Abstract
Samsoniella species have been often found in the forest habitat and rarely found in special karst eco-environments, such as Tiankeng, valleys and caves. In this research, eleven cordyceps specimens were collected from Mayao River Valley. A known species (S.haniana) and two new species (S.duyunensis and S.vallis) were established and described according to a multilocus phylogenetic analysis and morphological characteristics. Our results provide insight that the richness of Samsoniella species in karst eco-environments and further attention should be paid to entomopathogenic fungi in such habitats.
Key words: Entomopathogenic fungi, morphology, phylogenetic analysis, Sordariomycetes, valley
Introduction
The genus Samsoniella Mongkols., Noisrip., Thanakitp., Spatafora & Luangsa-ard was proposed based on the phylogenetic analysis of Isaria-like morphs in Cordycipitaceae and characterised by oval to fusiform conidia and bright red-orange teleomorphic stromata and anamorphic synnemata by Mongkolsamrit et al. (2018). One Isaria-like species, Penicilliumalboaurantium G. Sm. was transferred to Samsoniella and two new species, S.aurantia and S.inthanonensis were described.
Subsequently, Chen et al. (2020a) reported three new species, S.coleopterorum W.H. Chen, Y.F. Han & Z.Q. Liang, S.hymenopterorum W.H. Chen, et al. and S.lepidopterorum W.H. Chen, et al. from the forestry of Xishui and Rongjiang County, Guizhou Province, China. Those species have mononematous conidiophores rather than synnemata and associated with hymenopteran larvae, coleopteran larvae and lepidopteran pupae, respectively. Wang et al. (2020) described nine new species and a new combination from the forest habitats of Yunnan Province, China. The other thirteen new species were reported by Chen et al. (2021a, 2022), Wang et al. (2022) and Crous et al. (2023). Currently, there are twenty-nine species described in the genus of Samsoniella.
Additionally, it has been reported that Samsoniella species are found in the forest habitat. However, the other ecological habitats, especially the karst eco-environment which has special niches like Tiankeng, valleys and caves should have insects and entomopathogenic fungi. In this research, eleven cordyceps specimens were collected from Mayao River Valley, Guizhou, China. After detailed multiloci phylogenic analysis and morphological observation, two new species and one known species were identified.
Materials and methods
Specimen collection and identification
Eleven cordyceps specimens were collected from Mayao River Valley (26°22'8.3748"N, 107°23'16.96"E), Duyun City, Qiannan Buyei and Miao Autonomous Prefecture, Guizhou, on 4 September 2021 and 30 July 2022. The samples were placed in an ice box and brought to the laboratory and preserved in refrigerator at 4 °C before use. The surface of each arthropod body was rinsed with sterile water, followed by sterilisation with 75% ethanol for 3–5 s and rinsing again three times with sterilised water. After drying on sterilised filter paper, a piece of the synnemata, mycelium or the sclerotia was cut from the specimen and inoculated on agar plates of potato dextrose agar (PDA) or PDA modified by the addition of 1% w/v peptone containing 0.1 g/l streptomycin and 0.05 g/l tetracycline (Chen et al. 2019a). After fungal colonies emerged from the inoculated samples, a piece of mycelium from the colony edge was transferred onto new agar plated and cultured at 25 °C for 14 days under 12 h light/12 h dark conditions (Zou et al. 2010). The specimens and axenic cultures were deposited at the Institute of Fungus Resources, Guizhou University (formally Herbarium of Guizhou Agricultural College; code, GZAC), Guiyang City, Guizhou, China.
Colony morphology was determined on PDA cultures incubated at 25 °C for 14 days and the growth rate, the presence of octahedral crystals and the colony colours (surface and reverse) were observed. To investigate the microscopic characteristics, a little of the mycelia was picked up from the colony and mounted in lactophenol cotton blue or 20% lactate acid solution and the asexual morphological characteristics (e.g., conidiophores, phialides and conidia) were observed and measured under a Leica DM4 B microscope.
DNA extraction, polymerase chain reaction amplification and nucleotide sequencing
DNA extraction was carried out using a fungal genomic DNA extraction kit (DP2033, BioTeke Corporation) according to Liang et al. (2011). The extracted DNA was stored at −20 °C. Polymerase chain reaction (PCR) was used to amplify genetic markers using the following primer pairs: ITS4/ITS5 for the internal transcribed spacer (ITS) region (White et al. 1990), LR0R/LR5 for 28s large subunit ribosomal (LSU) (Vilgalys and Hester 1990), CRPB1/RPB1Cr for RNA polymerase II largest subunit (RPB1) (Castlebury et al. 2004), fRPB2-5F/fRPB2-7cR for RNA polymerase II second largest subunit (RPB2) (Liu et al. 1999) and 983F/2218R for translation elongation factor 1 alpha (TEF) (Castlebury et al. 2004). The thermal cycle of PCR amplification for these phylogenetic markers was set up following the procedure described by Chen et al. (2021). PCR products were purified and sequenced at Sangon Biotech (Shanghai) Co. The resulting sequences were submitted to GenBank (Table 1).
Table 1.
List of strains and GenBank accession numbers of sequences used in this study.
