Carbamoyl-phosphate synthase large chain |
| 1158 | 1237 | pyrA2 | Carbamoyl-phosphate synthase small chain |
| 1178 | 1257 | pyrE | Orotate phosphoribosyltransferase |
| Salvage and interconversion of nucleosides and nucleotides | |||
| 96 | 0104 | cpdB | 2′,3′-Cyclic nucleotide 2′-phosphodiesterase |
| 239 | 0255 | purA | Adenylosuccinate synthetase |
| 304 | 0321 | gmk | Guanylate kinase |
| 519 | 0572 | apt | Adenine phosphoribosyltransferase |
| 561 | 0618 | adk | Adenylate kinase |
| 672 | 0735 | gpt | Xanthine-guanine phosphoribosyltransferase |
| 679 | 0742 | prsA | Phosphoribosyl pyrophosphate synthetase |
| 768 | 0829 | guaB | Inosine-5′-monophosphate dehydrogenase |
| 790 | 0854 | guaC | GMP reductase |
| 972 | 0409 | guaA | GMP synthetase |
| 1104 | 1178 | deoD | Purine nucleoside phosphorylase |
| 1105 | 1179 | deoB | Phosphopentomutase |
| 1367 | 1474 | tmk | Thymidylate kinase |
| Sugar-nucleotide biosynthesis and conversions | |||
| 591 | 0646 | galU | UTP-glucose-1-phosphate uridylyltransferase |
| 624 | 0683 | glmU | UDP-N-acetylglucosamine pyrophosphorylase |
| 1420 | 1532 | glmS | Glutamine fructose-6-phosphate minotransferase |
| Regulatory functions | |||
| General | |||
| 41 | 0048 | Transcriptional regulator | |
| 81 | 0088 | rpoD | RNA polymerase sigma 70 factor |
| 151 | 0164, 0165 | Histidine kinase sensor protein | |
| 152 | 0166 | Transcriptional regulator | |
| 229 | 0244 | Histidine kinase sensor protein | |
| 263 | 0278 | gppA | Guanosine-5′-triphosphate,3′-diphosphate pyrophosphatase |
| 381 | 1043 | Transcriptional regulator | |
| 392 | 1032 | fliA | RNA polymerase sigma 28 factor |
| 397 | 1027 | fur | Ferric uptake regulation protein |
| 399 | 1025 | Transcriptional regulator | |
| 403 | 1021 | Transcriptional regulator | |
| 643 | 0703 | Transcriptional regulator | |
| 652 | 0714 | rpoN | RNA polymerase sigma-54 factor |
| 664 | 0727 | Transcriptional regulator | |
| 712 | 0775 | spoT | Guanosine-3′,5′-bis(diphosphate)-3′-pyrophosphohydrolase |
| 981 | 0400 | lytB | Lysis tolerance protein |
| 1207 | 1287 | Transcriptional regulator | |
| 1282 | 1364 | Histidine kinase sensor protein | |
| 1283 | 1365 | Transcriptional regulator | |
| 1335 | 1442 | csrA | Carbon storage regulator |
| 1443 | 1365 | Transcriptional regulator | |
| 1480 | 1572 | dniR | Regulatory protein |
| Chemotaxis and motility | |||
| 17 | 0019 | cheV1 | Chemotaxis protein |
| 75 | 0082 | MCP | |
| 91 | 0099 | MCP | |
| 95 | 0103 | MCP | |
| 358 | 1067 | cheY | Response regulator |
| 546 | 0599 | MCP | |
| 559 | 0616 | cheV2 | Chemotaxis protein |
| 988 | 0393 | cheV3 | Chemotaxis protein |
| 989 | 0392 | cheA | Histidine kinase |
| 990 | 0391 | cheW | Histidine kinase-MCP coupling protein |
| Transcription | |||
| Degradation of RNA | |||
| 1136 | 1213 | pnp | Polyribonucleotide nucleotidyltransferase |
| 1169 | 1248 | vacB | RNase II family protein |
| 1299 | 1407 | rbn | RNase N |
| DNA-dependent RNA polymerase | |||
| 1121 | 1198 | rpoB | DNA-directed RNA polymerase, beta subunit |
| 1213 | 1293 | rpoA | DNA-directed RNA polymerase, alpha subunit |
| 1458 | 1541 | mfd | Transcription-repair coupling factor |
| Transcription factors | |||
| 1 | 0001 | nusB | Transcription termination |
| 497 | 0550 | rho | Transcription termination factor |
| 800 | 0866 | greA | Transcription elongation factor (transcript cleavage factor) |
| 1126 | 1203 | nusG | Transcription antitermination protein |
| 1407 | 1514 | nusA | N utilization substance protein A |
| RNA processing | |||
| 583 | 0640 | pcnB | Polynucleotide adenylyltransferase |
| 607 | 0662 | rnc | RNase III |
| Translation | |||
| Aminoacyl-tRNA synthetases | |||
| 113 | 0123 | thrS | Threonyl-tRNA synthetase |
| 170 | 0182 | lysS | Lysyl-tRNA synthetase |
| 223 | 0238 | proS | Prolyl-tRNA synthetase |
| 302 | 0319 | argS | Arginyl-tRNA synthetase |
| 428 | 0476 | gltX | Glutamyl-tRNA synthetase |
| 560 | 0617 | aspS | Aspartyl-tRNA synthetase |
| 588 | 0643 | gltX | Glutamyl-tRNA synthetase |
| 711 | 0774 | tyrS | Tyrosyl-tRNA synthetase |
| 818 | 0886 | cysS | Cysteinyl-tRNA synthetase |
| 894 | 0960 | glyQ | Glycyl-tRNA synthetase alpha chain |
| 906 | 0972 | glyS | Glycyl-tRNA synthetase beta chain |
| 967 | 0417 | metG | Methionyl-tRNA synthetase |
| 978 | 0403 | pheS | Phenylalanyl-tRNA synthetase alpha chain |
| 979 | 0402 | pheT | Phenylalanyl-tRNA synthetase beta chain |
| 1080 | 1153 | valS | Valyl-tRNA synthetase |
| 1115 | 1190 | hisS | Histidyl-tRNA synthetase |
| 1162 | 1241 | alaS | Alanyl-tRNA synthetase |
| 1174 | 1253 | trpS | Tryptophanyl-tRNA synthetase |
| 1317 | 1422 | ileS | Isoleucyl-tRNA synthetase |
| 1373 | 1480 | serS | Seryl-tRNA synthetase |
| 1452 | 1547 | leuS | Leucyl-tRNA synthetase |
| Degradation of proteins, peptides and glycopeptides | |||
| 29 | 0033 | clpA | ATP-dependent protease, ATP-binding subunit |
| 155 | 0169 | Protease | |
| 249 | 0264 | clpB | Heat shock protein |
| 271 | 0286 | ftsH | ATP-dependent Zn metallopeptidase |
| 356 | 1069 | ftsH | ATP-dependent Zn metallopeptidase |
| 387 | 1037 | pepQ | Proline peptidase |
| 405 | 1019 | htrA | Protease DO |
| 411 | 1012 | Zn protease | |
| 422 | 0470 | pepF | Oligopeptidase |
| 464 | 0515 | hslV | Heat shock protein |
| 465 | 0516 | hslU | Heat shock protein |
| 517 | 0570 | pepA | Aminopeptidase |
| 602 | 0657 | Processing protease | |
| 603 | 0658 | gatB | Glu-tRNA amidotransferase, subunit B |
| 730 | 0794 | clpP | ATP-dependent protease, proteolytic subunit |
| 769 | 0830 | gatA | Glu-tRNA amidotransferase, subunit A |
| 909 | 0975 | gatC | Glu-tRNA amidotransferase, subunit C |
| 999 | 0382 | Zn-metallo protease | |
| 1269 | 1350 | prc | Carboxyl-terminal protease |
| 1288 | 1374 | clpX | ATP-dependent protease, ATP-binding subunit |
| 1293 | 1379 | lon | ATP-dependent protease 1a |
| 1328 | 1435 | sppA | Protease |
| 1491 | 1584 | ydiE | O-Sialoglycoprotein endopeptidase |
| Nucleoproteins | |||
| 774 | 0835 | DNA-binding protein HU | |
| 1052 | 1123 | slyD | FKBP-type peptidyl-prolyl cis-trans isomerase |
| Protein modification | |||
| 210 | 0224 | Peptide methionine sulfoxide reductase | |
| 729 | 0793 | def | Polypeptide deformylase |
| 1017 | 0363 | pcm | Protein-l-isoaspartate O-methyltransferase |
| 1219 | 1299 | map | Methionine aminopeptidase |
| 1334 | 1441 | ppiA | Peptidyl-prolyl cis-trans isomerase |
| Ribosomal proteins, synthesis and modification | |||
| 71 | 0076 | rpsT | 30S ribosomal protein S20 |
| 76 | 0083 | rpsI | 30S ribosomal protein S9 |
| 77 | 0084 | rplM | 50S ribosomal protein L13 |
| 115 | 0125 | rpmI | 50S ribosomal protein L35 |
| 116 | 0126 | rplT | 50S ribosomal protein L20 |
| 186 | 0200 | rpmF | 50S ribosomal protein L32 |
| 281 | 0296 | rplU | 50S ribosomal protein L21 |
| 282 | 0297 | rpmA | 50S ribosomal protein L27 |
| 357 | 1068 | prmA | Ribosomal protein L11 methyltransferase |
| 378 | 1047 | rbfA | Ribosome-binding factor A |
| 384 | 1040 | rpsO | 30S ribosomal protein S15 |
| 443 | 0491 | rpmB | 50S ribosomal protein L28 |
| 463 | 0514 | rplI | 50S ribosomal protein L9 |
| 498 | 0551 | rpmE | 50S ribosomal protein L31 |
| 509 | 0562 | rpsU | 30S ribosomal protein S21 |
| 982 | 0399 | rpsA | 30S ribosomal protein S1 |
| 1074 | 1147 | rplS | 50S ribosomal protein L19 |
| 1078 | 1151 | rpsP | 30S ribosomal protein S16 |
| 1119 | 1196 | rpsG | 30S ribosomal protein S7 |
| 1120 | 1197 | rpsL | 30S ribosomal protein S12 |
| 1122 | 1199 | rplL | 50S ribosomal protein L7/L12 |
| 1123 | 1200 | rplJ | 50S ribosomal protein L10 |
| 1124 | 1201 | rplA | 50S ribosomal protein L1 |
| 1125 | 1202 | rplK | 50S ribosomal protein L11 |
| 1127 | 1204 | rpmG | 50S ribosomal protein L33 |
| 1165 | 1244 | rpsR | 30S ribosomal protein S18 |
| 1167 | 1246 | rpsF | 30S ribosomal protein S6 |
| 1212 | 1292 | rplQ | 50S ribosomal protein L17 |
| 1214 | 1294 | rpsD | 30S ribosomal protein S4 |
| 1215 | 1295 | rpsK | 30S ribosomal protein S11 |
| 1217 | 1296 | rpsM | 30S ribosomal protein S13 |
| 1217 | 1297 | rpmJ | 50S ribosomal protein L36 |
| 1221 | 1301 | rplO | 50S ribosomal protein L15 |
| 1222 | 1302 | rpsE | 30S ribosomal protein S5 |
| 1223 | 1303 | rplR | 50S ribosomal protein L18 |
| 1224 | 1304 | rplF | 50S ribosomal protein L6 |
| 1225 | 1305 | rpsH | 30S ribosomal protein S8 |
| 1226 | 1306 | rpsN | 30S ribosomal protein S14 |
| 1227 | 1307 | rplE | 50S ribosomal protein L5 |
| 1228 | 1308 | rplX | 50S ribosomal protein L24 |
| 1229 | 1309 | rplN | 50S ribosomal protein L14 |
| 1230 | 1310 | rpsQ | 30S ribosomal protein S17 |
| 1231 | 1311 | rpmC | 50S ribosomal protein L29 |
| 1232 | 1312 | rplP | 50S ribosomal protein L16 |
| 1233 | 1313 | rpsC | 30S ribosomal protein S3 |
| 1234 | 1314 | rplV | 50S ribosomal protein L22 |
| 1235 | 1315 | rpsS | 30S ribosomal protein S19 |
| 1236 | 1316 | rplB | 50S ribosomal protein L2 |
| 1237 | 1317 | rplW | 50S ribosomal protein L23 |
| 1238 | 1318 | rplD | 50S ribosomal protein L4 |
| 1239 | 1319 | rplC | 50S ribosomal protein L3 |
| 1240 | 1320 | rpsJ | 30S ribosomal protein S10 |
| 1340 | 1447 | rpmH | 50S ribosomal protein L34 |
| 1389 | 1496 | rplY | 50S ribosomal protein L25 |
| 1445 | 1554 | rpsB | 30S ribosomal protein S2 |
| tRNA modification | |||
| 266 | 0281 | tgt | Queuine-tRNA-ribosyltransferase |
| 363 | 1062 | queA | S-Adenosylmethionine tRNA ribosyltransferase-isomerase |
| 1019 | 0361 | truA | Pseudouridylate synthase I |
| 1069 | 1141 | fmt | Methionyl-tRNA formyltransferase |
| 1075 | 1148 | trmD | tRNA (guanine-n1)-methyltransferase |
| 1254 | 1335 | trmU | tRNA(5-methylaminomethyl-2-thiouridylate)-methyltransferase |
| 1310 | 1415 | miaA | tRNA delta(2)-isopentenylpyrophosphate transferase |
| 1341 | 1448 | rnpA | RNase protein component |
| 1390 | 1497 | pth | Peptidyl-tRNA hydrolase |
| 1406 | 1513 | selA | l-Seryl-tRNA selenium transferase |
| Translation factors | |||
| 72 | 0077 | prfA | Peptide chain release factor 1 |
| 114 | 0124 | infC | Translation initiation factor IF-3 |
| 157 | 0171 | prfB | Peptide chain release factor 2 (RF-2) |
| 163 | 0177 | efp | Elongation factor P (EF-P) |
| 232 | 0247 | deaD | ATP-dependent RNA helicase dead |
| 377 | 1048 | infB | Translation initiation factor IF-2 |
| 1118 | 1195 | fusA | Elongation factor G (EF-G) |
| 1128 | 1205 | tufA | Elongation factor TU (EF-TU) |
| 1177 | 1256 | frr | Ribosome recycling factor (ribosome-releasing factor [RRF]) |
| 1218 | 1298 | infA | Translation initiation factor IF-1 |
| 1322 | 1431 | ksgA | Dimethyladenosine transferase |
| 1444 | 1555 | tfs | Elongation factor TS (EF-TS) |
| Transport and binding proteins | |||
| General | |||
| 66 | 0071 | ureI | Urea transporter |
| 167 | 0179 | ABC transporter, ATP-binding protein | |
| 200 | 0214 | Transporter | |
| 235 | 0250 | ABC transporter, ATP-binding protein | |
| 236 | 0251 | ABC transporter, permease | |
| 300 | 0613 | ABC transporter, ATP-binding protein | |
| 343 | 1082 | msbA | Multidrug resistance protein |
| 449 | 0497 | Transporter | |
| 450 | 0498 | Transporter | |
| 547 | 0600 | Secretion/efflux ABC transporter, ATP-binding protein | |
| 553 | 0606 | Efflux transporter | |
| 554 | 0607 | Efflux transporter | |
| 653 | 0715 | ABC transporter, ATP-binding protein | |
| 685 | 0748 | ABC transporter, ATP-binding protein | |
| 754 | 0818 | Osmoprotection binding protein | |
| 757 | 0818 | Osmoprotection binding protein | |
| 758 | 0819 | Osmoprotection ATP-binding protein | |
| 789 | 0853 | ABC transporter, ATP-binding protein | |
| 806 | 0872 | phnA | Alkylphosphonate uptake protein |
| 871 | 0936 | proP | Proline/betaine transporter |
| 1055 | 1126 | tolB | tonB-independent protein-uptake protein |
| 1057 | 1129 | exbD1 | Biopolymer transport protein |
| 1058 | 1130 | exbB1 | Biopolymer transport protein |
| 1107 | 1181 | Transporter | |
| 1129 | 1206 | ABC transporter, ATP-binding protein | |
| 1141 | 1220 | ABC transporter, ATP-binding protein | |
| 1320 | 1427 | Histidine-rich metal-binding protein | |
| 1321 | 1432 | Histidine- and glutamine-rich metal-binding protein | |
| 1338 | 1445 | exbB3 | Biopolymer transport protein |
| 1339 | 1446 | exbD3 | Biopolymer transport protein |
| 1484 | 1576 | ABC transporter, ATP-binding protein | |
| 1485 | 1577 | ABC transporter, permease | |
| Amino acids, peptides, and amines | |||
| 47 | 0055 | putP | Sodium/proline symporter |
| 121 | 0133 | sdaC | l-Serine transporter |
| 283 | 0298 | dppA | Periplasmic dipeptide transport substrate-binding protein |
| 284 | 0299 | dppB | Dipeptide transport system permease protein |
| 285 | 0300 | dppC | Dipeptide transport system permease protein |
| 286 | 0301 | dppD | Dipeptide transport system ATP-binding protein |
| 287 | 0302 | dppF | Dipeptide transport system ATP-binding protein |
| 406 | 1017 | Amino acid permease | |
| 874 | 0939 | Amino acid ABC transporter, permease protein | |
| 875 | 0940 | Amino acid ABC transporter, binding protein precursor | |
| 877 | 0942 | Sodium/alanine symporter | |
| 1096 | 1169 | Amino acid ABC transporter, permease protein | |
| 1097 | 1170 | Amino acid ABC transporter, permease protein | |
| 1098 | 1171 | Amino acid ABC transporter, ATP-binding protein | |
| 1099 | 1172 | Amino acid ABC transporter, binding protein precursor | |
| 1172 | 1251 | Peptide ABC transporter, ATP-binding protein | |
| 1358 | 1465 | Amino acid ABC transporter, ATP-binding protein | |
| 1399 | 1506 | gltS | Sodium/glutamate symporter |
| Anions | |||
| 425 | 0473 | modA | Molybdenum ABC transporter, periplasmic binding protein |
| 426 | 0474 | modB | Molybdenum ABC transporter, permease |
| 427 | 0475 | modC | Molybdenum ABC transporter, ATP-binding protein |
| 1384 | 1491 | Phosphate permease | |
| Carbohydrates, organic alcohols, and acids | |||
| 128 | 0140 | lldP | l-Lactate permease |
| 129 | 0141 | lldP | l-Lactate permease |
| 334 | 1091 | kgtP | α-Ketoglutarate permease |
| 635 | 0693 | atoE | Short-chain fatty acids transporter |
| 660 | 0724 | dcuA | Anaerobic C4-dicarboxylate membrane transporter |
| 1101 | 1174 | gluP | Glucose/galactose transporter |
| Cations | |||
| 124 | 0136 | bcp | Bacterioferritin comigratory protein |
| 348 | 1077 | nixA | High-affinity nickel transport protein |
| 352 | 1073 | copP | Copper-associated protein |
| 353 | 1072 | copA | Copper-transporting P-type ATPase |
| 423 | 0471 | kefB | Glutathione-regulated potassium efflux system protein |
| 442 | 0490 | Putative potassium channel protein | |
| 529 | 0582 | tonB1 | Siderophore-mediated iron transport protein |
| 598 | 0653 | pfr | Nonheme iron-containing ferritin |
| 626 | 0686 | fecA1 | Iron(III) dicitrate transport protein |
| 627 | 0687 | feoB | Ferrous iron transport protein B |
| 727 | 0791 | hmcT | Heavy-metal cation-transporting P-type ATPase |
| 743 | 0807 | fecA2 | Iron(III) dicitrate transport protein |
| 821 | 0888 | fecE | Iron(III) dicitrate transport system ATP-binding protein |
| 822 | 0889 | fecD | Iron(III) dicitrate transport system permease protein |
| 903 | 0969 | czcA1 | Cation efflux system protein |
| 904 | 0970 | czcB1 | Cation efflux system protein |
| 1109 | 1183 | Na+/H+ antiporter | |
| 1248 | 1328 | czcB2 | Cation efflux system protein |
| 1249 | 1329 | czcA2 | Cation efflux system protein |
| 1258 | 1339 | exbB2 | Biopolymer transport protein |
| 1259 | 1340 | exbD2 | Biopolymer transport protein |
| 1260 | 1341 | tonB2 | Siderophore-mediated iron transport protein |
| 1263 | 1344 | corA | Magnesium and cobalt transport protein |
| 1396 | 1503 | fixI | Component of cation transport for cbb3-type oxidase |
| 1426 | 1400 | fecA3 | Iron(III) dicitrate transport protein |
| 1447 | 1552 | nhaA | Na+/H+ antiporter I |
| Nucleosides, purines, and pyrimidines | |||
| 1106 | 1180 | Nucleoside transporter | |
| 1210 | 1290 | pnuC | Nicotinamide mononucleotide transporter |
Gene numbers correspond to those in Fig. 1.
