Abstract
Background
Ventricular arrhythmia in hypertrophic cardiomyopathy (HCM) relates to adverse structural change and genetic status. Cardiovascular magnetic resonance (CMR)–guided electrocardiographic imaging (ECGI) noninvasively maps cardiac structural and electrophysiological (EP) properties.
Objectives
The purpose of this study was to establish whether in subclinical HCM (genotype [G]+ left ventricular hypertrophy [LVH]−), ECGI detects early EP abnormality, and in overt HCM, whether the EP substrate relates to genetic status (G+/G−LVH+) and structural phenotype.
Methods
This was a prospective 211-participant CMR-ECGI multicenter study of 70 G+LVH−, 104 LVH+ (51 G+/53 G−), and 37 healthy volunteers (HVs). Local activation time (AT), corrected repolarization time, corrected activation-recovery interval, spatial gradients (GAT/GRTc), and signal fractionation were derived from 1,000 epicardial sites per participant. Maximal wall thickness and scar burden were derived from CMR. A support vector machine was built to discriminate G+LVH− from HV and low-risk HCM from those with intermediate/high-risk score or nonsustained ventricular tachycardia.
Results
Compared with HV, subclinical HCM showed mean AT prolongation (P = 0.008) even with normal 12-lead electrocardiograms (ECGs) (P = 0.009), and repolarization was more spatially heterogenous (GRTc: P = 0.005) (23% had normal ECGs). Corrected activation-recovery interval was prolonged in overt vs subclinical HCM (P < 0.001). Mean AT was associated with maximal wall thickness; spatial conduction heterogeneity (GAT) and fractionation were associated with scar (all P < 0.05), and G+LVH+ had more fractionation than G−LVH+ (P = 0.002). The support vector machine discriminated subclinical HCM from HV (10-fold cross-validation accuracy 80% [95% CI: 73%-85%]) and identified patients at higher risk of sudden cardiac death (accuracy 82% [95% CI: 78%-86%]).
Conclusions
In the absence of LVH or 12-lead ECG abnormalities, HCM sarcomere gene mutation carriers express an aberrant EP phenotype detected by ECGI. In overt HCM, abnormalities occur more severely with adverse structural change and positive genetic status.
Key Words: cardiac magnetic resonance imaging, ECG imaging, electrophysiology, hypertrophic cardiomyopathy
Central Illustration
Proarrhythmic substrate in hypertrophic cardiomyopathy (HCM) is related to adverse cardiac structural change (hypertrophy, fibrosis, disarray, small vessel disease), and those with a sarcomere gene mutation (genotype [G]+ left ventricular hypertrophy [LVH]+ vs G−LVH+) have a higher incidence of sudden cardiac death (SCD).1, 2, 3 Twelve-lead electrogram (ECG) abnormalities occur in those with sarcomeric mutations without hypertrophy (G+LVH− also known as subclinical HCM) and predict future LVH.4 Both the origins and mechanisms underlying ventricular arrhythmia remain poorly understood.
Abnormalities in ventricular activation and repolarization could be the link between structural changes and SCD.3,5 In this domain, electrocardiographic imaging (ECGI) provides detailed noninvasive electrophysiological (EP) assessment in intact hearts under physiological conditions6 and may be more sensitive to early changes (subclinical disease) than structural assessment by macroscopic imaging, ie, cardiovascular magnetic resonance (CMR) alone. ECGI is a high spatiotemporal resolution method of computing unipolar epicardial electrograms (UEGs) to panoramically map ventricular activation and repolarization.7 It has been validated ex vivo and in vivo, provided EP insights into multiple diseases, and shown potential to guide device and ablation therapies.7, 8, 9, 10, 11, 12, 13 Recent advances in this technique include our development of a fully washable and reuseable vest that has demonstrated high repeatability and allows noninvasive performance of ECGI at scale.14 Integration with advanced myocardial tissue characterization is therefore enabled by CMR. This approach, “CMR-ECGI,” now permits fully coregistered simultaneous interrogation of the myocardial structure, scar, and EP properties.
We hypothesized that CMR-guided ECGI would detect the following: 1) subtle EP abnormalities in subclinical HCM; and 2) EP abnormalities in overt disease related to genetic status (G+ vs G−LVH+) and structural changes (maximal wall thickness [MWT], late gadolinium enhancement [LGE]).
Methods
Ethical approval was obtained for each study site from the UK Research Ethics Committee (IRAS 227168), and all participants provided informed consent to participate in the study. An academic collaboration was formed for large-scale clinical deployment of ECGI technology (capturECGI Vest [Patent Application No. US 18/194 235] + body surface ECG + heart-torso geometry preprocessing14 led by University College London [UCL], United Kingdom; inverse solution led by Prof Rudy Lab, Washington University, St Louis, USA; digital 12-lead ECG analysis led by University of Glasgow, United Kingdom; and advanced artificial intelligence [AI]-powered CMR analysis + sequence optimization led by Barts Heart Centre, UCL, and National Institutes of Health, USA).
Study population
Adult participants (age 18-76 years) were recruited from cardiomyopathy clinics of 3 tertiary referral sites (Barts Heart Centre, St George’s University of London, Royal Free London, United Kingdom). Genotype classification was performed according to American College of Medical Genetics criteria.15 Hypertrophic cardiomyopathy was defined as per European Society of Cardiology (ESC)/American Heart Association guidelines (unexplained LVH: MWT >13 mm in any American Heart Association segment using either echocardiography or CMR in first-degree relatives of affected probands or else ≥15 mm).1,16 HCM was diagnosed by an inherited cardiomyopathy expert in a recruiting site. Overt HCM patients with intermediate/high SCD risk were identified based on the ESC risk score.17 Subclinical HCM (G+LVH−) was defined in individuals having a pathogenic/likely pathogenic sarcomere gene variant identified by cascade screening but without LVH. Healthy volunteers recruited under the same ethical protocol did not undergo genetic testing or Holter monitoring. Exclusion criteria were contraindication to CMR, needle-phobia, inability to consent, known or clinically suspected coronary artery disease, prior myectomy/alcohol septal ablation, implantable cardiac devices, persistent atrial fibrillation, high ventricular ectopic burden, poorly controlled hypertension (suboptimal control despite 2 antihypertensives) and significant valve disease. No patients were on cardiac myosin inhibitors. Recent (within 12 months) clinical Holter results and clinical echocardiograms identified participants with nonsustained ventricular tachycardia (≥3 beats of ventricular tachycardia at ≥120 beats/min) and LV outflow tract obstruction (gradient ≥30 mm Hg under any condition).
ECGI acquisition and analysis
A resting 256-lead ECG in the recumbent position was recorded for 5 minutes for signal averaging at a sampling frequency of 2,400 Hz using the fully reuseable, washable capturECGI vest and a high-performance amplifier system (g.HIamp 256 bundle GT-8016/USBamp GT-0216; g.tec medical engineering GmbH).14 The 5-minute body surface recording was quality controlled using in-house Matlab software (Mathworks) (Figure 1). Signal averaging enhanced the signal quality, and low-quality signals were removed. The vest was replaced with an identically sized “mirror-vest” using 256 identically positioned magnetic resonance imaging lucent fiducial markers for scanning. Heart-torso geometry was obtained by segmenting an anatomical transaxial stack (see following text) using Amira-Avizo software (Thermo Fisher, version 2021.1) for electrode positions and epicardial meshes. Body surface recordings and heart-torso geometry underwent the inverse solution of electrocardiography (defined as the reconstruction of cardiac electrical activity from given body surface electrocardiographic measurements) using previously validated protocols.7 Computed electrodes arising from valve-plane (nonconducting tissue) were excluded. A total of 1,000 UEGs (Figure 1) were computed per heart to obtain activation time (AT) (the time from the earliest activation in the epicardium to steepest point of the QRS downslope), repolarization time corrected for heart rate (RTc) (the steepest point of the T-wave upslope18), and activation-recovery interval (ARIc) (the difference of AT and RTc). Heart rate correction used the Fridericia formula.19 Dispersions of AT and ARIc (ΔAT, ΔARIc) were measured as the maximum-minimum of AT and ARIc, respectively, across the entire myocardium. Spatial gradients (gradient of activation time [GAT], gradient of repolarization time corrected [GRTc]) were computed for each epicardial site as the absolute value of the difference between neighboring sites (within 15 mm) divided by their distance, averaged across all neighbors. Signal amplitudes were defined as the peak to peak (maximum to minimum) of the QRS complexes. Parameters from all 1,000 electrodes were averaged per participant and maximal gradients were also recorded (max: GAT/GRTc). Subclinical HCM with max/mean GRTc >2 SDs above the mean were defined as having abnormal spatial repolarization. Signal fractionation was defined as number of UEGs with ≥2 negative deflections within the QRS complex.