| Species | Strain | Host/Substratum | GenBank accession number | Reference | ||||
|---|---|---|---|---|---|---|---|---|
| ITS | LSU | RPB1 | RPB2 | TEF | ||||
| Akanthomycesaraneosus | KY11341 | Araneae (Spider) | ON502826 | ON502832 | – | ON525442 | ON525443 | Chen et al. (2022) |
| KY11342 | Araneae (Spider) | ON502844 | ON502837 | – | ON525444 | ON525445 | Chen et al. (2022) | |
| Akanthomycesattenuatus | CBS 402.78 | Leaf litter; Acersaccharum | AJ292434 | AF339565 | EF468888 | EF468935 | EF468782 | Sung et al. (2007) |
| Akanthomyceslecanii | CBS 101247 | Hemiptera; Coccusviridis | JN049836 | AF339555 | DQ522407 | DQ522466 | DQ522359 | Spatafora et al. (2007) |
| Akanthomycestiankengensis | KY11571 | Araneae (Spider) | ON502848 | ON502825 | – | ON525446 | ON525447 | Chen et al. (2022) |
| KY11572 | Araneae (Spider) | ON502821 | ON502827 | – | ON525448 | ON525449 | Chen et al. (2022) | |
| Akanthomycestortricidarum | BCC72638 | Lepidoptera; tortricidae | MT356076 | MT356088 | MT477997 | MT477992 | MT478004 | Aini et al. (2020) |
| Beauveriabassiana | ARSEF 1564 | Lepidoptera; Arctiidae | HQ880761 | – | HQ880833 | HQ880905 | HQ880974 | Rehner et al. (2011) |
| Beauveriabrongniartii | ARSEF 617 | Coleoptera; Scarabaeidae | HQ880782 | – | HQ880854 | HQ880926 | HQ880991 | Rehner et al. (2011) |
| BCC 16585 | Coleoptera; Anomalacuprea (larva) | JN049867 | JF415967 | JN049885 | JF415991 | JF416009 | Kepler et al. (2012) | |
| Samsoniellaalboaurantia | CBS 240.32 | Lepidoptera (pupa) | – | JF415979 | JN049895 | JF415999 | JF416019 | Kepler et al. (2012) |
| CBS 262.58 | Soil | – | AB080087 | MF416654 | MF416448 | MF416497 | Kepler et al. (2012) | |
| Samsoniellaalpina | YFCC 5818 | Hepialidae (Hepialusbaimaensis) | – | MN576809 | MN576869 | MN576923 | MN576979 | Wang et al. (2020) |
| Samsoniellaalpina | YFCC 5831 |
Hepialidae (Hepialusbaimaensis) |
– | MN576810 | MN576870 | MN576924 | MN576980 | Wang et al. (2020) |
| Samsoniellaantleroides | YFCC 6016 | Noctuidae (Larvae) | – | MN576803 | MN576863 | MN576917 | MN576973 | Wang et al. (2020) |
| YFCC 6113 | Noctuidae (Larvae) | – | MN576804 | MN576864 | MN576918 | MN576974 | Wang et al. (2020) | |
| Samsoniellaaurantia | TBRC 7271 | Lepidoptera | – | MF140728 | MF140791 | MF140818 | MF140846 | Mongkolsamrit et al. (2018) |
| TBRC 7272 | Lepidoptera | – | MF140727 | – | MF140817 | MF140845 | Mongkolsamrit et al. (2018) | |
| Samsoniellacardinalis | YFCC 5830 | Limacodidae (Pupa) | – | MN576788 | MN576848 | MN576902 | MN576958 | Wang et al. (2020) |
| YFCC 6144 | Limacodidae (Pupa) | – | MN576786 | MN576846 | MN576900 | MN576956 | Wang et al. (2020) | |
| Samsoniellacoccinellidicola | YFCC 8772 | Coccinellidae | – | ON621670 | ON676502 | ON568685 | ON676514 | Wang et al. (2020) |
| YFCC 8773 | Coccinellidae | – | ON621671 | ON676503 | ON568686 | ON676515 | Wang et al. (2020) | |
| Samsoniellacoleopterorum | A19501 | Curculionidae (Snout beetle) | MT626376 | – | MT642600 | MN101585 | MN101586 | Chen et al. (2020) |
| Samsoniellacristata | YFCC 6021 | Saturniidae (Pupa) | – | MN576791 | MN576851 | MN576905 | MN576961 | Wang et al. (2020) |
| Samsoniellacristata | YFCC 6023 | Saturniidae (Pupa) | – | MN576792 | – | MN576906 | MN576962 | Wang et al. (2020) |
| Samsonielladuyunensis | DY09161 | Formicidae (Ant) | OQ379241 | OQ363112 | OR296698 | OQ397660 | OQ398145 | This study |
| DY09162 | Formicidae (Ant) | OQ379242 | OQ363114 | – | – | OQ398146 | This study | |
| DY07501 | Lepidoptera (Pupa) | OR263188 | OR263307 | OR282773 | OR282776 | OR282780 | This study | |
| DY07502 | Lepidoptera (Pupa) | OR263189 | OR263427 | – | OR282777 | OR282781 | This study | |
| Samsoniellaerucae | KY11121 | Lepidoptera (Caterpillar) | ON502828 | ON502835 | – | ON525424 | ON525425 | Chen et al. (2022) |
| Samsoniellaerucae | KY11122 | Lepidoptera (Caterpillar) | ON502847 | ON502822 | – | ON525426 | ON525427 | Chen et al. (2022) |
| Samsoniellafarinospora | YFCC 8774 | Araneae (Spider) | – | ON621672 | ON676504 | ON568687 | ON676516 | Wang et al. (2022) |
| YFCC 9051 | Lepidoptera: Hepialus | – | ON621673 | ON676505 | ON568688 | ON676517 | Wang et al. (2022) | |
| Samsoniellaformicae | KY11041 | Formicidae (Ant) | ON502852 | – | – | ON525420 | ON525421 | Chen et al. (2022) |
| KY11042 | Formicidae (Ant) | ON502842 | – | – | ON525422 | ON525423 | Chen et al. (2022) | |
| Samsoniellaguizhouensis | KY11161 | Lepidoptera (Pupa) | ON502823 | ON502830 | – | ON525428 | ON525429 | Chen et al. (2022) |
| KY11162 | Lepidoptera (Pupa) | ON502845 | ON502846 | – | ON525430 | ON525431 | Chen et al. (2022) | |
| Samsoniellahaniana | YFCC 8769 | Lepidoptera (Pupa) | – | ON621674 | ON676506 | ON568689 | ON676518 | Wang et al. (2022) |
| YFCC 8770 | Lepidoptera (Pupa) | – | ON621675 | ON676507 | ON568690 | ON676519 | Wang et al. (2022) | |
| YFCC 8771 | Lepidoptera (Pupa) | – | ON621676 | ON676508 | ON568691 | ON676520 | Wang et al. (2022) | |
| Samsoniellahaniana | DY091031 | Lepidoptera (Pupa) | OQ359979 | OQ363133 | – | – | OQ398149 | This study |
| DY091032 | Lepidoptera (Pupa) | OQ359978 | OQ363134 | – | – | OQ398150 | This study | |
| DY091021 | Coccinellidae (ladybug) | OQ379240 | OQ363115 | OR296699 | OQ397661 | OQ398147 | This study | |
| DY091022 | Coccinellidae (ladybug) | OQ359881 | OQ363117 | – | OQ397662 | OQ398148 | This study | |
| DY091151 | Lepidoptera (Pupa) | OQ360025 | OQ363136 | – | – | OQ398151 | This study | |
| DY091152 | Lepidoptera (Pupa) | OQ360053 | OQ363137 | – | – | OQ398152 | This study | |
| Samsoniellahepiali | ICMM 82–2 | Fungi (O.sinensis) | – | MN576794 | MN576854 | MN576908 | MN576964 | Wang et al. (2020) |
