Abstract
The WRAMP structure is a protein network associated with tail-end actomyosin contractility, membrane retraction, and directional persistence during cell migration. A marker of WRAMP structures is melanoma cell adhesion molecule (MCAM) which dynamically polarizes to the cell rear. However, factors that mediate MCAM polarization are still unknown. In this study, BioID using MCAM as bait identifies the ERM family proteins, moesin, ezrin, and radixin, as WRAMP structure components. We also present a novel image analysis pipeline, Protein Polarity by Percentile (“3P”), which classifies protein polarization using machine learning and facilitates quantitative analysis. Using 3P, we find that depletion of moesin, and to a lesser extent ezrin, decreases the proportion of cells with polarized MCAM. Furthermore, although copolarized MCAM and ERM proteins show high spatial overlap, 3P identifies subpopulations with ERM proteins closer to the cell periphery. Live-cell imaging confirms that MCAM and ERM protein polarization is tightly coordinated, but ERM proteins enrich at the cell edge first. Finally, deletion of a juxtamembrane segment in MCAM previously shown to promote ERM protein interactions impedes MCAM polarization. Our findings highlight the requirement for ERM proteins in recruitment of MCAM to WRAMP structures and an advanced computational tool to characterize protein polarization.
Melanoma cell adhesion molecule (MCAM) dynamically polarizes in melanoma cells as part of a protein network associated with persistent cell migration, but the proteins which mediate this polarization are unknown.
The authors used proteomics to identify proteins proximal to MCAM and developed a novel image analysis method to analyze protein polarity using machine learning. They found that the ezrin/radixin/moesin (ERM) family proteins copolarize with MCAM and promote its polarization.
This highlights a role for the ERM proteins in the assembly of polarized protein networks and advances computational methods for analyzing protein polarity.
INTRODUCTION
The spatial segregation of proteins underlies many cellular processes, including tissue morphogenesis, cell division, cell−cell communication, and migration (Mellman and Nelson, 2008; Allam et al., 2018). Disruption of protein polarization can lead to disease states such as birth defects (Butler and Wallingford, 2017), kidney and intestinal dysfunction (Schnell and Carroll, 2016; Klunder et al., 2017), and neurological disorders (Zhang and Wei, 2022). Importantly, the altered polarization of proteins involved in cell migration also underlies cancer cell invasion and metastasis (Etienne-Manneville, 2008). In migrating cells, actin-driven protrusion extends the leading edge membrane, while myosin activity at the cell rear drives membrane retraction and provides the contractile force for forward cell movement (Li et al., 2020; Shellard and Mayor, 2020; SenGupta et al., 2021). Although detailed models of cell migration have focused on events at the leading edge (Ridley et al., 2003; Petrie et al., 2009), symmetry breaking and movement can also be initiated at the rear of the cell (Mseka et al., 2007; Yam et al., 2007; Cramer, 2010), where spatial enrichment of myosin-II is important for initiating and sustaining cell polarity and directional migration (Vicente-Manzanares et al., 2007, 2008; Shutova et al., 2017; Tsai et al., 2019; Hadjitheodorou et al., 2021).
We previously described a network of rear-polarized proteins involved in directional migration, named the WRAMP (Wnt5a-Receptor-Actin-Myosin-Polarity) structure. Members of this network include cell adhesion receptors, such as the melanoma cell adhesion molecule glycoprotein (MCAM, aka MUC18 or CD146), together with F-actin and myosin-IIB (Witze et al., 2008). MCAM promotes metastasis and tumor progression in melanoma (Lehmann et al., 1989; Luca et al., 1993; McGary et al., 2002; Wu et al., 2008) through its functions as an adhesion protein and cell surface receptor (Lei et al., 2015; Wang et al., 2020). Its expression is also associated with malignancy and metastasis in prostate and breast cancers (Fritzsche et al., 2008; Wu et al., 2011; Zeng et al., 2012). In WRAMP structures, MCAM polarizes to the rear of cells during 2D and 3D cell migration, where its movement to the cell membrane is followed by membrane retraction (Witze et al., 2008, 2013; Connacher et al., 2017). WRAMP structures dynamically assemble and disassemble in cells, and migrating cells show increased directional persistence during periods when they are present. Moreover, these structures disassemble and then repolarize to new cell locations prior to directional changes, suggesting that MCAM may help establish the rear of the cell and determine the direction of migration (Connacher et al., 2017). WRAMP structures have been associated with other cytoskeletal and actin-binding proteins that copolarize with MCAM, F-actin, and myosin-II, and a role of cortical endoplasmic reticulum and localized Ca2+ influx in membrane detachment has been suggested (Mladinich and Huttenlocher, 2013; Witze et al., 2013).
These findings suggest a function for WRAMP structures in establishing the cell rear and promoting directional persistence during migration. However, much remains unknown about the protein components within WRAMP structures, their organization, and their order of assembly. Therefore, in this study we investigated the MCAM interactome using BioID proximity proteomics and identified moesin (MSN), ezrin (EZR), and radixin (RDX) as novel WRAMP structure components. Ezrin/radixin/moesin (ERM) family proteins are known to link cell adhesion proteins to the actin cytoskeleton. All copolarized with MCAM, and their siRNA depletion, particularly with siRNA-MSN, interfered with MCAM polarization, suggesting their functional role in WRAMP structure assembly. To facilitate this analysis, we developed a novel quantitative image analysis pipeline, named Protein Polarity by Percentile (3P), which couples custom metrics for cell polarity with machine learning (ML) to classify and describe protein polarization. Quantitative measurements of colocalization and polarization showed high spatiotemporal association of MCAM and the ERM proteins but identified cell subpopulations with closer apposition of ERM proteins to the cell periphery. Live imaging confirmed that ERM proteins can polarize to the cell edge before MCAM. Finally, deletion of a known ERM-binding site in the juxtamembrane region of MCAM led to decreased polarization of MCAM as well as ERM proteins. Together, these findings suggest a cooperative model for WRAMP structure assembly, where polarization of ERM proteins precedes the recruitment of MCAM, which in turn stabilizes the polarization of ERM proteins.
RESULTS
BioID identifies ERM proteins in the MCAM-proximal proteome
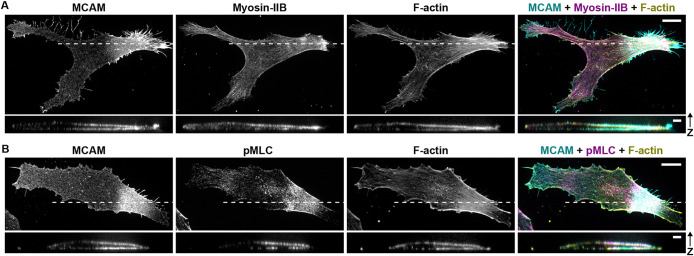
Human WM239a melanoma cells are relatively nonmotile but exhibit dynamic MCAM polarization, with 30−50% of cells in this study showing polarized MCAM at a fixed point in time, providing a tractable system for examining WRAMP structures. Figure 1A illustrates the copolarization of MCAM, F-actin, and myosin-IIB at WRAMP structures, where, as seen previously (Connacher et al., 2017), enriched myosin and F-actin showed filamentous morphologies with an overlapping but distinct pattern relative to MCAM. Cell imaging controls are shown in Supplemental Figure S1, A−C. Phosphorylated myosin light chain 2 (Ser19; pMLC), an indicator of myosin activity, was also enriched with polarized MCAM, supporting its association with membrane retraction through actomyosin contractility (Figure 1B). Examination of the 3D organization of MCAM, F-actin, and myosin markers by Z-stacking confocal imaging showed that polarized MCAM was largely enriched in basal and apical plasma membranes and decorated with myosin and F-actin localized at the cell cortex. The partial colocalization of MCAM with actomyosin suggests that MCAM might associate with actin filaments through indirect protein interactions.
FIGURE 1:
Copolarization of MCAM, myosin, and F-actin. Confocal imaging of WRAMP structures, illustrating MCAM polarization with enrichment of cortical F-actin and myosin-IIB. For fluorescence imaging experiments, WM239a melanoma cells were plated on untreated glass at low confluency to minimize contacts between cells and serum-starved overnight prior to fixation. (A) Cells show IF signal from mouse anti-MCAM (AF488), rabbit anti-myosin-IIB heavy chain (AF594), and F-actin (phalloidin, AFP405). (B) Cells show IF signal from anti-MCAM, rabbit anti-phosphorylated-Ser19 myosin light chain 2 (pMLC, AF594), and phalloidin. Confocal Z-stacks were collected with step size 200 nm. For each image channel, the full cell is shown as a maximum intensity projection. An enlarged orthogonal view (cross-section) generated along the white dashed line is shown below. Images are representative of 20−30 cells per condition across two to three biological replicates. Scale bars (full images) = 10 μm. Scale bars (cross-section) = 3 μm in both dimensions. Imaging controls are shown in Supplemental Figure S1.
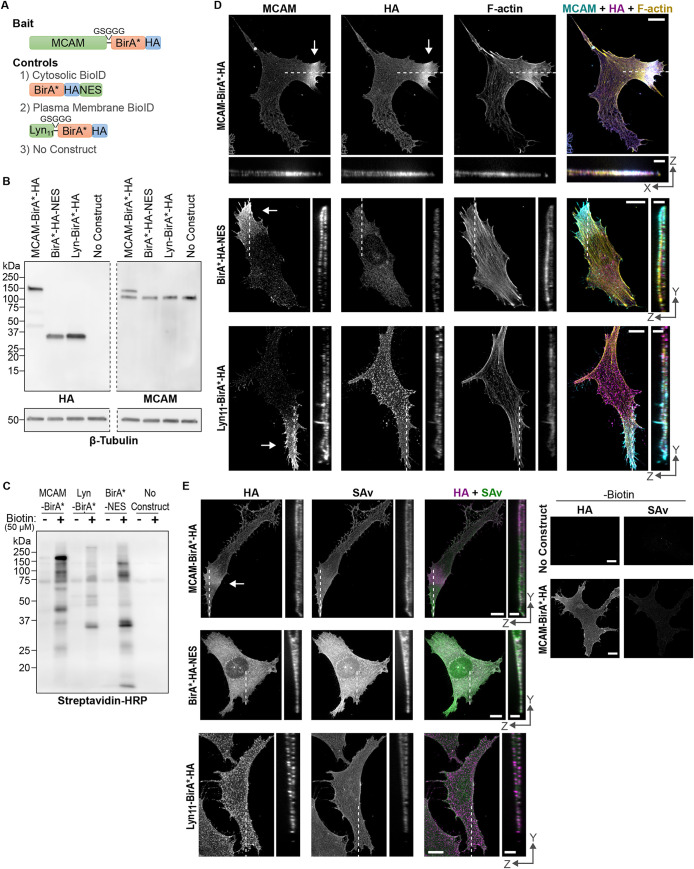
To identify proteins in the polarized MCAM interactome, we used BioID proximity proteomics (Roux et al., 2012; Sears et al., 2019) using a bait protein construct of full-length MCAM fused at its C-terminus with the biotin ligase mutant, BirA-R118G (BirA*), followed by a hemagglutinin tag (MCAM-BirA*-HA) (Figure 2A). Given the membrane enrichment of polarized MCAM, we designed a control BirA* tagged with a plasma membrane localization sequence from the Lyn tyrosine kinase (Lyn11-BirA*-HA) (Inoue et al., 2005). Additional controls included cytosolic BirA* fused to a nuclear export sequence (BirA*-HA-NES) (Wen et al., 1995) and parental WM239a cells lacking a BioID construct (Figure 2A).
FIGURE 2:
BioID experimental design. (A) Constructs used for MCAM bait and controls. (B) Western blots probed with anti-HA and anti-MCAM antibodies show stable expression of BioID constructs (left panel), and comparable expression of MCAM-BirA*-HA to endogenous MCAM (right panel). (C) Detection of protein biotinylation with HRP-coupled streptavidin, in stable cell lines grown for 72 h in biotin-free RPMI + 10% FBS followed by treatment with or without 50 µM biotin for 16 h. Streptavidin-HRP blots showing biotinylation activity and silver stains of samples used in MS experiments are shown in Supplemental Figure S3, A and B. Blot images were cropped to the right and left of the region shown to remove protein ladders and extra lanes, and along the dashed lines to remove a lane with ladder. (D) Confocal fluorescence images showing close alignment between MCAM-BirA*-HA (anti-HA) and total MCAM (anti-MCAM), which is absent with BirA*-HA-NES and Lyn11-BirA*-HA controls. White arrows indicate the location of polarized MCAM and MCAM-BirA*-HA. (E) Localization of biotinylated proteins detected using AlexaFluor 488 streptavidin (SAv) relative to HA-tagged BioID constructs. Cells were treated with 50 µM biotin for 16 h prior to fixation unless noted otherwise. Full X-Y images are shown as maximum intensity projections of confocal Z-stacks. Enlarged orthogonal views (cross-section) along the dashed white lines are shown below or at the right. Images are representative of approximately 20 cells per condition across two biological replicates for panel D and 10 cells in 1 replicate for panel (E). For (E), contrast was linearly scaled between the same lower and upper limits for each channel. Scale bars (full images) = 10 μm. Scale bars (cross-section) = 3 μm in both dimensions.
