Summary
Paediatric prospective coeliac disease studies with longitudinal collection of biologic samples and clinical data offer a unique perspective on disease risk. The present review highlights the wealth of information now available from international paediatric prospective studies on genetic and environmental risk factors for coeliac disease. In addition, recent omics studies have made it possible to study complex interactions between genetic and environmental factors and thereby further our insight in the etiopathogenesis of coeliac disease. In the future, paediatric prospective studies will be able to provide more detailed risk prediction models combining genes, the environment and biologic corroboration from multi-omics. They will also contribute to biomarker development and a better understanding of disease pathogenesis.
Introduction
Coeliac disease (CD) is an immune-mediated systemic disorder elicited by the ingestion of gluten-containing grains (wheat, rye, and barley).1 CD is characterized by the presence of tissue transglutaminase autoantibodies (tTGA) and endomysium autoantibodies (EMA) signifying an autoimmune process referred to as CD autoimmunity (CDA). Children with CDA may simultaneously have, or progress to characteristic mucosal lesions in the small bowel with shortening of the villi (e.g., villous atrophy), compensatory elongation of the crypts, and an increased number of intraepithelial lymphocytes which is indicative of CD.2 The variability of these mucosal changes and/or progression is exemplified in the instance of individuals with potential CD (positive tTG IgA and EMA IgA but with normal intestinal architecture defined by a Marsh score of 0 or 1 ).2,3 The largest prospective follow up study of such patients over 12 years demonstrated that a minority progress to villous atrophy, while a majority had transient, fluctuating, or even persistent autoantibody positivity without development of villous atrophy.4 According to the North American Society of Pediatric Gastroenterology, Hepatology, and Nutrition (NASPGHAN), diagnostic confirmation by intestinal biopsy is standard of care.5 However, diagnostic guidelines from the European Society of Paediatric Gastroenterology, Hepatology and Nutrition (ESPGHAN) give the option to confirm the diagnosis of CD in children with tTGA levels over ten times the upper limit of normal and a positive EMA on a separate blood draw.1,2 These highly positive serologies have been confirmed in European prospective studies to have a high positive predictive value of >99% for biopsy-proven CD.2 Although CD may clinically manifest at any age after gluten introduction, prospective birth cohort studies with longitudinal screening have consistently revealed the highest incidence between one and four years of age for both seroconversion to tTGA positivity (defined as CDA) and confirmed CD. 6–8 After this early peak, new CDA and CD cases continue to develop throughout the first decade, followed by a decline in subsequent years.9,10 In contrast, the rate of de novo seroconversion and confirmed CD cases in a large community based cohort of adults over a 10-year follow up period was 0.3% and 0.06%, respectively.11 Additional long-term prospective studies are needed to understand the natural history of seroconversion and confirmed CD diagnoses during adulthood.12
This article aims to provide a narrative review of the current evidence on the complex interactions of known environmental and genetic risk factors in CD development in children and adolescents through the lens of prospective studies. Prospective studies are the ideal platform to study early life exposures and their association with disease, in the background of genetic factors. However, the following points should be noted when considering the informative value of the different studies and interpretation of their results.
Firstly, some studies were designed primarily to explore risk factors for developing CD and were most rigorously designed to examine this outcome with respect to the number of included cases, known human leukocyte antigen (HLA) risk, and the length and intensity of follow up.
Secondly, many of the known prospective studies were initially designed to primarily evaluate environmental factors for other purposes, such as type 1 diabetes (T1D) with CD as a secondary outcome. While these are important studies with respect to the landscape of the current evidence, the inclusion criteria and outcome measures may differ from studies designed primarily to examine CD. Subsequently, certain HLA-DQ risk alleles for CD may be excluded potentially limiting the generalizability of the results.
Thirdly, with a peak incidence of CDA occurring at 1–4 years of age, potential environmental factors should be assessed from birth onwards, ideally also during pregnancy. Additionally, in order to properly determine the direction of an observed association, the study design should include prospective, frequent serological surveillance for tTGA particularly in the first years of life with timely guideline-directed CD diagnosis in those who screen positive. Relying solely on case finding based on history and/or clinical symptoms without prospective serological surveillance will leave many cases undiagnosed and misclassified as controls. Studies not designed for CD as a primary outcome may be more prone to these limitations.
Finally, results from screening studies with known CD permissive HLA-DQ genotypes can be extrapolated to the general population, acknowledging that 30–40% of the general population have a CD permissive genotype and the remaining 60–70% are HLA DQ2/8 negative and will not develop CD regardless of environmental exposures.
Considering these points, we aimed to review paediatric prospective studies which would be able to fulfil these conditions in the search for potential environmental triggers and predictive biologic markers in CD etiopathology in early childhood.
Overview of the Literature
Among 18 paediatric prospective cohort studies, five studies had the primary focus on CDA or CD as an outcome, with only four studies aiming to identify environmental triggering factors for CD; PreventCD13, CeliPrev14, Celiac Disease Genomic, Environmental, Microbiome, and Metabolomic (CDGemm)15, and Traditional Ethiopian Food study (TEF).16 (Table 1) Of note, PreventCD, CeliPrev, and CDGemm are the only studies that had a formal CD diagnosis included within the research protocol. The PROFICEL study investigated the microbiota and environmental factors in a small number of infants with known risk, without systematic screening for CDA or CD.17 Another eight studies primarily aimed to identify environmental factors in children with known risk for T1D, having CDA and/or CD as a secondary outcome. (Table 2) Six of those eight studies were informative regarding our question of environmental risk factors in CD: The Environmental Determinants of Diabetes in the Young (TEDDY)18, BabyDiab19, Diabetes Autoimmunity in the Young (DAISY)20, MIDIA21 (Norwegian acronym for environmental causes of T1D), DIABIMMUNE22 and T1D Prediction and Prevention (DIPP)23. On the other hand, Celiac Disease Prediction in Skane (CIPIS)24, 25, and BABYDIET26 did not routinely screen the whole cohort with tTGA and had a very small sample size of children with CD. Three additional population-based prospective observational birth cohorts that were not designed to examine CD, but reported factors related to CD outcomes at limited time points, were considered for the present review but with limitations (Table 3). Among those studies, the Generation R study performed a screening for CDA in the entire cohort at only around six years of age, with re-measurement of only those having positive tTGA results three years later and work up for CD if confirmed positive.27 The other cohort, the Norwegian Mother, Father, and Child Cohort Study (MoBa), followed a nationwide population-based cohort of pregnant women and their offspring, however, did not include standardized screening for CDA, and therefore, their analyses are limited by the inclusion of only clinically detected cases.28 The third cohort, All Babies In Southeast Sweden, ABIS, reported environmental factors in a large prospective birth cohort from the general population but also did not perform standardized screening for CDA, resulting in a very low prevalence of diagnosed CD in their cohort.29 Lastly, one randomized controlled intervention trial, the Enquiring About Tolerance (EAT) study, was designed to study the effect of early (age three to five months) versus later (after age six months) introduction of six different allergenic foods including wheat on the development of food allergies.30 However, tTGA was tested only at three years of age with risk of missing subsequent CDA cases and incomplete follow-up of those whom tested tTGA positive to determine whether they had confirmed CD. Regardless of the primary outcome of the study with respect to whether they were designed to study CDA and/or CD, T1D, or neither, very few studies had included a CD diagnosis protocol, leaving the decision to proceed to intestinal biopsy up to clinicians (Table 1–3). While not all children with CDA develop CD, CDA can still be used as a proxy to study risk factors that may ultimately lead to CD. This review did not include any studies examining risk factors for the development of CDA and CD during adulthood.