TABLE 2.
Annotation and classification of genes from H. pylori J99 and 26695
| Annotation category | No. of genes in:
|
||
|---|---|---|---|
| H. pylori J99 | H. pylori 26695 | Both strainsa | |
| Functionally classified | 877 | 898 | |
| Conserved with no known function | 275 | 290 | |
| H. pylori specific | 343 | 364 | |
| Total | 1,495 | 1,552 | |
| Amino acid biosynthesis | 44 | 44 | 44 |
| Biosynthesis of cofactors, etc. | 60b | 59 | 59 |
| Cell envelope | 160 | 164 | 156cde |
| Cellular processes | 96 | 113 | 92cde |
| DNA replication | 23 | 23 | 21ce |
| DNA restriction-modification, etc. | 66 | 68 | 51cde |
| Energy metabolism | 104 | 104 | 102cde |
| Fatty acid and phospholipid metabolism | 28 | 29 | 28e |
| Purine and pyrimidine biosynthesis | 34 | 34 | 34 |
| Regulatory functions | 32 | 32 | 31df |
| Transcription | 13 | 13 | 13 |
| Translation | 128 | 128 | 128 |
| Transport and binding proteins | 88g | 87 | 87 |
| Conserved with no known function | 275 | 290 | 267cde |
| H. pylori specific | 343 | 364 | 288cde |
| Total | 1,495 | 1,552 | 1,401h |
Using J99 genes as the basis for counting.
Includes the partial duplication of folE (JHP862).
Categories which include H. pylori J99-specific genes (see the text for details).
These numbers include the “split” genes based on the H. pylori J99 definition. There are 6 J99 genes which constitute 12 26695 genes in the cell envelope; 2 J99 genes which constitute 4 26695 genes in cellular processes; 1 J99 gene which constitutes 2 26695 genes in each of DNA restriction-modification, energy metabolism, and Regulatory functions; 5 J99 genes which constitute 7 26695 genes in conserved with no known function; and 24 J99 genes which constitute 33 26695 genes in H. pylori specific.
Categories which include H. pylori 26695-specific genes.
Does not include the duplication of the response regulator (JHP1283 and JHP1443).
Includes the partial duplication of proX (JHP754).
The remaining 94 genes represent the 89 J99-specific genes, tnpA/B from IS606, and the partial or complete duplications of three genes.
FIG. 1.
Linear representation of the H. pylori J99 chromosome, illustrating the location of each predicted protein-coding region, rRNA gene, tRNA gene, IS605 or IS606 element and related fragment, and NotI endonuclease site. The predicted protein-coding regions are color coded based on functional classification (see the bottom of the figure for the code), with the direction of transcription indicated by an arrowhead. H. pylori J99 ORFs are numbered sequentially in red, and the corresponding homologous gene, if it exists in strain 26695, is numbered in black. The positions of the NotI endonuclease sites in J99 are indicated with the number of conserved nucleotides in the recognition sequence (x/8) at the corresponding position in strain 26695. The numbers associated with the tRNA symbols (inverted triangles) represent the number of tRNA genes at a specific locus. Vertical hash marks, below the linear chromosome, are located every 20 kb.
Comparison of orthologous genes and their encoded products showed a high degree of conservation. Sequence variation between the two strains was significantly greater at the nucleotide level than at the amino acid level. Because the nucleotide variation occurred most commonly in the third position of a coding triplet, the primary sequence of the encoded protein was highly conserved (Table 3). The fact that many of the nucleotide differences are silent with respect to the protein sequence suggests that there is a strong selective pressure for functional conservation at the protein level.
TABLE 3.
Nucleotide and amino acid identity between genes common to H. pylori J99 and 26695
| % Identity | No. (%) of predicted ORFsa
|
|
|---|---|---|
| Nucleotide | Amino acid | |
| 100 | 0 (0) | 41 (2.9) |
| 98.0–99.9 | 8 (0.6) | 269 (19.3) |
| 96.0–97.9 | 249 (17.8) | 359 (25.7) |
| 94.0–95.9 | 566 (40.5) | 279 (20.0) |
| 92.0–93.9 | 306 (21.9) | 169 (12.1) |
| 90.0–91.9 | 89 (6.4) | 86 (6.2) |
| 85.0–89.9 | 81 (5.8) | 77 (5.5) |
| <85 | 97 (7.0) | 116 (8.3) |
| Total | 1,396 (100) | 1,396 (100) |
Genes that appear “split” by putative frameshifts in either strain have been classed as the larger ORF in the above analysis.
The nucleotide drift in the third position of a coding triplet is probably responsible for the majority of the DNA-based “diversity” reported for H. pylori (3, 4, 12, 54, 75, 77, 148). For example, pulsed-field gel electrophoresis mapping data have been interpreted to mean that the gene order and physical arrangement of the chromosome are highly variable from strain to strain (75, 148). By using this technique, strain J99 and strain 26695 would appear to be highly divergent in both the number of NotI fragments and gene location (6). This apparent genetic diversity is easily explained by two inversions in combination with the silent nucleotide drift, which is responsible for six of the seven additional NotI sites found in strain J99 compared to strain 26695 (6). Although the genomic content and the resulting physiological capabilities of the two strains are almost identical, these few differences in gene arrangement would have classified these strains as diverse. This example reveals the limitations of DNA-based methods when used to examine strain diversity.
NUTRITIONAL REQUIREMENTS
Amino Acids and Polyamines
Both sequenced strains of H. pylori have homologues to all the genes that would be needed to synthesize eight amino acids from central intermediary metabolites (Table 4). Studies of the growth requirements for several strains of H. pylori have shown an absolute need for arginine, histidine, leucine, isoleucine, valine, methionine, and phenylalanine (118, 132), a finding consistent with the genomic sequence information.
TABLE 4.
Predicted biosynthetic abilities and auxotrophies of H. pylori
| Category and Compound | No. of genes presenta | Predicted synthetic abilityb |
|---|---|---|
| Amino acids | ||
| Aspartic acid | 2 (2) | Y |
| Cysteine | 2 (3)c | Y |
| Glutamic acid | 1 (1) | Y |
| Glutamine | 1 (1) | Y |
| Glycine | 1 (1) | Y |
| Lysine | 6 (7)d | Y |
| Threonine | 5 (5) | Y |
| Tryptophan | 5 (5) | Y |
| Alanine | 1 (2) | N |
| Arginine | 0 (9) | N |
| Histidine | 0 (8) | N |
| Isoleucine | 2 (9) | N |
| Leucine | 1 (6) | N |
| Methionine | 1 (7) | N |
| Phenylalanine | 1 (3) | N |
| Proline | 1 (3) | N |
| Valine | 2 (9) | N |
| Asparagine | 0 (2)e | ? |
| Serine | 2 (3) | ? |
| Tyrosine | 1 (3) | ? |
| Cofactors and vitamins | ||
| Biotin | 4 (4) | Y |
| CoA | 0 (5)d | Y |
| Folate | 7 (10)d | Y |
| Molybdopterin | 8 (9)c | Y |
| Panothenate | 4 (4) | Y |
| Protoheme | 10 (10) | Y |
| Pyridine nucleotides | 3 (5)d | Y |
| Pyridoxial phosphate | 3 (?)d | Y |
| Riboflavin | 6 (7)c | Y |
| Thiamine | 3 (9)cd | Y |
| Thioredoxin | 2 (2) | Y |
| B12 | ||
| Glutathione | 0 (2) | N |
| Siroheme | 0 (1) | N |
| Ubiquinone | 3 (8) | ? |
| Menaquinone | 0 (6) | ?f |
| Polyamines | ||
| Agmatine | 1 (1) | Y |
| Putrescine | 1 (3) | N |
| Spermidine | 3 (5) | ?g |
| Pyrimidines | 11 (10)h | Y |
| Purines | 3 (10) | Ni |
Number of genes assigned in E. coli shown in parentheses. Data from reference 119.
Y, yes; N, no; ?, not clear.
H. pylori has a gene for each step in the biosynthetic pathway. The number of genes in E. coli is larger due to redundancy.
Not all genes in this pathway have been identified in E. coli.
Synthesis may occur via tRNA (see the text).
The presence of menaquinones in H. pylori has been demonstrated (see the text).
Spermidine synthesis may be mediated by NapC.
Two copies of pyrC were found in H. pylori.
Salvage pathway is present (see the text).
Although no homologues to the genes involved in the amidation of aspartate were detected in H. pylori, several strains have been reported to grow in the absence of asparagine (132). It is possible that asparagine is synthesized by an aspartyl-tRNA-asparagine amidotransferase, similar to what has been observed with glutaminyl-tRNA biosynthesis in Bacillus subtilis (see “Transcription and translation” below) (29).
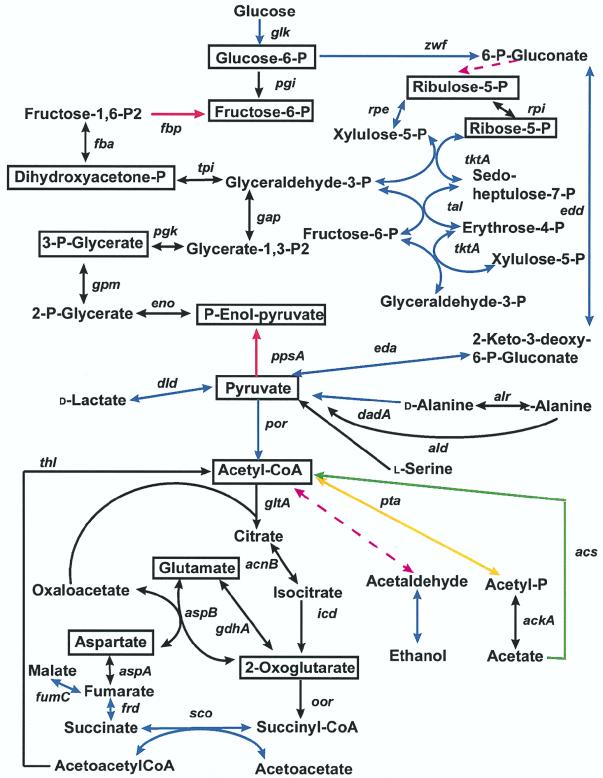
Both the serine and tyrosine biosynthetic pathways were complete except for a homologue to their respective specific transaminase. However, each of these reactions may be catalyzed by one of the several identified transaminases with undetermined substrate specificity (JHP206/HP220, JHP568/HP624, JHP673/HP736, and JHP976/HP405). Such an enzymatic activity would allow the de novo synthesis of these amino acids, as observed in some strains of H. pylori (132). Regardless of whether H. pylori can synthesize serine, it possesses a specific transporter which allows the acquisition of this amino acid from the environment. sdaC, which encodes the serine transporter, is contiguous with sdaB, whose protein product in Escherichia coli converts l-serine to pyruvate. Similarly, putP, which encodes a proline transporter, is adjacent to putA, which encodes a bifunctional enzyme that oxidizes proline to l-glutamate in E. coli. The alanine transporter gene (JHP877/HP0942) is clustered with two other genes (alr and dadA) which are involved in alanine metabolism (Fig. 2). A positive regulator of the dad operon is thought to be upstream of this gene cluster in E. coli. In H. pylori, a putative regulatory gene (JHP879/HP0944) has also been identified upstream of the dad gene cluster. This putative regulatory gene does not have homology to the putative E. coli regulator, a finding which may indicate that the regulation of alanine catabolism is different in these two species. The H. pylori gene clusters described above would allow for the uptake and utilization of serine, proline, and alanine as carbon and nitrogen sources. In addition, H. pylori has a transporter for the uptake of glutamate (JHP1399/HP1506), an amino acid abundant in gastric juice (82).
FIG. 2.
Central metabolic pathways of H. pylori. Boxed compounds are key central intermediates. Black lines are reactions predicted to occur from the genomic analysis. Blue lines represent reactions that have been reported in the literature and are consistent with genomic analysis. Green lines represent a predicted reaction occurring only in strain 26695. Yellow lines represent a predicted reaction occurring only in strain J99. Broken magenta lines represent reactions reported in the literature but for which no homologue to the enzyme has been identified in either genome. Red lines represent key steps regulating gluconeogenesis/glycolysis.