Western blotting verified similar levels of all three BioID proteins following stable lentiviral expression in WM239a cells (Figure 2B, left panel), with comparable expression across individual cells (Supplemental Figure S2). Importantly, the MCAM-BirA*-HA bait protein was expressed at a level comparable with endogenous MCAM (Figure 2B, right panel). Addition of biotin confirmed biotinylation activity for each BioID construct, as well as distinct patterns of protein biotinylation (Figure 2C). Confocal microscopy confirmed copolarization of the MCAM-BirA*-HA protein with F-actin and proper cytosolic and membrane localization of the BirA*-HA-NES and Lyn11-BirA*-HA controls, respectively (Figure 2D). We also examined the localization of biotinylated proteins in cells using AlexaFluor-coupled streptavidin and observed that the fluorescent signals corresponded well with the cellular compartments targeted by each BioID construct, although with patterns that appeared more diffuse (Figure 2E). The streptavidin signal was not always polarized in cells with polarized MCAM-BirA*-HA, which may reflect the long labeling period needed for BirA*-catalyzed biotinylation (Roux et al., 2012).
Biotinylated proteins were analyzed by mass spectrometry after treating the BioID or parental WM239a cell lines overnight with 50 µM biotin, isolating proteins from cell lysates using streptavidin affinity resin, and eluting in denaturing buffer with excess biotin. Western blotting confirmed that most biotinylated proteins were depleted from the cell lysates by affinity purification (Supplemental Figure S3A). From cells expressing various BioID-BirA* constructs, we detected distinct banding patterns of proteins eluted from the streptavidin resin by silver staining (Supplemental Figure S3B). This staining was more intense than that of naïve cells lacking BirA*, in support of the high-quality enrichment and interaction specificity of proteins isolated from BioID samples (Supplemental Figure S3B).
Proteins eluted from streptavidin resins were then identified by LC-MS/MS (Supplemental Tables S1 and S2). The distributions of protein abundances based on iBAQ intensity (intensity-Based Absolute Quantification; total precursor ion intensity divided by the number of theoretical peptides) (Schwanhüusser et al., 2011) were similar between the MCAM-BirA*-HA, BirA*-HA-NES, and Lyn11-BirA*-HA samples and higher than the parental WM239a controls (Supplemental Figure S3C). This indicated that the abundances of biotinylated proteins were similar between experiments and controls and the data were well suited for comparative analysis.
For initial analysis, MCAM-BirA*-HA samples were compared with the BirA*-HA-NES and parental WM239a controls, using the Statistical Analysis of INTeractome (SAINT) software (Choi et al., 2011, 2012; Nesvizhskii, 2015). SAINT is a computational method for proximity or affinity purification proteomics that compares the distribution of protein abundances between bait and control samples, using Bayesian statistics to estimate the likelihood that a protein is truly associated with the bait protein. This identified 416 proteins in the MCAM-BirA*-HA samples with a SAINT probability score corresponding to an FDR < 0.05 (Figure 3A, Supplemental Table S3). Comparison of these proteins to Gene Ontology (GO) gene sets (Ashburner et al., 2000; Carbon et al., 2021) showed statistically significant overlap with genes related to plasma membranes and organization of the cytoskeleton (Supplemental Table S4). Because MCAM is a transmembrane protein, enrichment of its prey over cytosolic BirA* might reflect their plasma membrane association rather than a specific association with MCAM. Therefore, we next compared MCAM-BirA*-HA to the plasma membrane−localized Lyn11-BirA*-HA.
FIGURE 3:
BioID identifies ERM proteins in the MCAM proximity proteome. (A) Proteins isolated from cells expressing MCAM-BirA*-HA were compared with cytosolic BioID (BirA*-HA-NES) and no-construct controls. SAINT scores were plotted against the log2 fold change of MCAM-BirA*-HA over 2 virtual controls (two highest spectral counts). Points in red identify 416 protein groups with SAINT scores corresponding to FDR < 0.05 (1,815 total protein groups). The protein group labeled “MCAM(BirA*)” combines sequences for endogenous MCAM and MCAM-BirA*-HA. (B) Volcano plot showing enrichment of MCAM-BirA*-HA samples over the plasma membrane BioID (Lyn11-BirA*-HA) control. P values were calculated for 1,502 high-confidence protein groups using a two-tailed Student’s t test for log2-transformed normalized iBAQ protein intensities as described in Supplemental Methods. Points in red correspond to those enriched in MCAM-BirA*-HA over the cytosolic BioID and no-construct controls, shown in panel (A). Log2 fold changes for proteins only detected in 1 Lyn11-BirA*-HA sample, which could not be compared by t test, are arbitrarily plotted at -0.5. (C) Log-ratio/log-average (MA) plot corresponding to data in panel B, showing log2 fold change vs. averaged mean iBAQ intensities for MCAM-BirA* and Lyn11-BirA*-HA. Points in orange are statistically significant after adjusting p-values using permutation-based correction (FDR < 0.05, s0 = 0 or s0 = 0.5). Proteins in blue highlight ERM proteins and a previously identified WRAMP component, NIBAN2. (D) Normalized iBAQ intensity values for ERM proteins detected with each BioID construct. (* indicates adjusted p value < 0.025, and ** indicates adjusted p-value < 0.005, for s0 = 0). (E) Gene symbols for proteins statistically significant in MCAM-BirA*-HA over Lyn11-BirA*-HA. Proteins are listed vertically by decreasing averaged mean iBAQ intensities for MCAM-BirA* and Lyn11-BirA*-HA. Those in bold were also significant in MCAM-BirA*-HA compared with BirA*-HA-NES and no-construct controls. Data in panels (A)−(E) represent three biological replicates of MCAM-BirA*-HA, Lyn11-BirA*-HA, and no construct samples, and two replicates of BirA*-HA-NES.
Spectral counts and iBAQ measurements showed greater similarity in the abundances of identified prey proteins between MCAM-BirA*-HA experiments and Lyn11-BirA*-HA controls (Supplemental Figure S3D). High-confidence identifications from the MCAM-BirA*-HA samples were compared with Lyn11-BirA*-HA based on Student’s t tests of normalized iBAQ intensities (Figure 3B; Supplemental Figure S3C; Supplemental Table S5). Proteins with differential abundances were evaluated using two stringencies of permutation-based estimation of FDR < 0.05, corresponding to a standard adjusted p-value (s0 = 0) or a parameter added for artificial within-groups variance (s0 = 0.5) to control the relative importance of p-value and mean fold change (Tusher et al., 2001). Proteins with significant enrichment using both analyses captured those with high abundance but small fold change enrichment, as well as proteins with large fold change but higher variance (Figure 3C; Supplemental Figure S4). This yielded 30 proteins that were enriched in the MCAM-BirA*-HA samples over the plasma membrane control, 13 of which were also enriched over the cytosolic and no-construct controls (Figure 3E). Reassuringly, MCAM or MCAM-BirA*-HA was strongly enriched over both cytosolic and plasma membrane controls. Enriched gene sets for the 30 proteins (Supplemental Table S6) showed a substantial number associated with ER, Golgi, and vesicle transport, suggesting that comparisons against the plasma membrane control were largely selective for proteins that were not integral components of the plasma membrane. Thus, the Lyn11-BirA*-HA control was highly stringent. Notably, NIBAN2/FAM129B, which we previously reported in WRAMP structures (Witze et al., 2013), was enriched relative to both the cytosolic and plasma membrane-localized controls (Figure 3A,B). This indicates that proteins known to copolarize with MCAM were successfully detected by the BioID screen.
Three of the 13 proteins that were enriched in MCAM-BirA*-HA samples over both plasma membrane and cytosolic BioID controls were moesin (MSN), ezrin (EZR), and radixin (RDX), which are members of the ERM protein family (Figure 3D). ERM proteins contain an N-terminal FERM domain connected by a flexible α-helical segment to a C-terminal C-ERMAD domain (Fehon et al., 2010). In their active conformation, ERM proteins bind membrane proteins at sites in the FERM domain and bind F-actin at a site in C-ERMAD, allowing ERM proteins to function as adaptors linking the plasma membrane to the actin cytoskeleton (Ivetic and Ridley, 2004; Fehon et al., 2010).
ERM proteins have been associated with cancer cell migration and invasion in multiple cancer types, including melanoma (Estecha et al., 2009; Lorentzen et al., 2011; Clucas and Valderrama, 2015; Hu et al., 2020), although little is known about their function and localization in these systems. Notably, ERM proteins have been shown to interact with MCAM by coimmunoprecipitation in melanoma cells (Luo et al., 2012). Our BioID results provided further support for their association in situ, and the role of ERM proteins as adaptors between plasma membrane and F-actin made them particularly appealing candidates for our system of MCAM polarization with actomyosin. Therefore, we focused on validating and characterizing the relationship between the ERM proteins and MCAM.
MSN, EZR, and RDX copolarize with MCAM
Because ERM proteins share many peptide sequences in common, bottom-up mass spectrometry did not clearly distinguish each protein’s association with MCAM. Therefore, we used indirect IF to evaluate the localization of endogenous ERM proteins. Western blotting HEK-293T cells transfected with MSN-, EZR-, or RDX-mCherry constructs confirmed that the antibodies raised against MSN, EZR, and RDX were specific for each protein with no apparent cross-reactivity (Supplemental Figure S5). We also confirmed insignificant bleed-through of IF signals between the emission channels used for MCAM and ERM proteins (Supplemental Figure S1, C and D).
Widefield imaging showed strong polarization of all three endogenous ERM proteins in the same regions of cells with polarized MCAM (Figure 4, A−C). For further validation, cells were transiently transfected to express each ERM protein fused to mCherry, followed by indirect IF to compare mCherry against endogenous MCAM and F-actin (Figure 4, E−G). The results confirmed copolarization of each ERM-mCherry construct with MCAM (Figure 4, E−G), which was absent with mCherry alone (Figure 4H). The MSN-, EZR-, and RDX-mCherry proteins showed similar localization to their cognate endogenous forms, except that RDX-mCherry lacked the nuclear signal that was present in anti-RDX immunostaining.
FIGURE 4:
Copolarization of ERM proteins and MCAM. (A−H) Widefield and (I−L) confocal immunofluorescence images showing the localization of ERM proteins relative to endogenous MCAM in WM239a cells. (A−C) Immunostaining of endogenous (A) MSN, (B) EZR, and (C) RDX against endogenous MCAM and F-actin/phalloidin. (D) Detection of phosphorylated ERM (pERM) using a phosphospecific antibody (T567/T564/T558 for EZR/RDX/MSN). For A−D, representative polarized cells were selected from several thousand cell images that were quantitatively analyzed in Figures 7 and 9. (E−G) Confirmation of ERM protein co-polarization with MCAM in WM239a cells expressing (E) MSN-mCherry, (F) EZR-mCherry, or (G) RDX-mCherry. Cells were fixed 48 h after transient transfection and stained with anti-MCAM antibody and phalloidin. Constructs were detected based on mCherry fluorescence which was preserved during fixation. Images are representative of at least 12 cells for each construct across two biological replicates. (H) Controls showing signal from cells transfected with mCherry empty vector or no plasmid (“mock” transfection, inset). White arrows denote regions with polarized MCAM. The mock transfection image is shown using the same linear contrast adjustment as the highest contrast mCherry construct. (I−L) Confocal Z-stacks shown as maximum intensity projections, with orthogonal views (cross-section) along dashed white lines. Cells were immunostained for endogenous MCAM and (I) MSN, (J) EZR, (K) RDX, (L) pERM, and F-actin/phalloidin. Images are representative of 16–24 cells per condition across 2 (I–K) or 3 (L) biological replicates. Scale bars (full images) = 10 μm. Scale bars (cross-section) = 3 μm in both dimensions.
Confocal microscopy was used to examine the 3D localization of endogenous ERM proteins with respect to endogenous MCAM and F-actin (Figure 4, I−K). The ERM proteins were enriched with polarized MCAM, in a manner that was closely apposed to the plasma membrane, consistent with their cortical localization. We also immunostained cells using a phosphospecific antibody against the activating C-ERMAD phosphorylation site in each ERM protein (pERM: phosphorylated at T567/T564/T558 in EZR/RDX/MSN) (Matsui et al., 1998). Cells with polarized MCAM also showed clear enrichment of pERM, similar to spatial patterns of MSN, EZR, and RDX (Figure 4, D and L). The cortical localization of pERM suggests that polarized ERM proteins are present in their activated state and is consistent with their potential role as adaptors between the plasma membrane and F-actin cytoskeleton.
Development of a quantitative image analysis pipeline to classify polarized cells
The copolarization of ERM proteins with MCAM prompted us to assess their spatial association and functional importance for MCAM polarization. Because MCAM polarization is a transient and dynamic process, examination of MCAM and ERM across thousands of cells was required. But manual analyses of cell images were subjective and time consuming, and tracking variations in phenotypes quickly became infeasible with increased sample size. Therefore, automated methods were needed to identify and characterize polarized cells. Existing tools often measure the ratios or differences in fluorescence intensity between the front and rear in cells moving directionally (Hetmanski et al., 2019; Moreau et al., 2019; Tsai et al., 2019; Bisaria et al., 2020; Zhang et al., 2020). These work well for cells migrating in microfluidic chambers or micropatterned strips, where morphology is constrained along a clearly defined axis of movement. But because cells in our system were not uniformly oriented and have heterogeneous, nonelliptical shapes, dividing them in half relative to their geometric major axis inadequately captured the locations of polarized proteins. Moreover, the area of spatial protein enrichment varied widely, likely due to the dynamic nature of WRAMP structures. This made it difficult to identify and characterize polarized cells using established tools. Therefore, we developed a new computational image analysis pipeline, named the “3P” pipeline that incorporated new metrics for protein polarity based on percentile-based thresholding with ML to facilitate the classification and quantitative description of polarized cells. In this strategy, single-cell features are extracted from images for use in ML. A subset of cells sampled across the dataset of interest is manually analyzed for polarization and used to train an ML model which is then applied to the entire dataset to identify polarized and nonpolarized cells.