| YFCC 661 | Fungi (O.sinensis) | – | MN576795 | MN576855 | MN576909 | MN576965 | Wang et al. (2020) | |
| Samsoniellahymenopterorum | A19521 | Vespidae (Bee) | MN128224 | – | MT642603 | MT642604 | MN101588 | Chen et al. (2020) |
| A19522 | Vespidae (Bee) | MN128081 | – | – | MN101590 | MN101591 | Chen et al. (2020) | |
| Samsoniellainthanonensis | TBRC 7915 | Lepidoptera (Pupa) | MF140761 | – | MF140790 | MF140815 | MF140849 | Mongkolsamrit et al. (2018) |
| TBRC 7916 | Lepidoptera (Pupa) | MF140760 | – | – | MF140814 | MF140848 | Mongkolsamrit et al. (2018) | |
| Samsoniellakunmingensis | YHH 16002 | Lepidoptera (Pupa) | – | MN576802 | MN576862 | MN576916 | MN576972 | Wang et al. (2020) |
| Samsoniellalanmaoa | YFCC 6148 | Lepidoptera (Pupa) | – | MN576789 | MN576849 | MN576903 | MN576959 | Wang et al. (2020) |
| Samsoniellalanmaoa | YFCC 6193 | Lepidoptera (Pupa) | – | MN576790 | MN576850 | MN576904 | MN576960 | Wang et al. (2020) |
| Samsoniellalepidopterorum | DL10071 | Lepidoptera (Pupa) | MN128076 | – | – | MN101593 | MN101594 | Chen et al. (2020) |
| DL10072 | Lepidoptera (Pupa) | MN128084 | – | – | MT642605 | MT642606 | Chen et al. (2020) | |
| Samsoniellaneopupicola | KY11321 | Lepidoptera (Pupa) | ON502843 | ON502839 | – | ON525432 | ON525433 | Chen et al. (2022) |
| KY11322 | Lepidoptera (Pupa) | ON502834 | ON502833 | – | ON525434 | ON525435 | Chen et al. (2022) | |
| Samsoniellapseudogunnii | GY407201 | Lepidoptera (Larvae) | MZ827470 | MZ827010 | – | MZ855239 | MZ855233 | Chen et al. (2021) |
| GY407202 | Lepidoptera (Larvae) | MZ831863 | MZ831865 | – | MZ855240 | MZ855234 | Chen et al. (2021) | |
| Samsoniellapseudotortricidae | YFCC 9052 | Lepidoptera (Pupa) | – | ON621677 | ON676509 | ON568692 | ON676521 | Wang et al. (2022) |
| YFCC 9053 | Lepidoptera (Pupa) | – | ON621678 | ON676510 | ON568693 | ON676522 | Wang et al. (2022) | |
| Samsoniellapupicola | DY101681 | Lepidoptera (Pupa) | MZ827085 | MZ827009 | – | MZ855237 | MZ855231 | Chen et al. (2021) |
| DY101682 | Lepidoptera (Pupa) | MZ827008 | MZ827635 | – | MZ855238 | MZ855232 | Chen et al. (2021) | |
| Samsoniellaramosa | YFCC 6020 | Limacodidae (Pupa) | – | MN576805 | MN576865 | MN576919 | MN576975 | Wang et al. (2020) |
| Samsoniellasinensis | YFCC 8766 | Lepidoptera (Larvae) | – | ON621679 | ON676511 | ON568694 | ON676523 | Wang et al. (2022) |
| YFCC 8767 | Dermaptera | – | ON621680 | ON676512 | ON568695 | ON676524 | Wang et al. (2022) | |
| YFCC 8768 | Dermaptera | – | ON621681 | ON676513 | ON568696 | ON676525 | Wang et al. (2022) | |
| Samsoniellatiankengensis | KY11741 | Lepidoptera (Pupa) | ON502840 | ON502838 | – | ON525436 | ON525437 | Chen et al. (2022) |
| KY11742 | Lepidoptera (Pupa) | ON502849 | ON502841 | – | ON525438 | ON525439 | Chen et al. (2022) | |
| Samsoniellatortricidae | YFCC 6013 | Tortricidae (Pupa) | – | MN576807 | MN576867 | MN576921 | MN576977 | Wang et al. (2020) |
| YFCC 6131 | Tortricidae (Pupa) | – | MN576806 | MN576866 | MN576920 | MN576976 | Wang et al. (2020) | |
| Samsoniellavallis | DY07241 | Lepidoptera (Pupa) | OR263159 | OR263306 | OR282772 | OR282774 | OR282778 | This study |
| DY07242 | Lepidoptera (Pupa) | OR263186 | OR263308 | – | OR282775 | OR282779 | This study | |
| DY091091 | Lepidoptera (Pupa) | OR263191 | OR263428 | – | – | OR282782 | This study | |
| DY091092 | Lepidoptera (Pupa) | OR263190 | OR263431 | – | – | OR282783 | This study | |
| Samsoniellawinandae | TBRC 17511 | Lepidoptera (Cocoon) | OM491228 | OM491231 | OM687901 | OM687899 | OM687896 | Crous et al. (2023) |
| TBRC 17512 | Limacodidae (Pupa) | OM491229 | OM491232 | OM687902 | OM687900 | OM687897 | Crous et al. (2023) | |
| Samsoniellayunnanensis | YFCC 1527 | Fungi (Cordycepscicadae) | – | MN576812 | MN576872 | MN576926 | MN576982 | Wang et al. (2020) |
| YFCC 1824 | Fungi (Cordycepscicadae) | – | MN576813 | MN576873 | MN576927 | MN576983 | Wang et al. (2020) | |
The new strains or species are in bold type.
Sequence alignment and phylogenetic analyses
DNASTAR Lasergene (version 6.0) was used to edit DNA sequences in this study. The ITS, LSU, RPB1, RPB2 and TEF sequences were downloaded from GenBank, based on Mongkolsamrit et al. (2018), Chen et al. (2020a, 2021a, 2022), Wang et al. (2020, 2022) and Crous et al. (2023) and others selected on the basis of BLASTn searches in GenBank. ITS sequences and other loci were aligned and edited by MAFFT v.7.037b (Katoh and Standley 2013) and MEGA6 (Tamura et al. 2013). Combined sequences of ITS, LSU, RPB1, RPB2 and TEF were obtained using SequenceMatrix v.1.7.8 (Vaidya et al. 2011). The model was selected for Bayesian analysis by ModelFinder (Kalyaanamoorthy et al. 2017) in PhyloSuite software (Zhang et al. 2020).
ITS sequences, other loci and the combined loci were analysed using Bayesian inference (BI) and maximum likelihood (ML) methods. For BI, a Markov chain Monte Carlo (MCMC) algorithm was used to generate phylogenetic trees with Bayesian probabilities using MrBayes v.3.2 (Ronquist et al. 2012) for the combined sequence datasets. The Bayesian analysis resulted in 20,001 trees after 10,000,000 generations. The first 4,000 trees, representing the burn-in phase of the analysis, were discarded, while the remaining 16,001 trees were used to calculate posterior probabilities in the majority rule consensus tree. After the analysis was finished, each run was examined using the programme Tracer v.1.5 (Drummond and Rambaut 2007) to determine burn-in and confirm that both runs had converged. ML analyses were constructed with IQ-TREE (Trifinopoulos et al. 2016), using an automatic selection of the model.