H. pylori also encodes homologues for four other amino acid uptake systems with unknown specificity. One of these systems consists of a gene cluster encoding a multisubunit periplasmic permease (JHP1096–1099/HP1169–1172). The putative operon encodes two permeases, an ATP-binding protein and a periplasmic binding protein. Based on sequence similarity, the ligand for this high-affinity transporter may be either glutamine, histidine, or arginine. H. pylori is unable to synthesize the last two amino acids and therefore requires transport systems for them. Furthermore, the apparent inability of H. pylori to synthesize phenylalanine, methionine, and the branched-chain amino acids necessitates specific transport of these amino acids. No such specific transport systems were identified, but they may be encoded by any of the transport systems with unassigned ligand specificity.
In addition to transporting amino acids, H. pylori may have the ability to transport the abundant peptides which are found in the stomach. Homologues to a dipeptide transport system are present and its five genes (JHP283–287/HP0298–0302) are arranged contiguously, similar to the organization of the dpp operon in E. coli (1). There is also a single gene (JHP1172/HP1251) which displays significant sequence similarity to an oligopeptide transporter.
The genomic sequence provides little information on the composition of polyamines in H. pylori, which are needed for optimal growth in most cells. The homologue of SpeA allows the conversion of arginine to agmatine in H. pylori. Although H. pylori has a homologue to speE, which encodes spermidine synthetase, it is unlikely that this enzyme can catalyze spermidine biosynthesis since no homologues for the enzymes that provide precursors for SpeE (SpeB to SpeD) were detected. However, H. pylori may be able to synthesize spermidine through nspC. The product of this gene can synthesize spermidine by decarboxylating carboxyspermidine (117). It is also possible that H. pylori uses nspC for the synthesis of norspermidine, a polyamine found in Vibrio alginolyticus.
Cofactors and Vitamins
Both sequenced strains of H. pylori have all the identified genes needed for the biosynthesis of biotin, folate, heme, molybdopterin, pantothenate, pyridoxial phosphate, riboflavin, and thioredoxin (Table 4). H. pylori has all the genes necessary for the synthesis of NAD with the exception of nadB, which encodes the aspartate oxidase subunit of quinolate synthetase in E. coli (138). This polypeptide is the oxygen-utilizing subunit of an enzyme which converts l-aspartate to iminoaspartate. The absence of NadB is not unexpected since the mechanism by which anaerobic or microaerophilic bacteria, such as H. pylori, synthesize iminoaspartate is unknown and is not likely to be oxygen dependent. In addition, nicotinamide mononucleotide transporter (JHP1210/HP1290) was identified. No homologues to enzymes involved in vitamin B12 and coenzyme A biosynthesis were identified. Vitamin B12 is an important cofactor for certain enzymes involved in anaerobic metabolism such as methionine synthase. H. pylori may not need to synthesize this vitamin, since homologues to B12-requiring enzymes were not identified. Bacteria synthesize coenzyme A de novo from pantothenate. However, no homologues to known enzymes involved in its biosynthesis were identified in H. pylori, making this pathway unique with respect to those previously reported. The pathway for thiamine biosynthesis has not been completely defined. Some of the genes believed to be involved in thiamine synthesis were found, suggesting that H. pylori can make this vitamin. However, it has been reported that H. pylori requires thiamine for growth (118).
H. pylori has homologues to all of the genes necessary to produce riboflavin. A single gene in H. pylori, homologous to both ribB and ribA (JHP740/HP0804), encodes a bifunctional enzyme with both GTP cyclohydrolase II and 3,4-dihydroxy-2-butanone 4-phosphate synthase activities (158). In addition, H. pylori has a separate GTP cyclohydrolase II (RibA) homologue (JHP738/HP802) downstream from the bifunctional RibAB. Worst et al. (158) have shown that expression of RibAB but not RibA is regulated by iron limitation in H. pylori. The significance of this enzymatic duplication and differential gene regulation is unknown.
Purine and Pyrimidine Biosynthesis, Salvage, and Interconversion
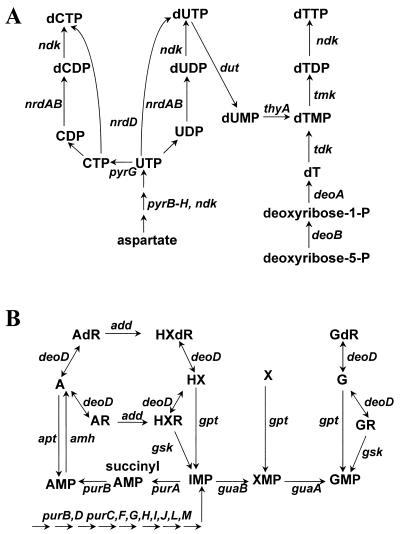
Enzymes for the de novo biosynthesis of purines are largely absent in both sequenced strains of H. pylori, which implies that this bacterium cannot synthesize purine nucleotides from formate, glycine, or serine. Genes that encode homologues for all of the purine salvage and interconversion enzymes are present (Fig. 3A). No homologues to a purine transporter were identified, although there is biochemical evidence for the transport of purine bases in H. pylori (107, 118). Based on similarity, it is likely that the putative transporter (JHP1106/HP1180) is specific for nucleosides, which would allow H. pylori to obtain purines via the salvage and interconversion pathways.
FIG. 3.
Pyrimidine salvage and interconversion (A) and purine salvage pathways (B). A, adenosine; G, guanine; HX, hypoxanthine; X, xanthine; R, ribonucleoside; dR, deoxyribonucleoside. Adapted from reference 119.
H. pylori possesses homologues to all of the genes necessary for the de novo synthesis of UTP and CTP, consistent with experimental results that show that radiolabeled pyrimidine nucleotide precursors are incorporated into DNA (108). Unlike E. coli, H. pylori possesses a second pyrC homologue encoding dihydroorotase, raising the possibility that the H. pylori enzyme exists as a heterodimer rather than as a homodimer as reported in E. coli. One of the two homologues (JHP528/HP0581) is more closely related to a PyrC in gram-negative organisms, whereas the other homologue (JHP251/HP0266) is more closely related to a PyrC in gram-positive organisms. Furthermore, H. pylori lacks a homologue for the regulatory chain of the aspartate transcarbamoylase (PyrI), suggesting that either a paralogous gene serves this function or PyrB functions in the absence of a regulatory subunit.
H. pylori has homologues for all of the enzymes used for the interconversion of pyrimidine deoxyribonucleotides (Fig. 3B) with the exception of thymidylate synthase (ThyA), which is required for the interconversion of dUMP to dTMP. In addition, homologues for all of the enzymes associated with the pyrimidine salvage pathway are absent except for DeoB. This absence is consistent with poor utilization of uracil and uridine and with the failure to detect incorporation of added thymine, cytosine, or deoxycytidine into DNA by H. pylori (108). Although thymidine kinase activity has been found in crude extracts of H. pylori (108), no gene encoding a homologue of thymidine kinase (tdk) was identified in H. pylori.
Inorganic Elements and Heavy Metals
Phosphorus is an essential element in bacteria. H. pylori possesses homologues to both polyphosphate kinase and inorganic pyrophosphatase. These enzymes confer the ability to synthesize and hydrolyze polyphosphate, in agreement with experimental evidence (17). H. pylori also possesses a phosphate transporter (JHP1384/HP1491).
Sulfur assimilation is restricted in H. pylori compared to E. coli. Homologues to the genes necessary for the assimilation of sulfide and cysteine (cysE and cysK) are present in both sequenced strains, whereas those for the assimilation of sulfate (cysA, cysC, cysD, cysH, and cysN), an energy-consuming process, are not. The absence of an identifiable sulfate permease supports the apparent inability of H. pylori to use sulfate. Whereas H. pylori can utilize only sulfide as a source of inorganic sulfur, the closely related bacterium Campylobacter jejuni has homologues to the genes necessary to assimilate sulfate, sulfite, and sulfide (based on analysis of the recently completed genome by the Sanger Centre). This difference in sulfur assimilation between H. pylori and Campylobacter spp. is also seen in sulfur dissimilation. Unlike many Campylobacter spp., H. pylori does not have the homologues necessary for the respiration of many sulfur compounds. The absence of these sulfur assimilatory and dissimilatory genes in H. pylori probably reflects the evolved physiology resulting from its unique gastric niche.
In E. coli and Salmonella typhimurium, nitrogen is derived mainly from the primary products of ammonia assimilation, i.e., glutamate and glutamine. Glutamine synthetase (GlnA) catalyzes the formation of glutamine, while either glutamate dehydrogenase (GdhA) or glutamate synthase (GltBD) catalyzes the formation of glutamate (131). Homologues to glnA and gdhA were identified, and the gene product of glnA has been characterized in H. pylori (45). Glutamate dehydrogenase and glutamine synthetase allow H. pylori to incorporate nitrogen from urea into amino acids, presumably via ammonia, as demonstrated previously (157). No homologue of GltBD was identified. In other bacteria, the absence of this enzyme results in the inability to grow in a medium with low levels of free ammonia (18, 32, 126). Presumably, H. pylori does not need GltBD because sufficient levels of free ammonia are generated by enzymes such as urease.
Homologues of genes belonging to several iron uptake systems were identified in both sequenced strains, indicating the importance of iron metabolism in H. pylori is similar to that in other pathogenic bacteria. H. pylori possesses homologues to some of the genes involved in the ferric citrate (Fec) transport system, but no identifiable homologues to FecB, a periplasmic protein, or FecC, a component of the cytoplasmic membrane channel, were found. Three homologues to genes encoding the outer membrane receptor FecA are present in H. pylori, one of which may be involved in iron uptake via the ferric citrate system. One of the three fecA homologues is adjacent to the feoB homologue, a gene encoding a cytoplasmic ferrous iron permease. These two genes may be involved in ferrous, rather than ferric, uptake. Another fecA homologue may be part of the TonB-dependent iron uptake system (55, 80). The tonB homologue is adjacent to homologues of the remaining two genes (exbB and exbD) comprising the TonB-dependent iron uptake system. In addition, two other sets of exbB and exbD genes were identified but are probably involved in other transport processes (13). H. pylori may also acquire iron through FrpB, for which four homologues were identified. In Neisseria spp., FrpB paralogs are outer membrane proteins that are induced at low iron concentrations, have homology to lactoferrin- and heme-binding proteins, and are postulated to be involved in iron acquisition. Under iron limitation, many bacteria use siderophores to acquire iron. Whether H. pylori produces siderophores is controversial (67, 70). No homologues to genes involved in siderophore biosynthesis were found. H. pylori may not need siderophores, because the amount of free inorganic iron present in the stomach should be sufficient to support bacterial growth.
In addition to iron transport, H. pylori possesses homologues to several other heavy-metal transporters, including NixA, which is responsible for Ni2+ uptake, a function necessary for urease activity (114), and CopAP, which is responsible for Cu2+ uptake (47). H. pylori also possesses homologues to a high-affinity, multisubunit molybdate uptake system, which is needed for the biosynthesis of molybdopterin.
Carbohydrates
Genomic analyses indicate that H. pylori has a limited capability to acquire sugars from the environment, which is in agreement with experimental findings (102). Only a homologue for a phosphoenolpyruvate-independent glucose and galactose transporter was identified, consistent with previous metabolic studies (98, 102). The apparent absence of other sugar uptake systems suggests a limited ability for sugar catabolism in H. pylori (see below). Several homologues to organic acid transporters were identified: two l-lactate permeases, a ketoglutarate permease, and a C4-dicarboxylate transporter that is used under anaerobic conditions in other bacteria. The presence of these transporters suggests that organic acids serve as important sources of carbon for H. pylori (see below).
In summary, based on the analysis of the sequence from both strains, H. pylori is capable of synthesizing the cofactors necessary for growth and acquiring important inorganic elements, although it is limited in its ability to use sulfur. H. pylori would be auxotrophic for at least nine amino acids and purines. Interestingly, no complex sugar transport or degradation homologues were found, suggesting that the bacterium does not acquire sugars from the environment and uses them as sources of energy or as sugar precursors.
CENTRAL INTERMEDIARY AND ENERGY METABOLISM
Central Intermediary Metabolism
Glycolysis and gluconeogenesis.
Genes involved in the metabolism of saccharides to simple sugars were not identified. This finding is consistent with metabolic studies which suggest that complex sugars are not a major energy source for H. pylori (102). Homologues for all enzymes, which carry out reversible steps in the pathway, are present in both sequenced strains of H. pylori. Since homologues of the two nonreversible gluconeogenic enzymes (fructose-1,6-bisphosphatase and pyruvate dikinase) are present and since homologues of the two nonreversible glycolytic enzymes (phosphofructose kinase and pyruvate kinase) were not identified, it appears that H. pylori uses the enzymes of the glycolytic/gluconeogenic pathway for anabolic biosynthesis rather than for catabolic energy production. The experimental evidence supports this hypothesis (59).
Entner-Doudoroff and phosphopentose pathways.
The genomic sequences show that H. pylori possesses homologues of all the genes involved in the Entner-Doudoroff pathway, suggesting that glucose can be used as a source of energy, which is consistent with published data (20, 103, 104). H. pylori has homologues of genes that encode all the enzymes in the phosphopentose shunt except gluconate-6-phosphate dehydrogenase. The presence of this enzymatic activity in crude extracts of H. pylori has been suggested (59, 100). However, the conversion of gluconate-6-phosphate to ribulose-5-phosphate could occur indirectly via the Entner-Doudoroff and the phosphopentose pathways (Fig. 2). The phosphopentose pathway enzymes that were identified in H. pylori allow for the generation of all the intermediates normally produced by this pathway.
Pyruvate metabolism.
The genomic analyses suggest that glucose or malate is not the primary source for production of pyruvate in H. pylori but, rather, that lactate, l-alanine, l-serine, and d-amino acids are the primary sources, which is consistent with the literature (106, 143). H. pylori appears only to convert pyruvate to acetyl coenzyme Δ (acetyl-6A) by pyruvate oxidoreductase (63, 64), lacking homologues for pyruvate dehydrogenase, pyruvate formate-lyase, and pyruvate oxidase. H. pylori dissimilates pyruvate to produce acetate, formate, succinate, and lactate (99, 106), an experimental observation consistent with the genomic analyses.
Fermentation.
H. pylori ferments pyruvate to acetate. Genetic analysis indicates that strain J99 can carry out this fermentation but that strain 26695 cannot do so due to a frameshift in its phosphotransacetylase gene. This mutation also implies that strain 26695 cannot convert acetate to acetyl-CoA by running the fermentative pathway in the reverse direction. However, in H. pylori 26695 the single, identifiable strain-specific gene involved in energy metabolism is an acetyl-CoA synthetase (HP1045) (149), which allows the direct conversion of acetate to acetyl-CoA.
The presence of alcohol dehydrogenase activity in H. pylori suggests that this bacterium can ferment pyruvate to ethanol (135, 136). Indeed, a homologue of this enzyme was identified in both strains. In addition, the single, identifiable strain-specific gene involved in energy metabolism in strain J99 (JHP1429) is a second alcohol dehydrogenase homologue. The role of alcohol dehydrogenase in H. pylori is unclear, because this organism apparently cannot ferment pyruvate to ethanol due to the absence of an identifiable acetaldehyde dehydrogenase homologue. The absence of such a homologue is consistent with biochemical studies (135, 136). The ability of H. pylori to produce acetaldehyde from ethanol (133) suggests that this bacterium is very sensitive to alcohols since it cannot detoxify the resulting acetaldehyde.
Tricarboxylic acid cycle.
The tricarboxylic acid (TCA) cycle of H. pylori in both sequenced strains is similar to the branched anaerobic TCA pathway used by E. coli (Fig. 2) with the following exceptions. Succinyl-CoA is generated from 2-oxoglutarate rather than from succinate (64). Furthermore, in H. pylori, fumarate is generated in the TCA pathway from aspartate rather than from malate. In contrast, genomic analysis suggests that in the closely related bacterium C. jejuni, succinyl-CoA is synthesized from succinate and that fumarate is synthesized from malate (based on analysis of the recently completed genome by the Sanger Centre). H. pylori possesses a homologue of fumarase, explaining the conversion of malate to fumarate observed in crude extracts (101). Unlike C. jejuni, no other malate-utilizing enzymes, such as malate dehydrogenase, malate synthase, or malate oxidoreductase, were identified in H. pylori.
Fatty acid degradation.
Both sequenced strains of H. pylori contain the genes necessary for C2 or short-chain fatty acid atabolism (25) and for a short-chain fatty acid transporter. No identifiable homologues were found to the genes involved in long-chain fatty acid β-oxidation. Together, these observations indicate that H. pylori may utilize acetoacetate and not acetobutyrate as a source for short chain fatty acid catabolites.
Electron Transport Chain
Electron donors.