Development of the 3P pipeline began with the creation of robust metrics to describe polarized MCAM using custom MATLAB scripts. First, individual cells were segmented based on MCAM and F-actin IF signals (Supplemental Figure S6), and a median background intensity was subtracted for each image. Next, regions with bright signal within cells were isolated by applying a threshold corresponding to the 80th percentile of all pixel intensities across the cell. Pixels above this threshold were segmented into individual objects, where an object is defined as a group of contiguously connected pixels (Figure 5A). In cells with polarized MCAM, the connected pixels were usually dominated by one large object encompassing the area of MCAM signal enrichment. To automatically identify this object, the integrated (summed) pixel intensity from the background-subtracted image was calculated for each object, and the object with the highest integrated intensity (the “brightest” object) was selected. Features were developed to characterize the region defined by the brightest object, based on its location and pixel intensity relative to the rest of the cell (Supplemental Table S7). Across the datasets presented in this study, the brightest object corresponded to the manually identified region of MCAM enrichment in 3,258 out of 3,337 (97.6%) polarized cells.
FIGURE 5:
Quantitative image analysis pipeline for classification and characterization of protein polarization. (A) Overview of the 3P image analysis pipeline. Cells are segmented based on MCAM IF signal, and regions with bright signal (> 80th percentile pixel intensity) are isolated as individual objects. The object with the greatest integrated intensity is selected as the brightest object. A subset of manually labeled cells is then used to train and test a ML classifier, based on selected features that are either standard from CellProfiler and MATLAB or custom-defined based on the brightest object. An iterative approach (gold box) is used for classifier training, by adding test images that are misclassified to the training set and repeating the cycle twice (see Supplemental Methods). (B) Violin plot showing the distributions and 25%, 50%, and 75% quantiles for the example of “median distance”, a custom feature that measures the median distance between the cell center and pixels in the brightest object. The results show overlapping distributions for polarized (N = 939) and nonpolarized (N = 1,238) cells based on manual scoring. (C) Examples of cells from three classes used to describe protein polarization. “Both Ends” and “None” classes were combined into a single “Nonpolarized” cell category after ML classification. Scale bars = 10 μm. (D) Confusion matrices after the second and final iteration using gbm (left) and extraTrees (center) classification models. Classifications from gbm and extraTrees were then combined to yield greater specificity, by requiring both classifiers to identify a cell as polarized (right). Classifications are shown for cells with a clear polarized or nonpolarized phenotype from a test set of cells separate from those used in iterative training. Cells with an ambiguous manual classification were omitted (Supplemental Methods). (B and D) The data are for experiments shown in Figure 6 and Supplemental Figures S7D−S9.
Our strategy uses supervised ML to leverage the combined predictive power of many features, because individual features which differed between polarized and nonpolarized cells showed overlapping distributions across populations of cells (Figure 5B). Thus, the 3P pipeline consists of a set of standardized image processing steps where images are segmented, the background subtracted, and features extracted (Figure 5A). Custom features based on the brightest object are combined with image classification features measured in MATLAB and CellProfiler (Carpenter et al., 2006; McQuin et al., 2018; Stirling et al., 2021), and individual features are scaled or converted to scale-invariant forms (Supplemental Figure S7A; see Supplemental Methods). Finally, a subset of cells from the dataset of interest are manually assigned to one of three classes, where the brightest MCAM object was (i) polarized at only one end of the cell (“OneEnd”); (ii) polarized at two or more peripheral regions (“BothEnds”); or (iii) distributed uniformly across the cell (“None”) (Figure 5C). In our datasets, only a small percentage (4%) of manually labeled cells under control conditions were classified as BothEnds, and likely reflect polarized MCAM disassembling in one region while accumulating in another region, as occasionally observed by live imaging. Therefore, we limited the characterization of MCAM polarization to cells classified as OneEnd and combined BothEnds and None into a nonpolarized category after ML classification.
The preprocessed data is then used to train a ML model. The pipeline is implemented with an iterative process to train ML models similar to that in CellProfiler Analyst (Jones et al., 2008). Here, the labeled data are split into an initial training and test set, and the training set is used to train a gradient boosting machine (gbm) (Greenwell et al., 2020) and an extremely randomized trees (extraTrees) (Simm et al., 2014) classifier on a set of 36 selected features (Supplemental Figure S7, B−D) using the R-package caret (Classification And Regression Training) (Kuhn, 2019, 2021). The resulting classification models are then applied to the test set, and a portion of the misclassified cells is added to a portion of the original training set to generate new training and test sets. These are used to train and test new models, and this cycle is repeated twice (Figure 5A; Supplemental Figure S8A). The gbm and extraTrees classification models from the final iteration are applied to the entire dataset to classify cells as polarized or nonpolarized, where both classifiers are required to agree on a “OneEnd” classification, as illustrated in the next section. For this study, a unique gbm model and a unique extraTrees model were trained for each dataset consisting of replicate experiments, with the annotated sample drawn across replicates (Supplemental Methods) to create a predictive model which accommodated the technical and biological variability of the dataset (Jones et al., 2009).
MSN promotes MCAM polarization
We applied the 3P pipeline to examine the requirement for ERM proteins in MCAM polarization. Cells were depleted in MSN, EZR, or RDX by siRNA transfection (Figure 6A), followed by immunostaining of MCAM and co-staining of ERM proteins, F-actin, or myosin-IIB. The performance of the classification model generated for this dataset was assessed on a test set consisting of cells separate from those used during iterative classifier training (see Supplemental Methods). The gbm and extraTrees models performed similarly, but their incorrectly classified cells were not completely identical. Therefore, to further increase specificity, cells were considered polarized only when both classifiers agreed on a OneEnd assignment; this resulted in overall sensitivity and specificity of 76.1% and 95.1%, respectively, when evaluated only on the cells in the test set with strong phenotypes (Figure 5D; see Supplemental Table S9 for sensitivity, specificity, and precision of individual replicates and conditions).
FIGURE 6:

Depletion of ERM proteins decreases MCAM polarization. (A) Western blots confirm specific knockdown of targeted ERM proteins. These blots are shown without vertical cropping in Supplemental Figure S9F. (B) Effects of siRNA knockdown of MSN, EZR, and RDX on the percentage of WM239a cells with polarized MCAM. The 3P pipeline was used to classify cells with polarized MCAM after treatment with each siRNA. For each replicate, the percentages of cells with polarized MCAM in each condition were normalized to cells treated with scrambled control siRNA by dividing by the percentage of polarized cells in the control siRNA condition. The average percentage of polarized cells for control replicates was 36%. Cells examined were from three biological replicates with at least two replicate coverslips, and totaled to 6,585 (scrambled siRNA control), 3,189 (siRNA-MSN), 3,227 (siRNA-EZR), and 4,316 (siRNA-RDX) cells (** indicates p-value < 0.025, * p-value < 0.05 for pairwise two-tailed Welch’s t tests after Holm-Bonferroni correction). For one replicate, one condition was an upper outlier based on Dixon’s Q test, and samples for all siRNAs were excluded. An additional MSN knockdown replicate was excluded due to differences in IF methods and a RDX knockdown with N < 200 cells (described further in Supplemental Methods). (C and D) The percentages of cells with polarized MCAM that also showed copolarization of (C) MSN or (D) EZR, based on coimmunostaining. Copolarization was scored manually by visible overlap of enriched MSN or EZR with MCAM, after classifying cells for polarized MCAM using 3P, as described in Supplemental Methods. Cells examined were gathered from two to three biological replicates and totaled 437 (control) and 147 (siRNA-MSN) cells in panel (C), and 258 (control) and 249 (siRNA-EZR) in panel (D). (ns indicates p-value > 0.05 for two-sample Wilcoxon test in panel (C); significance was not calculated in panel (D), because the control condition had only two replicates). Bars and errors show mean values and standard deviations for biological replicates.
Inspection of the gbm and extraTrees classification models showed that the custom measurements based on the brightest object were ranked as the most important features (examples in Supplemental Figure S8, B−F). Notably, when initially training on a larger number of features (indicated in Supplemental Table S8) which included the Haralick texture features (Haralick et al., 1973) from CellProfiler, overall sensitivity after combining gbm and extraTrees predictions did not exceed 50% for any iteration of training. Haralick features can describe variation in intensity across a cell and thus might be indicative of polarity; however, these features showed inconsistent ranges across replicate experiments (Supplemental Figure S7D). Thus, we found that feature selection and the new features for polarity added by our study turned out to be of greatest importance for robust classification of polarized cells across image sets.
We next assessed the effects of siRNA depletion of individual ERM proteins on MCAM polarization. Quantification of polarized and nonpolarized cells classified by 3P showed that MSN knockdown had the strongest effect on MCAM polarization, decreasing polarized cells to 40% of levels with control siRNA (Figure 6B). EZR knockdown resulted in a smaller decrease in polarized MCAM, and RDX depletion showed no consistent effect. The results demonstrate the functional importance of ERM proteins in promoting MCAM polarization, and suggests differences in their involvement, with MSN serving the most critical role.
We also examined cells that retained MCAM polarization after ERM protein knockdown. Although MSN knockdown appeared to decrease MCAM expression slightly (Supplemental Figure S9A), this had no effect on the relative intensity of MCAM polarization compared with the rest of the cell (Supplemental Figure S9B), nor the degree of MCAM copolarization with either myosin-IIB or F-actin (Supplemental Figure S9, C and D). Furthermore, cells depleted in each ERM protein did not block copolarization of MCAM with other ERM proteins (Supplemental Figure S9E). Interestingly, the residual MSN remaining after its knockdown could still consistently copolarize with MCAM, but EZR copolarization after its knockdown was reduced (Figure 6, C and D); this agrees with the suggestion that MSN is more indispensable for MCAM polarization.
ERM proteins spatially overlap with MCAM but arrive first at the cell periphery
To characterize the spatial overlap between polarized MCAM and ERM proteins, cells were costained for MCAM and F-actin, as well as MSN, EZR, RDX, or pERM. The 3P pipeline was then used to classify polarized MCAM, MSN, EZR, or pERM in an image dataset of several thousand cells. Of cells identified as polarized for MCAM, typically 85−90% were also classified as polarized for MSN, EZR, or pERM (Figure 7A). This subset was then inspected manually for copolarization. Cells costained with MCAM and RDX were only examined manually, due to the off-target nuclear signal of the anti-RDX antibody. Manual inspection showed MCAM copolarized with each ERM signal in nearly 100% of cells, and with F-actin in ∼75% of cells (Figure 7B). This confirmed that the polarization of ERM proteins and MCAM were highly correlated with respect to their spatial localization.
FIGURE 7:
Quantitative colocalization analysis identifies a cell subpopulation with ERM proteins closer to the cell periphery. Three-color IF widefield images of WM239a cells were labeled with anti-MCAM, phalloidin, and endogenous MSN, EZR, RDX, or pERM. The data are from experiments shown in Figure 4, A−D. (A) Venn diagrams showing the intersection of cells classified with polarized MCAM or ERM proteins using 3P. Percentages are shown relative to the total number of cells with polarized MCAM. (B) The percentages of cells with polarized MCAM that also show copolarized ERM proteins. Cells were first classified for polarized MCAM using 3P, and then manually assessed for copolarization of MSN, EZR, RDX or pERM, as described in Supplemental Methods. Numbers of cells examined totaled 1,291 cells costained for MCAM + MSN (5 replicates), 856 costained for MCAM + EZR (4 replicates), 419 costained for MCAM + RDX (2 replicates), and 1,096 costained for MCAM + pERM (4 replicates). (* indicates p-value < 0.05 for Kruskal-Wallis test followed by Dunn test with Holm-Bonferroni p-value adjustment). (C and D) Modified Manders’ Colocalization Coefficients for MCAM and MSN, EZR, or pERM in manually curated copolarized cells (see Supplemental Methods). Calculations of (C) MCC1 and (D) MCC2 defined MCAM as the first channel. The graphic illustrates MCC1 and MCC2 calculations in cells costained for MCAM + MSN. Boxplots show the distribution of single-cell values for each biological replicate, and individual data points are offset along the X-axis to show the distribution of cells across all replicates, where different colors correspond to different experiments. (E) Distributions of the distance between the weighted centroid of the brightest objects in MCAM and ERM channels. The distance was normalized by dividing by the maximum Feret diameter of each cell. Positive values indicate cells where ERM proteins were closer to the cell periphery than MCAM, relative to the cell center. Dashed lines indicate a normalized distance of ± 0.04, used to select outlier cells for manual evaluation. (F−H) Widefield images of cells showing separations between brightest object centroids, with (F) MSN, (G) EZR, or (H) pERM located closer to the cell periphery than MCAM. Scale bars = 10 µm.