Genealogical Concordance Phylogenetic Species Recognition (GCPSR) analysis
The Genealogical Concordance Phylogenetic Species Recognition model was applied to analyse the related species. The pairwise homoplasy index (PHI) (Bruen et al. 2006) is a model test based on the fact that multiple gene phylogenies will be concordant between species and discordant due to recombination and mutations within a species. The test was performed in SplitsTree4 (Huson and Bryant 2006) as described by Quaedvlieg et al. (2014) to determine the recombination level within phylogenetically closely-related species using a two-locus concatenated dataset. The new species and their closely-related species were analysed using this model. The relationships between closely-related species were visualised by constructing a split graph, using both the LogDet transformation and splits decomposition options.
Result
Phylogenetic analyses
In the phylogenetic tree, Beauveriabassiana (Bals.-Criv.) Vuill. (ARSEF 1564) and B.brongniartii (Sacc.) Petch (ARSEF 617 and BCC 16585) were used as the outgroups. The concatenated sequences (ITS, LSU, RPB1, RPB2 and TEF) included 36 species (81 strains) and consisted of 3,579 (ITS, 501; LSU, 775; RPB1, 641; RPB2, 770; and TEF, 892) characters with gaps.
The final value of the highest scoring tree was –15,629.246, which was obtained from the ML analysis of the dataset (ITS+LSU+RPB1+RPB2+TEF). The parameters of the GTR model used to analyse the dataset were estimated, based on the following frequencies: A = 0.235, C = 0.273, G = 0.270, T = 0.222; substitution rates AC = 1.00000, AG = 1.93319, AT = 1.00000, CG = 1.00000, CT = 4.27255 and GT = 1.00000; as well as the gamma distribution shape parameter α = 0.509. The selected models for BI analysis were SYM+G4 (ITS+LSU+RPB1+RPB2+TEF). The phylogenetic trees (Fig. 1), constructed using the ML and BI analyses were largely congruent and strongly supported in most branches. Strains DY091021, DY091022, DY091031, DY091032, DY091151, and DY091152 were clustered into an independent subclade and formed a subclade with Samsoniellahaniana Hong Yu bis, Yao Wang & Z.Q. Wang with high statistical support (100% ML /1 PP). Strains DY09161, DY09162, DY07501 and DY07502 were clustered into an independent clade with high statistical support (100% ML/1 PP). Strains DY07241, DY07242, DY091091 and DY091092 were clustered with S.aurantia in a clade with high statistical support in ML analysis (94% ML).
Figure 1.
Phylogenetic relationships amongst the new strains and their allies based on multigene dataset (ITS, LSU, RPB1, RPB2 and TEF). Statistical support values (≥ 70%/0.70) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
A two-locus concatenated dataset (LSU and TEF) was used to determine the recombination level within Samsonielladuyunensis (DY09161 and DY07501), Samsoniellavallis (DY07241 and DY091091), S.haniana (YFCC 8769, DY091031, DY091021 and DY091151) and S.aurantia (TBRC 7271). Chaiwan et al. (2022) noted that if the PHI is below the 0.05 threshold (Φw < 0.05), it indicates that there is significant recombination in the dataset. This means that related species in a group and recombination levels are not different. If the PHI is above the 0.05 threshold (Φw > 0.05), it indicates that it is not significant, which means the related species in a group level are different. The result of the pairwise homoplasy index (PHI) test of Samsoniellaaurantia, S.duyunensis, S.haniana and S.vallis was 1.0 and revealed that the four species were different (Fig. 2).
Figure 2.
Results of the pairwise homoplasy index (PHI) test of closely-related species using both LogDet transformation and splits decomposition. PHI test results (Փw) < 0.05 indicate significant recombination within the dataset. The new strains or species are in bold type.
Taxonomy
. Samsoniella duyunensis
W.H. Chen, Y.F. Han & J.D. Liang sp. nov.
90CA95AD-D8C9-5498-9FEC-6E2375C50DFA
847492
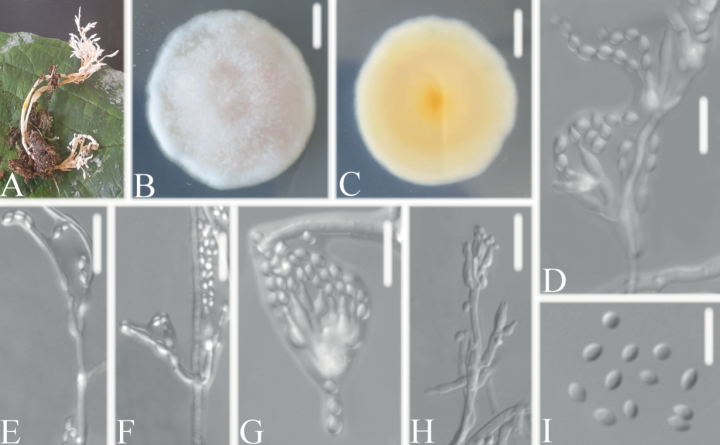
Figure 3.
SamsonielladuyunensisA infected ant (Formicidae) B, CPDA culture plate showing top (B) and reverse (C) sides of the colony D–J Phialides and conidia. Scale bars: 10 mm (B, C); 10 μm (D–J).
Type.
China, Guizhou, Qiannan Buyei and Miao Autonomous Prefecture, Duyun City, Mayao River Valley (26°22'8.3748"N, 107°23'16.96"E). On an ant (Formicidae), buried in soil, 4 September 2021, Wanhao Chen, GZAC DY0916 (holotype), ex-type living cultures, DY09161.
Description.
Synnemata arising from the host, irregularly branched, conidia in abundance at the apex. Colonies on PDA, attaining a diameter of 35–38 mm after 14 days at 25 °C, white, consisting of a basal felt, floccose hyphal overgrowth; reverse yellowish. Hyphae septate, hyaline, pale pink in the middle part, smooth-walled, 0.8–1.4 μm wide. Conidiophores hyaline, smooth-walled, with single phialide or whorls of 2–4 phialides or verticillium-like from hyphae directly, 10.0–21.3 × 1.7–1.9 μm. Phialides cylindrical to ellipsoidal, somewhat inflated base, 5.3–9.1 × 1.3–1.6 μm, tapering to a thin neck. Conidia hyaline, smooth-walled, fusiform to ellipsoidal, 2.1–2.9 × 1.1–1.7 μm, forming divergent and basipetal chains. Sexual state not observed.
Host.
Ant (Formicidae).
Etymology.
Referring to its location in Duyun City.
Additional material examined.
China, Guizhou, Qiannan Buyei and Miao Autonomous Prefecture, Duyun City, Mayao River Valley (26°22'8.3748"N, 107°23'16.96"E). On an ant (Formicidae), buried in soil, 4 September 2021, Wanhao Chen, DY09162 (living culture). On a pupa (Lepidoptera) clinging to fallen leaves, 30 July 2022, Wanhao Chen, GZAC DY0750 (specimen), DY07501 and DY07502 (living culture). On an ant (Formicidae) clinging to fallen leaves, 4 September 2021, Wanhao Chen, DY0906 (specimen).