Figure 1.
ECGI Acquisition and Postprocessing
(Top) A 256-lead electrocardiogram (ECG) is recorded using the fully re-useable capturECGI vest. The mirror vest is positioned to match the electrode vest and the participant wears this during the dark blood (HASTE [Half-Fourier Acquisition Single-shot Turbo spin Echo imaging]) anatomical stack acquisition. Heart-torso geometry is obtained using segmentation of the epicardium and the markers on the mirror vest using Amira-Avizo software (Thermo Fisher). The inverse solution is performed according to previously described protocols thereby obtaining 1,000 computed unipolar electrograms. (Bottom Left) Activation time (AT) is defined by the steepest QRS downslope. Repolarization time (RT) is defined by the steepest part of QRS upslope. Both are referenced to earliest epicardial activation. Activation recovery interval (ARI) is the difference between RT and AT. Three points (circle, triangle, square) on AT and ARIc maps are shown and their corresponding AT, RT, ARI are indicated on their computed unipolar electrograms (black arrows show ECGI intervals of the ‘circle’ UEG). RT (and ARI) are corrected for heart rate. (Bottom right) AT/RT gradients: (ΔAT/ ΔRT) between adjacent orange and blue unipolar electrograms (UEGs) are shown. Gradients are measured as the difference in AT/RT between neighboring electrograms divided by their inter-electrode distance (d).
CMR acquisition and analysis
All scans were performed at 3-T (Prisma, Siemens Germany) in UCL’s Bloomsbury Center for Clinical Phenotyping immediately after body surface ECG recordings. Heart-torso geometry was obtained from a free-breathing 100 thin-slice (4 mm contiguous) Half-Fourier Acquisition Single-shot Turbo spin Echo imaging (HASTE) sequence.14 Cine imaging was conventional with retrospectively gated breath-held balanced single-shot free precession short- and long-axis views. These were segmented for cardiac volume and wall thickness using fully automated AI-based algorithms shown to exceed human precision.20,21 Free-breathing motion-corrected LGE data were acquired 5 minutes following intravenous contrast (Dotarem, Guerbet). LGE was quantified using the 5-SD technique and expressed as absolute mass in grams. Precontrast and postcontrast modified Look-Locker inversion recovery (MOLLI) T1 with same-day hematocrit for extracellular volume (ECV) were acquired and septal ECV was computed.22
Digital 12-lead ECG
A 12-lead ECG was acquired at rest using the Beneheart R3 (Mindray), and recordings were sent to University of Glasgow Core-Lab for automated analysis of electrical intervals (PR, QRS, QTc), QT dispersion, amplitudes, and the presence of ECG abnormalities.23,24 ECG abnormalities known to be relevant to disease progression in HCM4,25 were defined as follows: abnormal Q waves (≥2 contiguous leads and with minimum amplitude 0.3 mV or ≥25% of the subsequent R-wave, or duration >40 ms); LVH criteria defined as Sokolow-Lyon or Cornell criteria and repolarization abnormalities (T-wave inversion of ≥0.1 mV in ≥2 contiguous leads and/or ST-segment depression ≥0.1 mV in ≥2 contiguous leads); and ECG fractionation defined as notching within the QRS complex (including R′, r′, S notching, R notching, or fragmentation) in 2 contiguous leads.
Statistical analysis
Statistical analysis was performed using Matlab (MathWorks, version May 11, 2022) and SPSS (IBM SPSS Statistics, version 28). The distribution of data was examined using histograms, Shapiro-Wilk tests and QQ plots. Normally distributed continuous variables were summarized as mean ± SD and compared using independent samples Student’s t-tests; non-normally distributed variables were presented as median (Q1-Q3) and compared using Mann-Whitney U tests. Multiple testing with Bonferroni was performed on a 3 comparisons per variable basis (P < 0.05/3). Categorical/binomial variables were expressed as absolute counts and percentages and compared using the chi-square/Fisher exact test as appropriate. Spearman’s correlations were used to examine simple linear/monotonic relationships between structural variables, risk markers (nonsustained ventricular tachycardia [NSVT]/left ventricular outflow tract obstruction [LVOTO]) and ECG/ECGI parameters. Multiple linear regression models were used to explore independent between-group differences by including a priori defined confounders as covariates, namely age, sex, LGE volume, and QRS duration for AT comparisons or QTc for ARIc comparisons. Additionally, differences in G+LVH− were adjusted for the presence vs absence of an abnormal resting 12-lead ECG. Variance inflation factor <3 excluded collinearity. Further details are provided in the Supplemental Methods. To discriminate subclinical HCM from health, a supervised machine learning approach using a support vector machine (SVM) classification was built. This incorporated all 12-ECGI biomarkers and was applied to the subclinical vs control data set followed by a 10-fold cross validation. Confusion matrices were generated to summarize model performance, and area under the receiver-operator curves (AUCs) were calculated using R package (ROCR). A similar SVM and receiver-operator curve approach explored whether in patients with overt HCM, those with high/intermediate risk (from the ESC Risk score) or those with prior NSVT could be discriminated from low-risk patients (Supplemental Methods).
Results
A total of 211 participants were prospectively studied: 70 G+LVH− subjects, 104 LVH+ patients (51 G+LVH+, 53 G−LVH+), and 37 healthy volunteers (HVs), of whom 26 were younger and used for age-matched comparisons with G+LVH−, whereas 23 (with overlap) were older and used for age-matched comparisons with G+LVH+ (reported in Supplemental Table 1). A comparison of Drug-Free HCM and matched healthy volunteers are shown in Supplemental Table 2. Relationships between LV morphology and ECGI parameters are shown in Supplemental Table 3. Likely pathogenic/pathogenic variants for G+LVH− were present in the following genes: 41 (59%) MYBPC3, 16 (23%) MYH7, 5 (7%) TNNI3, 4 (6%) TNNT2, 1 (1%) MYL2, 1 (1%) CSRP3, 1 (1%) ACTC1, 1 (1%) TPM1 and for G+LVH+: 28 (55%) MYBPC3, 13 (25%) MYH7, 4 (8%) TNNI3, 4 (8%) TNNT2, 1 (2%) TNNC1, and 1 (2%) CSRP3.
Clinical characteristics
Subclinical HCM (G+LVH−) vs HV
Compared with age-matched HVs, G+LVH− had similar age, sex, ethnicity, and body surface area (BSA). G+LVH− had similar LV cavity size and myocardium (MWT, LV mass, ECV, and LGE) but more hyperdynamic function. G+LVH− had a stronger family history of SCD.
G+LVH+ vs G+LVH−
Compared with G+LVH−, G+LVH+ had similar BSA but were older, were more often male and with smaller cavity sizes, had more abnormal myocardium (higher MWT, LV mass, ECV, and LGE), and had higher ejection fraction.