The cells verified for copolarization were next examined for colocalization between MCAM and ERM proteins. As part of the 3P pipeline, modified Manders’ Colocalization Coefficients (MCC) (Manders et al., 1993) were calculated to evaluate the signal overlap between the MCAM and ERM image channels within their brightest objects (Figure 7C). These were defined such that MCC1 equals the MCAM intensity integrated over the region of overlap normalized to the total integrated intensity over the brightest object for MCAM. Similarly, MCC2 is the fraction of integrated ERM intensity within the region of overlap normalized to its brightest object. The results showed median values of 0.97−0.98 for MCC1 and 0.93−0.95 for MCC2 (Figure 7, C and D). This suggested nearly complete overlap of polarized MCAM with ERM proteins and strong but less overlap of ERM proteins with MCAM.
As a complementary metric, we calculated the weighted centroids of the brightest segmented object for each image channel and plotted the distance between the centroids normalized to the maximum cell diameter. Values were reported as positive for cells where the ERM centroids were farther from the cell center and negative when the MCAM centroids were farther from the center. Most cells showed centroid distances close to zero (Figure 7E). However, outliers in each distribution suggested phenotypic differences in small subpopulations of cells. To examine this possibility, cells with normalized distances between centroids greater than 0.04 were examined manually. The results showed that in 50−60% of cells labeled with positive distances between MCAM-MSN, MCAM-EZR, or MCAM-pERM, the polarized ERM signals were extended further toward the cell periphery than MCAM (Figure 7, F–H; see Supplemental Methods for detailed quantification). By contrast, in cells labeled with negative distances, only 12−13% showed MCAM extended further toward the cell periphery than MSN or EZR and 24% showed MCAM extended further than pERM. Taken together, the results showed that polarized MCAM and ERM proteins occupy the same spatial regions in most cells, but that there is a biased tendency for ERM proteins to move closer to the cell periphery relative to MCAM. This suggested a temporal order where movement of ERM proteins to the cell edge precedes MCAM.
To examine this possibility, we performed live imaging on cells expressing MCAM-GFP, LifeAct-mTagBFP (Riedl et al., 2008), and each of the ERM proteins fused to mCherry. In total, 200 cells were evaluated over 3 h, of which 77 clearly showed dynamic polarization of MCAM-GFP. Kymographs were generated for 74 of these cells to monitor the order of movement to the cell periphery. The polarization of MCAM-GFP was accompanied by polarization of MSN-, ERM-, or RDX-mCherry in all cases but one (RDX-mCherry), in agreement with results from fixed cells showing these proteins strongly copolarized. The MCAM and ERM signals moved across the polarizing cells in waves followed by a small, transient membrane retraction (93% of cases, Supplemental Table S10) when they arrived at the cell edge. In most cells, MCAM-GFP and the ERM-mCherry constructs polarized to the edge simultaneously. However, in approximately a third of cases, the ERM protein reached the cell periphery at least 3 min before MCAM (Figure 8; Supplemental Figure S10; Supplemental Table S10). By contrast, MCAM-GFP appeared at the cell edge before the ERM protein in only one case. LifeAct-mTagBFP expression was detectable in 56 of the analyzed cells and copolarized with MCAM in all but one case (Figure 8). Typically, F-actin moved synchronously with MCAM, reaching the cell edge at the same time, in agreement with previous live imaging results (Connacher et al., 2017). Together, these results support an ordered assembly process, where the ERM proteins, MCAM, and F-actin show overlapping polarization dynamics, but ERM proteins may precede MCAM to the cell periphery.
FIGURE 8:
ERM polarization precedes MCAM at the cell periphery. (A−C) Live imaging was carried out using cells stably expressing MCAM-GFP and transiently transfected with LifeAct-mTagBFP + (A) MSN-mCherry, (B) EZR-mCherry, or (C) RDX-mCherry (Supplemental Movies S1−S3). Kymographs and selected movie frames show the dynamics of proteins polarizing to the cell periphery. Black arrowheads indicate frames shown below each kymograph. The cyan line in the full cell image shows the kymograph axis and the white box shows the region enlarged in frames below. White arrows indicate arrival of the ERM signal at the cell periphery, preceding the arrival of MCAM signal. Overlays of the MCAM and ERM signals for kymographs and selected movie frames are shown in Supplemental Figure S10. Movie frames were collected every two minutes, and vertical scale bars to the left of each kymograph indicate 6 min = 3 frames. Horizontal scale bars = 5 µm. A total of 22, 26, and 26 cells across two biological replicates were analyzed for MCAM-GFP + MSN-mCherry, EZR-mCherry, and RDX-mCherry, respectively.
Movie S1.
Movement of MCAM‐GFP, MSN‐mCherry, and LifeAct‐mTagBFP in a WM239a cell shows polarizing MSN preceding MCAM to the cell periphery at the region indicated by the white arrow. Time series corresponding to Figure 8A.
Movie S2.
Movement of MCAM‐GFP, EZR‐mCherry, and LifeAct‐mTagBFP in a WM239a cell shows polarizing EZR preceding MCAM to the cell periphery at the region indicated by the white arrow. Time series corresponding to Figure 8B.
Movie S3.
Movement of MCAM‐GFP, RDX‐mCherry, and LifeAct‐mTagBFP in a WM239a cell shows polarizing RDX preceding MCAM to the cell periphery at the region indicated by the white arrow. Time series corresponding to Figure 8C.
Deletion of a sequence motif for MCAM−ERM interactions disrupts polarization
Mutations of positively charged residues at the juxtamembrane region of the cell adhesion protein, ICAM-2, as well as similar regions in CD44, CD43, L-selectin, and β-dystroglycan are known to disrupt ERM binding (Legg and Isacke, 1998; Yonemura et al., 1998; Ivetic et al., 2002; Spence et al., 2004). In MCAM, deletion of this sequence motif (KKGK, res. 584-587) disrupts its binding interactions with MSN, as measured by coimmunoprecipitation (Luo et al., 2012). To ask if this motif was also important for polarization of either MCAM or ERM proteins, we used CRISPR-Cas9 to delete these four amino acids in the endogenous locus of MCAM. Two clonal cell lines with homozygous deletions of KKGK were selected (MCAMΔKKGK), and protein polarization was quantified and compared with naïve cells using the 3P pipeline. The MCAMΔKKGK deletion mutation decreased the proportion of cells with MCAM polarization to less than half that of controls (Figure 9A), suggesting that interactions with ERM proteins are important for recruitment. One complication was that the expression of MCAM in the mutant cell lines was also reduced, as shown by Western blotting and by the distribution of mean fluorescence intensity of MCAM in single cells (Figure 9, B−D). However, the performance of the ML classification was similar between cell lines, indicating that 3P was robust to these differences in signal (Supplemental Table S11). Thus, MCAM polarization was still detectable in naïve cells with comparably lower expression, and against these, the fractions of polarized cells in the MCAMΔKKGK-mutant cell lines were consistently lower (Figure 9E). Therefore, the decreased MCAM polarization in cells with MCAMΔKKGK could be ascribed to the mutation, and not the reduced expression.
FIGURE 9:
Deletion of the ERM binding motif in MCAM blocks copolarization of MCAM and ERM. Three-color IF widefield images of WM239a cells were labeled with anti-MCAM, phalloidin, and endogenous MSN, EZR, RDX or pERM. The data for the naïve cells are also shown in Figure 4, A−D and Figure 7. (A) Polarized MCAM quantified in naïve vs. clonal MCAMΔKKGK cell lines engineered using CRISPR/Cas9. The 3P pipeline was used to classify cells from five biological replicates, totaling 7,642 (Naïve), 6,154 (Clone 1), and 6,574 (Clone 2) cells. For each replicate, the percentages of cells with polarized MCAM in each cell line were normalized to the naïve control. The average percentage of polarized cells for naïve replicates was 50%. (B) Western blots showing expression of MCAM and ERM proteins in naïve and knock-in cell lines. The uncropped blots are shown in Supplemental Figure S11. (C) Quantification of Western blots show mean and standard deviations of four technical replicates each from two biological replicate lysates. Integrated band intensities were normalized to GAPDH loading controls and divided by the mean of the naïve condition. (D) Kernel density plot showing mean MCAM intensities in individual cells normalized by the median intensity of naïve cells. The results show reduced MCAM expression on a single-cell level. Data show results for one replicate, which are representative of 4 other replicates. Numbers of cells totaled 1,878 (Naïve), 1,253 (Clone 1), and 1,359 (Clone 2). The X-axis is cropped, and 56 cells with a normalized intensity > 3 are not shown. (E) Percentage of cells with polarized MCAM normalized to naïve cells vs. mean MCAM intensity. Each curve represents 1 biological replicate; colors correspond to cell lines as indicated in panel (D). Cells were binned by normalized mean MCAM intensities, and the percentage of MCAM-polarized cells in each bin was divided by the overall percentage of polarized cells in the naïve cells for that replicate. The result shows that polarization of MCAMΔKKGK is lower than wild type, independent of expression level. Cells examined total 5,663 (Naïve), 4,851 (Clone 1), and 5,594 (Clone 2). Bins with less than 50 cells were omitted from the analysis. One replicate was omitted from the plot because only 1 cell line had > 50 cells per bin. (F−H) Percentage of cells with polarization of (F) MSN, (G) EZR, and (H) pERM in each cell line, normalized to the naïve control. For naïve replicates, the average percentage of cells with polarized MSN, EZR, and pERM was 53%, 55%, and 54%, respectively. Protein polarization was classified by machine learning using the 3P pipeline. The results suggest that ERM polarization is stabilized by interactions with MCAM. Cells were examined in four to five biological replicates, totaling 2,936 (Naïve), 2,435 (Clone 1), and 2,476 (Clone 2), for MCAM + MSN; 1,688 (Naïve), 1,587 (Clone 1), 1,713 (Clone 2) for MCAM + EZR; and 2,224 (Naïve), 1,721 (Clone 1), 1,949 (Clone 2) for MCAM + pERM. Two replicates of the naïve condition in (G) and one replicate of Clones 1 and 2 were quantified from a single coverslip per condition. For panels (A) and (F−H), bars and errors show mean values and standard deviations for biological replicates, and ** indicates p-value < 0.002, and * indicates p-value < 0.05, for pairwise Welch’s two-tailed t-test with Holm-Bonferroni correction. (I) A model for copolarization of MCAM and ERM proteins, proposing ERM recruitment to the membrane followed by MCAM via interactions with the juxtamembrane (KKGK) ERM-binding motif.
Applying the 3P pipeline to the ERM proteins also revealed a 30−40% decrease in polarization of EZR, MSN, and pERM in the MCAMΔKKGK cell lines relative to the control, which appeared significant although somewhat variable between replicates (Figure 9, F−H). This suggests a cooperative dependence of ERM polarization on MCAM as well. Taken together, our data show that a mutation that is known to disrupt binding interactions between MCAM and MSN interferes with MCAM, and to some extent ERM protein, recruitment to the cell periphery. This supports an ordered process, where the polarization of ERM proteins precedes and guides the polarization of MCAM, with the possibility of positive feedback to maintain ERM polarization.
DISCUSSION
Our findings reveal a novel role for ERM proteins in the polarization of MCAM to WRAMP structures. Following their identification by proximity ligation, analysis of thousands of cells showed consistent copolarization of MSN, EZR, and RDX with MCAM, and extensive overlap in their regions of spatial enrichment. Interference with MCAM polarization was greatest following siRNA depletion of MSN, with lesser effect of EZR depletion. Importantly, in a subpopulation of cells, ERM proteins extended farther from the cell center than MCAM, suggesting their earlier movement toward the periphery. This was corroborated by live imaging, where MCAM and ERM proteins showed tightly coordinated spatiotemporal dynamics, but with ERM proteins arriving at the cell edge before MCAM. Deletion of juxtamembrane residues in MCAM that had been previously shown to be necessary for MCAM-ERM binding resulted in diminished MCAM and ERM polarization. Our results identify ERM proteins as novel components of WRAMP structures and suggest that their dynamic coordination with MCAM occurs via their protein−protein interactions. These findings add to our understanding of the dynamic assembly of polarized protein networks that control rear membrane retraction and directional migration.