Remarks.
Samsonielladuyunensis was easily identified as Samsoniella, based on the BLASTn result in NCBI and the phylogenetic analysis of the combined datasets (ITS, LSU, RPB1, RPB2 and TEF) (Fig. 1) and clustered into an independent clade. Comparing with the typical characteristics of the known species and the keys of Samsoniella species (Wang et al. 2022), S.duyunensis has a close relationship with S.coccinellidicola and S.sinensis by absence of sexual state, presence of synnemata and irregularly branched, moderately grow of colony. However, it is distinguished from S.coccinellidicola (phialides: 6.0–14.1 × 1.0–2.0 μm; conidia: fusiform or oval, 1.8–3.0 × 1.3–2.0 μm; host, adults of Coccinellidae) by shorter phialides, smaller conidia and its ant host and distinguished from S.sinensis (conidia: spherical, elliptical or fusiform; host: larva of Lepidoptera) by fusiform to ellipsoidal conidia and its ant host.
. Samsoniella vallis
W.H. Chen, Y.F. Han & J.D. Liang sp. nov.
72AC4F73-B0B2-5558-8567-BE546FF52AF1
847493
Figure 4.
SamsoniellavallisA infected pupa (Lepidoptera) B, CPDA culture plate showing top (B) and reverse (C) sides of the colony D–M phialides and conidia. Scale bars: 10 mm (B, C); 10 μm (D–M).
Type.
China, Guizhou, Qiannan Buyei and Miao Autonomous Prefecture, Duyun City, Mayao River Valley (26°22'8.3748"N, 107°23'16.96"E). On a pupa (Lepidoptera) clinging to fallen leaves, 30 July 2022, Wanhao Chen, GZAC DY0724 (holotype), ex-type living cultures, DY07241.
Description.
Synnemata arising from every part of the body of the pupa host. Synnemata erect, usually irregularly branched at the apex, conidia in abundance at the apex. Colonies on PDA, attaining a diameter of 31–37 mm after 14 days at 25 °C, white, consisting of a basal felt, floccose hyphal overgrowth; reverse yellowish. Hyphae septate, hyaline, smooth-walled, 2.1–3.0 μm wide. Conidiophores hyaline, smooth-walled, with single phialide or whorls of 2–4 phialides or verticillium-like from hyphae directly, 11.3–22.1 × 1.3–1.4 μm. Phialides cylindrical to ellipsoidal, somewhat inflated base, 7.2–8.1 × 2.8–3.2 μm, tapering to a thin neck. Conidia hyaline, smooth-walled, fusiform to ellipsoidal, 2.3–3.1 × 1.5–2.1 μm, forming divergent and basipetal chains. Sexual state not observed.
Host.
Pupa (Lepidoptera).
Etymology.
Referring to its location in Mayao River Valley.
Additional material examined.
China, Guizhou, Qiannan Buyei and Miao Autonomous Prefecture, Duyun City, Mayao River Valley (26°22'8.3748"N, 107°23'16.96"E). On a pupa (Lepidoptera) clinging to fallen leaves, 30 July 2022, Wanhao Chen, DY07242 (living culture); China, Guizhou, Qiannan Buyei and Miao Autonomous Prefecture, Duyun City, Mayao River Valley (26°22'8.3748"N, 107°23'16.96"E). On a pupa (Lepidoptera) clinging to fallen leaves, 4 September 2021, Wanhao Chen, GZAC DY09109 (specimen), DY091091 and DY091092 (living culture). On a pupa (Lepidoptera) clinging to fallen leaves, 4 September 2021, Wanhao Chen, GZAC DY0909 (specimen).
Remarks.
Samsoniellavallis was easily identified as Samsoniella, based on the BLASTn result in NCBI and the phylogenetic analysis of the combined datasets (ITS, LSU, RPB1, RPB2 and TEF) (Fig. 1) and clustered with S.aurantia in a clade. However, it is distinguished from S.aurantia (phialides: 5–13 × 2–3 μm; conidia: fusiform or oval, 2–4 × 1–2 μm) by shorter phialides, smaller fusiform to ellipsoidal conidia. Comparing with the typical characteristics of the known species and the keys of Samsoniella species (Wang et al. 2022), S.vallis has a close relationship with S.coccinellidicola and S.sinensis by absence of sexual state, presence of synnemata and irregularly branched, moderate growth of colony. However, it is distinguished from S.coccinellidicola (phialides: 6.0–14.1 × 1.0–2.0 μm; conidia: fusiform or oval; host, adults of Coccinellidae) by shorter phialides, fusiform to ellipsoidal conidia and its pupa host and distinguished from S.sinensis (phialides: 5.6–9.3 × 1.5–2.1 μm, conidia: spherical, elliptical or fusiform) by fusiform to ellipsoidal conidia and shorter phialides.
. Samsoniella haniana
Hong Yu bis, Yao Wang & Z.Q. Wang, in Wang, Wang, Dong, Fan, Dao & Yu, Journal of Fungi 8: 20, 2022
B9DF58D0-7DDF-5BEF-9C95-B07240D0727F
Figure 5.
SamsoniellahanianaA infected pupa (Lepidoptera) B, CPDA culture plate showing top (B) and reverse (C) sides of the colony D–I phialides and conidia. Scale bars: 10 mm (B, C); 10 μm (D–I).
Description.
Synnemata arising from every part of the body of the pupa host. Synnemata erect, usually irregularly branched at the apex, Isaria-like morph producing a mass of conidia at the branch apex, powdery and floccose. Colonies on PDA, attaining a diameter of 32–35 mm after 14 days at 25 °C, white, consisting of a basal felt, floccose hyphal overgrowth; reverse yellowish. Hyphae septate, hyaline, smooth-walled, 1.3–1.8 μm wide. Conidiophores hyaline, smooth-walled, with single phialide or whorls of 2–8 phialides or verticillium-like from hyphae directly, 16.1–23.9 × 1.7–2.2 μm. Phialides consisting of a cylindrical to ellipsoidal, somewhat inflated base, 5.0–6.9 × 1.8–2.5 μm, tapering to a thin neck. Conidia hyaline, smooth-walled, fusiform to subglobose, 1.7–3.4 × 1.7–2.1 μm, forming divergent and basipetal chains. Sexual state not observed.
Host.
Pupa (Lepidoptera).
Material examined.