G−LVH+ vs G+LVH+
Compared with G+LVH+, G−LVH+ had similar sex and BSA; were older; had similar LV cavity sizes, function, and MWT; had greater LV mass; had lower ECV; and had similar LGE. There was no difference in LV morphology. G+LVH+ had a stronger family history of sudden cardiac death (Table 1).
Table 1.
Demographics, CMR, and 12-Lead ECG Variables in Younger HVs and HCM Cohorts
| Younger HVa (n = 26) | G+LVH− (n = 70) | G+LVH+ (n = 51) | G−LVH+ (n = 53) |
P Value |
|||
|---|---|---|---|---|---|---|---|
| G+LVH− vs HV | G+LVH+ vs G+LVH− | G−LVH+ vs G+LVH+ | |||||
| Demographics | |||||||
| Age, y | 35 (25-38) | 36 (25-41) | 52 (37-59) | 59 (50-66) | 0.85 | <0.001 | 0.005 |
| Female | 14 (54) | 41 (59) | 17 (33) | 8 (15) | 0.68 | 0.003 | 0.09 |
| White | 21 (81) | 57 (81) | 43 (84) | 36 (68) | 0.94 | 0.68 | 0.051 |
| BSA, m2 | 1.8 ± 0.2 | 1.9 ± 0.2 | 1.9 ± 0.2 | 2.0 ± 0.2 | 0.45 | 0.21 | 0.42 |
| BMI, kg/m2 | 24 (22-27) | 25 (22-28) | 25 (24-28) | 26 (24-28) | 0.24 | 0.65 | 0.5 |
| Hypertension | 0 (0) | 4 (6) | 9 (18) | 19 (36) | 0.29 | 0.029 | 0.035 |
| Diabetes | 0 (0) | 0 (0) | 2 (4) | 0 (0) | >0.9 | 0.17 | 0.15 |
| On lipid lowering | 0 (0) | 0 (0) | 6 (12) | 16 (30) | >0.9 | 0.005 | 0.021 |
| On antiarrhythmic | 0 (0) | 3 (4) | 21 (41) | 25 (47) | 0.56 | <0.001 | 0.54 |
| Family history of SCD | 0 (0) | 12 (17) | 13 (25) | 5 (9) | 0.023 | 0.28 | 0.042 |
| Volumes and mass | |||||||
| LVEDV index, mL/m2 | 89.3 (74-109) | 81.4 (70-94) | 71.0 (63-80) | 77.7 (68-85) | 0.07 | <0.001 | 0.087 |
| LVEF, % | 65.9 (63-69) | 70.9 (66-75) | 77.7 (73-83) | 78.6 (75-82) | 0.001 | <0.001 | 0.79 |
| MWT, mm | 9.3 (8-11) | 9.8 (9-11) | 17.0 (15-21) | 17.5 (16-22) | 0.24 | <0.001 | 0.24 |
| LV mass index, g/m2 | 56.1 (47-61) | 51.5 (43-62) | 77.2 (67-92) | 101.6 (82-127) | 0.47 | <0.001 | <0.001 |
| LV morphology | |||||||
| Isolated basal septal LVH | — | — | 23 (45) | 29 (55) | — | — | 0.33 |
| Reverse septal curvature | — | — | 22 (43) | 15 (28) | — | — | 0.11 |
| Concentric | — | — | 0 (0) | 1 (2) | — | — | 0.32 |
| Mixed apical-ASH | — | — | 6 (12) | 8 (15) | — | — | 0.62 |
| Tissue characterization | |||||||
| ECV, % | 26.9 (24-29) | 27.1 (25-30) | 29.4 (27-34) | 26.8 (25-30) | 0.79 | <0.001 | 0.009 |
| LGE present | 0 (0) | 7 (10) | 47 (92) | 51 (96) | 0.09 | <0.001 | 0.43 |
| LGE mass, gb | 0 [0-0] | 0 [0-4] | 8.4 (3-15) | 5.9 (3-14) | 0.1 | <0.001 | 0.52 |
| LGE mass %b | 0 [0-0] | 0 [0-5] | 4.9 (2-11) | 3.5 (1-7) | 0.12 | <0.001 | 0.13 |
| 12-lead ECG: measures | |||||||
| HR, beats/min | 67 (60-75) | 65 (57-76) | 66 (56-72) | 63 (57-73) | 0.52 | 0.59 | 0.59 |
| QRS, ms | 90 (87-101) | 86 (80-94) | 96 (86-104) | 98 (92-108) | 0.005 | <0.001 | 0.029 |
| PR, ms | 160 (138-175) | 154 (140-170) | 165 (147-181) | 172 (152-196) | 0.58 | 0.065 | 0.13 |
| QTc, ms | 409 ± 19 | 397 ± 22 | 423 ± 29 | 437 ± 25 | 0.22 | <0.001 | 0.011 |
| QT dispersion, ms | 48 (24-62) | 43 (20-62) | 58 (38-77) | 52 (36-68) | 0.62 | 0.006 | 0.35 |
| Amp-Sokolow, mV | 2.0 (1.5-2.2) | 2.0 (1.6-2.7) | 2.2 (1.8-3.1) | 3.0 (2.3-4.5) | 0.25 | 0.17 | <0.001 |
| Amp-Cornell, mV | 1.2 (0.9-1.5) | 1.2 (1.0-1.6) | 2.1 (1.3-2.7) | 2.7 (1.9-3.1) | 0.55 | <0.001 | 0.001 |
| 12-lead ECG: qualitative | |||||||
| Abnormal ECG | 2 (8) | 22 (31) | 41 (80) | 48 (91) | 0.019 | <0.001 | 0.14 |
| Q waves | 1 (4) | 14 (20) | 14 (27) | 14 (26) | 0.07 | 0.36 | 0.91 |
| TWI | 0 (0) | 5 (7) | 34 (67) | 40 (75) | 0.33 | <0.001 | 0.32 |
| LVH voltage criteria | 0 (0) | 6 (9) | 19 (37) | 34 (64) | 0.33 | <0.001 | 0.006 |
| ST-segment depression | 0 (0) | 0 (0) | 20 (39) | 30 (57) | >0.9 | <0.001 | 0.076 |
| RBBB | 0 (0) | 0 (0) | 1 (2) | 3 (6) | >0.9 | 0.42 | 0.62 |
| LBBB | 0 (0) | 0 (0) | 0 (0) | 5 (9) | >0.9 | >0.9 | 0.06 |
| Fractionation | 1 (4) | 3 (4) | 10 (20) | 18 (34) | >0.9 | 0.007 | 0.1 |
Values are median (Q1-Q3), n (%), mean ± SD. BoldP values are statistically significant.
Amp = amplitude; ASH = asymmetric septal hypertrophy; BMI = body mass index; BSA = body surface area; ECG = electrocardiogram; ECV = extracellular volume; HR = heart rate; LBBB = left bundle branch block; LGE = late gadolinium enhancement; LVEDV = left ventricular end-diastolic volume; LVH = left ventricular hypertrophy; MWT = maximal wall thickness; QTc = corrected QT; RBBB = right bundle branch block; TWI = T-wave inversion.
The remaining 11 healthy volunteers (HVs) (not shown here) were older to permit comparisons with the G+LVH+ group.
Values of LGE mass and LGE mass % for Younger HV and G+LVH− are median [range].
12-lead ECG
Subclinical HCM (G+LVH−) vs HV
Compared with age-matched HVs, G+LVH− had shorter QRS intervals but otherwise similar electrical intervals, QRS amplitudes, and QT dispersion. G+LVH− had a higher prevalence of ECG abnormalities (31% [22] vs 8% [2]; P = 0.019, but not after correction for multiple comparisons).