Table 1:
Prospective Studies in which CD is the Primary Outcome
| Study | Location | Design | Inclusion | HLA Risk Known?1 | Years Recruiting | No. of Participants | CDA Screening?2 | Timepoints of Screening3 | CD Diagnosis Method4 | CD Protocol? | References |
|---|---|---|---|---|---|---|---|---|---|---|---|
| PreventCD | Croatia, Germany, Hungary, Italy, Israel, Netherlands, Poland, Spain | Double Blind Randomized Controlled Trial-early gluten introduction at 4 months or standard | Infant <3 mos, FDR with CD | Y | 2007–2013 | 963 | Y | Screened at 4,8,12,18,24, and 36 mos, annually thereafter | Biopsy (except 5 children diagnosed based on ESPGHAN serologic criteria) | Y-biopsies reviewed by expert pathologists and given Marsh grading | 7,13,42,43,44,60,68,83,84 |
| CeliPrev | Italy | Double Blind Randomized Controlled Trial-early gluten introduction at 6 months or 12 months | Infant <3 mos, FDR with CD | Y | 2003–2009 | 707 | Y | Screened at 15 mos, and 2,3,5,8, and 10 yrs Screened until age 5 yrs at intervals of every 6 mos until age 3 yrs, annually after | Biopsy (except 5 children that were symptomatic and improve d on GFD without biopsy) | Y-biopsies were reviewed at single center with Marsh grading | 8,14,50 |
| CDGEMM | US & Italy | Observational birth cohort study | Infants <6 mos FDR with CD, not yet introduced to solid foods | Y | 2014–2022 | 554 | Y | Screened until age 5 yrs at intervals of every 6 mos until age 3 yrs, annually after | Biopsy or ESPGHAN serologic criteria at discretion of their GI clinical provider | Y-Those who screened positive on research assays twice, were also screened with an EMA IgA and referred to GI for clinical evaluation | 15 |
| PROFICEL | Spain study | Observational birth cohort | Infants 4 mos, FDR for CD | Y | 2006–2010 | 164 | N | N/A | N | N | 17 |
| TEF Study | Ethiopia | Observational birth cohort study | Infants <6 wks | Y | 2018–2022 | 1,046 | Y | Screened at 24, 36, and 48 mos | If tTG IgA positive in 2 consecutive samples, referred to PCP for further work up | N | 16 |
Abbreviations:
CD coeliac disease; CDA coeliac disease autoimmunity; mos months; EMA IgA endomysial immunoglobulin A; ESPGHAN European Society for Pediatric Gastroenterology, Hepatology, and Nutrition; NASPGHAN North American Society for Pediatric Gastroenterology, Hepatology, and Nutrition; tTG IgA tissue transglutaminase autoantibody immunoglobulin A; wks weeks; yrs years
Refers to known HLA-DQ genotype risk.
Refers to presence of screening protocol for coeliac disease autoimmunity with autoantibodies.
Refers to the included timepoints of autoantibody screening.
NASPGHAN criteria for coeliac disease includes positive autoantibody screening and a duodenal biopsy with Marsh 2 (intraepithelial lymphocytes and crypt hyperplasia) or Marsh 3 (intraepithelial lymphocytes, crypt hyperplasia, and villous atrophy) pathology. ESPGHAN criteria refers to serologic criteria for coeliac disease including a tTG IgA over ten times the upper limit of normal and positive antiendomysial IgA on a separate blood sample.
Table 2:
Prospective studies in which type 1 diabetes is the primary outcome
| Study | Location | Design | Inclusion | HLA Risk Known?1 | Years Recruiting | No. of Participants | CDA Screening?2 | Timepoints of Screening3 | CD Diagnosis Method4 | CD Protocol? | References |
|---|---|---|---|---|---|---|---|---|---|---|---|
| TEDDY | US, Germany, Finland, Sweden | Observational birth cohort | Infants <3 months, general population & FDR with T1D; HLA screening for T1D HLA-DQ | Y | 2004–2010 | 6403 | Y | Every 3 mos from 3 mos to 4 yrs old and every 6 mos from 4 yrs-15 yrs | Those with confirmed positive tTG IgA were referred for clinical evaluation or by self-defined serologic criteria | Biopsy (Marsh 2 or greater) or persistent high tTG IgA levels (tTG IgA>10 x ULN for 2 visits) | 6,10,18,40,51,52,55,57,59,61,62,69,81 |
| DAISY | US | Observational birth cohort | Infants <3 months, general population & FDR with T1D; HLA screening for T1D HLA-DQ | Y | 1993–2004 | 1339 | Y | Screened at 9, 15, and 24 mos, annually thereafter | Those with confirmed positive tTG IgA were referred for clinical evaluation or by self-defined serologic criteria | Biopsy (Marsh 2 or greater) or persistent high tTG IgA levels (tTG IgA>10 x ULN for 2 visits) | 9,20,42,64,70 |
| BabyDiab | Germany | Observational birth cohort | At birth, FDR with T1D | Y | 1989–2000 | 1511 | Y | Screened at 9 mos, 2 yrs, 5 yrs, 8 yrs, 11 yrs, and 14 yrs | Those with confirmed positive tTG IgA were referred for clinical evaluation | Biopsy | 19 |
| MIDIA | Norway | Observational birth cohort | General population infants | Y-only DQ2/8 genotype enrolled | 2001–2007 | 912 | Y | Screened stored samples to find first positive and last negative-collected at 3, 6,9, and 12 mos, annually thereafter | Those with confirmed positive tTG IgA were referred for clinical evaluation | ESPG HAN 2012 criteria-Biopsy, HLA-DQ, and tTG IgA >10 x ULN | 21,71 |
| DIPP | Finland | Observational birth cohort | Infants; HLA screening for T1D HLA-DQ | Y | 1994–2005 | 1200 | Y | Annual CDA screening | Those with confirmed positive tTG IgA were referred for clinical evaluation | Biopsy | 23,72,85,86 |
| DIABIMMUNE | Finland, Estonia, Russia | Observational birth cohort | Infants and Children 3 yrs old; HLA screening for T1D HLA-DQ | Y-only DQ2/8, DQ2/X, and DQ8/X enrolled | 2008–2013 | 5819 | Y | Screened at 3, 6, 12, 18, 24 and 36 mos | Biopsy | Biopsy | 22,74 |
| DIPIS/CIPIS | Sweden | Observational birth cohort | Infants; HLA screening for T1D HLA-DQ | Y-based on B1*02 and B1*03 | 2001–2004 | 3435 | Y | Screened at 3 yrs old with repeat testing if positive | Biopsy (Marsh 1 or greater) with improvement in symptoms or serologies after treatment | Biopsy | 24,25 |
| BabyDiet | Germany | Observational birth cohort | Infants <2 months, FDR T1D; HLA screening for T1D HLA-DQ | Y | 2000–2006 | 120 | Y | Screened at 6,12,18,24,30, and 36 mos, annually thereafter | Biopsy (Marsh 1 or greater) with improvement in symptoms or serologies after treatment | Biopsy | 26 |
Abbreviations:
CD coeliac disease; CDA coeliac disease autoimmunity; mos months; EMA IgA endomysial immunoglobulin A; ESPGHAN European Society for Pediatric Gastroenterology, Hepatology, and Nutrition; NASPGHAN North American Society for Pediatric Gastroenterology, Hepatology, and Nutrition; tTG IgA tissue transglutaminase autoantibody immunoglobulin A; T1D type 1 diabetes; wks weeks; yrs years
Refers to known HLA-DQ genotype risk.
Refers to presence of screening protocol for coeliac disease autoimmunity with autoantibodies.
Refers to the included timepoints of autoantibody screening.
NASPGHAN criteria for coeliac disease includes positive autoantibody screening and a duodenal biopsy with Marsh 2 (intraepithelial lymphocytes and crypt hyperplasia) or Marsh 3 (intraepithelial lymphocytes, crypt hyperplasia, and villous atrophy) pathology. ESPGHAN criteria refers to serologic criteria for coeliac disease including a tTG IgA over ten times the upper limit of normal and positive antiendomysial IgA on a separate blood sample.
Table 3:
Other Prospective Paediatric Studies
| Study | Location | Design | Inclusion | HLA Risk Known?1 | Years Recruiting | No. of Participants | CDA Screening?2 | Timepoints of Screening3 | CD Diagnosis Method4 | CD Protocol? | References |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Prospective General Population Birth Cohort Studies Investigating Chronic Illnesses | |||||||||||
| Generation R | Netherlands | Birth Cohort | Infants | Y | 2002–2006 | 4,442 | Y | Screened at 6 yrs with a repeat screen at 9 yrs if positive | Referred for clinical evaluation if tTG IgA positive at 9 yrs old | Biopsy | 27,34,58,73 |
| MoBa | Norway | Birth Cohort | Pregnant women and their infants | N | 2000–2009 | 113,053 | N | NA | Clinical Case Finding | Unknown | 28,45,48,49,53,54,63,67 |
| ABIS | Sweden | Birth Cohort | Infants | N | 1997–1999 | 16,286 | N | NA | Clinical Case Finding | Biopsy | 29,47,66 |
| Randomized Controlled Trials for Food Allergy and Food Triggered Disease Prevention | |||||||||||
| EAT | UK | Randomized Clinical Trial-early high dose gluten introduction at 4–6 months compare d to after 6 months | Infants 3 months and exclusively breastfed | N | 2012–2015 | 1,004 | Y | 3 yrs | Clinical Referral to GI if tTG IgA higher than 3 times normal (moderate to high positive) | ESPG HAN 2020 serologic criteria | 30 |
Abbreviations:
CD coeliac disease; CDA coeliac disease autoimmunity; mos months; EMA IgA endomysial immunoglobulin A; ESPGHAN European Society for Pediatric Gastroenterology, Hepatology, and Nutrition; NASPGHAN North American Society for Pediatric Gastroenterology, Hepatology, and Nutrition; tTG IgA tissue transglutaminase autoantibody immunoglobulin A; T1D type 1 diabetes; wks weeks; yrs years
Refers to known HLA-DQ genotype risk.