China, Guizhou, Qiannan Buyei and Miao Autonomous Prefecture, Duyun City, Mayao River Valley (26°22'8.3748"N, 107°23'16.96"E). On a pupa (Lepidoptera), buried in soil, 4 September 2021, Wanhao Chen, GZAC DY09103 (specimen), DY091031, DY091032 (living culture); On a ladybug (Coccinellidae); On the moss, 4 September 2021, Wanhao Chen, GZAC DY09102 (specimen), DY091021, DY091022 (living culture); On a caterpillar (Lepidoptera), buried in soil, 4 September 2021, Wanhao Chen, GZAC DY09115 (specimen), DY091151, DY091152 (living culture); On a pupa (Lepidoptera), buried in soil, 4 September 2021, Wanhao Chen, GZAC DY0929, DY09158 (specimen).
Remarks.
Strains DY091021, DY091022, DY091031, DY091032, DY091151 and DY091152 were identified as belonging to Samsoniella, based on the BLASTn result and the phylogenetic analyses (Fig. 1) and clustered with S.haniana in a subclade with high statistical support (100% ML/ 1 PP). The characteristics of those strains were very closely linked with S.haniana, which had fusiform or oval conidia (2.3–3.7 × 1.2–2.8 μm), phialide (5.4–12.1 × 1.2–2.9 μm) and a pupa host. Thus, the molecular phylogenetic results and morphologically-based conclusions supported the idea that strains DY091021, DY091022, DY091031, DY091032, DY091151 and DY091152 were S.haniana.
Discussion
Samsoniella species are widely distributed and commonly isolated from soil, insects and spiders or as a fungicolous (Mongkolsamrit et al. 2018; Chen et al. 2020a, 2021a, 2022; Wang et al. 2020, 2022; Crous et al. 2023). Amongst 29 species, S.alboaurantia, S.alpina H. Yu et al., S.antleroides H. Yu et al., S.aurantia, S.cardinalis H. Yu et al., S.cristata H. Yu et al., S.erucae W.H. Chen et al., S.farinospora Hong Yu bis et al., S.guizhouensis W.H. Chen et al., S.haniana, S.hepiali (Q.T. Chen & R.Q. Dai ex R.Q. Dai, X.M. Li, A.J. Shao, Shu F. Lin, J.L. Lan, Wei H. Chen & C.Y. Shen) H. Yu et al., S.inthanonensis, S.kunmingensis H. Yu et al., S.lanmaoa H. Yu et al., S.lepidopterorum W.H. Chen et al., S.neopupicola W.H. Chen et al., S.pseudogunnii W.H. Chen et al., S.pseudotortricidae Hong Yu bis et al., S.pupicola W.H. Chen et al., S.ramosa H. Yu et al., S.sinensis Hong Yu bis et al., S.tiankengensis W.H. Chen et al., S.tortricidae H. Yu et al., S.winandae Mongkols., Noisrip. & Luangsa-ard and S.yunnanensis H. Yu et al. were reported as a fungal pathogen of lepidoptera insects. The host of S.coccinellidicola Hong Yu bis et al., S.coleopterorum W.H. Chen et al., S.formicae W.H. Chen et al. and S.hymenopterorum W.H. Chen et al. belonged to Coleoptera and Hymenoptera, respectively. In addition, the substrates of S.alboaurantia, S.farinospora and S.hepiali were soil, spider and fungi, respectively. Here, we reported Samsoniella species with two different hosts from the valley habitat. More Samsoniella species with different hosts or substrates will be reported from diverse habitats.
The taxonomic delimitation of Samsoniella was originally based on morphological characteristics and a multi-locus phylogenetic analysis. In the present study, the phylogenetic analysis of a single locus of an individual gene or gene fragment of ITS, LSU, RPB1, RPB2 and TEF was tested for the new species (Suppl. materials 1–5) and only the TEF could distinguish the new species. However, the new species S.vallis could not form an independent clade and clustered with S.aurantia as a subclade. A PHI test was added and could solve the taxonomic delimitation of S.vallis and S.aurantia. Thus, we recommend that the TEF locus should be used to distinguish the cryptic Samsoniella species and multiple approaches should be used for the further confirmation of a cryptic species.
Generally, species diversity of entomopathogenic fungi were mainly investigated in nature forest and grassland reservations and crop fields (Chen et al. 2019b, 2020b; He 2019; Fan 2020; Zhao et al. 2020, 2021; Zhang et al. 2021). Samsoniella species have often been reported from forests, but rarely found in special karst eco-environments, such as Tiankeng, valleys and caves. Chen et al. (2022) reported five new Samsoniella species from Monkey-Ear Tiankeng and provided new insights into the richness of species diversity of Samsoniella in such special habitat. This research provided further evidence of the richness of Samsoniella species in karst eco-environments. The Samsoniella species diversity should be extensively investigated in diverse habitats including karst.
Supplementary Material
Citation
Chen W-H, Liang J-D, Ren X-X, Zhao J-H, Han Y-F (2023) Two new species of Samsoniella (Cordycipitaceae, Hypocreales) from the Mayao River Valley, Guizhou, China. MycoKeys 99: 209–226. https://doi.org/10.3897/mycokeys.99.109961
Funding Statement
This work was funded by National Natural Science Foundation of China (31860002, 81960692), High-level Innovative Talents Training Object in Guizhou Province (Qiankehepingtairencai [2020]6005), Construction Program of Guizhou Engineering Research Center (Qian Fa Gai Gao Ji 2020-896).
Additional information
Conflict of interest
The authors have declared that no competing interests exist.
Ethical statement
No ethical statement was reported.
Funding
This work was funded by National Natural Science Foundation of China (31860002, 81960692), High-level Innovative Talents Training Object in Guizhou Province (Qiankehepingtairencai [2020]6005), Construction Program of Guizhou Engineering Research Center (Qian Fa Gai Gao Ji 2020-896).
Author contributions
Data curation: CW, XR, JL. Funding acquisition: JZ, YH, CW, JL. Writing – original draft: XR, CW, JL. Writing – review and editing: YH, JZ.
Author ORCIDs
Wan-Hao Chen https://orcid.org/0000-0001-7240-6841
Jian-Dong Liang https://orcid.org/0000-0002-3939-3900
Jie-Hong Zhao https://orcid.org/0000-0003-2972-382X
Yan-Feng Han https://orcid.org/0000-0002-8646-3975
Data availability
All of the data that support the findings of this study are available in the main text or Supplementary Information.