G+LVH+ vs G+LVH−
Compared with G+LVH−, G+LVH+ had similar heart rates, longer QRS and QTc intervals, greater QT dispersion, and greater Cornell amplitude (but similar Sokolow-Lyon). G+LVH+ also had a higher prevalence of ECG abnormalities including T-wave inversion (TWI), LVH by either ECG voltage criteria, ST depression, and fractionation.
G−LVH+ vs G+LVH+
Compared with G+LVH+, G−LVH+ had longer QTc intervals, higher QRS amplitudes and a higher prevalence of LVH by either ECG voltage criteria. Other 12-lead ECG intervals and prevalence of ECG abnormalities were similar (QRS duration was longer in G−LVH+ but not after correction for multiple comparisons) (Table 1).
ECGI
G+LVH− vs HV
Compared with age-matched HVs, G+LVH− had slower ventricular conduction (more prolonged mean AT) (Table 2, Figure 2). Differences persisted after adjusting for age, sex, QRS duration, LGE volume, and the presence of abnormal ECG (β = 0.42 [95% CI: 0.2–0.6]; P < 0.001). Compared with HVs, G+LVH− with a normal 12-lead ECG (n = 48) also had slower ventricular conduction (prolonged mean AT [39 ms (Q1-Q3: 35-44 ms) vs 35 ms (Q1-Q3: 31-41 ms)]; P = 0.009). There were no differences in repolarization duration (mean RTc/ARIc) or activation/repolarization dispersion (ΔAT/ΔRTc). G+LVH− had more spatially heterogenous repolarization (steeper GRTc, occurring in 23% [11 of 48] of those with normal ECGs) (Figure 3, Table 2) but similar GAT. There were no differences in signal amplitudes or fractionation.
Table 2.
ECGI Parameters in HV, Subclinical HCM, G+LVH+, and G−LVH+
| Younger HV (n = 26) | G+LVH− (n = 70) | G+LVH+ (n = 51) | G−LVH+ (n = 53) |
P Value |
|||
|---|---|---|---|---|---|---|---|
| G+LVH− vs HV | G+LVH+ vs G+LVH− | G−LVH+ vs G+LVH+ | |||||
| Conduction | |||||||
| Mean amplitude, mV | 1.4 (1.1-1.6) | 1.4 (1.2-1.7) | 1.6 (1.2-1.9) | 1.9 (1.4-2.6) | 0.4 | 0.42 | 0.12 |
| Fractionation (n/1,000) | 8 (0-22) | 8 (1-20) | 13 (2-27) | 2 (0-10) | 0.99 | 0.13 | 0.002 |
| Mean AT, ms | 35 (31-41) | 39 (35-45) | 41 (37-45) | 42 (38-47) | 0.008 | 0.23 | 0.27 |
| ΔAT, ms | 174 (153-196) | 177 (154-196) | 183 (162-208) | 188 (159-205) | 0.89 | 0.25 | 0.76 |
| Mean GAT, ms/mm | 0.42 (0.37-0.47) | 0.40 (0.31-0.49) | 0.40 (0.35-0.46) | 0.44 (0.30-0.56) | 0.38 | 0.82 | 0.33 |
| Max GAT, ms/mm | 4.7 (4.2-6.0) | 4.9 (4.2-5.6) | 5.1 (4.2-5.8) | 5.1 (4.3-6.4) | 0.74 | 0.54 | 0.27 |
| Repolarization | |||||||
| Mean ARIc, ms | 247 ± 18 | 245 ± 26 | 276 ± 29 | 281 ± 24 | 0.78 | <0.001 | 0.29 |
| ΔARIc, ms | 180 (159-197) | 181 (161-202) | 183 (162-208) | 193 (162-213) | 0.62 | 0.62 | 0.63 |
| Mean RTc, ms | 285 (274-297) | 283 (273-292) | 322 (296-340) | 327 (305-345) | 0.55 | <0.001 | 0.21 |
| ΔRTc, ms | 160 (145-179) | 164 (147-183) | 165 (150-178) | 170 (149-193) | 0.48 | 0.73 | 0.47 |
| Mean GRTc, ms/mm | 1.0 ± 0.2 | 1.1 ± 0.3 | 1.0 ± 0.3 | 1.0 ± 0.3 | 0.042 | 0.13 | 0.79 |
| Max GRTc, ms/mm | 9.5 (9.0-11) | 11.2 (10-12.9) | 11.3 (9.9-12.7) | 11.1 (9.3-14.0) | 0.005 | 0.79 | 0.92 |
Values are median (Q1-Q3) or mean ± SD. BoldP values are statistically significant.
Δ = dispersion; Amp = amplitude; ARIc = activation recovery interval corrected; AT = activation time; G = gradient; max = maximum; RTc = repolarization time corrected for heart rate; other abbreviations as in Table 1.
Figure 2.
Exemplar AT and ARIc Maps
Electrocardiographic imaging (ECGI) detects slowed ventricular conduction (prolongation of activation) even in the absence of left ventricular hypertrophy (LVH) in subclinical hypertrophic cardiomyopathy (HCM) (genotype [G]+ left ventricular hypertrophy [LVH]−). Prolongation of repolarization predominantly occurs in overt HCM (LVH+). ∗Statistically significant, P < 0.05. HV = healthy volunteer; LAO = left anterior oblique; NS = nonsignificant; RAO = right anterior oblique.
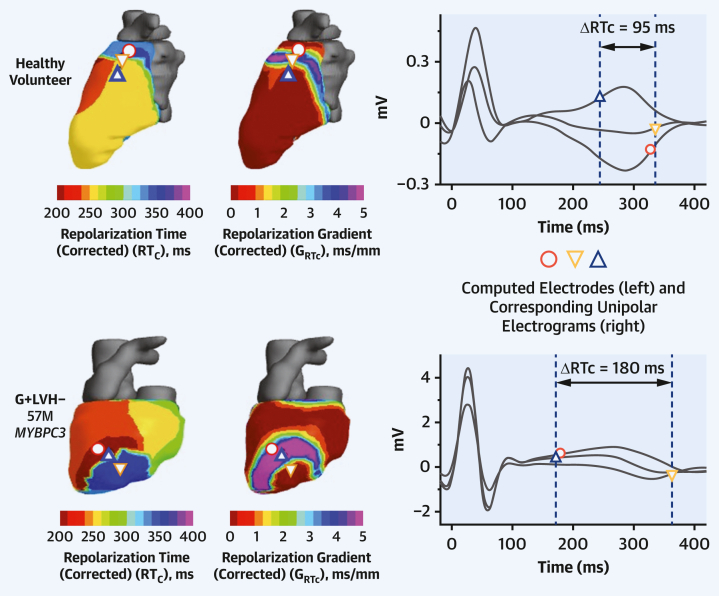
Figure 3.
Repolarization Gradient Maps
ECGI identifies concealed repolarization gradients (GRTc) occurring in a healthy volunteer and more steeply in subclinical HCM (G+LVH−). (Left) RTc maps and corresponding GRTc gradient maps are shown with superimposed shape labels showing computed electrodes. (Right) Corresponding unipolar electrograms are shown for each of the 3 computed unipolar electrodes annotated across the gradient. Abbreviations as in Figure 2.
G+LVH+ vs G+LVH−
Compared with G+LVH−, G+LVH+ had prolonged ventricular repolarization (elevated mean ARIc and RTc), and differences persisted after adjusting for age, sex, LGE volume, and QTc (ARIc: β = 0.31 [95% CI: 0.2-0.5]; P < 0.001; RTc: β = 0.28 [95% CI: 0.2-0.5]; P < 0.001). All other ECGI parameters were similar (Table 2, Figure 2). Further analyses comparing HVs matched to G+LVH+ are included in the Supplemental Results.