The initial transfer of electrons during the oxidation of d-lactate and NADH may be performed by homologues to d-lactate dehydrogenase (dld), NADH dehydrogenase I (nuoA-nouN), and hydrogenase (hyaA to hyaDD). The presence of these genes in both sequenced strains is consistent with measured activities in H. pylori cell membranes (22, 93, 101, 105). Although homologues to NuoE and NuoF were not identified in H. pylori, the nuo cluster contained two ORFs in an identical location and of similar size to the two E. coli genes. Thus, it is likely that these two H. pylori ORFs encode proteins with orthologous functions to NuoE and NuoF. In addition, homologues for the following electron-transferring systems were found: pyruvate ferredoxin oxidoreductase (porA, porB, porG, and porD), glycerol-3-phosphate dehydrogenase (glpC), and proline dehydrogenase (putA).
No genes encoding a succinate dehydrogenase homologue were identified, although such an activity has been observed in extracts from several H. pylori strains, including J99 (34, 59, 120). This apparent discrepancy can be explained by the observation that fumarate reductase can convert succinate to fumarate in vitro. The importance of fumarate reductase in respiration depends on environmental conditions. Under microaerophilic conditions, fumarate reductase (46) is not essential in H. pylori, which explains why high concentrations of fumarate reductase-specific antimicrobials are required to inhibit the growth of and kill H. pylori in vitro (105). Under these conditions, oxygenic respiration may be used by the bacterium and hence reduces the importance of fumarate reductase in metabolism. In the absence of oxygen, this enzyme may be essential. In the presence of oxygen and fumarate, H. pylori, like members of the related genus Wolinella, may prefer fumarate as a terminal electron acceptor over performing oxygenic respiration (61). The role fumarate reductase plays in respiration cannot be assessed until the microenvironment of H. pylori is better defined.
Quinones and cytochromes.
Analysis of the genomic sequence provides no clear indication to the composition of the H. pylori quinone pool. No homologues to genes involved in the biosynthesis of menaquinones, elements of anaerobic respiration, were identified despite the reported presence of menaquinone-6 and menaquinone-1 in the cell membranes of H. pylori (52, 94). The absence of identifiable homologues implies that H. pylori obtains menaquinones either by synthesis with genes that have yet to be identified or by uptake from its environment. H. pylori does contain homologues for three ubiquinone-biosynthetic enzymes (UbiA, UbiD, and UbiE), but no significant homologues for UbiB, UbiC, UbiF, UbiG, or UbiH were identified. UbiB, UbiF, and UbiH are oxygen-utilizing enzymes, and their apparent absence in H. pylori might be the result of the microaerophilic metabolism of the bacterium.
The biosynthesis of cytochromes by H. pylori has been reviewed in detail recently (51, 125). H. pylori uses a type II system for such biosynthesis, which is similar to that in many gram-positive bacteria and some members of the β-proteobacteria. Both strains possess homologues to all of the components needed for this biosynthetic system.
Terminal electron acceptors.
There appear to be three putative terminal electron acceptor systems in both sequenced strains of H. pylori, i.e., fumarate reductase, N-oxide reductase, and cytochrome c oxidase, suggesting that H. pylori may be able to use fumarate, N-oxides (i.e., dimethyl sulfoxide and trimethylamine-N-oxide), or oxygen as electron sinks. Whereas the presence of a cytochrome c oxidase in H. pylori would allow aerobic respiration, the presence of both fumarate reductase and an N-oxide reductase suggests that the bacterium may respire anaerobically as well. The terminal oxidase complex is similar to cbb3-type oxidase complexes and is encoded by a gene cluster composed of homologues to genes encoding the Rhizobium FixN, FixO, FixP, and FixQ subunits. The arrangement of these genes is identical to that found in Rhizobium spp. (130), and the presence of such a terminal oxidase in H. pylori is consistent with previous findings (116).
ATP-Proton Motive Force Conversion
The bacterial ATP synthase, a multisubunit enzyme, is composed of the F0 complex, which consists of three subunits that form a proton channel, and the F1 complex, which consists of five subunits that constitute the catalytic site for ATP synthesis. In E. coli, all eight subunits are encoded within the atp operon (76). All five subunits of the F1 complex and the F0 b subunit are contiguous on the H. pylori chromosome. The remaining two subunits of the F0 complex are encoded by genes present in other chromosomal regions. H. pylori has an additional subunit (JHP1065/HP1137), which is homologous to the ATPase b′, a diverged and duplicated form of the b subunit found among plants and photosynthetic bacteria. The gene encoding this homologue is located at one end of the ATP synthase gene cluster. Functionally, the H. pylori ATPase is similar to other bacterial ATPases in that it uses the proton motive force generated by the electron transport chain to synthesize ATP (97).
Detoxification
Organisms that come in contact with oxygen, like the microaerophilic H. pylori, must be able to protect themselves from the toxic products of oxygen metabolism, such as superoxide and hydrogen peroxide. Both sequenced strains of H. pylori possess a superoxide dismutase and a catalase, consistent with previous biochemical findings (57, 122, 140). Thus, H. pylori is a microaerophile not because of an inability to neutralize the toxic products of oxygen metabolism but, more probably, as a consequence of other metabolic limitations. H. pylori has two genes encoding peroxidases (JHP991/HP0390 and JHP1471/HP1563), one of which is located adjacent to the superoxide dismutase gene. The ability to isolate catalase-negative mutants of H. pylori (122, 156) suggests that at least one of the peroxidases can function as a catalase.
In summary, it would appear that H. pylori can use amino acids or simple carbohydrates as a major sources of energy. The bacterium is restricted with respect to pyruvate metabolism. Further, H. pylori possesses an electron transport chain that can use oxygen as a terminal electron acceptor, but homologues to fumarate reductase and N-oxide reductase suggest that the bacterium is capable of at least limited anaerobic metabolism.
MACROMOLECULE BIOSYNTHESIS AND MODIFICATION
DNA Replication, Recombination, and Restriction-Modification
H. pylori contains genes encoding homologues to the DnaE, DnaN, DnaX, DnaQ, and HolB subunits of the DNA polymerase III holoenzyme, which is responsible for DNA replication. While the H. pylori holoenzyme contains fewer than the 10 subunits that comprise DNA polymerase III in E. coli, the total number of subunits is consistent with that found in other bacterial genomes (24, 31, 43, 81, 84, 139). Indeed, the only common subunits among these different species are DnaE, DnaN, and DnaX. H. pylori contains homologues to all genes encoding enzymes involved in initiation and DNA chain elongation, except dnaC, which is also absent in several other bacterial genomes (24, 31, 43, 81, 84, 139).
H. pylori possesses homologues to several nucleases, including UvrABC endonuclease, UvrD, ExoA, and RecJ. Even though H. pylori contains homologues to xseA and mutS, it lacks recognizable homologues encoding the other subunits of exonuclease VII (XseB) and the MutHLS endonuclease repair complex, respectively. Recombinational repair in H. pylori appears to occur in a RecBC-independent manner (92), since the RecBCD exonuclease V is absent and a RecR homologue was identified. Other homologues identified in the recombination system include RecA and RecN, while no homologue to recE encoded exonuclease VIII was found. In addition, homologues of ruvABC and recG, whose products are involved in the branch migration and resolution of Holliday structures, were identified in H. pylori. Both H. pylori genomes also possess several homologues to the DNA topoisomerase I gene (topA), some of which are strain specific (6). There are genes encoding several other ATP-dependent helicases in H. pylori, including a homologue for pcrA and rep. Both sequenced H. pylori strains have numerous genes with similarity to DNA restriction and modification enzymes, many of which are strain specific. This finding suggests that H. pylori strains have their own unique complement of these genes (2, 6). While many of the products of these genes can be classified as type I, II, or III restriction or modification enzymes, their exact specificity remains to be determined.
Both H. pylori genomes appear to have three type I systems. In all three systems, the modification (HsdM) and restriction (HsdR) subunits are highly homologous between the strains but the specificity subunits (HsdS) have limited identity. Domains of HsdS proteins can be shuffled to produce new specificity, and it is this modular nature which allows the type I systems to evolve rapidly to a new DNA specificity.
Some of the type II systems in H. pylori 26695 and J99 appear to be functionally equivalent and may possess the same DNA specificity, while others have identity within the methyltransferase enzyme but differences in the restriction enzyme. There are nine type II modification methylases common to both genomes (6). In addition, H. pylori J99 possesses two unique type II restriction-modification systems. In H. pylori 26695 and several other strains, the M.HpyI gene, encoding a type II modification enzyme (159), flanks iceA, which encodes a putative type II restriction endonuclease (149). The absence of iceA in H. pylori J99, a recent clinical isolate from a patient diagnosed with a duodenal ulcer, suggests that iceA is not required for gastrointestinal disease and may not represent an informative epidemiological marker of pathogenicity and virulence, as previously hypothesized (15, 152).
Both H. pylori 26695 and J99 contain two type III restriction-modification systems, one of which is strain specific. Whereas in the other system (JHP1410/1411, HP1521/1522) the restriction gene product is 93% identical in the two strains, the modification gene product is conserved only at the N and C termini. This difference may result in unique specificity for each modification enzyme. The modification gene appears to be regulated by a slipped-strand repair mechanism and is frameshifted in both J99 and 26695 (6).
Transcription and Translation
Analysis of the H. pylori genomes identified only three sigma factors (RpoD, RpoN, and FliA). The presence of these three sigma factors had been suggested by putative promotors found upstream of H. pylori genes (83, 86, 89, 122, 141, 147). No homologue to the stationary-phase sigma factor (RpoS) or the heat shock-specific sigma factor (RpoH) was identified, implying that H. pylori responds to stress in different fashion from that described in other bacteria. Both sequenced H. pylori strains contain a fusion of rpoB and rpoC, which encode the β and β′ subunits of RNA polymerase (149, 160). H. pylori contains homologues to three termination factors (nusA, nusB, and rho) and lacks identifiable homologues to the tRNA maturation genes rnd, rph, and rnpB. The lack of identifiable transcriptional termination stem-loop structures suggests that in H. pylori termination is largely Rho dependent (6).
All the aminoacyl-tRNA synthetases are present in H. pylori, except glutaminyl- and asparaginyl-tRNA synthetases. Two copies of the gene encoding glutamyl-tRNA synthetase (gltX) are present in both H. pylori strains. It had been suggested that one of these copies may function as a glutaminyl-tRNA synthetase (149). However, the presence of homologues to the gatA, gatB, and gatC genes, which have been demonstrated to replace glutaminyl-tRNA synthetase activity in Bacillus subtilis (29, 146), makes it more likely that glutaminyl-tRNA synthetase activity in H. pylori is encoded by the gatABC homologues rather than gltX (6). Thus, the role of the second glutamyl-tRNA synthetase in H. pylori is unclear. A similar transamidation reaction, encoded by these three homologues or other genes, may function in H. pylori to synthesize asparaginyl-tRNA from aspartyl-tRNA, explaining the ability of H. pylori to grow without added asparagine despite its apparent inability to synthesize this amino acid.
Fatty Acid and Phospholipid
Identifiable homologues were found to many of the genes required for initiation and elongation of fatty acid biosynthesis. Both H. pylori strains contain the characteristically small acyl carrier protein (ACP) (78 amino acids) (JHP744/HP0808). H. pylori 26695 but not J99 possesses a second, significantly larger (153-amino-acid) homologue with an extended N-terminal domain (HP962). Genomic analysis indicates that H. pylori has a homologue to cyclopropane fatty acid synthase (JHP969/HP0416), consistent with the experimental evidence that H. pylori has a preponderance of C19:0 cyclopropane chains (73). No homologue to β-hydroxydecanoyl-ACP dehydrase, which catalyzes the formation of cis-3-decenoyl-ACP, an important intermediate in the biosynthesis of unsaturated fatty acids in E. coli, was found. The absence of this homologue is surprising, since unsaturated fatty acids are present in H. pylori (73).
The phospholipid composition of H. pylori consists mainly of phosphatidylethanolamine, cardiolipin, and phosphotidylglycerol, with smaller quantities of phosphatidylserine and phosphatidylcholine (73). The genome appears to encode all of the proteins necessary for the synthesis of these phospholipids except for cardiolipin synthase (cls), which catalyzes the final step of cardiolipin synthesis in E. coli. The H. pylori genome encodes at least two of the three enzymes necessary for phosphatidic acid synthesis (JHP895/HP0961 and JHP1267/HP1348). A homologue to the glycerol-3-phosphate acyltransferase was not identified in the H. pylori genome.
A characteristic feature of the lipid profile of H. pylori is the presence of cholesterol glucosides, which account for about 25% of the total lipid of the bacterium (56, 58). H. pylori does not appear to encode known enzymes for the synthesis of cholesterol and presumably scavenges this molecule from the environment. Little is known about glucoside-modified cholesterol synthesis, but the enzymes responsible are expected to be found among the species-specific genes with unknown function.
Peptidoglycan
The cytoplasmic synthesis of UDP-activated precursors of bacterial peptidoglycan assembly is well understood. The H. pylori genome encodes homologues to all of the enzymes in this pathway, beginning with the synthesis of UDP-N-acetylmuramic acid and ending with UDP-disaccharide pentapeptide linked to an undecaprenol lipid carrier (Fig. 4).
FIG. 4.
Peptidoglycan synthesis and recycling. GlcNAc, N-acetylglucosamine; MurNAc, N-acetylmuramic acid; Ala, alanine; Glu, glutamate, Dap, diaminopimelic acid. Adapted from reference 62.
After transport through the cytoplasmic membrane, peptidoglycan precursors are incorporated into the existing peptidoglycan layer by the penicillin-binding proteins (PBPs) (50). The genome of H. pylori encodes three proteins which show homology to PBPs, in agreement with a published gel electrophoresis study (68). The precise metabolic function of each PBP is uncertain. However, one of the PBPs (JHP544/HP597) has a transglycosylase motif. Since no additional genes with a similar motif were detected, this PBP may be the only enzyme in H. pylori that is involved in lengthening the glycan chain.
From the analysis of the genomes, it is uncertain if peptidoglycan fragments can be recycled by H. pylori. Although this bacterium has homologues to genes encoding a lytic amidase (JHP709/HP0772) and an N-acetylmuranoyl-l-alanine transglycosylase (JHP590/HP0645), it does not appear to have the other genes required for this recycling system (ampG, ampD, mpl) (127).
Outer Membrane
Approximately 4% of the coding capacity of both strains is devoted to outer membrane proteins. This amount is significantly larger than that of any other bacterial genome sequenced to date. The majority of these proteins belong to three paralogous families, the largest having 20 and 21 members in J99 and 26695, respectively (6). Several members of the largest paralogous family are porins (33, 41) or adhesins specific for the Lewis B carbohydrate moiety found on host cells (72). The sequence identity of orthologous members of this large family is high (greater than 95%), suggesting that strain-specific sequence differences play only a limited role in antigenic variation. Five orthologous pairs of this outer membrane protein family, including BabB (Lewis B adhesin), contain dinucleotide (CT) repeats in their signal sequences. Slipped-strand repair has been proposed to regulate the expression of these proteins (6, 149). Significantly, the predicted expression status (in frame or out of frame) of these five pairs is identical despite the presence of a different number of dinucleotide repeats in the two strains in each case (6).
Lipopolysaccharide
H. pylori has homologues to all enzymes required for 2-keto-3-deoxyoctulosonic acid (KDO)-lipid A biosynthesis. Compared to lipid A in E. coli, this moiety in H. pylori is underacylated, has longer fatty acid chains (C16 and C18), and lacks a phosphate group at the 4′-hydroxyl position on the nonreducing glucosamine of the disaccharide (115). Taken together, these findings suggest that in H. pylori lipid A is assembled as an acylated and diphosphorylated disaccharide, which is then modified by an unidentified phosphatase and esterase enzyme(s).
The chemical structures of the lipopolysaccharide cores from two H. pylori strains were shown to be identical heptasaccharides (9, 10). Synthesis of such a structure requires several glycosyltransferases. There are seven ORFs encoding these putative glycosyltransferases (JHP147/HP0159, JHP194/HP0208, JHP563/HP0619, JHP620/HP0679, JHP741/HP0805, JHP765/HP0826, and JHP1031/HP1105), which are common to both strains, three strain-specific ORFs in J99 (JHP562, JHP820, and JHP1032), and one strain-specific ORF in 26695 (HP1578). However, none of these ORFs can be assigned a substrate specificity. Whether the presence of these strain-specific glycosyltransferases in J99 and 26695 results in a different lipopolysaccharide core structure remains to be determined.