Supplementary materials
Phylogenetic relationships among the new strains and their allies based on ITS sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
Phylogenetic relationships among the new strains and their allies based on LSU sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
Phylogenetic relationships among the new strains and their allies based on RPB1 sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
Phylogenetic relationships among the new strains and their allies based on RPB2 sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
Phylogenetic relationships among the new strains and their allies based on TEF sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
References
- Aini AN, Mongkolsamrit S, Wijanarka W, Thanakitpipattana D, Luangsa-ard JJ, Budiharjo A. (2020) Diversity of Akanthomyces on moths (Lepidoptera) in Thailand. MycoKeys 71: 1–22. 10.3897/mycokeys.71.55126 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruen TC, Philippe H, Bryant D. (2006) A simple and robust statistical test for detecting the presence of recombination. Genetics 172(4): 2665–2681. 10.1534/genetics.105.048975 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castlebury LA, Rossman AY, Sung GH, Hyten AS, Spatafora JW. (2004) Multigene phylogeny reveals new lineage for Stachybotryschartarum, the indoor air fungus. Mycological Research 108(8): 864–872. 10.1017/S0953756204000607 [DOI] [PubMed] [Google Scholar]
- Chaiwan N, Jeewon R, Pem D, Jayawardena RS, Nazurally N, Mapook A, Promputtha I, Hyde KD. (2022) Fungal species from Rhododendron sp.: Discosiarhododendricola sp. nov., Neopestalotiopsisrhododendricola sp. nov. and Diaporthenobilis as a new host record. Journal of Fungi 8(9): е907. 10.3390/jof8090907 [DOI] [PMC free article] [PubMed]
- Chen WH, Liu C, Han YF, Liang JD, Tian WY, Liang ZQ. (2019) Three novel insect-associated species of Simplicillium (Cordycipitaceae, Hypocreales) from Southwest China. MycoKeys 58: 83–102. 10.3897/mycokeys.58.37176 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen ZH, Wang YB, Dai YD, Chen K, Xu L, He QC. (2019) Species diversity and seasonal fluctuation of entomogenous fungi of Ascomycota in Taibaoshan Forest Park in western Yunnan. Shengwu Duoyangxing 27(9): 993–1001. 10.17520/biods.2019135 [DOI] [Google Scholar]
- Chen MJ, Chen WY, Liu YJ, Hou NS, Lin Y. (2020) Species diversity of entomopathogenic fungi in Wanfo Mountain National Forest Park. Anhui Nongye Daxue Xuebao 47(2): 294–298. 10.13610/j.cnki.1672-352x.20200518.021 [DOI] [Google Scholar]
- Chen WH, Han YF, Liang JD, Tian WY, Liang ZQ. (2020) Morphological and phylogenetic characterisations reveal three new species of Samsoniella (Cordycipitaceae, Hypocreales) from Guizhou, China. MycoKeys 74: 1–15. 10.3897/mycokeys.74.56655 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen W, Liang J, Ren X, Zhao J, Han Y, Liang Z. (2021) Cryptic diversity of isaria-like species in Guizhou, China. Life 11(10): е1093. 10.3390/life11101093 [DOI] [PMC free article] [PubMed]
- Chen WH, Liang JD, Ren XX, Zhao JH, Han YF, Liang ZQ. (2022) Species diversity of Cordyceps-like fungi in the Tiankeng Karst Region of China. Microbiology Spectrum 10(5): e01975–e22. 10.1128/spectrum.01975-22 [DOI] [PMC free article] [PubMed]
- Crous PW, Osieck ER, Shivas RG, Tan YP, Bishop-Hurley SL, Esteve-Raventós F, Larsson E, Luangsa-ard JJ, Pancorbo F, Balashov S, Baseia IG, Boekhout T, Chandranayaka S, Cowan DA, Cruz RHSF, Czachura P, De la Peña-Lastra S, Dovana F, Drury B, Fell J, Flakus A, Fotedar R, Jurjević Ž, Kolecka A, Mack J, Maggs-Kölling G, Mahadevakumar S, Mateos A, Mongkolsamrit S, Noisripoom W, Plaza M, Overy DP, Piątek M, Sandoval-Denis M, Vauras J, Wingfield MJ, Abell SE, Ahmadpour A, Akulov A, Alavi F, Alavi Z, Altes A, Alvarado P, Anand G, Ashtekar N, Assyov B, Banc-Prandi G, Barbosa KD, Barreto GG, Bellanger JM, Bezerra JL, Bhat DJ, Bilanski P, Bose T, Bozok F, Chaves J, Costa-Rezende DH, Danteswari C, Darmostuk V, Delgado G, Denman S, Eichmeier A, Etayo J, Eyssartier G, Faulwetter S, Ganga KGG, Ghosta Y, Goh J, Góis JS, Gramaje D, Granit L, Groenewald M, Gulden G, Gusmão LFP, Hammerbacher A, Heidarian Z, Hywel-Jones N, Jankowiak R, Kaliyaperumal M, Kaygusuz O, Kezo K, Khonsanit A, Kumar S, Kuo CH, Læssøe T, Latha KPD, Loizides M, Luo SM, Maciá-Vicente JG, Manimohan P, Marbach PAS, Marinho P, Marney TS, Marques G, Martín MP, Miller AN, Mondello F, Moreno G, Mufeeda KT, Mun HY, Nau T, Nkomo T, Okrasińska A, Oliveira JPAF, Oliveira RL, Ortiz DA, Pawłowska J, Pérez-De-Gregorio MÀ, Podile AR, Portugal A, Privitera N, Rajeshkumar KC, Rauf I, Rian B, Rigueiro-Rodríguez A, Rivas-Torres GF, Rodriguez-Flakus P, Romero-Gordillo M, Saar I, Saba M, Santos CD, Sarma PVSRN, Siquier JL, Sleiman S, Spetik M, Sridhar KR, Stryjak-Bogacka M, Szczepańska K, Taşkın H, Tennakoon DS, Thanakitpipattana D, Trovao J, Türkekul İ, van Iperen AL, van’t Hof P, Vasquez G, Visagie CM, Wingfield BD, Wong PTW, Yang WX, Yarar M, Yarden O, Yilmaz N, Zhang N, Zhu YN, Groenewald JZ. (2023) Fungal Planet description sheets: 1478–1549. Persoonia 50(1): 158–310. 10.3767/persoonia.2023.50.05 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drummond A, Rambaut A. (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evolutionary Biology 7(1): e214. 10.1186/1471-2148-7-214 [DOI] [PMC free article] [PubMed]
- Fan Q. (2020) A preliminary study on taxonomy and phylogeny of Cordyceps sensu lato in Kunming. Thesis for Master’ s degree, Yunnan University.
- He RH. (2019) Studies on morphology, genetic differentiation and floristic fungi diversity of Ophiocordycepsnigrella. Thesis for Master’ s degree, Shanxi University.
