Abstract
The epidermis is the body’s first line of protection against dehydration and pathogens, continually regenerating the outermost protective skin layers throughout life. Both during embryonic development and wound healing, epidermal stem and progenitor cells must respond to external stimuli and insults to build, maintain, and repair the cutaneous barrier. Recent advances in CRISPR-based methods for cell lineage tracing have remarkably expanded the potential for experiments that track stem and progenitor cell proliferation and differentiation over the course of tissue and even organismal development. Additional tools for DNA-based recording of cellular signaling cues promise to deepen our understanding of the mechanisms driving normal skin morphogenesis and response to stressors, as well as the dysregulation of cell proliferation and differentiation in skin diseases and cancer. In this review, we highlight cutting-edge methods for cell lineage tracing, including in organoids and model organisms, and explore how cutaneous biology researchers might leverage these techniques to elucidate the developmental programs that support the regenerative capacity and plasticity of the skin.
Keywords: CRISPR, DNA barcoding, DNA typewriter, Genome editing, Prime editor
Introduction
As the external armor of the body, the skin plays a vital role in protecting terrestrial organisms from the assaults of diverse and swiftly changing environments to prevent dehydration, thermal instability, pathogen invasion, and UV damage (Sun et al. 2014a; Hsu and Fuchs 2022). To restore the body’s external barrier after wounding, the skin can rapidly rebuild the epidermis, but cutaneous appendages like sweat glands or hair follicles are lost in scarred adult tissue (Gay et al. 2013; Chen et al. 2020). Understanding the precise order in which skin stem cells divide during development and wound repair and tracking the cell type identities of their progeny would allow researchers to build a regenerative blueprint of the skin. Such a developmental roadmap of conserved morphogenetic programs, and how they are disrupted in dermatologic disease, could be harnessed to enhance healing of the epidermis but also to engineer fully functional skin appendages, a long-standing goal in the field of cutaneous biology.
The Promise and Limitations of Lineage Tracing Methods
Historical methods of lineage tracing in the skin include classic “pulse-chase” experiments in which cells or mice were injected with a labeled probe that can be tracked over time. These methods provided key insights into fundamental processes such as the cyclic stages of hair growth and regression and the organization of epidermal proliferating units (EPUs) (Potten and Allen 1975; Cotsarelis et al. 1990; Alcolea and Jones 2013). Later lineage tracing experiments utilized fluorescent markers (e.g., histone H2B-GFP) to pulse epidermal cells and calculate fluorescence dilution to estimate cell divisions (Tumbar et al. 2004). To add a further level of pulse control, investigators used Cre recombinase driven by tissue- and cell type-specific promoters to drive expression of a fluorescent reporter (e.g., tdTomato) in select cells that could then be visually chased over time (Alcolea and Jones 2013; Sada et al. 2016). However, important limitations of these methods include their low-throughput nature, eventual dilution of the marker used in “pulse-chase” experiments over many cell divisions, and leaky expression of Cre causing off-target labeling (Wang et al. 2023). More recently, optical imaging methods that rely on fluorescently labeled cells, advanced microscopy, and computational tracking algorithms have allowed researchers to follow individual cells in vivo to determine their fates (Rompolas et al. 2012; Cockburn et al. 2022). Live microscopy traces a continuous trajectory of cell fate, offering one of the highest temporal resolution methods of lineage tracing. However, light microscopy-based methods are limited in the number and type of cells they can track within an optical window using spectrally distinct fluorophores.
Approaching the challenge of lineage tracing from a different angle, scientists have developed computational tools that can approximate cellular differentiation trajectories through analysis of single-cell profiling methods. Transcriptional profiling via single-cell RNA sequencing (scRNA-seq) can richly describe a cell’s state and nominate specific biological pathways as important for its development via gene ontology analysis. By comparing transcriptional similarities, these methods can be used to define specific cell types, infer differentiation states, and estimate signaling pathway activation (Pollen et al. 2014; Herring et al. 2018; Calderon et al. 2022). Although computational methods for scRNA-seq can retrospectively infer developmental trajectories and intercellular communication networks, gene expression alone is a static measurement of cell state, does not directly reveal the routes taken to achieve a specific fate, and requires destruction of the sample - this is akin to taking a single high resolution photograph rather than a movie. Thus, laborious time-courses must be performed to map cellular trajectories, which are still marred by considerable uncertainty (Efremova et al. 2020; Lun and Bodenmiller 2020; Hu et al. 2021; Jin et al. 2021). While “RNA velocity” analysis methods can be used to generate pseudotime estimates (or “inferred trajectories”), including in murine skin (Joost et al. 2018), their conclusions are limited to inference rather than direct and prospective recording of lineage relationships (Haghverdi et al. 2016; Fleck et al. 2023; Heumos et al. 2023).