G−LVH+ vs G+LVH+
Compared with G−LVH+, G+LVH+ had more signal fractionation (P = 0.002) (Table 2, Figure 4), and differences persisted after adjusting for age, sex, LGE volume, and ECG fractionation (β = 0.39 [95% CI: 0.2-0.6]; P < 0.001). All other global and local ECGI parameters were similar (Table 2, Figure 2).
Figure 4.
Exemplar Fractionation Maps in HCM
G+LVH+ HCM expressing a large region of signal fractionation (A) despite no/minimal LGE (and mild LVH) by cardiovascular magnetic resonance imaging (B).
ECG vs ECGI
Across all participants, AT was associated with QRS duration (rs = 0.26 [95% CI: 0.1-0.4]; P < 0.001). Mean ARIc and RTc were both associated with QTc (mean ARIc: rs = 0.73 [95% CI: 0.7-0.8]; P < 0.001; mean RTc: rs = 0.73 [95% CI: 0.7-0.8]; P < 0.001) and QT dispersion (ARIc: rs = 0.16 [95% CI: 0.01-0.30]; P = 0.03; RTc: rs = 0.2 [95% CI: 0.05-0.30]; P = 0.008). ΔRTc was associated with QT dispersion (rs = 0.17 [95% CI: 0.01-0.30]; P = 0.03).
ECG structural relationships
12-lead ECG
In G+LVH−, MWT associated with QRS duration (rs = 0.42 [95% CI: 0.2-0.6]; P < 0.001), PR interval (rs = 0.31 [95% CI: 0.06-0.50]; P = 0.012), and Cornell but not Sokolow-Lyon amplitude (rs = 0.38 [95% CI: 0.1-0.6]; P = 0.002). In overt HCM (all LVH+), MWT associated with Cornell amplitude (rs = 0.34 [95% CI: 0.1-0.5]; P < 0.001), presence of an abnormal ECG (rs = 0.29 [95% CI: 0.1-0.5]; P = 0.019), and TWI (rs = 0.26 [95% CI: 0.07-0.40]; P = 0.007). Also, LGE associated with PR interval (rs = 0.26 [95% CI: 0.06-0.40]; P = 0.009), the presence of an abnormal ECG (rs = 0.39 [95% CI: 0.2-0.5]; P < 0.001), and TWI (rs = 0.40 [95% CI: 0.2-0.6]; P < 0.001).
ECGI relationships to risk markers
Maximal wall thickness
MWT associated with signal amplitude in HV (rs = 0.42 [95% CI: 0.3-0.7]; P = 0.032), G+LVH− (rs = 0.26 [95% CI: 0.02-0.50]; P = 0.028), and overt HCM (rs = 0.33 [95% CI: 0.1-0.5]; P < 0.001). In G+LVH−, MWT associated with mean ARIc (rs = 0.24 [95% CI: 0.003-0.50]; P = 0.046) and RTc (rs = 0.28 [95% CI: 0.04-0.50]; P = 0.019). In overt HCM, MWT associated with mean AT (rs = 0.25 [95% CI: 0.05-0.40]; P = 0.011), ΔAT (rs = 0.32 [95% CI: 0.1-0.5]; P = 0.001), ΔARIc (rs = 0.36 [95% CI: 0.2-0.5]; P < 0.001), and mean RTc (rs = 0.24 [95% CI: 0.04-0.40]; P = 0.015).
Scar burden
In overt HCM, LGE volume associated with fractionation (rs = 0.21 [95% CI: 0.01-0.40]; P = 0.032) and local AT gradients (GATmean: rs = 0.27 [95% CI: 0.08-0.40]; P = 0.005).
Left ventricular outflow tract obstruction
In total, 24 (23%) of overt HCM patients had LVOTO. No ECGI changes were associated with LVOTO.
SCD risk
To determine the relationship between ECGI abnormalities and surrogate markers of ventricular arrhythmia and risk in overt HCM, we describe a subgroup of 19 participants (18% of overt HCM) who had an intermediate/high SCD risk score or NSVT. Compared with overt HCM not meeting such criteria, these patients were similar in age (P = 0.29), sex (P = 0.74), and ethnicity (White: P = 0.80), but had a higher MWT (19.6 mm [Q1-Q3: 17-23 mm] vs 17.0 mm [Q1-Q3: 15-21 mm]; P = 0.005), more LGE (15.1 g [Q1-Q3: 8-28 g] vs 5.5 g [Q1-Q3: 2-13 g]; P < 0.001), and more spatially heterogenous conduction (GATmax P = 0.007). Other ECGI parameters were similar. After adjustment for MWT and LGE volume, those with NSVT/intermediate- or high-risk score continued to exhibit more spatially heterogenous conduction (ß = 0.29 [95% CI: 0.1–0.5]; P = 0.006). An exploratory analysis was performed for all those with NSVT (n = 16) showing similar findings (Supplemental Results).
Machine learning classification of ECGI HCM subtypes
The SVM differentiated subclinical HCM from HV with an AUC of 0.96 (bootstrap 95% CI: 94%-98%; sensitivity 100% [95% CI: 93%-100%], specificity 91% [95% CI: 76%-98%], positive predictive value 96% [95% CI: 87%-99%], negative predictive value 100% [95% CI: 87%-100%], balanced accuracy 95.7%) (Supplemental Tables 5 to 7, Supplemental Figure 1) and an accuracy of 80% after 10-fold cross-validation (95% CI: 73%-85%). This ECGI biomarker panel was able to identify HCM patients at greater risk of SCD (because of prior NSVT or intermediate/high ESC SCD risk status) compared with low-risk patients, with an AUC of 0.97 (bootstrap 95% CI: 96%-98%; sensitivity 94% [95% CI: 71%-99%], specificity 100% [95% CI: 95%-100%], positive predictive value 100% [95% CI: 77%-100%], negative predictive value 95% [95% CI: 93%-99%], and balanced accuracy 97.2%) (Supplemental Tables 5 to 8) and an accuracy of 82% after 10-fold cross-validation (95% CI: 78%-86%).
Discussion
This is a prospective multicenter study using CMR-guided ECGI to simultaneously characterize structural and EP parameters across the spectrum of genotyped HCM. The main findings are as follows: 1) EP abnormalities exist in individuals with subclinical HCM, including in those with normal resting 12-lead ECGs; and 2) in overt HCM, EP abnormalities occur more severely with adverse cardiac structural features and positive genetic status (Central Illustration). There is well-described evidence that such EP abnormalities, consisting of spatially heterogenous repolarization, slowed discontinuous ventricular conduction, and signal fractionation, are proarrhythmic,10,11,26,27 thus potentially affecting patient-specific risks for SCD in HCM. This is the largest clinical ECGI study ever performed and the only one to date to have studied subclinical HCM.
Central Illustration.
Electrophysiological Abnormalities in Hypertrophic Cardiomyopathy and Their Relationship to Risk Markers
Subclinical hypertrophic cardiomyopathy (HCM) individuals with pathogenic (P)/likely pathogenic (LP) sarcomeric variants are characterized by slowed ventricular conduction, even in the presence of a normal electrocardiogram (ECG), and spatially heterogenous repolarization. In overt HCM, conduction is regionally slowed and repolarization is prolonged and spatially heterogenous. In overt HCM, electrophysiological abnormalities relate to conventional risk markers: fractionation was worse in those with a sarcomeric mutation (G+LVH+) and more extensive scar, and spatially heterogenous conduction was related to scar extent and the presence of nonsustained ventricular tachycardia. Severity of left ventricular hypertrophy (LVH) associated with regionally slowed conduction and prolonged repolarization.