Refers to presence of screening protocol for coeliac disease autoimmunity with autoantibodies.
Refers to the included timepoints of autoantibody screening.
NASPGHAN criteria for coeliac disease includes positive autoantibody screening and a duodenal biopsy with Marsh 2 (intraepithelial lymphocytes and crypt hyperplasia) or Marsh 3 (intraepithelial lymphocytes, crypt hyperplasia, and villous atrophy) pathology. ESPGHAN criteria refers to serologic criteria for coeliac disease including a tTG IgA over ten times the upper limit of normal and positive antiendomysial IgA on a separate blood sample.
Genetic risk factors
HLA, family history of CD, female sex and geographic region are all important factors with variable genetic contributions to risk.18 Approximately 50% of the heritability of CD can be explained by genetic variation, with the HLA-DQ region attributing the most. Also contributing are non-MHC loci and additional independent loci within the MHC region identified through fine mapping and genome-wide association studies (GWAS).31,32 Prospective birth cohort studies enrolling newborns with HLA-DQ2 (which includes combinations of DQ2·5, 2·2, and 7·5) and DQ8 genotypes have identified children at highest risk and allowed risk stratification over a period of time.13, 14, 18 TEDDY enrolled children with CD-permissive HLA-DQ2 and DQ8 haplotypes and found that children homozygous for HLA-DQ2 were over five times as likely to develop CD compared to the lower risk HLA-DQ8 heterozygous or homozygous children. HLA-DQ2 heterozygosity conferred a more intermediate risk for CD. This risk stratification has also been found in other studies as well as the additional risk conferred for CD by HLA-DQ7·5/2·2 (equivalent to HLA-DQ2·5 in the trans confirmation) and the HLA-DQ2·2 allele more broadly.7,9, 14, 33
However, the heterogeneity in geographic risk for CD for any given HLA genotype underscores the importance of non-HLA genetic and epigenetic risk factors as well as environmental factors. CD risk among identical HLA-DQ genotypes were not equivalent in children from different geographic regions. In TEDDY, children from Sweden had the highest risk of developing CD independent of their HLA-DQ genotype and children from Colorado had the highest risk in the United States.10 Other prospective studies have similarly found differences in incidence based on region and ethnicity.22, 34
Genome-wide association studies and fine mapping of the MHC region have contributed greatly to the characterization of the non-HLA-DQ risk factors for CD and require large numbers of subjects that are not feasible in birth cohort studies.32, 35, 36 Over 40 loci outside the HLA-region have been shown to be associated with CD.37 In TEDDY, 6010 children were analysed using the Immunochip, and 54 single-nucleotide polymorphisms (SNPs) in 5 genes were associated with CD development over time (TAGAP, IL18R1, RGS21, PLEK, and CCR9).38 SNPs that mapped to the PKIA and PFKFB3 regions met genome-wide significance only in Sweden suggesting that non-HLA risk may be at least in part accountable for regional variability in CD incidence. The identified genes have important functions in T-cell activation and homing, IL-2 regulation, and interferon gamma production. Interestingly, in other studies leveraging pathway analyses, genetic variation in interferon gamma signalling has also been found to be associated with CD development.39 Other non-HLA genes have also been implicated in other autoimmune diseases like rheumatoid arthritis, T1D, and multiple sclerosis and offer a potential mechanism for co-occurring autoimmune conditions.
While these analyses have highlighted risk factors for CD development, other studies have also shed light on potential protective factors. A nested case-control study found that the presence of HLA-DPB1*04:01 may protect genetically susceptible children, specifically those with HLA-DQ2·5, from developing CDA.40 This underscores the importance of understanding the biological interactions of the HLA-DQ region with HLA non-DQ variants for better prospective risk prediction of CD in genetically susceptible children. Birth cohort studies can also be used to help validate genetic risk scores, leveraging genetic data from prior GWAS studies. The addition of celiac risk SNPs improves disease prediction superior to HLA risk stratification alone.41,42 Longitudinal cohort studies in CD can also serve to identify biologic events that occur prior to disease onset. In PreventCD, circulating microRNA (miRNA) were measured in CD patients versus controls, which identified 53 significant miRNAs in circulation that represent potential biomarker candidates, including eight miRNA detected in circulation even before tTGA detection. These findings require confirmation and future implementation strategies to see how such data could be used for disease prediction in birth cohort studies.43 In a separate study utilizing an Italian cohort of newborns from at-risk families and including some children from the PreventCD cohort, gene expression was analyzed from 9 celiac disease-candidate genes at time points prior to diagnosis, at the time of CD diagnosis and at least 1 year after gluten-free diet. From this study, three genes (KIAA, TAGAP, and SH2B3) were found to be overexpressed months before CD diagnosis. This was a small cohort number, with only a single highly variable pre-disease time point and a limited number of genes investigated. This is an example of how longitudinal birth cohort studies can begin to develop a molecular-based disease prediction model.44 The further use of omics is discussed later in this review as part of future studies highlighting the potential of prospective studies.
Environmental risk factors
Prospective studies offer a unique perspective on the complex interplay of genetic and environmental risk factors (Figure 1). The detailed collection of early life exposures and frequent serial biologic testing allow researchers to analyse the relationship between specific risk factors and the development of disease.
Figure 1: Published Risk Factors From Prospective Coeliac Disease Studies Impacting Later Coeliac Disease Development.
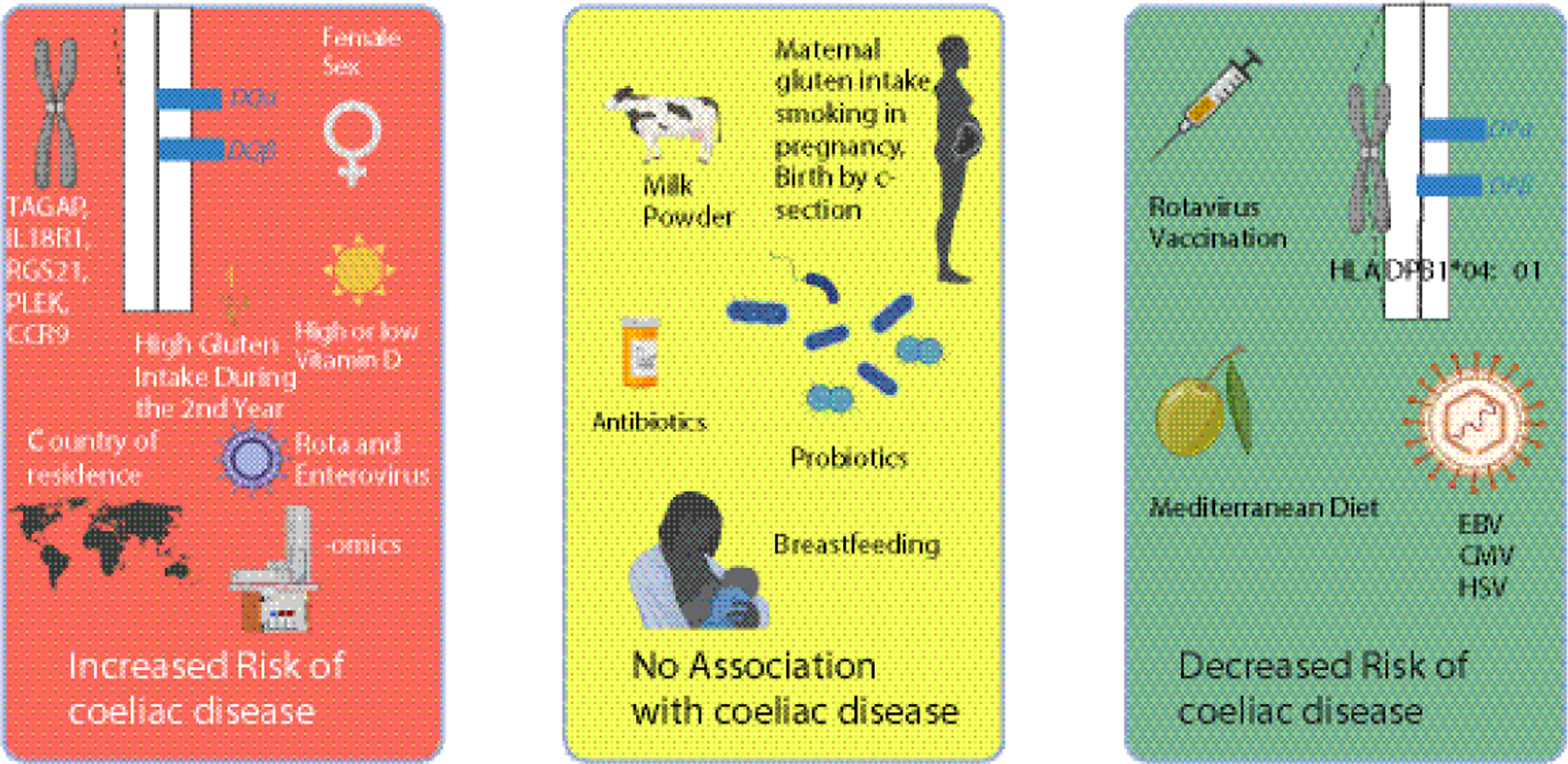
This figure highlights genetic and environmental risk factors discussed in this review from prospective coeliac disease studies that increase the risk of coeliac disease (red box), decrease the risk of coeliac disease (green box), or do not impact coeliac disease risk (yellow box).