Key to this study was our development of the 3P pipeline, which accurately classified protein polarization in cells that were heterogeneous with respect to morphology, marker expression, and location of polarized objects. This advances quantitative image analysis over previous methods that are more effective in cells with uniform shape, defined orientation, and direction of movement (Hetmanski et al., 2019; Moreau et al., 2019; Bisaria et al., 2020; Zhang et al., 2020; Xie et al., 2021). Facilitated by object-based image analysis, 3P uses a percentile threshold to segment the brightest region of the cell and collect information about its location and intensity. Object-based approaches have been used to measure the polarity or orientation of discrete structures in cells, including protrusions (Diz-Muñoz et al., 2013; Yolland et al., 2019), focal adhesions (Zhang et al., 2014), filaments (Sorrentino et al., 2021; Nunes Vicente et al., 2022), and Golgi (Khuntia et al., 2022). By defining new metrics and adapting existing ones, 3P combines multiple measurements using ML to successfully classify polarization in randomly oriented cells with heterogeneous shapes and varying grades of protein enrichment. ML is an established method to classify and profile cellular phenotypes using standardized sets of features extracted with CellProfiler and other software (Carpenter et al., 2006; Caicedo et al., 2017; Piccinini et al., 2017; Smith et al., 2018). But the new custom features in 3P turned out to be necessary to achieve robust classification across multiple experiments and in cells with altered expression levels, such as the MCAMΔKKGK cell lines. In fact, for our datasets, standard Zernike (Zernike, 1934) and Haralick texture features used by CellProfiler to describe different distributions of intensity across cells usually had to be removed. The results show that ML can be a powerful tool to quantify cell polarization when feature sets are adapted or augmented with new measurements, such as the ones we present here. An additional strength of the ML approach used by 3P is that it allowed classification of polarity for multiple proteins, even though differences existed in their patterns of localization. Because 3P not only identifies polarized cells but also measures the location and colocalization of polarized proteins, we were able to sort cells based on these features and identify outlier cells where regions enriched in MCAM and ERM proteins were offset. Thus, the pipeline provided a powerful exploratory tool for discovering rare phenotypes in a large population of cells. We anticipate that 3P will be effective for classifying many polarization phenotypes in different cell types.
The copolarization between ERM proteins and MCAM in our system resembles other cellular features involved in migration. Uropods are knob-like structures described in Dictyostelium discoideum and normal and malignant leukocytes (Sánchez-Madrid and Del Pozo, 1999; Borset et al., 2000; Xu et al., 2003; Van Haastert and Devreotes, 2004; Dampmann et al., 2020; Bein et al., 2022), as well as in the Walker 256 carcinosarcoma (Rossy et al., 2007), 11A squamous cell carcinoma (Faber et al., 2013), and MDA-MB-231 breast cancer cell lines (Poincloux et al., 2011). Uropods are stably polarized and include adhesion proteins, ERM proteins, myosin-II, pMLC, and PI(4,5)P2 (Rosenman et al., 1993; Del Pozo et al., 1995; Serrador et al., 1997; Sánchez-Madrid and Del Pozo, 1999; Eddy et al., 2000; Fais and Malorni, 2003; Lee et al., 2004; Lacalle et al., 2007; Lokuta et al., 2007; Sánchez-Madrid and Serrador, 2009; Martinelli et al., 2013; Liu et al., 2015; Hind et al., 2016; García-Ortiz and Serrador, 2020). They form at the rear of cells, where myosin activity generates a strong contractile force that lifts the tail membrane during migration and propels cells forward. Other EZR-containing structures have been identified that share uropod components but are morphologically distinct. A similar EZR-rich uropod-like structure (ERULS) has been described at the rear of melanoma cells moving through a blebbing mode of amoeboid migration (Lorentzen et al., 2011). Finally, a polarized protein cap in nonadhered cells referred to as a single-cell (sc) pole has similar composition to uropods and ERULS and has been described in multiple melanoma and other cancer cell lines, where it is associated with increased invasion and metastasis (Lorentzen et al., 2018).
WRAMP structures share many features with uropods, ERULS, and sc poles, in that all contain EZR, CD44 (Wang et al., 2015), actomyosin, pMLC, and PI(4,5)P2. Uropods and ERULS also polarize to the rear of migrating cells. Moreover, MSN polarization at the rear of neutrophil-like HL-60 cells makes them resistant to switching directions during confined 1D migration (Prentice-Mott et al., 2016). This is reminiscent of our previous observation that MCAM polarization is associated with directional persistence (Connacher et al., 2017). MCAM is also found in sc poles, along with all three ERM proteins, and MCAM overexpression or depletion respectively increases or decreases EZR polarization (Lorentzen et al., 2018). Nevertheless, each of these structures have unique characteristics. Uropods and ERULSs are associated with amoeboid modes of migration, while WRAMP structures form during mesenchymal migration on 2D substrates as well as amoeboid migration in 3D hydrogels (Witze et al., 2013; Connacher et al., 2017). In addition, uropods are highly stable, and migrating cells can make U-turns rather than disassembling the uropod (Ramsey, 1972; Gerisch and Keller, 1981; Zigmond et al., 1981; Swanson and Lansing Taylor, 1982), whereas WRAMP structures disassemble and then reassemble in new locations during directional changes (Connacher et al., 2017). Furthermore, EZR knockdown decreases MCAM polarization in WRAMP structures but not in sc poles (Lorentzen et al., 2018). Overall, our findings support the likelihood that while different polarity structures share common components for cell movement, they are functionally distinct (Lorentzen et al., 2018).
Our findings offer new insight into the coordinated dynamics of WRAMP structure assembly. We previously showed by live imaging that recruitment of MCAM to WRAMP structures was highly correlated with F-actin, while myosin-IIB typically lagged behind by 2-3 min (Connacher et al., 2017). Cortical ER was also recruited to WRAMP structures, followed by elevation of cytosolic Ca2+ up to ∼0.8 µM and calpain-dependent membrane retraction (Witze et al., 2013). Inhibition of plasma membrane and ER calcium channels blocked membrane retraction but not MCAM recruitment, suggesting that Ca2+ influx into the cytosol is a later step in WRAMP structure formation (Witze et al., 2013). Thus, the ordered dynamics of WRAMP structure components suggests stepwise events that may ultimately promote rear membrane retraction. In this study, most polarized cells showed high spatial overlap between MCAM and ERM signals at any fixed moment in time. But the ability of 3P to identify subgroups of cells with incompletely overlapping signals revealed at least 4% of polarized cells with visible enrichment of MSN, EZR, or pERM further toward the cell periphery relative to MCAM. Live imaging showed slightly earlier recruitment of ERM proteins to the end of the cell in approximately 30% of cells undergoing polarization. Such subtle differences support a model for coordinated recruitment of ERM proteins and MCAM through protein−protein interactions. This is consistent with the reduced polarization of MCAM and ERM proteins seen in the cell lines expressing MCAM∆KKGK. Although this KKGK deletion mutant was shown to disrupt MCAM-ERM coimmunoprecipitation, it did not completely block MCAM−ERM interactions (Luo et al. 2012). This may explain why in our hands, edited cell lines still retained some copolarization of MCAM and ERM. The greater reduction in MCAM polarization relative to MSN and EZR polarization and the small but significant antecedence of ERM proteins to the cell edge suggests that ERM proteins likely initiate MCAM polarization and that MCAM promotes additional ERM recruitment. ERM proteins are known to exchange between inactive and active conformations, where they localize in the cytosol in an inactive state with their FERM and C-ERMAD domains associated in a closed conformation. They are activated by FERM domain binding to plasma membrane PI(4,5)P2 and C-ERMAD phosphorylation, which disrupts the interactions between FERM and C-ERMAD to favor an open conformation. This permits binding with membrane proteins at the FERM domain and F-actin at the C-ERMAD (Fievet et al., 2004; Ivetic and Ridley, 2004; Fehon et al., 2010). Formation of a ternary complex between MCAM, MSN, and the Rho inhibitor, RhoGDI, has also been shown to increase RhoA-GTP (Luo et al., 2012). Based on this, we propose a working model (Figure 9I) in which the initial signal for WRAMP structure formation involves recruitment of ERM proteins to membranes, possibly in response to a membrane phosphoinositide signal. Direct protein interactions in turn promote the coordinated recruitment of ERM proteins and MCAM. MCAM−ERM interactions may also serve to tether the actin cytoskeleton to the plasma membrane and promote activation of Rho family GTPases, potentially through localized loss of available RhoGDI.
Our findings also support growing evidence for distinct context- or cell type−dependent functions of different ERM proteins in migration. Estecha et al., (2009) observed that MSN depletion reduced the invasion of melanoma cell lines into collagen, whereas EZR depletion did not. In invading cells, MSN localized strongly to the rear while EZR remained nonpolarized. In contrast, EZR localized to the rear in ERULS in melanoma cells, and its depletion reduced amoeboid invasion into collagen (Lorentzen et al., 2011). In our system, all three ERM proteins polarize with similar dynamics relative to MCAM, yet only MSN depletion strongly inhibited MCAM polarization. Thus, while all three proteins have been shown to bind MCAM (Luo et al., 2012), their contributions to MCAM polarization do not seem to be equivalent. Much remains unclear about how homologous proteins that share common activation mechanisms and protein interactions can have different functions within the same cell. Although MSN, EZR, and RDX share approximately 75% sequence homology, MSN lacks a proline-rich region and EZR contains unique tyrosine phosphorylation sites that function in distinct regulatory pathways for migration (Funayama et al., 1991; Lankes and Furthmayr, 1991; Krieg and Hunter, 1992; Heiska and Carpén, 2005; García-Ortiz and Serrador, 2020). Furthermore, binding of ERM proteins to different cell adhesion proteins show varying Kd on the order of 1−100 nM, often dependent on the presence of phosphoinositides (Hirao et al., 1996; Heiska et al., 1998; Ivetic et al., 2002; Hamada, 2003); however, measurements of different ERM proteins binding to the same membrane protein are limited. Thus, differences in regulation and possibly binding affinity and kinetics may account for variations in function, although additional comparative studies of direct ERM−protein interactions in vitro and in vivo are needed.
MATERIALS AND METHODS
Antibodies, plasmids, and siRNA
Primary antibodies used for IF imaging and Western blotting (WB) were: anti-MCAM (mouse, sc-18837, Santa Cruz Biotechnology), anti-myosin-IIB (rabbit, M7939, Sigma-Aldrich), anti-EZR (rabbit, 3145S, Cell Signaling Technology), anti-MSN (rabbit, 3150S, Cell Signaling Technology), anti-RDX (rabbit, 2636S, Cell Signaling Technology), anti-pMLC2 (Ser19) (rabbit, 3671S, Cell Signaling Technology), anti-pERM (EZR-Thr567)/RDX-Thr564/MSN-Thr558) (3726T, Cell Signaling Technology), anti-HA (WB: mouse, 16B12, BioLegend; IF: rabbit, 3724S, Cell Signaling Technology, anti-mCherry (rabbit, 5993 BioVision, Inc.), anti-GAPDH (mouse, MA5-15738, Invitrogen), and anti-β-tubulin (rabbit, 2146S, Cell Signaling Technology). Secondary antibodies for WB were anti-mouse IgG HRP-linked (7076S, Cell Signaling Technology), and anti-rabbit IgG HRP-linked (7074S, Cell Signaling Technology); and for IF were AlexaFluor 488 goat anti-mouse IgG (H+L) and AlexaFluor 594 goat anti-rabbit IgG (H+L) (A11001, A11012, Invitrogen). For IF, AlexaFluor Plus 405 Phalloidin (A30104, Invitrogen) and AlexaFluor 488 streptavidin (S11223, Invitrogen) were used to detect F-actin and biotin, respectively. For Western blotting, biotinylated proteins were detected using Streptavidin-HRP (RPN1231-2ML from GE Healthcare or 3999S from Cell Signaling Technology).
Plasmids used in this study and cloning primer sequences are summarized in Supplemental Table S12. Expression plasmids will be made available on AddGene. Internal sequencing primers and entry and destination vector sequences are in Supplemental Data File S1. All open reading frames of plasmids were confirmed by Sanger sequencing (Quintara Biosciences). Protein depletion experiments used siGENOME Human EZR siRNA SMARTPool (M-017370-02-0005), ON-TARGETplus Human RDX siRNA SMARTPool (L-011762-00-0005), ON-TARGETplus Human MSN siRNA SMARTPool (L-011087-00-0005), and ON-TARGETplus Non-targeting Control Pool (D-001810-10-05), all from Horizon Discovery.
Details of cDNA plasmids, cell culture, lentivirus and siRNA transfection, fluorescence cell imaging, Western blotting, methods for validating ERM antibody specificity, generation of CRISPR/Cas9 cell lines, the 3P pipeline, kymographs, and statistical analysis of image data, including sample inclusion and exclusion criteria, are presented in Supplemental Materials and Methods.
BioID experiments and data analysis
Detailed methods are presented in Supplemental Materials. WM239a cells were transfected with lentiviruses for expression of BirA(R118G) (BirA*), MCAM-BirA*-HA, Lyn11-BirA*-HA and BirA*-HA-NES, followed by 14 d selection of stable cell lines in Hygromycin-B. To test for protein biotinylation by Western blotting, cells were grown for 72 h in biotin-free RPMI (MBS653376, MyBioSource, with 25 mM HEPES) + 10% FBS, treated overnight with serum-free, biotin-free RPMI with or without supplementation with 50 µM biotin (B4639, Sigma), and lysates were harvested for analysis. For mass spectrometry (MS) experiments, cells for each condition were cultured in biotin-free RPMI + 10% FBS for 72 h prior to treatment with biotin, replated in two 15-cm dishes per sample each with 2.2 × 106 cells, and treated in serum-free RPMI + 50 µM biotin, 0.005% dimethyl sulfoxide (DMSO). After 16.5 h, cells were washed twice with PBS, and harvested in 850 µl EDTA-free RIPA buffer (50 mM Tris, pH 7.5, 150 mM NaCl, 0.1% [wt/vol] SDS, 0.5% [wt/vol] sodium deoxycholate) with 1% (vol/vol) Triton X-100 and 1X EDTA-free cOmplete protease inhibitor (Roche). Lysates were sonicated on ice, clarified by centrifugation at 15,000 × g at 4°C for 20 min, and stored at −70°C. Three biological replicates were collected for all constructs, except BirA*-HA-NES (two replicates).