As novel HCM therapies show promise in disease modification with some suggestion that early expression of sarcomeric gene variants are potentially reversible,28,29 the discovery of sensitive biomarkers for early disease is an emerging priority. Structural changes adopted in modern-day HCM risk stratification (LA dilation, severe LVH, LVOTO, high LGE burden) tend to occur late. Furthermore, NSVT is the only EP characteristic considered in current risk stratification algorithms but has the highest predictive value for SCD; it is, however, uncommon in early disease and is a dichotomous risk marker.17 Here we show quantifiable EP abnormalities occurring in the absence of standard 12-lead ECG changes or measurable hypertrophy or fibrosis. Although cascade genetic screening allows the identification of a larger number of individuals at risk, HCM is characterized by variable penetrance and disease expression so detecting phenotype development is a priority. Due to the complexity of a 12-biomarker ECGI panel, a machine learning approach was used for discrimination between patient groups. This showed the potential of ECGI to discriminate subclinical HCM from healthy individuals and identify patients with HCM at higher risk of SCD. Future work will need to externally validate these discovered ECGI biomarker panels to understand their real-world diagnostic and prognostic potential.
Notably, EP abnormalities occurring in G+LVH− and G+LVH+ differed, likely reflecting disease heterogeneity and different phenotypic stages of disease. Longitudinal studies relating EP abnormalities to events are now possible using this more sustainable, time and cost-effective, and reuseable ECGI methodology. This may lay the foundation for ECGI-based risk stratification and provide support for decisions surrounding exercise prescription.
Ventricular conduction was slower in G+LVH− compared with HVs (increased AT) despite the absence of CMR markers of fibrosis and LVH (scar, ECV and MWT were similar to health). Importantly this occurred in a cohort predominantly without bundle branch block (2% in G+LVH+, none in G+LVH−). Furthermore, slowed ventricular conduction is missed by the 12-lead ECG occurring in both G+LVH− with normal and abnormal 12-lead ECG. In overt disease, ventricular conduction was not globally slower compared with matched HV (similar AT) but more regionally slowed (resulting in increased ΔAT) (Supplemental Results). This finding is consistent with invasive endocardial mapping findings in overt HCM which showed lateral LV activation before the septum, longer action potential durations, prolonged stimulus-to-V times and reduced action potential upstroke velocity on patch-clamp tests of HCM myocardium.5,30 Mechanisms of ventricular conduction slowing in HCM are likely to include decreased electrical coupling, discontinuous propagation, and reduced conduction velocity. Our discovery that these occur in the absence of hypertrophy and fibrosis in subclinical HCM challenges the LVH-centric view of EP abnormalities and raises the possibility that electrical changes may be more closely related to myocyte disarray, ischemia, or electromechanical factors than actual LVH.28,31 However, a compounding effect of LVH is observed in overt disease, as demonstrated by the association of MWT and both activation time and dispersion. Slowed discontinuous ventricular conduction and local conduction disturbances are a well-known proarrhythmic prerequisite for ventricular arrhythmia initiation.10,11 Our findings provide a possible reason behind the increased incidence of ventricular arrhythmia observed with greater degrees of LVH.1,17,32 Slower ventricular conduction has been detected by ECGI in other disease states with primary structural substrate changes including arrhythmogenic cardiomyopathy (AC) and cardiac amyloidosis, and overlying the right ventricular outflow tract epicardium in Brugada syndrome (although this is primarily a channelopathy, epicardial focal fibrosis has also been observed on autopsy).8, 9, 10 Furthermore, in aborted SCD survivors and Brugada syndrome, ECGI has demonstrated greater conduction heterogeneity postexercise, further demonstrating its ability to detect abnormal pro-arrhythmic rate adaptation.33
Repolarization in subclinical HCM was more spatially heterogenous (steeper repolarization gradients) when detected by ECGI, but globally, repolarization was largely normal on ECG (normal QTc, low prevalence of TWI/no ST-segment depression). Relationships between early LVH and prolonged repolarization can be observed in subclinical HCM where a positive relationship existed between MWT and ARIc. In overt disease where LVH is established, prolonged repolarization is detected in both ECG (elevated QTc) and ECGI (ARIc, RTc). In HCM mechanisms of repolarization abnormalities are likely related to ionic remodeling, which can be heterogenous because of LVH, calcium handling and sensitivity changes, which can result in action potential prolongation with early and late afterdepolarization leading to ventricular arrhythmia in structurally remodeled (hypertrophy, fibrosis, disarray) and therefore susceptible myocardium.30,31,34 The exaggerated spatially heterogenous repolarization noted in the absence of hypertrophy could be evidence of cellular and molecular changes occurring in response to sarcomeric mutation or prehypertrophic myocardial substrate changes.28,31 Repolarization gradients as seen in subclinical HCM have been observed in ventricular fibrillation survivors with structurally normal hearts.27,33 Spatially heterogenous repolarization is known to be proarrhythmic by supporting the conditions for asymmetric excitability and propagation of re-entry. This abnormality has also been demonstrated by ECGI in multiple arrhythmogenic diseases, including Brugada, AC, long QT syndrome, and early repolarization, and hypothesized in heart failure.9,10,26,35 Other ECG techniques quantifying repolarization abnormality in overt HCM include dynamic QTc changes on Holter monitoring, prolonged QTc as a predictor of ICD discharge, and QT dispersion; however, none have been especially useful in HCM, limited by lack of spatial information, technical limitations, and bias toward more advanced disease.36,37 With QTc being normal in subclinical disease, it is unlikely to afford benefits as an early disease biomarker.
Scar volume was related to spatial conduction heterogeneity (activation gradients and fractionation). This demonstrates the ability of ECGI to detect and quantify preclinical conduction discontinuities resulting from scar (not detected solely by LGE CMR). Conduction discontinuities most likely occur because of islands of viable tissue trapped within fibrotic tissue. Indeed, the fact that myocytes are electrotonically uncoupled will also serve to exaggerate cellular repolarization differences and increase repolarization gradients.38 Spatial conduction heterogeneity also associated with NSVT and SCD risk score independent of scar, showing the incremental value of ECGI for detecting proarrhythmic features. This EP abnormality has potential for arrhythmia formation3,9,11,38 through asymmetric loading on a propagating wave front favoring unidirectional block, creating dispersion of repolarization and facilitating re-entry. Both spatial conduction heterogeneity and fractionation are observed in AC scar, and fractionation is observed in both infarct and AC scar.9,11
Despite similar MWT and LGE burden and a more benign 12-lead ECG phenotype, ECGI-detected fractionation was greater in genotype-positive HCM patients compared with G−LVH+. Greater fractionation in G+LVH+ could not be accounted for by bundle branch block, suggesting a genuinely greater area of tissue with fractionated electrograms. Exaggerated spatially heterogenous ventricular conduction is another plausible contributor to the greater ventricular arrhythmia seen in G+LVH+ HCM compared with G−LVH+.2 Unexpectedly, we found more severe 12-lead ECG abnormalities in G−LVH+ compared with G+LVH+ HCM (longer QRS duration, longer QTc, and greater QRS amplitudes). This mirrors data from a prior study showing G−LVH+ was characterized by greater lateral T-wave inversion.39 Overall differences in ECG and ECGI between G− and G+ HCM could be in part related to different LV morphologies (greater LV mass despite similar MWT) and different tissue characterization (lower ECV) as described by large registry studies.2,40
Study limitations
ECGI provides epicardial EP mapping data only and does not therefore measure potentially important transmural electrophysiology; it also misses EP phenomena occurring within the septum. As expected, participants with subclinical HCM were 16 years younger than overt disease counterparts and medication use affects electrophysiology and may influence results. However, our sensitivity analyses (Supplemental Results) showed consistent EP changes even when comparing overt HCM and drug-free HCM to age-matched HVs. Although ECGI-detected EP abnormalities found in our cross-sectional study have proven associations with ventricular arrhythmia in other diseases and show the potential to identify patients at greater risk of malignant arrhythmias, hold-out data set internal validation of the SVM was not possible because of the limited sample size. To mitigate this, we undertook 10-fold cross validation, which showed good accuracy of the combined panel. Longitudinal multicenter studies are now needed to externally validate these results and confirm their generalizability. No HV underwent genetic testing. G+LVH− subjects here were recruited in the context of familial evaluation, and these findings do not apply to those with pathogenic sarcomere variants as secondary genomic findings.