Perinatal exposures
Perinatal factors, which include events and conditions occurring during pregnancy and early infancy, have been studied in several registry-based and prospective birth cohorts with emphasis on how maternal and infant characteristics and mode of delivery influence the risk of CD development later in childhood. Maternal factors have been extensively examined without conclusive results. Data linked to medical birth registries have indicated that risk of CD in children increase with maternal age as well as level of education and maternal CD, 45,46 but this may also be confounded by factors such as maternal hospitalization, medication during pregnancy and mode of delivery, which could not be accounted for in these retrospective studies. In the MoBa cohort, neither maternal infections nor maternal antibiotics during pregnancy were associated with CD in children.47,48 Smoking during pregnancy also did not influence CD risk.49 It has previously been hypothesized that the mode of delivery may impact CD risk due to alterations in the intestinal microbiome. This has extensively been investigated in birth registry studies and cohorts and it has been consistently found that caesarean section delivery does not increase CD risk. 50,51 Thus, there is little evidence from prospective birth cohorts that these perinatal factors may be involved in CD pathogenesis.
Maternal dietary exposures during pregnancy have been suggested to influence a child’s CD risk. Gluten consumption during pregnancy, particularly fiber-rich foods, and whole-grain cereals, may modify gut microbiome diversity. So far, evidence is limited and findings contradictory. One birth cohort study of children with known genetic CD risk found no association with maternal gluten consumption, while another general population birth cohort study found that higher gluten intake during pregnancy was associated with increased CD risk for children.28,52 Data about maternal dietary supplementation during pregnancy (specifically vitamin D, omega-3 fatty acids, and iron) and risk of CD in the offspring is also scarce and disparate due to differences in inclusion criteria and study design.53–55
Dietary exposures during early infancy
Breastmilk contains bioactive and immunological factors that protect against infections and contribute to healthy microbial colonization that in turn promote the growth of beneficial bacteria in the infant’s gut. Breastfeeding has been proposed to be potentially protective against CD development and the impact of breastfeeding on CD development has been extensively studied in both RCTs and cohort studies. These studies have consistently demonstrated that breastfeeding duration (exclusive or any) does not change the risk of developing CD.56
Similar to common food allergens, it was suggested that early gluten introduction may promote tolerance and protect from CD development.20 Subsequent observational and two RCTs have shown that timing of gluten introduction (both early and late) is not associated with a cumulative risk of CD development.13, 14, 56 EAT reported that early introduction with high amounts of gluten would decrease the risk of CD by the age of three years.30 However, this British RCT was not designed to test CD as an outcome and had major limitations limiting its generalizability and caution should be taken in interpretation of the results.
It has been hypothesized that vitamin D deficiency can also play an important role in CD pathogenesis in children due to dysregulation of the immune system. Low maternal vitamin D levels during pregnancy, due to lack of UV light exposures and consumption of vitamin D rich foods may impact CD risk and in childhood, through an association with the timing of gluten introduction and seasonal viral infections. A nested case-control study of the TEDDY cohort found that both low and high vitamin D concentrations were associated with increased risk of CDA. This points to the complexity of Vitamin D effects on the immune system as both low and high levels likely impact CD risk through immune sequelae.57
Dietary exposures during childhood
Dietary patterns may have both inflammatory and anti-inflammatory properties. Western diets are known to have intestinal inflammatory effects that could in turn be associated with CD. A Mediterranean-like diet, rich in vegetables, grains, and vegetable oils has been associated with lower risk of CDA, pointing towards the importance of studying the whole diet.58 High gluten-containing sources such as bread and milk-based cereal drinks during the second year of life was associated with increased risk of CDA and CD.59 In Italian participants of PreventCD, children who developed CD reported dietary intake that contained higher amounts of refined sugars and lower intake of legumes, vegetables, and fruits compared to those without CD. 60
While the timing of gluten introduction does not seem to impact CD risk, a higher compared to a lower amount of ingested gluten during the second year of life was consistently associated with both subsequent CDA and CD development in three prospective studies (TEDDY, PreventCD, and DAISY).6, 56, 60–64
Role of infections
Several studies have investigated the association between infections and CD, with a particular focus on respiratory and gastrointestinal infections. Many of these studies are limited by a cross-sectional study design and the subsequent conclusions are limited by risk of reverse causality (i.e., infections may actually occur after seroconversion to CDA).65 Results from prospective birth cohorts with longitudinal testing for CDA have also demonstrated an association between infections and the risk of developing CDA and/or CD.66 However, there has been interstudy variability amongst birth cohorts regarding the timing and nature of the questionnaires, biologic samples utilized to determine the presence of infection and the period of follow up for the study outcomes (Supplementary Appendix, p. 3–7). Both PreventCD and MoBa found an association between frequent parent-reported early life respiratory infections and subsequent CD.67,68 In contrast, the TEDDY study found no such association between parent-reported respiratory infection and subsequent CDA.69
Interestingly, both MoBa and TEDDY found an association between parent-reported gastrointestinal infections and subsequent risk of CDA and CD.67,69 It follows that in TEDDY, rotavirus vaccination was associated with a decreased incidence of CDA. This reduction was only observed in children introduced to gluten before six months of age, highlighting potential complex interactions between infections and diet in CD.69 Similarly, DAISY demonstrated that frequent rotavirus infections determined through the presence of serum rotavirus antibodies was also associated with subsequent CDA.70 Varied results from multiple cohort studies and complex interactions with dietary factors suggests that multiple viruses may be implicated and that many environmental risk factors may modify subsequent CDA and/or CD development.
In contrast to parent-reported infections only, there are a few prospective study designs that include longitudinal biological samples for more objective evidence of viral infection. Three independent studies (MIDIA, TEDDY, and DIPP) demonstrate an association between enterovirus and subsequent CDA and/or CD through stool samples collected before age 3 years(TEDDY and MIDIA) and serum samples collected 24 months prior to seroconversion (DIPP).62, 71, 72 The Norwegian MIDIA study also found an association between frequent parechovirus infection and subsequent CD.21 Interestingly, TEDDY found an interaction between enterovirus infection between 1–2 years of age and early life cumulative gluten intake on subsequent CDA development.62 While early studies suggested a possible association with adenovirus serotype 12, this association was not confirmed in two later prospective cohort studies.71, 72
There are some prospective studies that have found no association between CD risk and viral infections or even a decreased risk with certain viral infections based on biologic samples. In the Generation R study, cytomegalovirus (CMV) or combined CMV, Epstein-Barr virus (EBV) and/or herpes virus infection was associated with subsequent decreased incidence of CD.73 However, this protective association should be interpreted with caution as Generation R screened for CD and obtained viral serology at just a single time point of age six years. DIABIMMUNE analysed nasal swabs and stool samples for common viruses and virus-specific serum antibodies and found no evidence of an association with CD and viral infection including both CMV and EBV.74
While mechanistic studies to causally implicate viral infections in CD pathogenesis are largely lacking and the exact mechanisms by which viruses contribute to the development of autoimmune disease are still not fully understood, different pathways have been proposed as potential mechanisms.75 These include molecular mimicry between viral antigens and self-antigens, viral-mediated destruction of tissue and subsequent release of self-antigens, mucosal barrier dysfunction, and chronic viral infection and shedding which results in the release of self-antigens (bystander and epitope spreading mechanisms). Moreover, it is plausible that distinct viruses launch a type 1 interferon response leading to an inflammatory T cell response and inhibition of regulatory T cells as has been shown for reovirus-mediated abrogation of gluten tolerance in a murine model.76
Thus, several of the prospective studies indicate that viral infections, particularly enterovirus and rotavirus infections, may be involved in the pathogenesis of CD. The exact role of viral infections in CD pathogenesis remains unclear, and there is a lack of human studies on underlying mechanisms.