The O chain from lipopolysaccharides of H. pylori is composed of Lewis acids [Lex and Ley; mono- and difucosylated repeating disaccharides of β-(1,4)-linked galactose and N-acetylglucosamine, respectively] (9, 10). These carbohydrate moieties, which are identical to those found on host tissues, have been implicated in colonization and persistence and may also play a role in autoimmunity (7, 8). The biosynthetic pathway of these O chains has not been determined. One enzyme, α-(1,3)-fucosyltransferase (JHP596/HP0651), is to be involved in this pathway (21, 96). Each genome has two α-(1,3)-fucosyltransferases which differ in the number of a 7-amino-acid sequence repeat (YDDLRVN). Regulation of these genes appears to occur through slipped-strand repair at two distinct polynucleotide repeats (6).
Flagella
Flagellar biosynthesis in gram-negative bacteria has been extensively studied. The assembly of a functional flagellum requires numerous proteins and is a highly regulated process. Homologues to all of the required genes involved in flagellar biosynthesis were identified in both sequenced strains. In other gram-negative bacteria, including C. jejuni, inactivation of the biosynthetic pathway disrupts the expression of the system as a whole. However, in H. pylori, inactivation of the hook protein does not result in suppression of flagellin expression (124), indicating that flagellar biosynthesis is not as highly regulated as in other bacteria. This difference can be explained by the absence of FlgM in H. pylori, a protein that in other bacteria controls feedback regulation of the flagellar biosynthetic cascade.
The flagellar filament of H. pylori is composed of two flagellin subunits, FlaA and FlaB, and genes encoding both are present. Despite reports suggesting that the genes for these flagellin subunits exhibit strain variation (66, 123), in strains J99 and 26695 the FlaA protein sequences are identical and the FlaB protein sequences differ by only a single amino acid. In addition, the protein sequences reported for FlaA and FlaB from another strain are nearly identical to those from strains J99 and 26695 (89).
The flagellar filament of H. pylori is encased within a sheath that is continuous with the outer membrane (90, 91, 128). The sheath, whose composition has yet to be defined, probably protects the polymeric filament from dissociation in the low pH of the stomach. Likewise, other polymeric structures, such as fimbriae, would also be subject to dissociation at low pH, thus explaining why adhesion to the host epithelium by H. pylori appears to be mediated by integral outer membrane proteins (71, 72).
In summary, H. pylori has the necessary homologues for DNA, RNA, and protein synthesis.
CELLULAR PROCESSES
Protein Secretion
H. pylori has homologues to two leader peptidases, enzymes required for protein secretion. The type I leader peptidase, LepB, is responsible for cleavage of the signal sequence from most periplasmic and outer membrane proteins. The lspA (previously called ureD) gene product is the leader peptidase responsible for processing prelipoproteins.
The secretion of proteins through the cytoplasmic membrane utilizes specific machinery which consists of several sec gene products. H. pylori contains all the known Sec proteins except SecB and SecG. Mutation of secB, which encodes a chaperone, affects the secretion of only a limited number of proteins in E. coli, and other preproteins may utilize alternative chaperones. Indeed, it has been demonstrated that groESL mutants in E. coli have significant effects on the secretion of some proteins (85). Thus, it is likely that in H. pylori, other cytoplasmic chaperones are used to usher proteins, in conjunction with SecA, to the cytoplasmic membrane in preparation for secretion. SecE is proposed to be integral to the translocation machinery (121). Initial analysis of both genomes did not identify a SecE orthology. However, a region between JHP1126/HP1203 and JHP1127/HP1204 contains a 59-amino-acid ORF that has similarity to the functionally important region of E. coli SecE.
Cag Pathogenicity Island
H. pylori strains associated with clinically severe gastric disease (peptic ulcers) more commonly possess an approximately 40-kb pathogenicity island containing cytotoxin-associated genes (Cag pathogenicity island [cagPAI]) than do strains isolated from patients with uncomplicated gastritis (26). Type I strains have been defined as H. pylori isolates which have the entire cagPAI, express the cytotoxicity-associated immunodominant antigen (CagA) and an active vacuolating cytotoxin (VacA), and induce interleukin-8 (IL-8) secretion by gastric epithelial cells. In addition to cagA, it has been demonstrated by mutational analysis that several of the genes in the cagPAI are required for wild-type induction of IL-8 secretion by gastric epithelial cells. In contrast, Type II strains do not express CagA, have no vacuolating activity even though a truncated VacA may be produced, and do not induce IL-8 secretion at levels comparable to that induced by type I strains.
The initial cagPAI sequence was produced in two parts by two different laboratories, each of which sequenced clones from the same ordered cosmid library that had been constructed from H. pylori NCTC 11638 (2, 19). These groups showed that the cagPAI was separated into the cagI and cagII segments by an intervening stretch of DNA that was itself bordered on each side by a newly identified insertion sequence (IS605) element. In contrast, the cagPAI in H. pylori J99 and 26695, both type I strains, consists of the cagI and cagII segments fused as a single unit without the stretch of intervening DNA flanked on each side by IS605. In all three strains, the cagI and cagII segments are essentially the same and the cagPAI is found in the same relative location.
Many of the genes in the cagPAI are required for specific host cell responses to infection by H. pylori, including induction of (i) IL-8 secretion by gastric epithelial cells (2, 19, 137), (ii) tyrosine phosphorylation in host proteins (137), and (iii) cytoskeletal rearrangements during actin pedestal formation at the host cell surface (137). The manner in which cagPAI genes cause these host cell responses is unknown. However, five ORFs show significant sequence similarity to genes encoding the VirB protein family, which is responsible for DNA transfer in Agrobacterium tumefaciens (type IV secretion system) (23). These genes also have sequence similarity to genes involved in conjugative transfer of plasmids in E. coli and protein export in Bordetella pertussis (155). It is unknown how these components of conjugative and protein transport systems function in H. pylori. It has been suggested that the cagPAI encodes a type of contact-mediated secretion apparatus analogous to the type III secretion systems identified and characterized in several enteric pathogens such as Yersinia, Salmonella, Shigella, and pathogenic E. coli (88). It is therefore likely that the virB homologues of the cagPAI region encode components of the secretion apparatus which may deliver effector molecules (DNA or protein) directly to the host cell to elicit the responses mentioned above.
Insertion Elements
The IS element IS605 contains genes encoding two previously identified transposases flanked by a short nucleotide sequence with dyadic symmetry and a common central core sequence (19). IS605 transposes as a single unit in E. coli, suggesting that it could also be functional in H. pylori (79). One of the transposase genes (tnpA) is related to IS200 found in gram-negative bacteria, and the other (tnpB) is related to IS1341 found in a gram-positive thermophylic bacterium. It is unusual to find an insertion element with transposases from apparently two different origins. The IS605 element was first described within the cagPAI of NCTC 11638. Strain 26695 has five full copies of IS605 and one copy of a related insertion element, IS606, none of which is located within the cagPAI. Strain J99 has no complete copies of IS605 but has one copy of IS606 on its chromosome. The short flanking sequence of IS605, without the transposases (is605), is present on both ends of the cagPAI in strain NCTC 11638. These sequences flanking cagPAI are thought to be remnants of a recent transposition or of another type of recombinational event. The dyadic repeats of IS605 and IS606 are also found within the J99 and 26695 genomes, at both common and distinct locations. IS605 dyadic repeats are coincident, in one or the other genomes, with several of the major organizational differences between the two sequenced strains. H. pylori plasmid pHPM186 (GenBank accession no. AF077006) contains an IS605 element which is adjacent to three genes that flank an IS605 element in the plasticity zone of strain 26695 (6). This finding suggests that plasmid integration plays a role in generating the limited genomic diversity in H. pylori. Thus, insertion elements, such as IS605, may have been involved in the acquisition of cagPAI and the plasticity zone by H. pylori via horizontal transfer, as has also been postulated for other pathogenicity islands found in a wide range of pathogenic bacteria.
Transformation
Many H. pylori strains are naturally competent for DNA transformation, and the efficiency of this process varies from strain to strain. Besides the conjugative DNA transfer/protein export homologues (VirB proteins) encoded by the cagPAI, both strains contain additional members of this family, some of which are strain specific. Recently, some members of the VirB family have been shown to play a role in DNA transformation in H. pylori (60). The existence of strain-specific VirB family homologues may explain the variation in DNA transformation efficiency seen between strains. One additional transformation-associated gene (comEC), which is required for uptake of DNA into B. subtilis, was also identified.
Chemotaxis
Chemotaxis, the sensory adaptation mechanism by which motile bacteria recognize and react to environmental conditions, has been found in H. pylori (110). Three homologues of the chemotaxis pathway in E. coli (CheW, CheA, and CheY), as well as four methyl-accepting chemotaxis proteins (MCPs), which mediate specificity for ligands, were identified in both sequenced strains of H. pylori. Proteins with similarity to MCPs are not necessarily involved in flagellar chemotaxis (5, 30). Neither strain contains identifiable homologues to CheR or CheB, enzymes which, respectively, add methyl groups to or remove methyl groups from the MCPs, precisely modulating the chemotactic response. By contrast, C. jejuni does possess homologues to both CheR and CheB. Therefore, the chemotaxis observed in H. pylori may occur by a CheB- and CheR-independent mechanism, similar to that seen in CheB CheR mutants of E. coli (145). The apparent inability of H. pylori to precisely modulate chemotaxis may reflect its limited but unique gastric niche.
Both H. pylori strains have three homologues to CheV, another chemotaxis protein, which has an N-terminal domain similar to CheW and a C-terminal response regulator domain similar to CheY (44). The CheW protein and the N-terminal domain of CheV are both capable of modulating the CheA-MCP interaction (134). The three CheV orthologous pairs have greater than 97% identity, whereas none of the paralogues have more than 40% identity at the amino acid level, suggesting that each orthologous pair has functional similarity and that each paralogue has a different specific function.
Cell Division and Morphology
Several genes implicated in bacterial cell division were found in both sequenced strains of H. pylori. Among these are ftsZ and ftsA, which are adjacent to each other, as is generally observed (154). H. pylori does not possess an unambiguous homologue to FtsW, but it does possess two homologues to RodA, a protein with significant sequence similarity to FtsW (69). It is possible that one of the RodA homologues actually functions as FtsW. In E. coli, at least one of the peptidoglycan-synthesizing enzymes is specifically involved in cell division (142). H. pylori has three homologues to peptidoglycan-synthesizing enzymes, one of which could act specifically during cell division. Two homologues of the metalloprotease FtsH, phenotypically associated with cell division, were identified. Interestingly, mutagenesis studies have shown that one of the FtsH homologues (JHP356/HP1069) is essential for growth in vitro (48). This suggests that the second homologue (JHP271/HP0286) is unable to functionally replace the first homologue and probably has a different function. In addition, homologues of ftsK and fic were identified.
No homologues to ZipA, which is believed to initiate formation of the FtsZ ring (53), SulA, which is thought to inhibit the formation of the FtsZ ring (14), FtsQ, FtsN, or FtsL were identified. Also, no homologue to MinC, a cell division inhibitor, was found, whereas homologues to the activator (MinD) and cofactor (MinE) of MinC were identified.
Two homologues of cell morphology-determining proteins, RodA and MreB, were found. RodA is required for the catalytic activity of a PBP during elongation in E. coli (74), whereas the function of MreB is unknown.
Virulence Factors
A number of H. pylori proteins have been implicated in pathogenesis (for reviews, see references 36 and 95). All of the reported genes encoding potential virulence factors in H. pylori were identified in both strains, with the exception of iceA, which is missing from strain J99 (see “DNA replication, recombination, and restriction-modification” above).
Bacterial motility has been suggested to be required for colonization of the gastric mucosa, since H. pylori mutants unable to synthesize either of the flagellar subunits (FlaA or FlaB) cannot colonize gnotobiotic piglets (38, 40). Whether motility is needed for the persistence of an infection is not known, but owing to the rapid gastric epithelium and mucous turnover, it would probably be required.
The cytotoxin VacA induces vacuole formation in cultured epithelial cells and may be an important component in the induction of gastric cell lesions by H. pylori (27, 49). Different alleles of the vacA gene have been reported (11, 28). Strains J99 and 26695 possess different vacA alleles (s1b/ml and s1a/ml, respectively) and were isolated from patients with H. pylori-related disease of different severity. Epidemiological studies with humans have correlated the level of expression of the vacA gene with disease outcome (42), and particular vacA genotypes are associated with more severe disease symptoms (109, 144, 151). Interestingly, in gnotobiotic piglets, a mutant with a knockout mutation of vacA had no discernible effect on colonization, epithelial vacuolation, or gastritis, suggesting that VacA is not a virulence factor in this animal model (37).
The urease enzyme of H. pylori has been extensively studied (for reviews, see references 111 to 113) and has been shown to be a colonization factor (39, 78, 150). Urease is found in the cytoplasm as well as on the surface of H. pylori. The mechanism by which this enzyme is translocated is controversial (129, 153).
Other components which have been implicated in virulence, include the cagPAI, lipopolysaccharide, outer membrane proteins, and a number of enzymes, such as phospholipases, catalase, superoxide dismutase, and a mucinase homologue (36). The involvement of these components in pathogenesis remains to be elucidated. Homologues to the genes encoding all these products were present in both strains.
In summary, H. pylori possess homologues to the genes necessary for DNA replication, transcription, translation, and cell division, as well for as the synthesis of other important cellular macromolecules such as lipids and peptidoglycan. Each strain appears to possess a number of unique restriction enzymes that would result in selectivity to transformation with foreign DNA. Further features of this bacterium are the large number of related outer membrane proteins and the presence of a putative pathogenicity island.
CONCLUSION
The genomic analysis suggests that both strains have essentially identical metabolic potential. The strain-specific genes that encode proteins with an assigned function are predicted to have little impact on the physiology of H. pylori. For the most part, the reported experimental data for biochemical activities present or absent in H. pylori are in agreement with the predicted metabolic capabilities. Our genomic analysis reveals nutritional requirements which would restrict the environments in which H. pylori can survive and, in part, explains its limited niche. H. pylori has broad catabolic capabilities. While it can perform oxygenic respiration, anaerobic respiration, and fermentation, H. pylori is limited in what it can use as a carbon source. Carbohydrate utilization is restricted to glucose, via the Entner-Doudoroff pathway, and to sugars with shorter carbon backbones than C6, for which transport systems are present. In addition, H. pylori possesses numerous transporter systems for the uptake of amino acids, which must be available in the gastric environment. Other than carbohydrates, H. pylori apparently uses amino acids as an important source of carbon which would yield less energy than using hexoses and could result in slow growth as occurs in vitro. Further, the deamination of amino acids, together with the action of urease, requires a mechanism to deal with increased levels of ammonia.
Colonization by H. pylori involves an interaction between the outer membrane of the bacterium and the gastric epithelium of the host. The outer membrane composition of H. pylori is unique in its protein content and lipopolysaccharide structure, which are consistent with the persistence of H. pylori in a restricted niche. Compared to other bacteria, H. pylori devotes a significantly higher percentage of its coding capacity to that of outer membrane proteins, further emphasizing the importance of these proteins. A number of genes involved in determining the composition of the outer membrane are differentially regulated by slipped-strand repair (6). This differential regulation and strain-specific outer membrane-related genes may play a role in the severity of H. pylori-related disease and the ability of H. pylori to persist chronically in its host. The other strain-specific genes may also play a role in these aspects of pathogenesis. None of these strain-specific genes with an assigned function are predicted to have a significant impact on the pathophysiological capabilities of H. pylori, although those with an unassigned function may be important. However, there is evidence that host factors are involved in H. pylori-related diseases (35), and our genomic analyses suggest that these host factors may play a more significant role than was previously appreciated.
The analysis presented here, like previous analyses of other sequenced bacterial genomes, has found several biochemical pathways for which we were unable to identify all of the genes which should be present for that pathway to be functional. Some of these genes may not be present in the genome. In other cases, the biochemical activity may be present within the organism but the gene responsible for this activity may be unidentifiable by current in silico techniques. The apparent incompleteness of the same metabolic pathways in two unrelated strains may suggest that these pathways are functional in H. pylori and that the unidentified genes are different from previously described orthologues. These genes would be among the 40% of the genome to which no function has been assigned.
For the first time, the genomes of two strains from the same bacterial species have been compared (6). This publication provided an opportunity to begin defining the physiology of H. pylori, a globally important pathogen. Future genomic comparisons of multiple strains, carefully correlated with epidemiological data, will identify the minimal genomic complement of this species and the genes required for virulence. Such an approach will be applied to other pathogenic bacteria, and the genes identified from these studies will become candidates for therapeutic intervention.