Biotinylated proteins were isolated by adding lysates to High Performance Streptavidin Sepharose (17-5113-01, Cytiva Life Sciences) and rotating overnight at 4°C. Beads were washed as described (Hesketh et al., 2017) in 500 µl BioID buffer (50 mM Tris, pH 7.5, 150 mM NaCl, 0.4% (wt/vol) SDS, 1% (vol/vol) IGEPAL CA-630, 1.5 mM MgCl2, 1 mM EGTA) and samples from replicate 15-cm dishes were combined. Beads were pelleted, washed once with 2% (wt/vol) SDS in 50 mM Tris, pH 7.5, twice with BioID buffer, and stored at −70°C in a 90% aliquot for MS and 10% aliquot for Western blotting and silver staining.
For MS, samples were eluted, reduced, and alkylated using 10% (wt/vol) SDS, 10 mM TCEP, 40 mM 2-chloroacetamide, 2 mM biotin, 50 mM Tris-HCl, pH 8.5, with boiling for 10 min and shaking at 37°C for 30 min. Proteins were digested using the SP3 method (Hughes et al., 2014), with adsorption to 200 µg carboxylate-functionalized speedbeads (Cytiva Life Sciences) and proteolysis at 37°C overnight in 50 mM Tris-HCl, pH 8.5, 0.5 µg Lys-C/trypsin (Promega). Peptides were desalted by binding beads in 95−100% acetonitrile (ACN) and washing once with 100% ACN. Peptides were then eluted twice with 1% trifluoroacetic acid (TFA), 3% ACN, dried by speedvac, and stored at −20°C. Peptides were then suspended in 3% (vol/vol) ACN, 0.1% (vol/vol) TFA and analyzed by LC/MS/MS using a Q-Exactive HF-X mass spectrometer interfaced with an Ultimate 3000 nanoUPLC system (Thermo Fisher Scientific). Two replicate injections were performed per sample. Separations were performed at 300 nl/min on a reversed-phase C18 M-class column (1.7 µm, 130 Å, 75 mm × 250 mm, Waters), with a gradient of 2−20% ACN in 100 min and 20−32% ACN in 20 min. Precursor mass spectra (MS1) were acquired at resolution 120,000 from 380−1580 m/z with automatic gain control (AGC) target of 3 × 106 and maximum injection time of 45 ms. Precursor peptide ion isolation width for MS2 scans was 1.4 m/z, and the top 12 most intense ions were sequenced. All MS2 spectra were acquired at resolution 15,000 with higher energy collision dissociation (HCD) at 27% normalized collision energy, AGC target of 1 × 105, and 100 ms maximum injection time. Dynamic exclusion was 20 s with mass tolerance ±10 ppm. Raw files were searched against the Uniprot Human database UP000005640 (downloaded 11/2/2020) using MaxQuant v.1.6.14.0 (Cox and Mann, 2008). Cys carbamidomethylation was a fixed modification, while Met oxidation, protein N-terminal acetylation, and Lys biotinylation were variable modifications. All peptide and protein identifications were thresholded at 1% false discovery rate (FDR).
Values for protein-level quantification summed peptide spectral counts (MS/MS counts) or MS1 peak intensities from MaxQuant output files, matching evidence file IDs to protein groups. All peptides were used; thus, razor peptides were assigned to more than one group. Protein intensities were converted to intensity-based absolute quantification (iBAQ) intensities by dividing by the number of iBAQ peptides from the proteinGroups file (Schwanhüusser et al., 2011). For SAINT analysis (Choi et al., 2011), nonhuman contaminant protein groups were removed, technical replicates were summed, and spectral counts were used to calculate SAINT scores using the “Resource for Evaluation of Protein Interaction Networks” (REPRINT) online platform (Nesvizhskii, 2015). MCAM-BirA*-HA samples, with MCAM assigned as the bait protein, were compared with BirA*-HA-NES and no-construct control samples collapsed to 2 virtual controls (2 highest spectral counts). Samples with a SAINT score corresponding to an FDR < 0.05 were considered significant. Student’s t tests comparing MCAM-BirA*-HA to Lyn11-BirA*-HA or BirA*-HA-NES used iBAQ intensities in each biological replicate (summed technical replicates), divided by the median iBAQ intensity of all proteins in that replicate and multiplied by the global median (the median of replicate medians). Two-sample two-tailed Student’s t tests of log2-transformed iBAQ intensities were implemented in Perseus v.2.0.9.0 (Tyanova et al., 2016; Tyanova and Cox, 2018). Proteins with FDR < 0.05 for s0 = 0 or s0 = 0.5 were pooled to generate the final set of enriched proteins (Supplemental Figure S4). Identification of significantly overlapping gene sets was performed using the Human Molecular Signatures Database (MSigDB, v2023.1.Hs) (Subramanian et al., 2005; Liberzon et al., 2011, Broad Institute, Inc. et al., 2023) with GO: biological process (GOBP) and GO: cellular component (GOCC) collections (Ashburner et al., 2000; Carbon et al., 2021). Significant overlap was determined using a hypergeometric distribution, with FDR q-value < 0.05.
Development of the 3P Image Analysis Pipeline
Detailed methods are presented in Supplemental Materials. Briefly, image segmentation was based on MCAM and F-actin image channels and performed using a custom script in MATLAB (v.9.8.0, R2020a) including the Image Processing Toolbox (v.11.1, 2020) and the Statistics and Machine Learning Toolbox (v.11.7, 2020). Extraction of standard single-cell features for classification of image data were measured using a CellProfiler pipeline (version 4.2.1 or earlier; Carpenter et al., 2006; McQuin et al., 2018; Stirling et al., 2021) and MATLAB. Extraction of custom single-cell features was performed using MATLAB. Features are summarized in Supplemental Table S7. Machine learning was performed in R (R Core Team, 2021) (version R4.2.2) using the CARET (Classification And REgression Training) package (Kuhn, 2019, 2021).
Code availability
Code for analysis of BioID data, kymograph analysis, and the 3P pipeline are available at https://github.com/Suzannah-Miller/Miller-et-al_ProteinPolarity.git.
Data availability
Peptide and protein identification and quantification, SAINT, Perseus, and enriched gene sets results are in Supplemental Tables, S1−S6. Training sets for ML are in Supplemental Table S13. Supplemental Table S1 and the classified 3P pipeline feature data and manual annotation are reposited at https://github.com/Suzannah-Miller/Miller-et-al_ProteinPolarity/tree/main/Supplemental-Data-Tables to accommodate large file sizes. These data are described in Supplemental Table S14. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE (Perez-Riverol et al., 2022) partner repository with the dataset identifier PXD047722 and 10.6019/PXD047722.
Supplementary Material
Acknowledgments
We are indebted to Joseph Dragavon for training and consultation in the BioFrontiers Advanced Light Microscopy Core facility, and Theresa Nahreini for assistance with flow cytometry. We also thank Xuedong Liu for sharing lentivirus plasmids and protocols, Molishree Joshi for assistance with gene editing experiments, Mary Katherine Connacher, Thomas Lee, and Holly Gates-Meyer for advice on protocols, and Andrew Kavran for advice on machine learning. Resources that supported this work include the Advanced Light Microscopy Core facility (RRID: SCR_018302), the Central Analytical Lab Mass Spectrometry facility (RRID:SCR_018992), the Flow Cytometry Shared Core, the Biochemistry Cell Culture Facility, and the Shared Instruments Pool in the Department Of Biochemistry at University of Colorado Boulder, and the Functional Genomics Shared Resource facility (RRID:SCR_021987) at University of Colorado Anschutz Medical Campus. This work was supported by NIH awards R35GM136392 (N.G.A.), R01GM127986 (N.G.A.), T32GM008769 (S.G.M.); S10OD025267 (Orbitrap HFX MS), R01CA107098S1 (Nikon Ti-E Widefield microscope), S10ODO21601 (FACSAria Fusion Cell Sorter), and P30CA046934 (Functional Genomics Core), as well as the BioFrontiers Institute and Howard Hughes Medical Institute (Nikon Ti-E spinning-disk confocal microscope). This material is also based upon work supported by the National Science Foundation Graduate Research Fellowship Program (S.G.M.) under Grant No. 1650115. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Abbreviations used:
- 3P
Protein Polarity by Percentile
- CARET
Classification And Regression Training
- ERM
ezrin/radixin/moesin
- ERULS
ezrin-rich uropod-like structure
- iBAQ
intensity-based absolute quantification
- IF
immunofluorescence
- MCAM
melanoma cell adhesion molecule
- ML
machine learning
- pERM
phosphorylated ezrin/radixin/moesin
- SAINT
significance analysis of interactome
- WRAMP structure
Wnt5a-receptor-actin-myosin-polarity structure.
Footnotes
This article was published online ahead of print in MBoC in Press (http://www.molbiolcell.org/cgi/doi/10.1091/mbc.E23-06-0255) on December 20, 2023.
REFERENCES
- Allam AH, Charnley M, Russell SM (2018). Context-specific mechanisms of cell polarity regulation. J Mol Biol 430, 3457–3471. [DOI] [PubMed] [Google Scholar]
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. (2000). Gene Ontology: tool for the unification of biology. Nat Genet 25, 25–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bein J, Flinner N, Häupl B, Mathur A, Schneider O, Abu-Ayyad M, Hansmann ML, Piel M, Oellerich T, Hartmann S (2022). T-cell-derived Hodgkin lymphoma has motility characteristics intermediate between Hodgkin and anaplastic large cell lymphoma. J Cell Mol Med 26, 3495–3505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bisaria A, Hayer A, Garbett D, Cohen D, Meyer T (2020). Membrane-proximal F-actin restricts local membrane protrusions and directs cell migration. Science 368, 1205–1210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borset M, Hjertner O, Yaccoby S, Epstein J, Sanderson RD (2000). Syndecan-1 is targeted to the uropods of polarized myeloma cells where it promotes adhesion and sequesters heparin-binding proteins. Blood 96, 2528–2536. [PubMed] [Google Scholar]
- Broad Institute, Inc., Massachusetts Institute of Technology, Regents of the University of California (2023). Investigate Human Gene Sets. Available at: https://www.gsea-msigdb.org/gsea/msigdb/annotate.jsp. (accessed July 22, 2022). [Google Scholar]
- Butler MT, Wallingford JB (2017). Planar cell polarity in development and disease. Nat Rev Mol Cell Biol 18, 375–388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caicedo JC, Cooper S, Heigwer F, Warchal S, Qiu P, Molnar C, Vasilevich AS, Barry JD, Bansal HS, Kraus O, et al. (2017). Data-analysis strategies for image-based cell profiling. Nat Methods 14, 849–863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carbon S, Douglass E, Good BM, Unni DR, Harris NL, Mungall CJ, Basu S, Chisholm RL, Dodson RJ, Hartline E, et al. (2021). The Gene Ontology resource: enriching a GOld mine. Nucleic Acids Res 49, D325–D334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carpenter AE, Jones TR, Lamprecht MR, Clarke C, Kang IH, Friman O, Guertin DA, Chang JH, Lindquist RA, Moffat J, et al. (2006). CellProfiler: Image analysis software for identifying and quantifying cell phenotypes. Genome Biol 7, R100.1-R100.10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi H, Larsen B, Lin ZY, Breitkreutz A, Mellacheruvu D, Fermin D, Qin ZS, Tyers M, Gingras AC, Nesvizhskii AI (2011). SAINT: probabilistic scoring of affinity purification-mass spectrometry data. Nat Methods 8, 70–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi H, Liu G, Mellacheruvu D, Tyers M, Gingras AC, Nesvizhskii AI (2012). Analyzing protein-protein interactions from affinity purification-mass spectrometry data with SAINT. Curr Protoc Bioinform 39, 8.15.8-8.15.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clucas J, Valderrama F (2015). ERM proteins in cancer progression. J Cell Sci 128, 1253–1253. [DOI] [PubMed] [Google Scholar]
- Connacher MK, Tay JW, Ahn NG (2017). Rear-polarized Wnt5a-receptor-actin-myosin-polarity (WRAMP) structures promote the speed and persistence of directional cell migration. Mol Biol Cell 28, 1924–1936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cox J, Mann M (2008). MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26, 1367–1372. [DOI] [PubMed] [Google Scholar]
- Cramer LP (2010). Forming the cell rear first: breaking cell symmetry to trigger directed cell migration. Nat Cell Biol 12, 628–632. [DOI] [PubMed] [Google Scholar]
- Dampmann M, Görgens A, Möllmann M, Murke F, Dührsen U, Giebel B, Dürig J (2020). CpG stimulation of chronic lymphocytic leukemia cells induces a polarized cell shape and promotes migration in vitro and in vivo. PLoS One 15, 1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diz-Muñoz A, Fletcher DA, Weiner OD (2013). Use the force: Membrane tension as an organizer of cell shape and motility. Trends Cell Biol 23, 47–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eddy RJ, Pierini LM, Matsumura F, Maxfield FR (2000). Ca2+-dependent myosin II activation is required for uropod retraction during neutrophil migration. J Cell Sci 113, 1287–1298. [DOI] [PubMed] [Google Scholar]
- Estecha A, Sánchez-Martín L, Puig-Kröger A, Bartolomé RA, Teixidó J, Samaniego R, Sánchez-Mateos P (2009). Moesin orchestrates cortical polarity of melanoma tumour cells to initiate 3D invasion. J Cell Sci 122, 3492–3501. [DOI] [PubMed] [Google Scholar]
- Etienne-Manneville S (2008). Polarity proteins in migration and invasion. Oncogene 27, 6970–6980. [DOI] [PubMed] [Google Scholar]
- Faber A, Hoermann K, Stern-Straeter J, Schultz DJ, Goessler UR (2013). Functional effects of SDF-1α on a CD44+ CXCR4+ squamous cell carcinoma cell line as a model for interactions in the cancer stem cell niche. Oncol Rep 29, 579–584. [DOI] [PubMed] [Google Scholar]
- Fais S, Malorni W (2003). Leukocyte uropod formation and membrane/cytoskeleton linkage in immune interactions. J Leukoc Biol 73, 556–563. [DOI] [PubMed] [Google Scholar]
- Fehon RG, McClatchey AI, Bretscher A (2010). Organizing the cell cortex: the role of ERM proteins. Nat Rev Mol Cell Biol 11, 276–287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fievet BT, Gautreau A, Roy C, Del Maestro L, Mangeat P, Louvard D, Arpin M (2004). Phosphoinositide binding and phosphorylation act sequentially in the activation mechanism of ezrin. J Cell Biol 164, 653–659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fritzsche FR, Wassermann K, Rabien A, Schicktanz H, Dankof A, Loening SA, Dietel M, Jung K, Kristiansen G (2008). CD146 protein in prostate cancer: Revisited with two different antibodies. Pathology 40, 457–464. [DOI] [PubMed] [Google Scholar]
- Funayama N, Nagafuchi A, Sato N, Tsukita S, Tsukita S (1991). Radixin is a novel member of the band 4.1 family. J Cell Biol 115, 1039–1048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- García-Ortiz A, Serrador JM (2020). ERM Proteins at the crossroad of leukocyte polarization, migration and intercellular adhesion. Int J Mol Sci 21, 1502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerisch G, Keller HU (1981). Chemotactic reorientation of granulocytes stimulated with micropipettes containing fMet-Leu-Phe. J Cell Sci 52, 1–10. [DOI] [PubMed] [Google Scholar]
- Greenwell B, Boehmke B, Cunningham J(2020). gbm: Generalized Boosted Regression Models. R package version 2.1.8 or later. Available at: https://CRAN.Rproject.org/package=gbm.