Other environmental factors
Other factors during infancy include season of birth and antibiotic and probiotic exposures. There is a moderately, albeit significant increased CD risk in individuals born in the spring or summer months.77,78 It has been hypothesized that exposure to certain seasonal viral infections during a time window when the infantś immune system develops tolerance to food antigens may influence CD risk. This is further supported by the interactions observed between season of birth, early infectious exposures, timing of gluten and CDA development in the TEDDY cohort.69
Antibiotics could also disrupt the gut microbiota, leading to dysregulation of the immune system and potentially triggering CD in genetically susceptible individuals. While registry-based cohort studies have suggested antibiotic usage as a risk factor for CD,46, 79, 80 a prospective birth cohort could not confirm this; there was no association seen between cumulative antibiotic use and CDA risk in TEDDY.69 Similarly, while it is thought that probiotics can help lead to a restoration of the microbiota balance and potentially protect against autoimmunity development, early probiotic use in the first year of life was not associated with decreased risk of CDA in the TEDDY cohort.81
Role of omics in the prediction of CD
Omics span many different modalities such as genomics, epigenomics, transcriptomics, proteomics, metabolomics and microbiome that collectively aim at characterization and quantification of pools of biological molecules that translate into the structure, function, and dynamics of cell, tissue or an organism to holistically study complex biological processes and disease mechanisms. These high-throughput technologies have been exploited in CD research although mostly in cross-sectional studies and certainly not to their full potential in prospective study settings. In order to harness the role of omics in CD prediction, prospective studies that allow analysis of samples collected prior to disease development are essential.
Metabolomics entails the large-scale analysis of small molecules including amino acids, carbohydrates, nucleotides, lipids, coenzymes, and cofactors within a given compartment. There is a fair number of studies exploiting metabolomics in CD that vary considerably in terms of the used sample matrix (serum, plasma, urine, stool), analysis methodology and most of the studies have been conducted in cross-sectional designs.82 Only PreventCD and DIPP have leveraged samples collected in prospective birth cohort studies to study metabolomics. PreventCD initially reported no difference in serum metabolites between individuals who later developed CD and those who did not based on serum amino acids and polar lipids.83 However, a subsequent sub-cohort study found lysophostidylcholines, ether-linked phosphatidylcholine and some phospatidylcholines differed between the two groups.84 The study did not correct for confounding factors and the analysis was targeted to only a limited number of lipids in different groups therefore limiting the conclusions.
DIPP conducted longitudinal lipidomics profiling of plasma samples and reported increased amounts of triacylglycerols and a decreased level of phosphatidylcholines in CD progressors as early as three months of age, even before the introduction of gluten to the diet.85 A further study found that perfluorinated alkyl substances (PFAs), while not a natural metabolite, were elevated significantly from birth to three months of age only in the CD group, and that distinct PFA profiles correlated with triacylglycerol concentrations particularly of those with saturated fatty acids.86 Subsequently, exposure to environmental toxicants such as PFA exposure may modulate lipid metabolism, which also correlates with later CD development.
Although the microbiome entails all micro-organisms including bacteria, fungi, archaea, and even viruses in a given anatomical site, this section will focus on bacteria, as viral infections were previously covered. Conducted studies have significant differences in the applied methodology (high-throughput vs low throughput) and sampling site (duodenum vs feces).87 Two individual studies focusing solely on fecal microbiome in paediatric CD cases followed prospectively from birth have been published. A Finnish study that analysed stool microbiome from samples collected at nine and 12 months of age, revealed no differences between those who later developed CD and those who did not.88 On the other hand, fecal microbiome profiling of samples collected at four and six months of age as a part of the prospective PROFICEL study identified differences for those who subsequently developed CD.17 Children who did not develop CD during the follow up had an increase in bacterial diversity over time, characterized by increases in particular bacterial strains.
Since the gut microbiome is an independent metabolically active “organ” with profound effects on host metabolism, it is particularly meaningful to integrate microbiome and metabolomic datasets. Few studies have combined intestinal microbiome characterization with metabolomics in prospective cohorts. Some studies have been limited by very small sample sizes (i.e. only one CD case) and poorly timed sample collection (i.e. the samples were collected after CD development).89 CDGEMM analysed fecal microbiota, functional pathways, and metabolites in prospectively collected samples including samples obtained prior to disease development.15 The analysis comprised samples collected at the time of diagnosis as well as 3, 6, 9, 12, and 18 months prior to CD development. Intrasubject longitudinal analysis in CD cases revealed increased abundance of Dialister invisus, Parabacteroides sp., Lachnospiraceae bacterium and serine and threonine metabolites, which have also been linked to other autoimmune and inflammatory conditions and suggest a role of the microbiota and resulting metabolites in CD development. In the future, multi-omics approaches with models that also include environmental exposures and clinical data should also be undertaken. Such multi-level data integration would ensure the most robust model for CD development. Longitudinal omics-studies with larger samples sizes are also needed to validate the referenced findings.
Conclusion
Genetic factors including HLA-DQ haplotypes, non-HLA SNPs, female sex, and to some degree country of residence play a significant role in disease risk. Prospective studies have identified environmental factors including vitamin D levels, early life infections, specific infant dietary patterns and in particular, early life cumulative gluten intake in the first two years of life to be associated with increased risk for CD development. Other factors including HLA-DPB*04:01, rotavirus vaccination, and other dietary patterns have been associated with decreased risk. Finally, many other previously-reported risk factors have been disproven through these prospective studies and additional exposomes including toxic exposures, epigenetics, and biologic perinatal exposures remain unexplored.
It remains to be determined how much of an impact these environmental factors affect risk in real-world settings and whether they could be exploited to prevent disease. It is more likely that these factors will need to be accounted for however, in clinical trials of young children. Of particular interest should be the role of repeated early life gastrointestinal infections (especially enterovirus) and cumulative gluten intake in the background of genetic risk. The complexities of genetic and environmental interactions supported by biologic elements will require machine learning to develop robust disease models of prediction.90 Herein lies the true potential of prospective cohorts; setting the stage for risk modeling and biomarker discovery that may also aid in disease pathogenesis and development of clinical intervention trials in order to prevent CD in at-risk individuals in the near future.
Supplementary Material
Search Strategy and Selection Criteria:
References for this review were identified through searches of PubMed with the MeSH search terms “coeliac disease” and “cohort” from January 1993 until March 2023. This yielded 2,329 articles from which articles that included prospective studies and risk factors for CD development were selected. Only papers published in English were reviewed. Articles deemed important for describing potential associations later examined in birth cohort studies that included national registries and other retrospective data were also included. Overall, we identified 18 prospective cohort studies which were reviewed in depth.
Key Messages.
Prospective paediatric birth cohorts enable platforms for the study of risk factors and biologic markers of coeliac disease autoimmunity (CDA) and coeliac disease (CD).
Eighteen paediatric prospective studies have identified or confirmed several key risk factors and biologic markers associated with CD development during childhood.
This includes genetic and demographic factors such as HLA-DQ haplotypes, non-HLA SNPs, and female sex; and environmental factors including serum vitamin D, early life infections (particularly Enterovirus), a high early life cumulative gluten intake, and to some degree country of residence.
The study of and use of omic measures in these studies are still somewhat preliminary, but may provide important biologic insight into CD development.
Future studies should combine these risk and protective factors contributing to CD development for improved risk prediction models and application to future CD prevention trials.
Acknowledgements:
The TEDDY Study is funded by U01 DK63829, U01 DK63861, U01 DK63821, U01 DK63865, U01 DK63863, U01 DK63836, U01 DK63790, UC4 DK63829, UC4 DK63861, UC4 DK63821, UC4 DK63865, UC4 DK63863, UC4 DK63836, UC4 DK95300, UC4 DK100238, UC4 DK106955, UC4 DK112243, UC4 DK117483, U01 DK124166, U01 DK128847, and Contract No. HHSN267200700014C from the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK), National Institute of Allergy and Infectious Diseases (NIAID), Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD), National Institute of Environmental Health Sciences (NIEHS), Centers for Disease Control and Prevention (CDC), and JDRF. This work is supported in part by the NIH/NCATS Clinical and Translational Science Awards to the University of Florida (UL1 TR000064) and the University of Colorado (UL1 TR002535). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Declarations: MS is a consultant for Pfizer (DSMB for celiac disease clinical trial) and Takeda (celiac disease advisory board).