REFERENCES
- 1.Abouhamad W N, Manson M, Gibson M M, Higgins C F. Peptide transport and chemotaxis in Escherichia coli and Salmonella typhimurium: characterization of the dipeptide permease (Dpp) and the dipeptide-binding protein. Mol Microbiol. 1991;5:1035–1047. doi: 10.1111/j.1365-2958.1991.tb01876.x. [DOI] [PubMed] [Google Scholar]
- 2.Akopyants N, Clifton S, Kersulyte D, Crabtree J, Youree B, Reece C, Bukanov N, Drazek E, Roe B, Berg D. Analyses of the cag pathogenicity island of Helicobacter pylori. Mol Microbiol. 1998;28:37–53. doi: 10.1046/j.1365-2958.1998.00770.x. [DOI] [PubMed] [Google Scholar]
- 3.Akopyanz N, Bukanov N, Westblom T, Berg D. PCR-based RFLP analysis of DNA sequence diversity in the gastric pathogen Helicobacter pylori. Nucleic Acids Res. 1992;20:6221–6225. doi: 10.1093/nar/20.23.6221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Akopyanz N, Bukanov N, Westblom T, Kresovich S, Berg D. DNA diversity among clinical isolates of Helicobacter pylori detected by PCR-based RAPD fingerprinting. Nucleic Acids Res. 1992;20:5137–5142. doi: 10.1093/nar/20.19.5137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Alm R, Manning P. Characterization of the hlyB gene and its role in the production of the El Tor haemolysin of Vibrio cholerae O1. Mol Microbiol. 1990;4:413–425. doi: 10.1111/j.1365-2958.1990.tb00608.x. [DOI] [PubMed] [Google Scholar]
- 6.Alm R A, Lee L-S, Moir D T, King B L, Brown E D, Doig P C, Smith D R, Noonan B, Guild B C, deJonge B L, Carmel G, Tummino P J, Caruso A, Uria-Nickelsen M, Mills D M, Ives C, Gibson R, Merberg D, Mills S D, Jiang Q, Taylor D E, Vovis G F, Trust T J. Genomic sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature. 1999;397:176–180. doi: 10.1038/16495. [DOI] [PubMed] [Google Scholar]
- 7.Appelmelk B, Negrini R, Moran A, Kuipers E. Molecular mimicry between Helicobacter pylori and the host. Trends Microbiol. 1997;5:70–73. doi: 10.1016/S0966-842X(96)10084-6. [DOI] [PubMed] [Google Scholar]
- 8.Appelmelk B, Simoons-Smit I, Negrini R, Moran A, Aspinall G, Forte J, deVries T, Quan H, Verboom T, Maaskant J, Ghiara P, Kuipers E, Bloemena E, Tadema T, Townsend R, Tyagarajan K, Jr J C, Monteiro M, Savio A, deGraaff J. Potential role of molecular mimicry between Helicobacter pylori lipopolysaccharide and host Lewis blood group antigens in autoimmunity. Infect Immun. 1996;64:2031–2040. doi: 10.1128/iai.64.6.2031-2040.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Aspinall G O, Monteiro M A. Lipopolysaccharides of Helicobacter pylori strains P466 and Mo19—structures of the O antigen and core oligosaccharide regions. Biochemistry. 1996;35:2498–2504. doi: 10.1021/bi951853k. [DOI] [PubMed] [Google Scholar]
- 10.Aspinall G O, Monteiro M A, Pang H, Walsh E J, Moran A P. Lipopolysaccharide of the Helicobacter pylori type strain NCTC 11637 (Atcc 43504)—structure of the O antigen chain and core oligosaccharide regions. Biochemistry. 1996;35:2489–2497. doi: 10.1021/bi951852s. [DOI] [PubMed] [Google Scholar]
- 11.Atherton J, Cao P, Jr R P, Tummuru M, Blaser M, Cover T. Mosaicism in vacuolating cytotoxin alleles of Helicobacter pylori. J Biol Chem. 1995;270:17771–17777. doi: 10.1074/jbc.270.30.17771. [DOI] [PubMed] [Google Scholar]
- 12.Berg D, Gilman R, Lelwala-Guruge J, Srivastava K, Valdez Y, Watanabe J, Miyagi J, Akopyants N, Ramirez-Ramos A, Yoshiwara T, Recavarren S, Leon-Barua R. Helicobacter pylori populations in Peruvian patients. Clin Infect Dis. 1997;25:996–1002. doi: 10.1086/516081. [DOI] [PubMed] [Google Scholar]
- 13.Berg D E, Hoffman P S, Appelmelk B J, Kusters J G. The Helicobacter pylori genome sequence: genetic factors for long life in the gastric mucosa. Trends Microbiol. 1997;5:468–474. doi: 10.1016/s0966-842x(97)01164-5. [DOI] [PubMed] [Google Scholar]
- 14.Bi E, Lutkenhaus J. Cell division inhibitors SulA and MinCD prevent formation of the FtsZ ring. J Bacteriol. 1993;175:1118–1125. doi: 10.1128/jb.175.4.1118-1125.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Blaser M. Heterogeneity of Helicobacter pylori. Eur J Gastroenterol Hepatol. 1997;9:S3–S6. doi: 10.1007/978-94-009-1792-7_3. [DOI] [PubMed] [Google Scholar]
- 16.Blaser M. Not all Helicobacter pylori strains are created equal: should all be eliminated? Lancet. 1997;349:1020–1022. doi: 10.1016/S0140-6736(96)09133-7. [DOI] [PubMed] [Google Scholar]
- 17.Bode G, Mauch F, Ditschuneit H, Malfertheiner P. Identification of structures containing polyphosphate in Helicobacter pylori. J Gen Microbiol. 1993;139:3029–3033. doi: 10.1099/00221287-139-12-3029. [DOI] [PubMed] [Google Scholar]
- 18.Brenchley J E, Magasanik B. Mutants of Klebsiella aerogenes lacking glutamate dehydrogenase. J Bacteriol. 1974;117:544–550. doi: 10.1128/jb.117.2.544-550.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Censini S, Lange C, Xiang Z, Crabtree J, Ghiara P, Borodovsky M, Rappuoli R, Covacci A. cag, a pathogenicity island of Helicobacter pylori, encodes type I-specific and disease-associated virulence factors. Proc Natl Acad Sci USA. 1996;93:14648–14653. doi: 10.1073/pnas.93.25.14648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Chalk P A, Roberts A D, Blows W M. Metabolism of pyruvate and glucose by intact cells of Helicobacter pylori studied by 13C NMR spectroscopy. Microbiology. 1994;140:2085–2092. doi: 10.1099/13500872-140-8-2085. [DOI] [PubMed] [Google Scholar]
- 21.Chan G Z, Palcic M M, Taylor D E. Cloning and heterologous expression of an α(1,3)-fucosyltransferase from the gastric pathogen Helicobacter pylori. J Biol Chem. 1997;272:21357–21363. doi: 10.1074/jbc.272.34.21357. [DOI] [PubMed] [Google Scholar]
- 22.Chang H T, Marcelli S W, Davison A A, Chalk P A, Poole R K, Miles R J. Kinetics of substrate oxidation by whole cells and cell membranes of Helicobacter pylori. FEMS Microbiol Lett. 1995;129:33–38. doi: 10.1016/0378-1097(95)00130-W. [DOI] [PubMed] [Google Scholar]
- 23.Christie P J. Agrobacterium tumefaciens T-complex transport apparatus: a paradigm for a new family of multifunctional transporters in eubacteria. J Bacteriol. 1997;179:3085–3094. doi: 10.1128/jb.179.10.3085-3094.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Cole S, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Gordon S, Eiglmeier K, Gas S, III C B, Tekaia F, Badcock K, Basham D, Brown D, Chillingworth T, Connor R, Davies R, Devlin K, Feltwell T, Gentles S, Hamlin N, Holroyd S, Hornsby T, Jagels K, Krogh A, McLean J, Moule S, Murphy L, Oliver K, Osborne J, Quail M, Rajandream M-A, Rogers J, Rutter S, Seeger K, Skelton J, Squares R, Squares S, Sulston J, Taylor K, Whitehead S, Barrell B. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature. 1998;393:537–544. doi: 10.1038/31159. [DOI] [PubMed] [Google Scholar]
- 25.Corthesy-Theulaz I E, Bergonzelli G E, Henry H, Bachmann D, Schorderet D F, Blum A L, Ornston L N. Cloning and characterization of Helicobacter pylori succinyl CoA:acetoacetate CoA-transferase, a novel prokaryotic member of the CoA-transferase family. J Biol Chem. 1997;272:25659–25667. doi: 10.1074/jbc.272.41.25659. [DOI] [PubMed] [Google Scholar]
- 26.Covacci A, Falkow S, Berg D, Rappuoli R. Did the inheritance of a pathogenicity island modify the virulence of Helicobacter pylori? Trends Microbiol. 1997;5:205–208. doi: 10.1016/S0966-842X(97)01035-4. [DOI] [PubMed] [Google Scholar]
- 27.Cover T, Blaser M. Helicobacter pylori and gastroduodenal disease. Annu Rev Med. 1992;42:135–145. doi: 10.1146/annurev.me.43.020192.001031. [DOI] [PubMed] [Google Scholar]
- 28.Cover T L, Tummuru M K, Cao P, Thompson S A, Blaser M J. Divergence of genetic sequences for the vacuolating cytotoxin among Helicobacter pylori strains. J Biol Chem. 1994;269:10566–10573. [PubMed] [Google Scholar]
- 29.Curnow A, Hong K, Yuan R, Kim S, Martins O, Winkler W, Henkin T, Soll D. Glu-tRNAGln amidotransferase: a novel heterotrimeric enzyme required for correct decoding of glutamine codons during translation. Proc Natl Acad Sci USA. 1997;94:11819–11826. doi: 10.1073/pnas.94.22.11819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Darzins A. Characterization of a Pseudomonas aeruginosa gene cluster involved in pilus biosynthesis and twitching motility: sequence similarity to the chemotaxis proteins of enterics and the gliding bacterium Myxococcus xanthus. Mol Microbiol. 1994;11:137–153. doi: 10.1111/j.1365-2958.1994.tb00296.x. [DOI] [PubMed] [Google Scholar]
- 31.Deckert G, Warren P V, Gaasterland T, Young W G, Lenox A L, Graham D E, Overbeek R, Snead M A, Keller M, Aujay M, Huber R, Feldman R A, Short J M, Olsen G J, Swanson R V. The complete genome of the hyperthermophilic bacterium Aquifex aeolicus. Nature. 1998;392:353–358. doi: 10.1038/32831. [DOI] [PubMed] [Google Scholar]
- 32.Dendinger S M, Patil L G, Brenchley J E. Salmonella thyphimurium mutants with altered glutamate dehydrogenase and glutamate synthase activities. J Bacteriol. 1980;141:140–148. doi: 10.1128/jb.141.1.190-198.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Doig P, Exner M, Hancock R, Trust T. Isolation and characterization of a conserved porin protein from Helicobacter pylori. J Bacteriol. 1995;177:5447–5452. doi: 10.1128/jb.177.19.5447-5452.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Doig P, Trust T. Identification of surface-exposed outer membrane antigens of Helicobacter pylori. Infect Immun. 1994;62:4526–4533. doi: 10.1128/iai.62.10.4526-4533.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Dubois A, Berg D, Incecik E, Fiala N, Heman-Ackah L, Perez-Perez G, Blaser M. Transient and persistent infection of nonhuman primates with Helicobacter pylori: implications for human disease. Infect Immun. 1996;64:2885–2891. doi: 10.1128/iai.64.8.2885-2891.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Dunn B, Cohen H, Blaser M. Helicobacter pylori. Clin Microbiol Rev. 1997;10:720–741. doi: 10.1128/cmr.10.4.720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Eaton K, Cover T, Tummuru M, Blaser M, Krakowka S. Role of vacuolating cytotoxin in gastritis due to Helicobacter pylori in gnotobiotic piglets. Infect Immun. 1997;65:3462–3464. doi: 10.1128/iai.65.8.3462-3464.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Eaton K, Morgan D, Krakowka S. Motility as a factor in the colonisation of gnotobiotic piglets by Helicobacter pylori. J Med Microbiol. 1992;37:123–127. doi: 10.1099/00222615-37-2-123. [DOI] [PubMed] [Google Scholar]
- 39.Eaton K A, Brooks C L, Morgan D R, Krakowka S. Essential role of urease in pathogenesis of gastritis induced by Helicobacter pylori in gnotobiotic piglets. Infect Immun. 1991;59:2470–2475. doi: 10.1128/iai.59.7.2470-2475.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Eaton K A, Suerbaum S, Josenhans C, Krakowka S. Colonization of gnotobiotic piglets by Helicobacter pylori deficient in two flagellin genes. Infect Immun. 1996;64:2445–2448. doi: 10.1128/iai.64.7.2445-2448.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Exner M, Doig P, Trust T, Hancock R. Isolation and characterization of a family of porin proteins from Helicobacter pylori. Infect Immun. 1995;63:1567–1572. doi: 10.1128/iai.63.4.1567-1572.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Forsyth M H, Atherton J C, Blaser M J, Cover T L. Heterogeneity in levels of vacuolating cytotoxin gene (vacA) transcription among Helicobacter pylori strains. Infect Immun. 1998;66:3088–3094. doi: 10.1128/iai.66.7.3088-3094.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Fraser C M, Norris S J, Weinstock G M, White O, Sutton G G, Dodson R, Gwinn M, Hickey E K, Clayton R, Ketchum K A, Sodergren E, Hardham J M, McLeod M P, Salzberg S, Peterson J, Khalak H, Richardson D, Howell J K, Chidambaram M, Utterback T, McDonald L, Artiach P, Bowman C, Cotton M D, Venter J C, et al. Complete genome sequence of Treponema pallidum, the syphilis spirochete. Science. 1998;281:375–388. doi: 10.1126/science.281.5375.375. [DOI] [PubMed] [Google Scholar]
- 44.Fredrick K, Helmann J. Dual chemotaxis signaling pathways in Bacillus subtilis: a sigma D-dependent gene encodes a novel protein with both CheW and CheY homologous domains. J Bacteriol. 1994;176:2727–2735. doi: 10.1128/jb.176.9.2727-2735.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Garner R M, Fulkerson J J, Mobley H L. Helicobacter pylori glutamine synthetase lacks features associated with transcriptional and posttranslational regulation. Infect Immun. 1998;66:1839–1847. doi: 10.1128/iai.66.5.1839-1847.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Ge Z, Jiang Q, Kalisiak M S, Taylor D E. Cloning and functional characterization of Helicobacter pylori fumarate reductase operon comprising three structural genes coding for subunits C, A and B. Gene. 1997;204:227–234. doi: 10.1016/s0378-1119(97)00550-7. [DOI] [PubMed] [Google Scholar]
- 47.Ge Z, Taylor D E. Helicobacter pylori genes hpcopA and hpcopP constitute a cop operon involved in copper export. FEMS Microbiol Lett. 1996;145:181–188. doi: 10.1111/j.1574-6968.1996.tb08575.x. [DOI] [PubMed] [Google Scholar]
- 48.Ge Z, Taylor D E. Sequencing, expression, and genetic characterization of the Helicobacter pylori ftsH gene encoding a protein homologous to members of a novel putative ATPase family. J Bacteriol. 1996;178:6151–6157. doi: 10.1128/jb.178.21.6151-6157.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Ghiara P, Marchetti M, Blaser M J, Tummuru M K, Cover T L, Segal E D, Tompkins L S, Rappuoli R. Role of the Helicobacter pylori virulence factors vacuolating cytotoxin, CagA, and urease in a mouse model of disease. Infect Immun. 1995;63:4154–4160. doi: 10.1128/iai.63.10.4154-4160.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Ghuyssen J-M. Serine β-lactamases and penicillin-binding proteins. Annu Rev Microbiol. 1994;45:37–67. doi: 10.1146/annurev.mi.45.100191.000345. [DOI] [PubMed] [Google Scholar]
- 51.Goldman B S, Kranz R G. Evolution and horizontal transfer of an entire biosynthetic pathway for cytochrome C biogenesis—Helicobacter, Deinococcus, Archaea and more. Mol Microbiol. 1998;27:871–873. doi: 10.1046/j.1365-2958.1998.00708.x. [DOI] [PubMed] [Google Scholar]
- 52.Goodwin C S, Collins M D, Blincow E. The absence of thermoplasmaquinones in Campylobacter pyloridis, and its temperature and pH growth range. Microbiol Lett. 1986;32:137–140. [Google Scholar]