- Hadjitheodorou A, Bell GRR, Ellett F, Shastry S, Irimia D, Collins SR, Theriot JA (2021). Directional reorientation of migrating neutrophils is limited by suppression of receptor input signaling at the cell rear through myosin II activity. Nat Commun 12, 6619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamada K (2003). Structural basis of adhesion-molecule recognition by ERM proteins revealed by the crystal structure of the radixin-ICAM-2 complex. EMBO J 22, 502–514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haralick RM, Shanmugam K, Dinstein I (1973). Textural features for image classification. IEEE Trans Syst Man Cybern SMC-3, 610–621. [Google Scholar]
- Heiska L, Alfthan K, Grönholm M, Vilja P, Vaheri A, Carpén O (1998). Association of Ezrin with intercellular adhesion molecule-1 and -2 (ICAM-1 and ICAM-2). J Biol Chem 273, 21893–21900. [DOI] [PubMed] [Google Scholar]
- Heiska L, Carpén O (2005). Src phosphorylates ezrin at tyrosine 477 and induces a phosphospecific association between ezrin and a kelch-repeat protein family member. J Biol Chem 280, 10244–10252. [DOI] [PubMed] [Google Scholar]
- Hesketh GG, Youn JY, Samavarchi-Tehrani P, Raught B, Gingras AC (2017). Parallel exploration of interaction space by BioID and affinity purification coupled to mass spectrometry. In: Proteomics: Methods and Protocols, ed. Comai L, Katz JE, Mallick P, New York, NY: Humana, 115–136. [DOI] [PubMed] [Google Scholar]
- Hetmanski JHR, de Belly H, Busnelli I, Waring T, Nair RV, Sokleva OD, Cameron A, Gauthier N, Lamaze C, Swift J, et al. (2019). Membrane tension orchestrates rear retraction in matrix-directed cell migration. Dev Cell 51, 460–475.e10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hind LE, Vincent WJB, Huttenlocher A (2016). Leading from the back: the role of the Uropod in neutrophil polarization and migration. Dev Cell 38, 161–169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirao M, Sato N, Kondo T, Yonemura S, Monden M, Sasaki T, Takai Y, Tsukita S, Tsukita S (1996). Regulation mechanism of ERM (ezrin/radixin/moesin) protein/plasma membrane association: possible involvement of phosphatidylinositol turnover and Rho-dependent signaling pathway. J Cell Biol 135, 37–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu S, Shi X, Liu Y, He Y, Du Y, Zhang G, Yang C, Gao F (2020). CD44 cross-linking increases malignancy of breast cancer via upregulation of p-Moesin. Cancer Cell Int 20, 1–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes CS, Foehr S, Garfield DA, Furlong EE, Steinmetz LM, Krijgsveld J (2014). Ultrasensitive proteome analysis using paramagnetic bead technology. Mol Syst Biol 10, 757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inoue T, Heo WD, Grimley JS, Wandless TJ, Meyer T (2005). An inducible translocation strategy to rapidly activate and inhibit small GTPase signaling pathways. Nat Methods 2, 415–418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ivetic A, Deka J, Ridley A, Ager A (2002). The cytoplasmic tail of L-selectin interacts with members of the Ezrin-Radixin-Moesin (ERM) family of proteins. J Biol Chem 277, 2321–2329. [DOI] [PubMed] [Google Scholar]
- Ivetic A, Ridley AJ (2004). Ezrin/radixin/moesin proteins and Rho GTPase signalling in leucocytes. Immunology 112, 165–176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones TR, Carpenter AE, Lamprecht MR, Moffat J, Silver SJ, Grenier JK, Castoreno AB, Eggert US, Root DE, Golland P, et al. (2009). Scoring diverse cellular morphologies in image-based screens with iterative feedback and machine learning. Proc Natl Acad Sci USA 106, 1826–1831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones TR, Kang IH, Wheeler DB, Lindquist RA, Papallo A, Sabatini DM, Golland P, Carpenter AE (2008). CellProfiler Analyst: data exploration and analysis software for complex image-based screens. BMC Bioinformatics 9, 482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khuntia P, Rawal S, Marwaha R, Das T (2022). Actin-driven Golgi apparatus dispersal during collective migration of epithelial cells. Proc Natl Acad Sci 119, e2204808119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klunder LJ, Faber KN, Dijkstra G, Van IJzendoorn SCD (2017). Mechanisms of cell polarity - Controlled epithelial homeostasis and immunity in the intestine. Cold Spring Harb Perspect Biol 9, a027888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krieg J, Hunter T (1992). Identification of the two major epidermal growth factor-induced tyrosine phosphorylation sites in the microvillar core protein ezrin. J Biol Chem 267, 19258–19265. [PubMed] [Google Scholar]
- Kuhn M (2019). The caret package. Package documentation. Available at: https://topepo.github.io/caret/. Accessed Nov 11, 2022.
- Kuhn M (2021). caret: Classification and Regression Training. R package version 6.0-90 or later. Available at: https://CRAN.R-project.org/package=caret.
- Lacalle RA, Peregil RM, Albar JP, Merino E, Martínez-A C, Mérida I, Mañes S (2007). Type I phosphatidylinositol 4-phosphate 5-kinase controls neutrophil polarity and directional movement. J Cell Biol 179, 1539–1553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lankes WT, Furthmayr H (1991). Moesin: A member of the protein 4.1-talin-ezrin family of proteins. Proc Natl Acad Sci 88, 8297–8301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee JH, Katakai T, Hara T, Gonda H, Sugai M, Shimizu A (2004). Roles of p-ERM and Rho-ROCK signalling in lymphocyte polarity and uropod formation. J Cell Biol 167, 327–337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Legg JW, Isacke CM (1998). Identification and functional analysis of the ezrin-binding site in the hyaluronan receptor, CD44. Curr Biol 8, 705–708. [DOI] [PubMed] [Google Scholar]
- Lehmann JM, Riethmuller G, Johnson JP (1989). MUC18, a marker of tumor progression in human melanoma, shows sequence similarity to the neural cell adhesion molecules of the immunoglobulin superfamily. Proc Natl Acad Sci 86, 9891–9895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lei X, Guan CW, Song Y, Wang H (2015). The multifaceted role of CD146/MCAM in the promotion of melanoma progression. Cancer Cell Int 15, 3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X, Miao Y, Pal DS, Devreotes PN (2020). Excitable networks controlling cell migration during development and disease. Semin Cell Dev Biol 100, 133–142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liberzon A, Subramanian A, Pinchback R, Thorvaldsdottir H, Tamayo P, Mesirov JP (2011). Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu X, Yang T, Suzuki K, Tsukita S, Ishii M, Zhou S, Wang G, Cao L, Qian F, Taylor S, et al. (2015). Moesin and myosin phosphatase confine neutrophil orientation in a chemotactic gradient. J Exp Med 212, 267–280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lokuta MA, Senetar MA, Bennin DA, Nuzzi PA, Chan KT, Ott VL, Huttenlocher A (2007). Type Iγ PIP kinase is a novel uropod component that regulates rear retraction during neutrophil chemotaxis. Mol Biol Cell 18, 5069–5080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorentzen A, Becker PF, Kosla J, Saini M, Weidelle K, Ronchi P, Klein C, Wolf MJ, Geist F, Seubert B, et al. (2018). Single cell polarity in liquid phase facilitates tumour metastasis. Nat Commun 9, 887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorentzen A, Bamber J, Sadok A, Elson-Schwab I, Marshall CJ (2011). An ezrin-rich, rigid uropod-like structure directs movement of amoeboid blebbing cells. J Cell Sci 124, 1256–1267. [DOI] [PubMed] [Google Scholar]
- Luca M, Hunt B, Bucana C, Johnson J, Fidler I, Bar-Eli M (1993). Direct correlation between MUC18 expression and metastatic potential of human melanoma cells. Melanoma Res 3, 35–41. [DOI] [PubMed] [Google Scholar]
- Luo Y, Zheng C, Zhang J, Lu D, Zhuang J, Xing S, Feng J, Yang D, Yan X (2012). Recognition of CD146 as an ERM-binding protein offers novel mechanisms for melanoma cell migration. Oncogene 31, 306–321. [DOI] [PubMed] [Google Scholar]
- Manders EMM, Verbeek FJ, Aten JA (1993). Measurement of co-localization of objects in dual-colour confocal images. J Microsc 169, 375–382. [DOI] [PubMed] [Google Scholar]
- Martinelli S, Chen EJH, Clarke F, Lyck R, Affentranger S, Burkhardt JK, Niggli V (2013). Ezrin/Radixin/Moesin proteins and flotillins cooperate to promote uropod formation in T cells. Front Immunol 4, 1–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsui T, Maeda M, Doi Y, Yonemura S, Amano M, Kaibuchi K, Tsukita S, Tsukita S (1998). Rho-Kinase phosphorylates COOH-terminal threonines of Ezrin/Radixin/Moesin (ERM) proteins and regulates their head-to-tail association. J Cell Biol 140, 647–657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGary EC, Lev DC, Bar-Eli M (2002). Cellular adhesion pathways and metastatic potential of human melanoma. Cancer Biol Ther 1, 454–465. [DOI] [PubMed] [Google Scholar]
- McQuin C, Goodman A, Chernyshev V, Kamentsky L, Cimini BA, Karhohs KW, Doan M, Ding L, Rafelski SM, Thirstrup D, et al. (2018). CellProfiler 3.0: Next-generation image processing for biology. PLoS Biol 16, e2005970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mellman I, Nelson WJ (2008). Coordinated protein sorting, targeting and distribution in polarized cells. Nat Rev Mol Cell Biol 9, 833–845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mladinich KM, Huttenlocher A (2013). WRAMPing up calcium in migrating cells by localized ER transport. Dev Cell 26, 560–561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moreau HD, Blanch-Mercader C, Attia R, Maurin M, Alraies Z, Sanseàu D, Malbec O, Delgado MG, Bousso P, Joanny JF, et al. (2019). Macropinocytosis overcomes directional bias in dendritic cells due to hydraulic resistance and facilitates space exploration. Dev Cell 49, 171–188.e5. [DOI] [PubMed] [Google Scholar]
- Mseka T, Bamburg JR, Cramer LP (2007). ADF/cofilin family proteins control formation of oriented actin-filament bundles in the cell body to trigger fibroblast polarization. J Cell Sci 120, 4332–4344. [DOI] [PubMed] [Google Scholar]
- Nesvizhskii AI (2015). Resource for evaluation of protein interaction networks. Available at: https://reprint-apms.org. Accessed April 11, 2022.