SK is a consultant for Danone, has received honoraria from Mead Johnson, Takeda, Nestle and Pfizer, and has participated on advisory boards for Takeda, Sanofi, Danone, GlaxoKlineSmith, and Janssen.
CAA has no conflict of interest.
KL serves on the Scientific Advisory Board of The Finnish Celiac Society and serves as Treasurer of the Multi Society Celiac Disease Consortium.
EL is a consultant for Takeda and serves on the Scientific Advisory Board for the Celiac Disease Foundation and Beyond Celiac.
DA is a consultant for Takeda and serves on the Scientific Advisory Board for Allero Therapeutics and the Swedish Celiac Disease Research Foundation.
Guarantor of the article: Daniel Agardh
RESOURCES
- 1.Husby S, Koletzko S, Korponay-Szabó IR, Mearin ML, Phillips A, Shamir R, et al. European Society for Pediatric Gastroenterology, Hepatology, and Nutrition guidelines for the diagnosis of coeliac disease. J Pediatr Gastroenterol Nutr. 2012;54(1):136–60. [DOI] [PubMed] [Google Scholar]
- 2.Husby S, Koletzko S, Korponay-Szabó I, Kurppa K, Mearin ML, Ribes-Koninckx C, et al. European Society Paediatric Gastroenterology, Hepatology and Nutrition Guidelines for Diagnosing Coeliac Disease 2020. J Pediatr Gastroenterol Nutr. 2020;70(1):141–56. [DOI] [PubMed] [Google Scholar]
- 3.Ludvigsson JF, Leffler DA, Bai JC, Biagi F, Fasano A, Green PH, et al. The Oslo definitions for coeliac disease and related terms. Gut. 2013;62(1):43–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Auricchio R, Mandile R, Del Vecchio MR, Scapaticci S, Galatola M, Maglio M, et al. Progression of Celiac Disease in Children With Antibodies Against Tissue Transglutaminase and Normal Duodenal Architecture. Gastroenterology. 2019;157(2):413–20.e3. [DOI] [PubMed] [Google Scholar]
- 5.Hill ID, Fasano A, Guandalini S, Hoffenberg E, Levy J, Reilly N, et al. NASPGHAN Clinical Report on the Diagnosis and Treatment of Gluten-related Disorders. J Pediatr Gastroenterol Nutr. 2016;63(1):156–65. [DOI] [PubMed] [Google Scholar]
- 6.Andrén Aronsson C, Lee HS, Hård Af Segerstad EM, Uusitalo U, Yang J, Koletzko S, et al. Association of Gluten Intake During the First 5 Years of Life With Incidence of Celiac Disease Autoimmunity and Celiac Disease Among Children at Increased Risk. JAMA. 2019;322(6):514–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Meijer CR, Auricchio R, Putter H, Castillejo G, Crespo P, Gyimesi J, et al. Prediction Models for Celiac Disease Development in Children From High-Risk Families: Data From the PreventCD Cohort. Gastroenterology. 2022;163(2):426–36. [DOI] [PubMed] [Google Scholar]
- 8.Lionetti E, Castellaneta S, Francavilla R, Pulvirenti A, Naspi Catassi G, Catassi C. Long-Term Outcome of Potential Celiac Disease in Genetically at-Risk Children: The Prospective CELIPREV Cohort Study. J Clin Med. 2019;8(2). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Liu E, Dong F, Barón AE, Taki I, Norris JM, Frohnert BI, et al. High Incidence of Celiac Disease in a Long-term Study of Adolescents With Susceptibility Genotypes. Gastroenterology. 2017;152(6):1329–36.e1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Stahl M, Li Q, Lynch K, Koletzko S, Mehta P, Gragert L, et al. Incidence of Pediatric Celiac Disease Varies by Region. Am J Gastroenterol. 2023;118(3):539–45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Choung RS, Khaleghi S, Cartee AK, Marietta EV, Larson JJ, King KS, et al. Community-Based Study of Celiac Disease Autoimmunity Progression in Adults. Gastroenterology. 2020;158(1):151–9.e3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Catassi C, Kryszak D, Bhatti B, Sturgeon C, Helzlsouer K, Clipp SL, et al. Natural history of celiac disease autoimmunity in a USA cohort followed since 1974. Ann Med. 2010;42(7):530–8. [DOI] [PubMed] [Google Scholar]
- 13.Vriezinga SL, Auricchio R, Bravi E, Castillejo G, Chmielewska A, Crespo Escobar P, et al. Randomized feeding intervention in infants at high risk for celiac disease. N Engl J Med. 2014;371(14):1304–15. [DOI] [PubMed] [Google Scholar]
- 14.Lionetti E, Castellaneta S, Francavilla R, Pulvirenti A, Tonutti E, Amarri S, et al. Introduction of gluten, HLA status, and the risk of celiac disease in children. N Engl J Med. 2014;371(14):1295–303. [DOI] [PubMed] [Google Scholar]
- 15.Leonard MM, Valitutti F, Karathia H, Pujolassos M, Kenyon V, Fanelli B, et al. Microbiome signatures of progression toward celiac disease onset in at-risk children in a longitudinal prospective cohort study. Proc Natl Acad Sci U S A. 2021;118(29). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gudeta AN, Andrén Aronsson C, Binagdie BB, Girma A, Agardh D. Incidence of celiac disease autoimmunity and associations with maternal tuberculosis and pediatric Helicobacter pylori infections in 4-year-old Ethiopian children followed up in an HLA genotyped birth cohort. Front Pediatr. 2022;10:999287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Olivares M, Walker AW, Capilla A, Benítez-Páez A, Palau F, Parkhill J, et al. Gut microbiota trajectory in early life may predict development of celiac disease. Microbiome. 2018;6(1):36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Liu E, Lee HS, Aronsson CA, Hagopian WA, Koletzko S, Rewers MJ, et al. Risk of pediatric celiac disease according to HLA haplotype and country. N Engl J Med. 2014;371(1):42–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hummel S, Hummel M, Banholzer J, Hanak D, Mollenhauer U, Bonifacio E, et al. Development of autoimmunity to transglutaminase C in children of patients with type 1 diabetes: relationship to islet autoantibodies and infant feeding. Diabetologia. 2007;50(2):390–4. [DOI] [PubMed] [Google Scholar]
- 20.Norris JM, Barriga K, Hoffenberg EJ, Taki I, Miao D, Haas JE, et al. Risk of celiac disease autoimmunity and timing of gluten introduction in the diet of infants at increased risk of disease. JAMA. 2005;293(19):2343–51. [DOI] [PubMed] [Google Scholar]
- 21.Tapia G, Chudá K, Kahrs CR, Stene LC, Kramna L, Mårild K, et al. Parechovirus Infection in Early Childhood and Association With Subsequent Celiac Disease. Am J Gastroenterol. 2021;116(4):788–95. [DOI] [PubMed] [Google Scholar]
- 22.Simre K, Uibo O, Peet A, Tillmann V, Kool P, Hämäläinen AM, et al. Exploring the risk factors for differences in the cumulative incidence of coeliac disease in two neighboring countries: the prospective DIABIMMUNE study. Dig Liver Dis. 2016;48(11):1296–301. [DOI] [PubMed] [Google Scholar]
- 23.Simell S, Hoppu S, Simell T, Ståhlberg MR, Viander M, Routi T, et al. Age at development of type 1 diabetes- and celiac disease-associated antibodies and clinical disease in genetically susceptible children observed from birth. Diabetes Care. 2010;33(4):774–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Björck S, Brundin C, Lörinc E, Lynch KF, Agardh D. Screening detects a high proportion of celiac disease in young HLA-genotyped children. J Pediatr Gastroenterol Nutr. 2010;50(1):49–53. [DOI] [PubMed] [Google Scholar]
- 25.Björck S, Lynch K, Brundin C, Agardh D. Repeated Screening Can Be Restricted to At-Genetic-Risk Birth Cohorts. J Pediatr Gastroenterol Nutr. 2016;62(2):271–5. [DOI] [PubMed] [Google Scholar]
- 26.Hummel S, Pflüger M, Hummel M, Bonifacio E, Ziegler AG. Primary dietary intervention study to reduce the risk of islet autoimmunity in children at increased risk for type 1 diabetes: the BABYDIET study. Diabetes Care. 2011;34(6):1301–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Jansen M, van Zelm M, Groeneweg M, Jaddoe V, Dik W, Schreurs M, et al. The identification of celiac disease in asymptomatic children: the Generation R Study. J Gastroenterol. 2018;53(3):377–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lund-Blix NA, Tapia G, Mårild K, Brantsæter AL, Eggesbø M, Mandal S, et al. Maternal fibre and gluten intake during pregnancy and risk of childhood celiac disease: the MoBa study. Sci Rep. 2020;10(1):16439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ludvigsson JF, Ludvigsson J. Stressful life events, social support and confidence in the pregnant woman and risk of coeliac disease in the offspring. Scand J Gastroenterol. 2003;38(5):516–21. [PubMed] [Google Scholar]