Conclusions
In the absence of LVH or abnormalities on the 12-lead ECG, HCM sarcomere gene mutation carriers express an aberrant electrophysiological phenotype detected by ECGI. In overt HCM, electrophysiological abnormalities occur more severely with adverse structural change and positive genetic status.
Perspectives.
COMPETENCY IN PATIENT CARE AND PROCEDURAL SKILLS: In patients with HCM, CMR-guided electrocardiographic mapping can detect pathogenic sarcomere variants even in the absence of myocardial hypertrophy or abnormalities on the standard 12-lead ECG.
TRANSLATIONAL OUTLOOK: Abnormalities detected by this method could identify patients with subclinical disease who might benefit from therapeutic intervention.
Funding Support and Author Disclosures
Drs Joy, Webber, Guger, Orini, and Captur, and Prof Lambiase are coinventors of the patented capturECGI vest. Dr Joy is funded by a British Heart Foundation Clinical Research Training Fellowship (FS/CRTF/21/2469). Dr Lopes is supported by a Medical Research Council UK Research and Innovation Clinical Academic Research Partnership award (MR/T005181/1). Dr Dall’Armellina has received funding from a BHF Intermediate Clinical Research Fellowship (FS/13/71/30378). Prof Manisty receives funding directly and indirectly from the National Institutes of Health Research Biomedical Research Centres at University College London Hospitals and Barts Health NHS Trusts. Prof Rudy is the inventor of ECGI and receives royalties from Case Western Reserve University and Washington University in St Louis. Prof Lambiase is funded from University College London/University College London Hospitals Biomedicine National Institute for Health and Care Research, Barts Biomedical Research Centre; and has received educational grants from Abbott and Boston Scientific. Prof Moon receives funding directly and indirectly from the National Institutes of Health and Care Research Biomedical Research Centres at University College London Hospitals and Barts Health NHS trusts; is the chief executive officer of MyCardium AI Ltd; and has served on advisory boards for Sanofi and Genzyme. Dr Captur is supported by the SCMR Seed Grant Programme, the British Heart Foundation special project grant (SP/20/2/34841), the National Institute for Health and Care Research innovation for innovation iFAST grant, and the National Institute for Health and Care Research University College London Hospitals Biomedical Research Centre. All other authors have reported that they have no relationships relevant to the contents of this paper to disclose.
Acknowledgment
Source code for ML is available via GitHub (see Supplemental Statistical Methods).
Footnotes
Listen to this manuscript's audio summary by Editor-in-Chief Dr Valentin Fuster onwww.jacc.org/journal/jacc.
The authors attest they are in compliance with human studies committees and animal welfare regulations of the authors’ institutions and Food and Drug Administration guidelines, including patient consent where appropriate. For more information, visit the Author Center.
Appendix
For expanded Methods and Results section, as well as supplemental tables and a figure, please see the online version of this paper.
Appendix
References
- 1.Ommen S.R., Mital S., Burke M.A., et al. 2020 AHA/ACC guideline for the diagnosis and treatment of patients with hypertrophic cardiomyopathy. J Am Coll Cardiol. 2020;76(25):e159–e240. doi: 10.1016/j.jacc.2020.08.045. [DOI] [PubMed] [Google Scholar]
- 2.Ho C.Y., Day S.M., Ashley E.A., et al. Genotype and lifetime burden of disease in hypertrophic cardiomyopathy. Circulation. 2018;138(14):1387–1398. doi: 10.1161/CIRCULATIONAHA.117.033200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Saumarez R.C., Pytkowski M., Sterlinski M., et al. Paced ventricular electrogram fractionation predicts sudden cardiac death in hypertrophic cardiomyopathy. Eur Heart J. 2008;29(13):1653–1661. doi: 10.1093/eurheartj/ehn111. [DOI] [PubMed] [Google Scholar]
- 4.Lorenzini M., Norrish G., Field E., et al. Penetrance of hypertrophic cardiomyopathy in sarcomere protein mutation carriers. J Am Coll Cardiol. 2020;76(5):550–559. doi: 10.1016/j.jacc.2020.06.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Schumacher B., Gietzen F.H., Neuser H., et al. Electrophysiological characteristics of septal hypertrophy in patients with hypertrophic obstructive cardiomyopathy and moderate to severe symptoms. Circulation. 2005;112(14):2096–2101. doi: 10.1161/CIRCULATIONAHA.104.515643. [DOI] [PubMed] [Google Scholar]
- 6.Rudy Y. Noninvasive mapping of repolarization with electrocardiographic imaging. J Am Heart Assoc. 2021;10(9) doi: 10.1161/JAHA.121.021396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ramanathan C., Ghanem R.N., Jia P., Ryu K., Rudy Y. Noninvasive electrocardiographic imaging for cardiac electrophysiology and arrhythmia. Nat Med. 2004;10(4):422–428. doi: 10.1038/nm1011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Orini M., Graham A.J., Martinez-Naharro A., et al. Noninvasive mapping of the electrophysiological substrate in cardiac amyloidosis and its relationship to structural abnormalities. J Am Heart Assoc. 2019;8(18) doi: 10.1161/JAHA.119.012097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Andrews C.M., Srinivasan N.T., Rosmini S., et al. Electrical and structural substrate of arrhythmogenic right ventricular cardiomyopathy determined using noninvasive electrocardiographic imaging and late gadolinium magnetic resonance imaging. Circ Arrhythm Electrophysiol. 2017;10(7) doi: 10.1161/CIRCEP.116.005105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhang J., Sacher F., Hoffmayer K., et al. Cardiac electrophysiological substrate underlying the ecg phenotype and electrogram abnormalities in Brugada syndrome patients. Circulation. 2015;131(22):1950–1959. doi: 10.1161/CIRCULATIONAHA.114.013698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cuculich P.S., Zhang J., Wang Y., et al. The electrophysiological cardiac ventricular substrate in patients after myocardial infarction: noninvasive characterization with electrocardiographic imaging. J Am Coll Cardiol. 2011;58(18):1893–1902. doi: 10.1016/j.jacc.2011.07.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Graham A.J., Orini M., Zacur E., et al. Simultaneous comparison of electrocardiographic imaging and epicardial contact mapping in structural heart disease. Circ Arrhythm Electrophysiol. 2019;12(4) doi: 10.1161/CIRCEP.118.007120. [DOI] [PubMed] [Google Scholar]
- 13.Graham A.J., Orini M., Zacur E., et al. Evaluation of ECG imaging to map hemodynamically stable and unstable ventricular arrhythmias. Circ Arrhythm Electrophysiol. 2020;13(2) doi: 10.1161/CIRCEP.119.007377. [DOI] [PubMed] [Google Scholar]