In recent years, enormous strides in genome engineering methods as well as in nucleotide sequencing capacity, efficiency, and cost have enabled the development of DNA sequence-based lineage tracing methods that can more directly capture the lineage relationships among thousands or potentially millions of cells (Kris A. Wetterstrand 2019). The improved accuracy of high-throughput DNA sequencing has also made it feasible for scientists to track cell lineage through naturally occuring somatic mutations in the nuclear or mitochondrial genome. The phylogenetic concepts used to derive lineage trees from natural and engineered mutations are similar; however, locus-specific engineered single mutations or short nucleotide sequence insertions (barcodes) only require sequencing a defined locus or set of loci rather than many scattered loci to extract sufficient information to build complete lineage maps. Methods to directly track cell lineage using engineered barcodes have emerged through creative applications of serine and tyrosine recombinases, CRISPR/Cas9 editors, base editors, and prime editors to alter endogenous DNA sequences in a defined manner (Frieda et al. 2017; Hwang et al. 2019; Chow et al. 2021; Loveless et al. 2021a; Choi et al. 2022).
Importantly, these lineage tracing technologies can still be multiplexed with single-cell and spatial mRNA sequencing at the time of tissue or organism harvest to add historical context, not only capturing the end-fate of cells, but also revealing where they came from and how they arrived at their final destination. Moreover, adaptation of these methods have been used to engineer DNA-based recording systems (akin to a tape recorder). These catalog biologically meaningful data, such as signaling cascade activation, in a sequential manner within intact tissues or organisms over time to document the type and order of intrinsic and extrinsic cues received by cells (Frieda et al. 2017; Tang and Liu 2018; Chen et al. 2021b). Here, we summarize how novel DNA editing tools work, how they have been utilized to map cell lineage in other organ systems, and how they could be leveraged to draft a comprehensive morphogenetic blueprint of normal skin, to engineer fully functional grafts and appendages, and to understand cutaneous malformations, disease pathology, and carcinogenesis.
Principles of DNA Sequencing-Based Platforms for Lineage Tracing
Sophisticated tools for genome engineering have increasingly allowed scientists to precisely insert, delete, and modify nucleotides at specific loci (Doudna and Charpentier 2014; Komor et al. 2016; Anzalone et al. 2019). While the proposed use of these tools has primarily been focused on editing native DNA sequences, such as the correction of disease-associated-mutations, they have also enabled completely new methods for lineage tracing. By inserting DNA barcodes, researchers can now edit the genome to distinctly label cells and their progeny in a manner that can be deciphered through routine nucleotide sequencing (Masuyama et al. 2019; Frangoul et al. 2021; Urnov 2021). In methods that apply static barcodes, lineage relationships (similar to family trees) can be retroactively constructed by labeling individual cells in a founder population with a unique marker (Figure 1A), for example, through lentiviral delivery of a single, unique DNA barcode or a fluorophore (Livet et al. 2007; Hampel et al. 2011; Bhang et al. 2015). In the Brainbow method, founder cells undergo fluorescent labeling using stochastic Cre/loxP recombinase activity to randomize the expression of genetically encoded fluorophores in individual cells, thus marking a founder and its progeny with a distinct color combination (Livet et al. 2007). However, because the label remains unchanged (i.e., static) in a founder clone’s descendents across generations, one can link all progeny only by their shared founder. This is analogous to mapping one’s ancestry using only a surname that was passed down, which restricts the ability to reconstruct the more complicated branching of family trees. In contrast, lineage tracing methods that utilize dynamic generation of barcodes to cumulatively label cells over multiple generations are able to accurately define intricate branching lineage hierarchies (Figure 1B). Herein, we primarily discuss dynamic barcoding and the genome editing tools commonly used to generate cumulative barcode diversity.
Figure 1: Dynamic DNA barcoding enhances lineage tracing capacity and can be multiplexed with single-cell transcriptomics.