- 30.Logan K, Perkin MR, Marrs T, Radulovic S, Craven J, Flohr C, et al. Early Gluten Introduction and Celiac Disease in the EAT Study: A Prespecified Analysis of the EAT Randomized Clinical Trial. JAMA Pediatr. 2020;174(11):1041–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kumar V, Wijmenga C, Withoff S. From genome-wide association studies to disease mechanisms: celiac disease as a model for autoimmune diseases. Semin Immunopathol. 2012;34(4):567–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Gutierrez-Achury J, Zhernakova A, Pulit SL, Trynka G, Hunt KA, Romanos J, et al. Fine mapping in the MHC region accounts for 18% additional genetic risk for celiac disease. Nat Genet. 2015;47(6):577–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Fernández-Fernández S, Borrell B, Cilleruelo ML, Tabares A, Jiménez-Jiménez J, Rayo AI, et al. Prevalence of Celiac Disease in a Long-term Study of a Spanish At-genetic-risk Cohort From the General Population. J Pediatr Gastroenterol Nutr. 2019;68(3):364–70. [DOI] [PubMed] [Google Scholar]
- 34.Jansen MAE, Beth SA, van den Heuvel D, Kiefte-de Jong JC, Raat H, Jaddoe VWV, et al. Ethnic differences in coeliac disease autoimmunity in childhood: the Generation R Study. Arch Dis Child. 2017;102(6):529–34. [DOI] [PubMed] [Google Scholar]
- 35.Dubois PC, Trynka G, Franke L, Hunt KA, Romanos J, Curtotti A, et al. Multiple common variants for celiac disease influencing immune gene expression. Nat Genet. 2010;42(4):295–302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Trynka G, Hunt KA, Bockett NA, Romanos J, Mistry V, Szperl A, et al. Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nat Genet. 2011;43(12):1193–201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Withoff S, Li Y, Jonkers I, Wijmenga C. Understanding Celiac Disease by Genomics. Trends Genet. 2016;32(5):295–308. [DOI] [PubMed] [Google Scholar]
- 38.Sharma A, Liu X, Hadley D, Hagopian W, Liu E, Chen WM, et al. Identification of Non-HLA Genes Associated with Celiac Disease and Country-Specific Differences in a Large, International Pediatric Cohort. PLoS One. 2016;11(3):e0152476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kumar V, Gutierrez-Achury J, Kanduri K, Almeida R, Hrdlickova B, Zhernakova DV, et al. Systematic annotation of celiac disease loci refines pathological pathways and suggests a genetic explanation for increased interferon-gamma levels. Hum Mol Genet. 2015;24(2):397–409. [DOI] [PubMed] [Google Scholar]
- 40.Hadley D, Hagopian W, Liu E, She JX, Simell O, Akolkar B, et al. HLA-DPB1*04:01 Protects Genetically Susceptible Children from Celiac Disease Autoimmunity in the TEDDY Study. Am J Gastroenterol. 2015;110(6):915–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sharp SA, Jones SE, Kimmitt RA, Weedon MN, Halpin AM, Wood AR, et al. A single nucleotide polymorphism genetic risk score to aid diagnosis of coeliac disease: a pilot study in clinical care. Aliment Pharmacol Ther. 2020;52(7):1165–73. [DOI] [PubMed] [Google Scholar]
- 42.Romanos J, Rosén A, Kumar V, Trynka G, Franke L, Szperl A, et al. Improving coeliac disease risk prediction by testing non-HLA variants additional to HLA variants. Gut. 2014;63(3):415–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Tan IL, Coutinho de Almeida R, Modderman R, Stachurska A, Dekens J, Barisani D, et al. Circulating miRNAs as Potential Biomarkers for Celiac Disease Development. Front Immunol. 2021;12:734763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Galatola M, Cielo D, Panico C, Stellato P, Malamisura B, Carbone L, et al. Presymptomatic Diagnosis of Celiac Disease in Predisposed Children: The Role of Gene Expression Profile. J Pediatr Gastroenterol Nutr. 2017;65(3):314–20. [DOI] [PubMed] [Google Scholar]
- 45.Emilsson L, Magnus MC, Størdal K. Perinatal risk factors for development of celiac disease in children, based on the prospective Norwegian Mother and Child Cohort Study. Clin Gastroenterol Hepatol. 2015;13(5):921–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Canova C, Zabeo V, Pitter G, Romor P, Baldovin T, Zanotti R, et al. Association of maternal education, early infections, and antibiotic use with celiac disease: a population-based birth cohort study in northeastern Italy. Am J Epidemiol. 2014;180(1):76–85. [DOI] [PubMed] [Google Scholar]
- 47.Mårild K, Ludvigsson J, Sanz Y, Ludvigsson JF. Antibiotic exposure in pregnancy and risk of coeliac disease in offspring: a cohort study. BMC Gastroenterol. 2014;14:75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Mårild K, Kahrs CR, Tapia G, Stene LC, Størdal K. Maternal Infections, Antibiotics, and Paracetamol in Pregnancy and Offspring Celiac Disease: A Cohort Study. J Pediatr Gastroenterol Nutr. 2017;64(5):730–6. [DOI] [PubMed] [Google Scholar]
- 49.Mårild K, Tapia G, Midttun Ø, Ueland PM, Magnus MC, Rewers M, et al. Smoking in pregnancy, cord blood cotinine and risk of celiac disease diagnosis in offspring. Eur J Epidemiol. 2019;34(7):637–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lionetti E, Castellaneta S, Francavilla R, Pulvirenti A, Catassi C. Mode of Delivery and Risk of Celiac Disease: Risk of Celiac Disease and Age at Gluten Introduction Cohort Study. J Pediatr. 2017;184:81–6.e2. [DOI] [PubMed] [Google Scholar]
- 51.Koletzko S, Lee HS, Beyerlein A, Aronsson CA, Hummel M, Liu E, et al. Cesarean Section on the Risk of Celiac Disease in the Offspring: The Teddy Study. J Pediatr Gastroenterol Nutr. 2018;66(3):417–24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Uusitalo U, Lee HS, Aronsson CA, Yang J, Virtanen SM, Norris J, et al. Gluten consumption during late pregnancy and risk of celiac disease in the offspring: the TEDDY birth cohort. Am J Clin Nutr. 2015;102(5):1216–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Mårild K, Tapia G, Haugen M, Dahl SR, Cohen AS, Lundqvist M, et al. Maternal and neonatal vitamin D status, genotype and childhood celiac disease. PLoS One. 2017;12(7):e0179080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Størdal K, Haugen M, Brantsæter AL, Lundin KE, Stene LC. Association between maternal iron supplementation during pregnancy and risk of celiac disease in children. Clin Gastroenterol Hepatol. 2014;12(4):624–31.e1–2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Yang J, Tamura RN, Aronsson CA, Uusitalo UM, Lernmark Å, Rewers M, et al. Maternal use of dietary supplements during pregnancy is not associated with coeliac disease in the offspring: The Environmental Determinants of Diabetes in the Young (TEDDY) study. Br J Nutr. 2017;117(3):466–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Szajewska H, Shamir R, Stróżyk A, Chmielewska A, Zalewski BM, Auricchio R, et al. Systematic review: early feeding practices and the risk of coeliac disease. A 2022 update and revision. Aliment Pharmacol Ther. 2023;57(1):8–22. [DOI] [PubMed] [Google Scholar]
- 57.Andrén Aronsson C, Liu X, Norris JM, Uusitalo U, Butterworth MD, Koletzko S, et al. 25(OH)D Levels in Infancy Is Associated With Celiac Disease Autoimmunity in At-Risk Children: A Case-Control Study. Front Nutr. 2021;8:720041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Barroso M, Beth SA, Voortman T, Jaddoe VWV, van Zelm MC, Moll HA, et al. Dietary Patterns After the Weaning and Lactation Period Are Associated With Celiac Disease Autoimmunity in Children. Gastroenterology. 2018;154(8):2087–96.e7. [DOI] [PubMed] [Google Scholar]