- 53.Hale C A, deBoer P A J. Direct binding of FtsZ to ZipA, an essential component of the septal ring structure that mediates cell division in Escherichia coli. Cell. 1997;96:1–20. doi: 10.1016/s0092-8674(00)81838-3. [DOI] [PubMed] [Google Scholar]
- 54.Han J, Yu E, Lee I, Lee Y. Diversity among clinical isolates of Helicobacter pylori in Korea. Mol Cells. 1997;7:544–547. [PubMed] [Google Scholar]
- 55.Hancock R E, Hantke K, Braun V. Iron transport in Escherichia coli K-12. 2,3-Dihydroxybenzoate-promoted iron uptake. Arch Microbiol. 1997;114:231–239. doi: 10.1007/BF00446867. [DOI] [PubMed] [Google Scholar]
- 56.Haque M, Hirai Y, Yokota K, Mori N, Jahan I, Ito H, Hotta H, Yano I, Kanemasa Y, Oguma K. Lipid profile of Helicobacter spp.: presence of cholesterol glucoside as a characteristic feature. J Bacteriol. 1996;178:2065–2070. doi: 10.1128/jb.178.7.2065-2070.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Hazell S L, Evans D J, Jr, Graham D Y. Helicobacter pylori catalase. J Gen Microbiol. 1991;137:57–61. doi: 10.1099/00221287-137-1-57. [DOI] [PubMed] [Google Scholar]
- 58.Hirai Y, Haque M, Yoshida T, Yokota K, Yasuda T, Oguma K. Unique cholesteryl glucosides in Helicobacter pylori: composition and structural analysis. J Bacteriol. 1995;177:5327–5333. doi: 10.1128/jb.177.18.5327-5333.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Hoffman P S, Goodwin A, Johnsen J, Magee K, Veldhuyzen van Zanten S J. Metabolic activities of metronidazole-sensitive and -resistant strains of Helicobacter pylori: repression of pyruvate oxidoreductase and expression of isocitrate lyase activity correlate with resistance. J Bacteriol. 1996;178:4822–4829. doi: 10.1128/jb.178.16.4822-4829.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Hofreuter D, Odenbreit S, Henke G, Haas R. Natural competence for DNA transformation in Helicobacter pylori: identification and genetic characterization of the comB locus. Mol Microbiol. 1998;28:1027–1038. doi: 10.1046/j.1365-2958.1998.00879.x. [DOI] [PubMed] [Google Scholar]
- 61.Holt J G, Krieg N R, Sneath P H A, Staley J T, Williams S T. Bergey’s manual of determinative bacteriology. 9th ed. Baltimore, Md: The Williams & Wilkins Co.; 1994. [Google Scholar]
- 62.Holtje J V. Growth of the stress-bearing and shape-maintaining murine sacculus of Escherichia coli. Microbiol Mol Biol Rev. 1998;62:181–203. doi: 10.1128/mmbr.62.1.181-203.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Hughes N J, Chalk P A, Clayton C L, Kelly D J. Identification of carboxylation enzymes and characterization of a novel four-subunit pyruvate:flavodoxin oxidoreductase from Helicobacter pylori. J Bacteriol. 1995;177:3953–3959. doi: 10.1128/jb.177.14.3953-3959.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hughes N J, Clayton C L, Chalk P A, Kelly D J. Helicobacter pylori porCDAB and oorDABC genes encode distinct pyruvate:flavodoxin and 2-oxoglutarate:acceptor oxidoreductases which mediate electron transport to NADP. J Bacteriol. 1998;180:1119–1128. doi: 10.1128/jb.180.5.1119-1128.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Hunt R. The role of Helicobacter pylori in pathogenesis: the spectrum of clinical outcomes. Scand J Gastroenterol Suppl. 1996;220:3–9. [PubMed] [Google Scholar]
- 66.Hurtado A, Owen R J, Desai M. Flagellin gene profiling of Helicobacter pylori infecting symptomatic and asymptomatic individuals. Res Microbiol. 1994;145:585–594. doi: 10.1016/0923-2508(94)90075-2. [DOI] [PubMed] [Google Scholar]
- 67.Husson M O, Legrand D, Spik G, Leclerc H. Iron acquisition by Helicobacter pylori: importance of human lactoferrin. Infect Immun. 1993;61:2694–2697. doi: 10.1128/iai.61.6.2694-2697.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Ikeda F, Yokota Y, Mine Y, Tatsuta M. Activity of cefixime against Helicobacter pylori and affinities for the penicillin-binding proteins. Antimicrob Agents Chemother. 1990;34:2426–2428. doi: 10.1128/aac.34.12.2426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Ikeda M, Sato T, Wachi M, Jung H K, Ishino F, Kobayashi Y, Matsuhashi M. Structural similarity among Escherichia coli FtsW and RodA proteins and Bacillus subtilis SpoVE protein, which function in cell division, cell elongation, and spore formation, respectively. J Bacteriol. 1989;171:6375–6378. doi: 10.1128/jb.171.11.6375-6378.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Illingworth D S, Walter K S, Griffiths P L, Barclay R. Siderophore production and iron-regulated envelope proteins of Helicobacter pylori. Int J Med Microbiol Virol Parasitol Infect Dis. 1993;280:113–119. [PubMed] [Google Scholar]
- 71.Ilver D, Arnquist A, Frick I-M, Engstrand L, Covacci A, Magnusson K-E, Boren T. The Helicobacter pylori blood group antigen binding adhesin. Gut. 1996;39:A55. . (Abstract.) [Google Scholar]
- 72.Ilver D, Arnqvist A, Ogren J, Frick I, Kersulyte D, Incecik E, Berg D, Covacci A, Engstrand L, Boren T. Helicobacter pylori adhesin binding fucosylated histo-blood group antigens revealed by retagging. Science. 1998;279:373–377. doi: 10.1126/science.279.5349.373. [DOI] [PubMed] [Google Scholar]
- 73.Inamoto Y, Ariyama S, Hamanaka Y, Okita K, Kaneta Y, Nagate T, Kondou I, Takemoto T. Lipid analysis of Helicobacter pylori. J Clin Gastroenterol. 1993;17:S136–S139. doi: 10.1097/00004836-199312001-00024. [DOI] [PubMed] [Google Scholar]
- 74.Ishino F, Park W, Tomioka S, Tamaki S, Takase I. Peptidoglycan synthetic activities in membranes of Escherichia coli caused by overproduction of penicillin-binding protein 2 and RodA protein. J Biol Chem. 1986;261:7024–7031. [PubMed] [Google Scholar]
- 75.Jiang Q, Hiratsuka K, Taylor D. Variability of gene order in different Helicobacter pylori strains contributes to genome diversity. Mol Microbiol. 1996;20:833–842. doi: 10.1111/j.1365-2958.1996.tb02521.x. [DOI] [PubMed] [Google Scholar]
- 76.Kadner R. Cytoplasmic membrane. In: Neidhardt F C, Curtiss III R, Ingraham J L, Lin E C C, Low K B, Magasanik B, Reznikoff W S, Riley M, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella: cellular and molecular biology. Vol. 1. Washington, D.C: ASM Press; 1996. pp. 58–87. [Google Scholar]
- 77.Kansau I, Raymond J, Bingen E, Courcoux P, Kalach N, Bergeret M, Braimi N, Dupont C, Labigne A. Genotyping of Helicobacter pylori isolates by sequencing of PCR products and comparison with the RAPD technique. Res Microbiol. 1996;147:661–669. doi: 10.1016/0923-2508(96)84023-x. [DOI] [PubMed] [Google Scholar]
- 78.Karita M, Tsuda M, Nakazawa T. Essential role of urease in vitro and in vivo Helicobacter pylori colonization study using a wild-type and isogenic urease mutant strain. J Clin Gastroenterol. 1995;21:S160–163. [PubMed] [Google Scholar]
- 79.Kersulyte D, Akopyants N S, Clifton S, Roe B S, Berg D E. Novel sequence organization and insertion specificity of IS605 and IS606: chimeric transposable elements of Helicobacter pylori. Gene. 1998;223:175–186. doi: 10.1016/s0378-1119(98)00164-4. [DOI] [PubMed] [Google Scholar]
- 80.Klebba P E, Rutz J M, Liu J, Murphy C K. Mechanisms of TonB-catalyzed iron transport through the enteric bacterial cell envelope. J Bioenerg Biomembr. 1993;25:603–611. doi: 10.1007/BF00770247. [DOI] [PubMed] [Google Scholar]
- 81.Klenk H P, Clayton R A, Tomb J F, White O, Nelson K E, Ketchum K A, Dodson R J, Gwinn M, Hickey E K, Peterson J D, Richardson D L, Kerlavage A R, Graham D E, Kyrpides N C, Fleischmann R D, Quackenbush J, Lee N H, Sutton G G, Gill S, Kirkness E F, Dougherty B A, McKenney K, Adams M D, Loftus B, Venter J C, et al. The complete genome sequence of the hyperthermophilic, sulphate-reducing archaeon Archaeoglobus fulgidus. Nature. 1997;390:364–370. doi: 10.1038/37052. [DOI] [PubMed] [Google Scholar]
- 82.Komorowska M, Szafran H, Popiela T, Szafran Z. Free amino acids in gastric juice. Acta Physiol Pol. 1981;32:559–567. [PubMed] [Google Scholar]
- 83.Kostrzynska M, O’Toole P W, Taylor D E, Trust T J. Molecular characterization of a conserved 20-kilodalton membrane-associated lipoprotein antigen of Helicobacter pylori. J Bacteriol. 1994;176:5938–5948. doi: 10.1128/jb.176.19.5938-5948.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Kunst F, Ogasawara N, Moszer I, Albertini A, Alloni G, Azevedo V, Bertero M, Bessieres P, Bolotin A, Borchert S, Borriss R, Boursier L, Brans A, Braun M, Brignell S, Bron S, Brouillet S, Bruschi C, Caldwell B, Capuano V, Carter N, Choi S-K, Codani J-J, Connerton I, Cummings N, Daniel R, Denizot F, Devine K, Dusterhoft A, Ehrlich S, Emmerson P, Entian K, Errington J, Fabret C, Ferrari E, Foulger D, Fritz C, Fujita M, Fujita Y, Fuma S, Galizzi A, Galleron N, Ghim S-Y, Glaser P, Goffeau A, Golightly E, Grandi G, Guiseppi G, Guy B, Haga K, Haiech J, Harwood C, Henaut A, Hilbert H, Holsappel S, Hosono S, Hullo M-F, Itaya M, Jones L, Joris B, Karamata D, Kasahara Y, Klaerr-Blanchard M, Klein C, Kobayashi Y, Loetter P, Koningstein G, Krogh S, Kumano M, Kurita K, Lapidus A, Lardinois S, Lauber J, Lazarevic V, Lee S-M, Levine A, Liu H, Masuda S, Mauel C, Medigue C, Median N, Mellado R, Mizuno M, Moestl D, Nakai S, Noback M, Noone D, O’Reilly M, Ogawa K, Ogiwara A, Oudega B, Park S-H, Parro V, Pohl T, Portetelle D, Porwollik S, Prescott A, Presecan E, Pujic P, Purnelle B, et al. The complete genome sequence of the Gram-positive bacterium Bacillus subtilis. Nature. 1997;390:249–256. doi: 10.1038/36786. [DOI] [PubMed] [Google Scholar]
- 85.Kusukawa N, Yura T, Ueguchi C, Akiyama Y, Ito K. Effects of mutations in heat-shock genes groES and groEL on protein export in Escherichia coli. EMBO J. 1989;8:3517–3521. doi: 10.1002/j.1460-2075.1989.tb08517.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Labigne A, Cussac V, Courcoux P. Shuttle cloning and nucleotide sequences of Helicobacter pylori genes responsible for urease activity. J Bacteriol. 1991;173:1920–1931. doi: 10.1128/jb.173.6.1920-1931.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Labigne A, deReuse H. Determinants of Helicobacter pylori pathogenicity. Infect Agents Dis. 1996;5:191–202. [PubMed] [Google Scholar]
- 88.Lee C A. Type III secretion systems: machines to deliver bacterial proteins into eukaryotic cells? Trends Microbiol. 1997;5:148–156. doi: 10.1016/S0966-842X(97)01029-9. [DOI] [PubMed] [Google Scholar]
- 89.Leying H, Suerbaum S, Geis G, Haas R. Cloning and genetic characterization of a Helicobacter pylori flagellin gene. Mol Microbiol. 1992;6:2863–2874. doi: 10.1111/j.1365-2958.1992.tb01466.x. [DOI] [PubMed] [Google Scholar]
- 90.Luke C, Penn C. Identification of a 29kDa flagellar sheath protein in Helicobacter pylori using a murine monoclonal antibody. Microbiology. 1995;141:597–604. doi: 10.1099/13500872-141-3-597. [DOI] [PubMed] [Google Scholar]
- 91.Luke C J, Kubiak E, Cockayne A, Elliott T S, Penn C W. Identification of flagellar and associated polypeptides of Helicobacter (formerly Campylobacter) pylori. FEMS Microbiol Lett. 1990;59:225–230. doi: 10.1016/0378-1097(90)90061-t. [DOI] [PubMed] [Google Scholar]
- 92.Mahdi A A, Lloyd R G. Identification of the recR locus of Escherichia coli K-12 and analysis of its role in recombination and DNA repair. Mol Gen Genet. 1989;216:503–510. doi: 10.1007/BF00334397. [DOI] [PubMed] [Google Scholar]
- 93.Maier R J, Fu C, Gilbert J, Moshiri F, Olson J, Plaut A G. Hydrogen uptake hydrogenase in Helicobacter pylori. FEMS Microbiol Lett. 1996;141:71–76. doi: 10.1111/j.1574-6968.1996.tb08365.x. [DOI] [PubMed] [Google Scholar]
- 94.Marcelli S W, Chang H T, Chapman T, Chalk P A, Miles R J, Poole R K. The respiratory chain of Helicobacter pylori: identification of cytochromes and the effects of oxygen on cytochrome and menaquinone levels. FEMS Microbiol Lett. 1996;138:59–64. doi: 10.1111/j.1574-6968.1996.tb08135.x. [DOI] [PubMed] [Google Scholar]
- 95.Marshall D G, Dundon W G, Beesley S M, Smyth C J. Helicobacter pylori—a conundrum of genetic diversity. Microbiology. 1998;144:2925–2939. doi: 10.1099/00221287-144-11-2925. [DOI] [PubMed] [Google Scholar]
- 96.Martin S, Edbrooke M, Hodgman T, van der Eijnden D, Bird M. Lewis X biosynthesis in Helicobacter pylori. Molecular cloning of an α(1,3)-fucosyltransferase gene. J Biol Chem. 1997;272:21349–21356. doi: 10.1074/jbc.272.34.21349. [DOI] [PubMed] [Google Scholar]
- 97.Matin A, Zychlinsky E, Keyhan M, Sachs G. Capacity of Helicobacter pylori to generate ionic gradients at low pH is similar to that of bacteria which grow under strongly acidic conditions. Infect Immun. 1996;64:1434–1436. doi: 10.1128/iai.64.4.1434-1436.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Mendz G L, Burns B P, Hazell S L. Characterisation of glucose transport in Helicobacter pylori. Biochim Biophys Acta. 1995;1244:269–276. doi: 10.1016/0304-4165(95)00018-7. [DOI] [PubMed] [Google Scholar]
- 99.Mendz G L, Hazell S L. Amino acid utilization by Helicobacter pylori. Int J Biochem Cell Biol. 1995;27:1085–1093. doi: 10.1016/1357-2725(95)00069-2. [DOI] [PubMed] [Google Scholar]
- 100.Mendz G L, Hazell S L. Evidence for a pentose phosphate pathway in Helicobacter pylori. FEMS Lett. 1991;84:331–336. [Google Scholar]
- 101.Mendz G L, Hazell S L. Fumarate catabolism in Helicobacter pylori. Biochem Mol Biol Int. 1993;31:325–332. [PubMed] [Google Scholar]
- 102.Mendz G L, Hazell S L. Glucose phosphorylation in Helicobacter pylori. Arch Biochem Biophys. 1993;300:522–525. doi: 10.1006/abbi.1993.1071. [DOI] [PubMed] [Google Scholar]
- 103.Mendz G L, Hazell S L, Burns B P. The Entner-Doudoroff pathway in Helicobacter pylori. Arch Biochem Biophys. 1994;312:349–356. doi: 10.1006/abbi.1994.1319. [DOI] [PubMed] [Google Scholar]
- 104.Mendz G L, Hazell S L, Burns B P. Glucose utilization and lactate production by Helicobacter pylori. J Gen Microbiol. 1993;139:3023–3028. doi: 10.1099/00221287-139-12-3023. [DOI] [PubMed] [Google Scholar]
- 105.Mendz G L, Hazell S L, Srinivasan S. Fumarate reductase: a target for therapeutic intervention against Helicobacter pylori. Arch Biochem Biophys. 1995;321:153–159. doi: 10.1006/abbi.1995.1380. [DOI] [PubMed] [Google Scholar]
- 106.Mendz G L, Hazell S L, van Gorkom L. Pyruvate metabolism in Helicobacter pylori. Arch Microbiol. 1994;162:187–192. doi: 10.1007/BF00314473. [DOI] [PubMed] [Google Scholar]