- Nunes Vicente F, Lelek M, Tinevez JY, Tran QD, Pehau-Arnaudet G, Zimmer C, Etienne-Manneville S, Giannone G, Leduc C (2022). Molecular organization and mechanics of single vimentin filaments revealed by super-resolution imaging. Sci Adv 8, 2696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perez-Riverol Y, Bai J, Bandla C, Hewapathirana S, García-Seisdedos D, Kamatchinathan S, Kundu D, Prakash A, Frericks-Zipper A, Eisenacher M, et al. (2022). The PRIDE database resources in 2022: A Hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res 50, D543–D552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petrie RJ, Doyle AD, Yamada KM (2009). Random versus directionally persistent cell migration. Nat Rev Mol Cell Biol 10, 538–549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piccinini F, Balassa T, Szkalisity A, Molnar C, Paavolainen L, Kujala K, Buzas K, Sarazova M, Pietiainen V, Kutay U, et al. (2017). Advanced cell classifier: user-friendly machine-learning-based software for discovering phenotypes in high-content imaging data. Cell Syst 4, 651–655. [DOI] [PubMed] [Google Scholar]
- Poincloux R, Collin O, Lizárraga F, Romao M, Debray M, Piel M, Chavrier P (2011). Contractility of the cell rear drives invasion of breast tumor cells in 3D Matrigel. Proc Natl Acad Sci USA 108, 1943–1948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Del Pozo MA, Sánchez-Mateos P, Nieto M, Sánchez-Madrid F (1995). Chemokines regulate cellular polarization and adhesion receptor redistribution during lymphocyte interaction with endothelium and extracellular matrix. Involvement of cAMP signaling pathway. J Cell Biol 131, 495–508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- R Core Team (2021). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. Vienna, Austria. [Google Scholar]
- Ramsey WS (1972). Analysis of individual leucocyte behavior during chemotaxis. Exp Cell Res 70, 129–139. [DOI] [PubMed] [Google Scholar]
- Riedl J, Crevenna A, Kessenbrock K, Yu JH, Nuikirchen D, Bista M, Bradke F, Jenne D, Holak TA, Werb Z, et al. (2008). Lifeact: A versatile marker to visualize F-actin. Nat Methods 5, 605–607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ridley AJ, Schwartz MA, Burridge K, Firtel RA, Ginsberg MH, Borisy G, Parsons JT, Horwitz AR (2003). Cell migration: integrating signals from front to back. Science 302, 1704–1709. [DOI] [PubMed] [Google Scholar]
- Rosenman SJ, Ganji AA, Tedder TF, Gallatin WM (1993). Syn-capping of human T lymphocyte adhesion/activation molecules and their redistribution during interaction with endothelial cells. J Leukoc Biol 53, 1–10. [DOI] [PubMed] [Google Scholar]
- Rossy J, Gutjahr MC, Blaser N, Schlicht D, Niggli V (2007). Ezrin/moesin in motile Walker 256 carcinosarcoma cells: Signal-dependent relocalization and role in migration. Exp Cell Res 313, 1106–1120. [DOI] [PubMed] [Google Scholar]
- Roux KJ, Kim DI, Raida M, Burke B (2012). A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J Cell Biol 196, 801–810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sánchez-Madrid F, Del Pozo MA (1999). Leukocyte polarization in cell migration and immune interactions. EMBO J 18, 501–511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sánchez-Madrid F, Serrador JM (2009). Bringing up the rear: defining the roles of the uropod. Nat Rev Mol Cell Biol 10, 353–359. [DOI] [PubMed] [Google Scholar]
- Schnell U, Carroll TJ (2016). Planar cell polarity of the kidney. Exp Cell Res 343, 258–266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwanhüusser B, Busse D, Li N, Dittmar G, Schuchhardt J, Wolf J, Chen W, Selbach M (2011). Global quantification of mammalian gene expression control. Nature 473, 337–342. [DOI] [PubMed] [Google Scholar]
- Sears RM, May DG, Roux KJ (2019). BioID as a tool for protein-proximity labeling in living cells. In: Enzyme-Mediated Ligation Methods, ed. Nuijens T, Schmidt M, New York, NY: Humana, 299–313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- SenGupta S, Parent CA, Bear JE (2021). The principles of directed cell migration. Nat Rev Mol Cell Biol 22, 529–547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Serrador JM, Alonso-Lebrero JL, del Pozo MA, Furthmayr H, Schwartz-Albiez R, Calvo J, Lozano F, Sánchez-Madrid F (1997). Moesin interacts with the cytoplasmic region of intercellular adhesion molecule-3 and is redistributed to the uropod of T lymphocytes during cell polarization. J Cell Biol 138, 1409–1423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shellard A, Mayor R (2020). All roads lead to directional cell migration. Trends Cell Biol 30, 852–868. [DOI] [PubMed] [Google Scholar]
- Shutova MS, Asokan SB, Talwar S, Assoian RK, Bear JE, Svitkina TM (2017). Self-sorting of nonmuscle myosins IIA and IIB polarizes the cytoskeleton and modulates cell motility. J Cell Biol 216, 2877–2889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simm J, de Abril IM, Sugiyama M (2014). Tree-based ensemble multi-task learning method for classification and regression. IEICE Trans Inf Syst 97, 1677–1681. [Google Scholar]
- Smith K, Piccinini F, Balassa T, Koos K, Danka T, Azizpour H, Horvath P (2018). Phenotypic image analysis software tools for exploring and understanding big image data from cell-based assays. Cell Syst 6, 636–653. [DOI] [PubMed] [Google Scholar]
- Sorrentino S, Conesa JJ, Cuervo A, Melero R, Martins B, Fernandez-Gimenez E, de Isidro-Gomez FP, de la Moreno J, Studt JD, Sorzano COS, et al. (2021). Structural analysis of receptors and actin polarity in platelet protrusions. Proc Natl Acad Sci 118, e2105004118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spence HJ, Chen YJ, Batchelor CL, Higginson JR, Suila H, Carpen O, Winder SJ (2004). Ezrin-dependent regulation of the actin cytoskeleton by β -dystroglycan. Hum Mol Genet 13, 1657–1668. [DOI] [PubMed] [Google Scholar]
- Stirling DR, Swain-Bowden MJ, Lucas AM, Carpenter AE, Cimini BA, Goodman A (2021). CellProfiler 4: improvements in speed, utility and usability. BMC Bioinformatics 22, 433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Gouloub TR, Lander ES, et al. (2005). Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102, 15545–15550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swanson JA, Taylor DL (1982). Local and spatially coordinated movements in Dictyostelium discoideum amoebae during chemotaxis. Cell 28, 225–232. [DOI] [PubMed] [Google Scholar]
- Tsai TYC, Collins SR, Chan CK, Hadjitheodorou A, Lam PY, Lou SS, Yang HW, Jorgensen J, Ellett F, Irimia D, et al. (2019). Efficient front-rear coupling in neutrophil chemotaxis by dynamic myosin II localization. Dev Cell 49, 189–205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- The MathWorks, Inc (2020). Image Processing Toolbox version 11.1. Natick, MA: The MathWorks, Inc. [Google Scholar]
- The MathWorks, Inc (2020). MATLAB version R2020a. Natick, MA: The MathWorks, Inc. [Google Scholar]
- The MathWorks, Inc (2020). Statistics and Machine Learning Toolbox version 11.7. Natick, MA: The MathWorks, Inc. [Google Scholar]
- Tusher VG, Tibshirani R, Chu G (2001). Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 98, 5116–5121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tyanova S, Cox J (2018). Perseus: A Bioinformatics Platform for Integrative Analysis of Proteomics Data in Cancer Research. In: Cancer Systems Biology, ed. von Stechow L, New York, NY: Humana, 133–148. [DOI] [PubMed] [Google Scholar]
- Tyanova S, Temu T, Sinitcyn P, Carlson A, Hein MY, Geiger T, Mann M, Cox J (2016). The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods 13, 731–740. [DOI] [PubMed] [Google Scholar]
- Van Haastert PJM, Devreotes PN (2004). Chemotaxis: signalling the way forward. Nat Rev Mol Cell Biol 5, 626–634. [DOI] [PubMed] [Google Scholar]
- Vicente-Manzanares M, Koach MA, Whitmore L, Lamers ML, Horwitz AF (2008). Segregation and activation of myosin IIB creates a rear in migrating cells. J Cell Biol 183, 543–554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vicente-Manzanares M, Zareno J, Whitmore L, Choi CK, Horwitz AF (2007). Regulation of protrusion, adhesion dynamics, and polarity by myosins IIA and IIB in migrating cells. J Cell Biol 176, 573–580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang W, Runkle KB, Terkowski SM, Ekaireb RI, Witze ES (2015). Protein depalmitoylation is induced by Wnt5a and promotes polarized cell behavior. J Biol Chem 290, 15707–15716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Xu Q, Zhang N, Du X, Xu G, Yan X (2020). CD146, from a melanoma cell adhesion molecule to a signaling receptor. Signal Transduct Target Ther 5, 148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wen W, Meinkotht JL, Tsien RY, Taylor SS (1995). Identification of a signal for rapid export of proteins from the nucleus. Cell 82, 463–473. [DOI] [PubMed] [Google Scholar]
- Witze ES, Connacher MK, Houel S, Schwartz MP, Morphew MK, Reid L, Sacks DB, Anseth KS, Ahn NG (2013). Wnt5a directs polarized calcium gradients by recruiting cortical endoplasmic reticulum to the cell trailing edge. Dev Cell 26, 645–657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Witze ES, Litman ES, Argast GM, Moon RT, Ahn NG (2008). Wnt5a control of cell polarity and directional movement by polarized redistribution of adhesion receptors. Science 320, 365–369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu GJ, Wu MWH, Wang C, Liu Y (2011). Enforced expression of METCAM/MUC18 increases tumorigenesis of human prostate cancer LNCaP cells in nude mice. J Urol 185, 1504–1512. [DOI] [PubMed] [Google Scholar]
- Wu GJ, Fu P, Wang SW, Wu MWH (2008). Enforced expression of MCAM/MUC18 increases in vitro motility and invasiveness and in vivo metastasis of two mouse melanoma K1735 sublines in a syngeneic mouse model. Mol Cancer Res 6, 1666–1677. [DOI] [PubMed] [Google Scholar]
- Xie C, Jiang Y, Zhu Z, Huang S, Li W, Ou G (2021). Actin filament debranching regulates cell polarity during cell migration and asymmetric cell division. Proc Natl Acad Sci 118, e2100805118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu J, Wang F, Van Keymeulen A, Herzmark P, Straight A, Kelly K, Takuwa Y, Sugimoto N, Mitchison T, Bourne HR (2003). Divergent signals and cytoskeletal assemblies regulate self-organizing polarity in neutrophils. Cell 114, 201–214. [DOI] [PubMed] [Google Scholar]
- Yam PT, Wilson CA, Ji L, Hebert B, Barnhart EL, Dye NA, Wiseman PW, Danuser G, Theriot JA (2007). Actin-myosin network reorganization breaks symmetry at the cell rear to spontaneously initiate polarized cell motility. J Cell Biol 178, 1207–1221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yolland L, Burki M, Marcotti S, Luchici A, Kenny FN, Davis JR, Serna-Morales E, Müller J, Sixt M, Davidson A, et al. (2019). Persistent and polarized global actin flow is essential for directionality during cell migration. Nat Cell Biol 21, 1370–1381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yonemura S, Hirao M, Doi Y, Takahashi N, Kondo T, Tsukita S, Tsukita S (1998). Ezrin/Radixin/Moesin (ERM) proteins bind to a positively charged amino acid cluster in the Juxta-Membrane cytoplasmic domain of CD44, CD43, and ICAM-2. J Cell Biol 140, 885–895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeng Q, Li W, Lu D, Wu Z, Duan H, Luo Y, Feng J, Yang D, Fu L, Yan X (2012). CD146, an epithelial-mesenchymal transition inducer, is associated with triple-negative breast cancer. Proc Natl Acad Sci 109, 1127–1132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zernike F (1934). Beugungstheorie des schneidenver-fahrens und seiner verbesserten form, der phasenkontrastmethode. Physica 1, 689–704. [Google Scholar]
- Zhang J, Guo WH, Wang YL (2014). Microtubules stabilize cell polarity by localizing rear signals. Proc Natl Acad Sci 111, 16383–16388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L, Wei X (2022). The roles of Par3, Par6, and aPKC polarity proteins in normal neurodevelopment and in neurodegenerative and neuropsychiatric disorders. J Neurosci 42, 4774–4793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Flores LR, Keeling MC, Sliogeryt K, Gavara N (2020). Ezrin phosphorylation at T567 modulates cell migration, mechanical properties, and cytoskeletal organization. Int J Mol Sci 21, 435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zigmond SH, Levitsky HI, Kreel BJ (1981). Cell polarity: An examination of its behavioral expression and its consequences for polymorphonuclear leukocyte chemotaxis. J Cell Biol 89, 585–592. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Peptide and protein identification and quantification, SAINT, Perseus, and enriched gene sets results are in Supplemental Tables, S1−S6. Training sets for ML are in Supplemental Table S13. Supplemental Table S1 and the classified 3P pipeline feature data and manual annotation are reposited at https://github.com/Suzannah-Miller/Miller-et-al_ProteinPolarity/tree/main/Supplemental-Data-Tables to accommodate large file sizes. These data are described in Supplemental Table S14. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE (Perez-Riverol et al., 2022) partner repository with the dataset identifier PXD047722 and 10.6019/PXD047722.