- Huson DH, Bryant D. (2006) Application of phylogenetic networks in evolutionary studies. Molecular Biology and Evolution 23(2): 254–267. 10.1093/molbev/msj030 [DOI] [PubMed] [Google Scholar]
- Kalyaanamoorthy S, Minh BQ, Wong TK, Von Haeseler A, Jermiin LS. (2017) ModelFinder: Fast model selection for accurate phylogenetic estimates. Nature Methods 14(6): 587–589. 10.1038/nmeth.4285 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katoh K, Standley DM. (2013) MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Molecular Biology and Evolution 30(4): 772–780. 10.1093/molbev/mst010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kepler RM, Sung GH, Ban S, Nakagiri A, Chen MJ, Huang B, Li Z, Spatafora JW. (2012) New teleomorph combinations in the entomopathogenic genus Metacordyceps. Mycologia 104(1): 182–197. 10.3852/11-070 [DOI] [PubMed] [Google Scholar]
- Liang JD, Han YF, Zhang JW, Du W, Liang ZQ, Li ZZ. (2011) Optimal culture conditions for keratinase production by a novel thermophilic Myceliophthorathermophila strain GZUIFR-H49-1. Journal of Applied Microbiology 110(4): 871–880. 10.1111/j.1365-2672.2011.04949.x [DOI] [PubMed] [Google Scholar]
- Liu YJ, Whelen S, Hall BD. (1999) Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerase II subunit. Molecular Biology and Evolution 16(12): 1799–1808. 10.1093/oxfordjournals.molbev.a026092 [DOI] [PubMed] [Google Scholar]
- Mongkolsamrit S, Noisripoom W, Thanakitpipattana D, Wuthikun T, Spatafora JW, Luangsa-ard JJ. (2018) Disentangling cryptic species with isaria-like morphs in Cordycipitaceae. Mycologia 110(1): 230–257. 10.1080/00275514.2018.1446651 [DOI] [PubMed] [Google Scholar]
- Quaedvlieg W, Binder M, Groenewald JZ, Summerell BA, Carnegie AJ, Burgess TI, Crous W. (2014) Introducing the consolidated species concept to resolve species in the Teratosphaeriaceae. Persoonia 33(1): 1–40. 10.3767/003158514X681981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rehner SA, Minnis AM, Sung GH, Luangsa-ard JJ, Devotto L, Humber RA. (2011) Phylogeny and systematics of the anamorphic, entomopathogenic genus Beauveria. Mycologia 103(5): 1055–1073. 10.3852/10-302 [DOI] [PubMed] [Google Scholar]
- Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP. (2012) MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology 61(3): 539–542. 10.1093/sysbio/sys029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spatafora JW, Sung GH, Sung JM, Hywel-Jones NL, White Jr JF. (2007) Phylogenetic evidence for an animal pathogen origin of ergot and the grass endophytes. Molecular Ecology 16(8): 1701–1711. 10.1111/j.1365-294X.2007.03225.x [DOI] [PubMed] [Google Scholar]
- Sung GH, Hywel-Jones NL, Sung JM, Luangsa-ard JJ, Shrestha B, Spatafora JW. (2007) Phylogenetic classification of Cordyceps and the clavicipitaceous fungi. Studies in Mycology 57: 5–59. 10.3114/sim.2007.57.01 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. (2013) MEGA6: Molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution 30(12): 2725–2729. 10.1093/molbev/mst197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trifinopoulos J, Nguyen LT, von Haeseler A, Minh BQ. (2016) W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Research 44(W1): W232–W235. 10.1093/nar/gkw256 [DOI] [PMC free article] [PubMed]
- Vaidya G, Lohman DJ, Meier R. (2011) SequenceMatrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 27(2): 171–180. 10.1111/j.1096-0031.2010.00329.x [DOI] [PubMed] [Google Scholar]
- Vilgalys R, Hester M. (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. Journal of Bacteriology 172(8): 4238–4246. 10.1128/jb.172.8.4238-4246.1990 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang YB, Wang Y, Fan Q, Duan DE, Zhang GD, Dai RQ, Dai YD, Zeng WB, Chen ZH, Li DD, Tang DX, Xu ZH, Sun T, Nguyen T, Tran N, Dao V, Zhang CM, Huang LD, Liu YJ, Zhang XM, Yang DR, Sanjuan T, Liu XZ, Yang ZL, Yu H. (2020) Multigene phylogeny of the family Cordycipitaceae (Hypocreales): New taxa and the new systematic position of the Chinese cordycipitoid fungus Paecilomyceshepiali. Fungal Diversity 103(1): 1–46. 10.1007/s13225-020-00457-3 [DOI] [Google Scholar]
- Wang Z, Wang Y, Dong Q, Fan Q, Dao VM, Yu H. (2022) Morphological and phylogenetic characterization reveals five new species of Samsoniella (Cordycipitaceae, Hypocreales). Journal of Fungi 8(7): е747. 10.3390/jof8070747 [DOI] [PMC free article] [PubMed]
- White TJ, Bruns T, Lee S, Taylor J. (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ. (Eds) PCR protocols: a guide to methods and applications.Academic Press, New York, 315–322. 10.1016/B978-0-12-372180-8.50042-1 [DOI]
- Zhang D, Gao F, Jakovlic I, Zou H, Zhang J, Li WX, Wang GT. (2020) PhyloSuite: An integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Molecular Ecology Resources 20(1): 348–355. 10.1111/1755-0998.13096 [DOI] [PubMed] [Google Scholar]
- Zhang J, Lu JJ, Zhao XC, Kang C, Wu ZJ, Deng CY, Li Y. (2021) A preliminary report on species diversity of Cordyceps s. I. in Doupeng Mountain area. Guizhou Province. Acta Edulis Fungi 28(6): 174–187. 10.16488/j.cnki.1005-9873.2021.06.022 [DOI] [Google Scholar]
- Zhao ZY, Wang YB, Wang ZQ, Tang DX, Geng YP, Yu H. (2020) Species diversity of Cordyceps sensu lato in Weishan, Yunnan. Redai Yaredai Zhiwu Xuebao 28(5): 455–462. [Google Scholar]
- Zhao ZY, Wang ZQ, Fan Q, Tang DX, Sun T, Wang YB, Geng YP, Yu H. (2021) Species diversity of Cordyceps sensu lato in the Taiji Mountains Nature Reserve of Dali, Yunnan. Ecologic Science 40(3): 128–135. 10.14108/j.cnki.1008-8873.2021.03.016 [DOI] [Google Scholar]
- Zou X, Liu AY, Liang ZQ, Han YF, Yang M. (2010) Hirsutellaliboensis, a new entomopathogenic species affecting Cossidae (Lepidoptera) in China. Mycotaxon 111(1): 39–44. 10.5248/111.39 [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Phylogenetic relationships among the new strains and their allies based on ITS sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
Phylogenetic relationships among the new strains and their allies based on LSU sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
Phylogenetic relationships among the new strains and their allies based on RPB1 sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
Phylogenetic relationships among the new strains and their allies based on RPB2 sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
Phylogenetic relationships among the new strains and their allies based on TEF sequence
This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.
Wan-Hao Chen, Jian-Dong Liang, Xiu-Xiu Ren, Jie-Hong Zhao, Yan-Feng Han
Data type
tiff
Explanation note
Statistical support values (≥ 50%/0.50) are shown at the nodes for ML bootstrap support/BI posterior probabilities. The new strains or species are in bold type.
Data Availability Statement
All of the data that support the findings of this study are available in the main text or Supplementary Information.