- 107.Mendz G L, Jimenez B M, Hazell S L, Gero A M, O’Sullivan W J. Salvage synthesis of purine nucleotides by Helicobacter pylori. J Appl Bacteriol. 1994;77:674–681. doi: 10.1111/j.1365-2672.1994.tb02818.x. [DOI] [PubMed] [Google Scholar]
- 108.Mendz G L, Jiminez B M, Hazell S L, Gero A M, O’Sullivan W J. De novo synthesis of pyrimidine nucleotides by Helicobacter pylori. J Appl Bacteriol. 1994;77:1–8. doi: 10.1111/j.1365-2672.1994.tb03036.x. [DOI] [PubMed] [Google Scholar]
- 109.Miehlke S, Meining A, Morgner A, Bayerdorffer E, Lehn N, Stolte M, Graham D Y, Go M F. Frequency of vacA genotypes and cytotoxin activity in Helicobacter pylori associated with low-grade gastric mucosa-associated lymphoid tissue lymphoma. J Clin Microbiol. 1998;36:2369–2370. doi: 10.1128/jcm.36.8.2369-2370.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Mizote T, Yoshiyama H, Nakazawa T. Urease-independent chemotactic responses of Helicobacter pylori to urea, urease inhibitors, and sodium bicarbonate. Infect Immun. 1997;65:1519–1521. doi: 10.1128/iai.65.4.1519-1521.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Mobley H L. Defining Helicobacter pylori as a pathogen: strain heterogeneity and virulence. Am J Med. 1996;100:2S–9S. doi: 10.1016/s0002-9343(96)80223-3. [DOI] [PubMed] [Google Scholar]
- 112.Mobley H L. Helicobacter pylori factors associated with disease development. Gastroenterology. 1997;113:S21–S28. doi: 10.1016/s0016-5085(97)80006-6. [DOI] [PubMed] [Google Scholar]
- 113.Mobley H L, Island M D, Hausinger R P. Molecular biology of microbial ureases. Microbiol Rev. 1995;59:451–480. doi: 10.1128/mr.59.3.451-480.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Mobley H L T, Garner R M, Bauerfeind P. Helicobacter pylori nickel-transport gene nixA: synthesis of catalytically active urease in Escherichia coli independent of growth conditions. Mol Microbiol. 1995;16:97–109. doi: 10.1111/j.1365-2958.1995.tb02395.x. [DOI] [PubMed] [Google Scholar]
- 115.Moran A P, Lindner B, Walsh E. Structural characterization of the lipid A component of Helicobacter pylori rough- and smooth-form lipopolysaccharides. J Bacteriol. 1997;179:6453–6463. doi: 10.1128/jb.179.20.6453-6463.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Nagata K, Tsukita S, Tamura T, Sone N. A cb-type cytochrome-c oxidase terminates the respiratory chain in Helicobacter pylori. Microbiology. 1996;142:1757–1763. doi: 10.1099/13500872-142-7-1757. [DOI] [PubMed] [Google Scholar]
- 117.Nakao H, Shioda S, Yamamoto S. Purification and properties of carboxynorspermidine decarboxylase, a novel enzyme involved in norspermidine biosynthesis from Vibrio alginolyticus. J Gen Microbiol. 1990;136:1699–1704. [Google Scholar]
- 118.Nedeskov P. Nutritional requirements for growth of Helicobacter pylori. Appl Environ Microbiol. 1994;60:3450–3453. doi: 10.1128/aem.60.9.3450-3453.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Neidhardt F C, Curtiss III R, Ingraham J L, Lin E C C, Low K B, Magasanik B, Reznikoff W S, Riley M, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella: cellular and molecular biology. 2nd ed. Washington, D.C: ASM Press; 1996. [Google Scholar]
- 120.Newell D. Identification of the outer membrane proteins of Campylobacter pyloridis and antigenic cross-reactivity between C. pyloridis and C. jejuni. J Gen Microbiol. 1987;133:163–170. doi: 10.1099/00221287-133-1-163. [DOI] [PubMed] [Google Scholar]
- 121.Nishiyama K, Mizushima S, Tokuda H. A novel membrane protein involved in protein translocation across the cytoplasmic membrane of Escherichia coli. EMBO J. 1993;12:3409–3415. doi: 10.1002/j.1460-2075.1993.tb06015.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Odenbreit S, Wieland B, Haas R. Cloning and genetic characterization of Helicobacter pylori catalase and construction of a catalase-deficient mutant strain. J Bacteriol. 1996;178:6960–6967. doi: 10.1128/jb.178.23.6960-6967.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Ohta-Tada U, Takagi A, Koga Y, Kamiya S, Miwa T. Flagellin gene diversity among Helicobacter pylori strains and IL-8 secretion from gastric epithelial cells. Scand J Gastroenterol. 1997;32:455–459. doi: 10.3109/00365529709025080. [DOI] [PubMed] [Google Scholar]
- 124.O’Toole P W, Kostrzynska M, Trust T J. Non-motile mutants of Helicobacter pylori and Helicobacter mustelae defective in flagellar hook production. Mol Microbiol. 1994;14:691–703. doi: 10.1111/j.1365-2958.1994.tb01307.x. [DOI] [PubMed] [Google Scholar]
- 125.Page M D, Tomlinson E, Ferguson S J. Unexpected implications from the Helicobacter pylori genome for understanding periplasmic c-type cytochrome assembly in gram-negative bacteria in coexistence with disulphide bond formation. Mol Microbiol. 1997;26:413–415. doi: 10.1046/j.1365-2958.1997.6061962.x. [DOI] [PubMed] [Google Scholar]
- 126.Pahel G, Zelenetz A D, Tyler B M. gltB gene and regulation of nitrogen metabolism by glutamine synthetase in Escherichia coli. J Bacteriol. 1978;133:139–148. doi: 10.1128/jb.133.1.139-148.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Park J T. Why does Escherichia coli recycle its cell wall peptides? Mol Microbiol. 1995;17:421–426. doi: 10.1111/j.1365-2958.1995.mmi_17030421.x. [DOI] [PubMed] [Google Scholar]
- 128.Penn C W, Luke C J. Bacterial flagellar diversity and significance in pathogenesis. FEMS Microbiol Lett. 1992;79:331–336. doi: 10.1111/j.1574-6968.1992.tb14060.x. [DOI] [PubMed] [Google Scholar]
- 129.Phadnis S H, Parlow M H, Levy M, Ilver D, Caulkins C M, Connors J B, Dunn B E. Surface localization of Helicobacter pylori urease and a heat shock protein homolog requires bacterial autolysis. Infect Immun. 1996;64:905–912. doi: 10.1128/iai.64.3.905-912.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Preisig O, Anthamatten D, Hennecke H. Genes for a microaerobically induced oxidase complex in Bradyrhizobium japonicum are essential for a nitrogen-fixing endosymbiosis. Proc Nat Acad Sci USA. 1993;90:3309–3013. doi: 10.1073/pnas.90.8.3309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Reitzer L. Ammonia assimilation and the biosynthesis of glutamine, glutamate, aspartate, asparagine, l-alanine, and d-alanine. In: Neidhardt F C, Curtiss III R, Ingraham J L, Lin E C C, Low K B, Magasanik B, Reznikoff W S, Riley M, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella: cellular and molecular biology. Vol. 1. Washington, D.C: ASM Press; 1996. pp. 391–407. [Google Scholar]
- 132.Reynolds D J, Penn C W. Characteristics of Helicobacter pylori growth in a defined medium and determination of its amino acid requirements. Microbiology. 1994;140:2649–2656. doi: 10.1099/00221287-140-10-2649. [DOI] [PubMed] [Google Scholar]
- 133.Roine R P, Salmela K S, Salaspuro M. Alcohol metabolism in Helicobacter pylori-infected stomach. Ann Med. 1995;27:583–588. doi: 10.3109/07853899509002473. [DOI] [PubMed] [Google Scholar]
- 134.Rosario M M, Fredrick K L, Ordal G W, Helmann J D. Chemotaxis in Bacillus subtilis requires either of two functionally redundant CheW homologs. J Bacteriol. 1994;176:2736–2739. doi: 10.1128/jb.176.9.2736-2739.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Salmela K S, Roine R P, Hook-Nikanne J, Kosunen T U, Salaspuro M. Acetaldehyde and ethanol production by Helicobacter pylori. Scand J Gastroenterol. 1994;29:309–312. doi: 10.3109/00365529409094841. [DOI] [PubMed] [Google Scholar]
- 136.Salmela K S, Roine R P, Koivisto T, Hook-Nikanne J, Kosunen T U, Salaspuro M. Characteristics of Helicobacter pylori alcohol dehydrogenase. Gastroenterology. 1993;105:325–330. doi: 10.1016/0016-5085(93)90704-g. [DOI] [PubMed] [Google Scholar]
- 137.Segal E D, Lange C, Covacci A, Tompkins L S, Falkow S. Induction of host signal transduction pathways by Helicobacter pylori. Proc Natl Acad Sci USA. 1997;94:7595–7599. doi: 10.1073/pnas.94.14.7595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Seifert J, Kunz N, Flachmann R, Laufer A, Jany K D, Gassen H G. Expression of the E. coli nadB gene and characterization of the gene product l-aspartate oxidase. Biol Chem Hoppe-Seyler. 1990;371:239–248. [PubMed] [Google Scholar]
- 139.Smith D, Douchette-Stamm L, Deloughery C, Lee H, Dubois J, Aldredge T, Bashirzadeh R, Blakely D, Cook R, Gilbert K, Harrison D, Hoang L, Keagle P, Lumm W, Pothier B, Qiu D, Spadafors R, Vicaire R, Wang Y, Wierzbowski J, Gibson R, Jiwani N, Caruso A, Bush D, Safer H, Patwell D, Prabhakar S, McDougall S, Shimer G, Goyal A, Pietrokovski S, Church G, Daniels C, Mao J-I, Rice P, Nolling J, Reeve J. Complete genome sequence of Methanobacterium thermoautotrophicum deltaH: functional analysis and comparative genomics. J Bacteriol. 1997;179:7135–7155. doi: 10.1128/jb.179.22.7135-7155.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Spiegelhalder C, Gerstenecker B, Kersten A, Schiltz E, Kist M. Purification of Helicobacter pylori superoxide dismutase and cloning and sequencing of the gene. Infect Immun. 1993;61:5315–5325. doi: 10.1128/iai.61.12.5315-5325.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Spohn G, Beier D, Rappuoli R, Scarlato V. Transcriptional analysis of the divergent cagAB genes encoded by the pathogenecity island of Helicobacter pylori. Mol Microbiol. 1997;26:361–372. doi: 10.1046/j.1365-2958.1997.5831949.x. [DOI] [PubMed] [Google Scholar]
- 142.Spratt B G. Properties of the penicillin-binding proteins of Escherichia coli K12. Eur J Biochem. 1977;72:341–352. doi: 10.1111/j.1432-1033.1977.tb11258.x. [DOI] [PubMed] [Google Scholar]
- 143.Stark R M, Suleiman M S, Hassan I J, Greenman J, Millar M R. Amino acid utilisation and deamination of glutamine and asparagine by Helicobacter pylori. J Med Microbiol. 1997;46:793–800. doi: 10.1099/00222615-46-9-793. [DOI] [PubMed] [Google Scholar]
- 144.Stephens J C, Stewart J A, Folwell A M, Rathbone B J. Helicobacter pylori cagA status, vacA genotypes and ulcer disease. Eur J Gastroenterol Hepatol. 1998;10:381–384. doi: 10.1097/00042737-199805000-00005. [DOI] [PubMed] [Google Scholar]
- 145.Stock A, Schaeffer E, Koshland D E, Stock J. A second type of protein methylation reaction in bacterial chemotaxis. J Biol Chem. 1987;262:8011–8014. [PubMed] [Google Scholar]
- 146.Strauch M A, Zalkin H, Aronson A I. Characterization of the glutamyl-tRNA(Gln)-to-glutaminyl-tRNA(Gln) amidotransferase reaction of Bacillus subtilis. J Bacteriol. 1988;170:916–920. doi: 10.1128/jb.170.2.916-920.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Suerbaum S, Josenhans C, Labigne A. Cloning and genetic characterization of the Helicobacter pylori and Helicobacter mustellae flaB genes and construction of H. pylori flaA- and flaB-negative mutants by allelic exchange. J Bacteriol. 1993;175:3278–3288. doi: 10.1128/jb.175.11.3278-3288.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Taylor D, Eaton M, Chang N, Salama S. Construction of a Helicobacter pylori genome map and demonstration of diversity at the genome level. J Bacteriol. 1992;174:6800–6806. doi: 10.1128/jb.174.21.6800-6806.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Tomb J-F, White O, Kerlavage A, Clayton R, Sutton G, Fleischmann R, Ketchum K, Klenk H, Gill S, Dougherty B, Nelson K, Quackenbush J, Zhou L, Kirkness E, Peterson S, Loftus B, Richardson D, Dodson R, Khalak H, Glodek A, McKenney K, Fitzegerald L, Lee N, Adams M, Hickey E, Berg D, Gocayne J, Utterback T, Peterson J, Kelley J, Cotton M, Weidman J, Fujii C, Bowman C, Watthey L, Wallin E, Hayes W, Borodovsky M, Karp P, Smith H, Fraser C, Venter J. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature. 1997;388:539–547. doi: 10.1038/41483. [DOI] [PubMed] [Google Scholar]
- 150.Tsuda M, Karita M, Morshed M, Okita K, Nakazawa T. A urease-negative mutant of Helicobacter pylori constructed by allelic exchange mutagenesis lacks the ability to colonize the nude mouse stomach. Infect Immun. 1994;62:3586–3589. doi: 10.1128/iai.62.8.3586-3589.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.van Doorn L J, Figueiredo C, Sanna R, Pena S, Midolo P, Ng E K, Atherton J C, Blaser M J, Quint W G. Expanding allelic diversity of Helicobacter pylori vacA. J Clin Microbiol. 1998;36:2597–2603. doi: 10.1128/jcm.36.9.2597-2603.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.van Doorn L J, Figueiredo C, Sanna R, Plaisier A, Schneeberger P, de Boer W, Quint W. Clinical relevance of the cagA, vacA, and iceA status of Helicobacter pylori. Gastroenterology. 1998;115:58–66. doi: 10.1016/s0016-5085(98)70365-8. [DOI] [PubMed] [Google Scholar]
- 153.Vanet A, Labigne A. Evidence for specific secretion rather than autolysis in the release of some Helicobacter pylori proteins. Infect Immun. 1998;66:1023–1027. doi: 10.1128/iai.66.3.1023-1027.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Vincente M, Gomez M J, Agata J A. Regulation of transcription of cell division genes in the Escherichia coli dcw cluster. Cell Mol Life Sci. 1990;59:317–324. doi: 10.1007/s000180050158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Weiss A A, Johnson F D, Burns D L. Molecular characterization of an operon required for pertussis toxin secretion. Proc Natl Acad Sci USA. 1993;90:2970–2974. doi: 10.1073/pnas.90.7.2970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Westblom T U, Phadnis S, Langenberg W, Yoneda K, Madan E, Midkiff B R. Catalase negative mutants of Helicobacter pylori. Eur J Clin Microbiol Infect Dis. 1992;11:522–526. doi: 10.1007/BF01960807. [DOI] [PubMed] [Google Scholar]
- 157.Williams C L, Preston T, Hossack M, Slater C, McColl K E. Helicobacter pylori utilises urea for amino acid synthesis. FEMS Immunol Med Microbiol. 1996;13:87–94. doi: 10.1111/j.1574-695X.1996.tb00220.x. [DOI] [PubMed] [Google Scholar]
- 158.Worst D J, Gerrits M M, Vandenbroucke-Grauls C M, Kusters J G. Helicobacter pylori ribBA-mediated riboflavin production is involved in iron acquisition. J Bacteriol. 1998;180:1473–1479. doi: 10.1128/jb.180.6.1473-1479.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Xu Q, Peek R M J, Miller G G, Blaser M J. The Helicobacter pylori genome is modified at CATG by the product of hpyIM. J Bacteriol. 1997;179:6807–6815. doi: 10.1128/jb.179.21.6807-6815.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Zakharova N, Hoffman P S, Berg D E, Severinov K. The largest subunits of RNA polymerase from gastric helicobacters are tethered. J Biol Chem. 1998;273:19371–19374. doi: 10.1074/jbc.273.31.19371. [DOI] [PubMed] [Google Scholar]