- 59.Hård Af Segerstad EM, Liu X, Uusitalo U, Agardh D, Andrén Aronsson C. Sources of dietary gluten in the first 2 years of life and associations with celiac disease autoimmunity and celiac disease in Swedish genetically predisposed children: The Environmental Determinants of Diabetes in the Young (TEDDY) study. Am J Clin Nutr. 2022;116(2):394–403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Auricchio R, Calabrese I, Galatola M, Cielo D, Carbone F, Mancuso M, et al. Gluten consumption and inflammation affect the development of celiac disease in at-risk children. Sci Rep. 2022;12(1):5396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Andrén Aronsson C, Lee HS, Koletzko S, Uusitalo U, Yang J, Virtanen SM, et al. Effects of Gluten Intake on Risk of Celiac Disease: A Case-Control Study on a Swedish Birth Cohort. Clin Gastroenterol Hepatol. 2016;14(3):403–9.e3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Lindfors K, Lin J, Lee HS, Hyöty H, Nykter M, Kurppa K, et al. Metagenomics of the faecal virome indicate a cumulative effect of enterovirus and gluten amount on the risk of coeliac disease autoimmunity in genetically at risk children: the TEDDY study. Gut. 2020;69(8):1416–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Lund-Blix NA, Mårild K, Tapia G, Norris JM, Stene LC, Størdal K. Gluten Intake in Early Childhood and Risk of Celiac Disease in Childhood: A Nationwide Cohort Study. Am J Gastroenterol. 2019;114(8):1299–306. [DOI] [PubMed] [Google Scholar]
- 64.Mårild K, Dong F, Lund-Blix NA, Seifert J, Barón AE, Waugh KC, et al. Gluten Intake and Risk of Celiac Disease: Long-Term Follow-up of an At-Risk Birth Cohort. Am J Gastroenterol. 2019;114(8):1307–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Størdal K, Kahrs C, Tapia G, Agardh D, Kurppa K, Stene LC. Review article: exposure to microbes and risk of coeliac disease. Aliment Pharmacol Ther. 2021;53(1):43–62. [DOI] [PubMed] [Google Scholar]
- 66.Welander A, Tjernberg AR, Montgomery SM, Ludvigsson J, Ludvigsson JF. Infectious disease and risk of later celiac disease in childhood. Pediatrics. 2010;125(3):e530–6. [DOI] [PubMed] [Google Scholar]
- 67.Mårild K, Kahrs CR, Tapia G, Stene LC, Størdal K. Infections and risk of celiac disease in childhood: a prospective nationwide cohort study. Am J Gastroenterol. 2015;110(10):1475–84. [DOI] [PubMed] [Google Scholar]
- 68.Auricchio R, Cielo D, de Falco R, Galatola M, Bruno V, Malamisura B, et al. Respiratory Infections and the Risk of Celiac Disease. Pediatrics. 2017;140(4). [DOI] [PubMed] [Google Scholar]
- 69.Kemppainen KM, Lynch KF, Liu E, Lönnrot M, Simell V, Briese T, et al. Factors That Increase Risk of Celiac Disease Autoimmunity After a Gastrointestinal Infection in Early Life. Clin Gastroenterol Hepatol. 2017;15(5):694–702.e5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Stene LC, Honeyman MC, Hoffenberg EJ, Haas JE, Sokol RJ, Emery L, et al. Rotavirus infection frequency and risk of celiac disease autoimmunity in early childhood: a longitudinal study. Am J Gastroenterol. 2006;101(10):2333–40. [DOI] [PubMed] [Google Scholar]
- 71.Kahrs CR, Chuda K, Tapia G, Stene LC, Mårild K, Rasmussen T, et al. Enterovirus as trigger of coeliac disease: nested case-control study within prospective birth cohort. BMJ. 2019;364:l231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Oikarinen M, Puustinen L, Lehtonen J, Hakola L, Simell S, Toppari J, et al. Enterovirus Infections Are Associated With the Development of Celiac Disease in a Birth Cohort Study. Front Immunol. 2020;11:604529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Jansen MA, van den Heuvel D, van der Zwet KV, Jaddoe VW, Hofman A, Escher JC, et al. Herpesvirus Infections and Transglutaminase Type 2 Antibody Positivity in Childhood: The Generation R Study. J Pediatr Gastroenterol Nutr. 2016;63(4):423–30. [DOI] [PubMed] [Google Scholar]
- 74.Simre K, Uibo O, Peet A, Puustinen L, Oikarainen S, Tamminen K, et al. Early-life exposure to common virus infections did not differ between coeliac disease patients and controls. Acta Paediatr. 2019;108(9):1709–16. [DOI] [PubMed] [Google Scholar]
- 75.Smatti MK, Cyprian FS, Nasrallah GK, Al Thani AA, Almishal RO, Yassine HM. Viruses and Autoimmunity: A Review on the Potential Interaction and Molecular Mechanisms. Viruses. 2019;11(8). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Bouziat R, Hinterleitner R, Brown JJ, Stencel-Baerenwald JE, Ikizler M, Mayassi T, et al. Reovirus infection triggers inflammatory responses to dietary antigens and development of celiac disease. Science. 2017;356(6333):44–50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Ivarsson A, Hernell O, Nyström L, Persson LA. Children born in the summer have increased risk for coeliac disease. J Epidemiol Community Health. 2003;57(1):36–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Lebwohl B, Green PH, Murray JA, Ludvigsson JF. Season of birth in a nationwide cohort of coeliac disease patients. Arch Dis Child. 2013;98(1):48–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Mårild K, Ye W, Lebwohl B, Green PH, Blaser MJ, Card T, et al. Antibiotic exposure and the development of coeliac disease: a nationwide case-control study. BMC Gastroenterol. 2013;13:109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Dydensborg Sander S, Nybo Andersen AM, Murray JA, Karlstad Ø, Husby S, Størdal K. Association Between Antibiotics in the First Year of Life and Celiac Disease. Gastroenterology. 2019;156(8):2217–29. [DOI] [PubMed] [Google Scholar]
- 81.Uusitalo U, Andren Aronsson C, Liu X, Kurppa K, Yang J, Liu E, et al. Early Probiotic Supplementation and the Risk of Celiac Disease in Children at Genetic Risk. Nutrients. 2019;11(8). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Vacca M, Porrelli A, Calabrese FM, Lippolis T, Iacobellis I, Celano G, et al. How Metabolomics Provides Novel Insights on Celiac Disease and Gluten-Free Diet: A Narrative Review. Front Microbiol. 2022;13:859467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Kirchberg FF, Werkstetter KJ, Uhl O, Auricchio R, Castillejo G, Korponay-Szabo IR, et al. Investigating the early metabolic fingerprint of celiac disease - a prospective approach. J Autoimmun. 2016;72:95–101. [DOI] [PubMed] [Google Scholar]
- 84.Auricchio R, Galatola M, Cielo D, Amoresano A, Caterino M, De Vita E, et al. A Phospholipid Profile at 4 Months Predicts the Onset of Celiac Disease in at-Risk Infants. Sci Rep. 2019;9(1):14303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Sen P, Carlsson C, Virtanen SM, Simell S, Hyöty H, Ilonen J, et al. Persistent Alterations in Plasma Lipid Profiles Before Introduction of Gluten in the Diet Associated With Progression to Celiac Disease. Clin Transl Gastroenterol. 2019;10(5):1–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Sinisalu L, Sen P, Salihović S, Virtanen SM, Hyöty H, Ilonen J, et al. Early-life exposure to perfluorinated alkyl substances modulates lipid metabolism in progression to celiac disease. Environ Res. 2020;188:109864. [DOI] [PubMed] [Google Scholar]
- 87.Verdu EF, Schuppan D. Co-factors, Microbes, and Immunogenetics in Celiac Disease to Guide Novel Approaches for Diagnosis and Treatment. Gastroenterology. 2021;161(5):1395–411.e4. [DOI] [PubMed] [Google Scholar]
- 88.Rintala A, Riikonen I, Toivonen A, Pietilä S, Munukka E, Pursiheimo JP, et al. Early fecal microbiota composition in children who later develop celiac disease and associated autoimmunity. Scand J Gastroenterol. 2018;53(4):403–9. [DOI] [PubMed] [Google Scholar]
- 89.Sellitto M, Bai G, Serena G, Fricke WF, Sturgeon C, Gajer P, et al. Proof of concept of microbiome-metabolome analysis and delayed gluten exposure on celiac disease autoimmunity in genetically at-risk infants. PLoS One. 2012;7(3):e33387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.MacEachern SJ, Forkert ND. Machine learning for precision medicine. Genome. 2021;64(4):416–25. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