- 14.Webber M., Joy G., Bennett J., et al. Technical development and feasibility of a reusable vest to integrate cardiovascular magnetic resonance with electrocardiographic imaging. J Cardiovasc Magn Reson. 2023;25:73. doi: 10.1186/s12968-023-00980-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Richards S., Aziz N., Bale S., et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17(5):405–424. doi: 10.1038/gim.2015.30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Elliott P.M., Anastasakis A., Borger M.A., et al. 2014 ESC Guidelines on diagnosis and management of hypertrophic cardiomyopathy: The Task Force for the Diagnosis and Management of Hypertrophic Cardiomyopathy of the European Society of Cardiology (ESC) Eur Heart J. 2014;35(39):2733–2779. doi: 10.1093/eurheartj/ehu284. [DOI] [PubMed] [Google Scholar]
- 17.O’Mahony C., Jichi F., Pavlou M., et al. A novel clinical risk prediction model for sudden cardiac death in hypertrophic cardiomyopathy (HCM Risk-SCD) Eur Heart J. 2014;35(30):2010–2020. doi: 10.1093/eurheartj/eht439. [DOI] [PubMed] [Google Scholar]
- 18.Orini M., Srinivasan N., Graham A.J., et al. Further evidence on how to measure local repolarization time using intracardiac unipolar electrograms in the intact human heart. Circ Arrhythm Electrophysiol. 2019;12(11) doi: 10.1161/CIRCEP.119.007733. [DOI] [PubMed] [Google Scholar]
- 19.Andršová I., Hnatkova K., Šišáková M., et al. Influence of heart rate correction formulas on QTc interval stability. Sci Rep. 2021;11(1) doi: 10.1038/s41598-021-93774-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Davies R.H., Augusto J.B., Bhuva A., et al. Precision measurement of cardiac structure and function in cardiovascular magnetic resonance using machine learning. J Cardiovasc Magn Reson. 2022;24(1):16. doi: 10.1186/s12968-022-00846-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Augusto J.B., Davies R.H., Bhuva A.N., et al. Diagnosis and risk stratification in hypertrophic cardiomyopathy using machine learning wall thickness measurement: a comparison with human test-retest performance. Lancet Digit Health. 2021;3(1):e20–e28. doi: 10.1016/S2589-7500(20)30267-3. [DOI] [PubMed] [Google Scholar]
- 22.Messroghli D.R., Moon J.C., Ferreira V.M., et al. Clinical recommendations for cardiovascular magnetic resonance mapping of T1, T2, T2∗ and extracellular volume: A consensus statement by the Society for Cardiovascular Magnetic Resonance (SCMR) endorsed by the European Association for Cardiovascular Imaging (EACVI) J Cardiovasc Magn Reson. 2017;19(1):75. doi: 10.1186/s12968-017-0389-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Macfarlane P.W., Devine B., Clark E. The University of Glasgow (Uni-G) ECG analysis program. Computers in Cardiology. 2005:451–454. [Google Scholar]
- 24.Macfarlane P.W., McLaughlin S.C., Rodger J.C. Influence of lead selection and population on automated measurement of QT dispersion. Circulation. 1998;98(20):2160–2167. doi: 10.1161/01.cir.98.20.2160. [DOI] [PubMed] [Google Scholar]
- 25.Ho C.Y., Cirino A.L., Lakdawala N.K., et al. Evolution of hypertrophic cardiomyopathy in sarcomere mutation carriers. Heart. 2016;102(22):1805. doi: 10.1136/heartjnl-2016-310015. [DOI] [PubMed] [Google Scholar]
- 26.Vijayakumar R., Silva J.N.A., Desouza K.A., et al. Electrophysiologic substrate in congenital long QT syndrome. Circulation. 2014;130(22):1936–1943. doi: 10.1161/CIRCULATIONAHA.114.011359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Cluitmans M.J.M., Bear L.R., Nguyên U.C., et al. Noninvasive detection of spatiotemporal activation-repolarization interactions that prime idiopathic ventricular fibrillation. Sci Transl Med. 2021;13(620):eabi9317. doi: 10.1126/scitranslmed.abi9317. [DOI] [PubMed] [Google Scholar]
- 28.Joy G., Moon J.C., Lopes L.R. Detection of subclinical hypertrophic cardiomyopathy. Nat Rev Cardiol. 2023;20(6):369–370. doi: 10.1038/s41569-023-00853-7. [DOI] [PubMed] [Google Scholar]
- 29.Ho C.Y., Mealiffe M.E., Bach R.G., et al. Evaluation of mavacamten in symptomatic patients with nonobstructive hypertrophic cardiomyopathy. J Am Coll Cardiol. 2020;75(21):2649–2660. doi: 10.1016/j.jacc.2020.03.064. [DOI] [PubMed] [Google Scholar]
- 30.Coppini R., Ferrantini C., Yao L., et al. Late sodium current inhibition reverses electromechanical dysfunction in human hypertrophic cardiomyopathy. Circulation. 2013;127(5):575–584. doi: 10.1161/CIRCULATIONAHA.112.134932. [DOI] [PubMed] [Google Scholar]
- 31.Joy G., Kelly C.I., Webber M., et al. Microstructural and microvascular phenotype of sarcomere mutation carriers and overt hypertrophic cardiomyopathy. Circulation. 2023;148:808–818. doi: 10.1161/CIRCULATIONAHA.123.063835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Monserrat L., Elliott P.M., Gimeno J.R., et al. Non-sustained ventricular tachycardia in hypertrophic cardiomyopathy: an independent marker of sudden death risk in young patients. J Am Coll Cardiol. 2003;42(5):873–879. doi: 10.1016/s0735-1097(03)00827-1. [DOI] [PubMed] [Google Scholar]
- 33.Shun-Shin M.J., Leong K.M.W., Ng F.S., et al. Ventricular conduction stability test: a method to identify and quantify changes in whole heart activation patterns during physiological stress. EP Eur. 2019;21(9):1422–1431. doi: 10.1093/europace/euz015. [DOI] [PubMed] [Google Scholar]
- 34.Finocchiaro G., Sheikh N., Leone, et al. Arrhythmogenic potential of myocardial disarray in hypertrophic cardiomyopathy: genetic basis, functional consequences and relation to sudden cardiac death. EP Eur. 2021;23(7):985–995. doi: 10.1093/europace/euaa348. [DOI] [PubMed] [Google Scholar]
- 35.Zhang J., Hocini M., Strom M., et al. The electrophysiological substrate of early repolarization syndrome: noninvasive mapping in patients. J Am Coll Cardiol EP. 2017;3(8):894–904. doi: 10.1016/j.jacep.2016.12.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Jalanko M., Väänänen H., Tarkiainen M., et al. Fibrosis and wall thickness affect ventricular repolarization dynamics in hypertrophic cardiomyopathy. Ann Noninvasive Electrocardiol. 2018;23(6) doi: 10.1111/anec.12582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Maron B.J., Leyhe M.J., III, Casey S.A., et al. Assessment of QT dispersion as a prognostic marker for sudden death in a regional nonreferred hypertrophic cardiomyopathy cohort. Am J Cardiol. 2001;87(1):114–115. doi: 10.1016/s0002-9149(00)01285-6. [DOI] [PubMed] [Google Scholar]
- 38.Gardner P.I., Ursell P.C., Fenoglio J.J., et al. Electrophysiologic and anatomic basis for fractionated electrograms recorded from healed myocardial infarcts. Circulation. 1985;72(3):596–611. doi: 10.1161/01.cir.72.3.596. [DOI] [PubMed] [Google Scholar]
- 39.Robyns T., Breckpot J., Nuyens D., et al. Clinical and ECG variables to predict the outcome of genetic testing in hypertrophic cardiomyopathy. Eur J Med Genet. 2020;63(3) doi: 10.1016/j.ejmg.2019.103754. [DOI] [PubMed] [Google Scholar]
- 40.Neubauer S., Kolm P., Ho C.Y., et al. Distinct subgroups in hypertrophic cardiomyopathy in NHLBI HCM Registry. J Am Coll Cardiol. 2019;74(19):2333–2345. doi: 10.1016/j.jacc.2019.08.1057. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.