(a) Static barcoding introduces a unique barcode per cell in a population during a single labeling event (e.g., lentiviral transduction); the founder cell’s barcode is then stably passed on to all its progeny. (b) Dynamic barcoding continually diversifies a DNA barcode over multiple generations of cells to permit reconstruction of a more complex lineage tree. (c) Dynamic lineage tracing in tissues like the epidermis requires introduction of a genome editor (e.g., Cas9), an array of DNA targets that become the edited barcode, and guide RNA(s) to direct the genome editor to the targets; the lineage of all cells within the tissue can be deciphered by the similarity of their final edited barcode. (d) By embedding a DNA barcode (BC) under the control of a promoter that recruits RNA polymerase-II (e.g., placing the barcode within the 5’ or 3’ UTR of a fluorescent protein or other selection marker driven by a constitutive promoter), the barcode itself is transcribed into mRNA including a poly-A tail; thus, when capturing the entire transcriptome with a poly-dT primer in scRNAseq, the barcode is read along with all other mRNAs so that a cell’s transcriptional state can be captured along with its lineage.
Although nested recombinase systems and multiple rounds of lentiviral barcode delivery have both been utilized to generate more complex lineage trees (He et al. 2017; Kong et al. 2020; Jindal et al. 2023), most dynamic barcoding methods heavily rely on CRISPR/Cas9 and its derivative editor systems. CRISPR/Cas9 requires guide RNAs (gRNA) to direct DNA-cutting enzymes to a specific genomic locus, where Cas9 cleaves both strands of DNA (Doudna and Charpentier 2014). The resulting double-stranded break (DSB) will usually be corrected by endogenous repair processes like non-homologous end-joining (NHEJ), but DNA repair will occasionally generate mutations at the site of the DSB, typically nucleotide insertions and/or deletions (collectively referred to as “indels”). These changes typically remain stable since they no longer match the gRNA and are not targeted again by Cas9; indels that disrupt a gene’s function are commonly used in a knock-out strategy. Instead of relying on random indels, CRISPR/Cas9 users can add a donor DNA template that includes the desired changes from the host sequence. The donor DNA contains complementary regions on both sides of the host target sequence, which allows homology directed repair (HDR) machinery to use the donor DNA as the template for repair, thus incorporating the desired changes to the host DNA sequence. For a more thorough review of basic CRISPR gene editing methods, we direct the reader to a previous article in this series in the Journal of Investigative Dermatology (Guitart et al. 2016).
Subsequent iterations of genome engineering technologies allow investigators to insert, delete, or modify specific short nucleotide sequences in a chosen genomic location. When fused to DNA modifiers such as transfer RNA (tRNA) adenosine deaminases, cytidine deaminases, or reverse transcriptases, the CRISPR-Cas9 system acts as a navigator to direct these enzymes to a specific locus, which allows targeted adenine base editing, cytosine base editing, and prime editing, respectively. Prime editing was developed by fusing a reverse-transcriptase domain to a Cas9-nickase (Anzalone et al. 2019), which “nicks” DNA, generating a single-stranded DNA break within the genomic target site that is encoded by the prime editing guide RNA (pegRNA). The pegRNA also contains a homology arm, which helps guide the insertion to the correct location in the genomic DNA, and a reverse transcriptase template (RTT) that contains a primer binding site, which primes the enzyme to reverse transcribe the pegRNA template into the nicked DNA strand including the desired insertion or deletion. Thus, the endogenous target locus and the desired nucleotide edits are both encoded in the same pegRNA. Endogenous repair mechanisms then incorporate the edited DNA strand into both native strands of genomic DNA, thus maintaining the desired nucleotide edits in subsequent rounds of DNA replication during mitosis. Later iterations of prime editing have offered increasing editing efficiency (Chen et al. 2021a; Doman et al. 2023).
DNA sequencing-based lineage tracing methods capitalize on the stochastic nature of target site editing by enzymes like CRISPR/Cas9 or more precise insertion of specific DNA sequences with prime editors, both of which result in diverse DNA edits among progenitor cells that are then passed down and can accumulate additional DNA edits in subsequent generations (Figure 1C). The first Cas9-based methods to generate dynamic DNA barcodes for tracking cell lineages relied on NHEJ to accumulate random deletions and/or insertions in an engineered DNA barcode over many cellular generations (McKenna et al. 2016; Perli et al. 2016; Kalhor et al. 2017; Schmidt et al. 2017; Spanjaard et al. 2018; Chan et al. 2019; Loveless et al. 2021b) (Figure 1D). Relationships between cells could then be computationally inferred from how similar or different the resultant barcode sequences are to one another. These methods require that a tandem array of Cas9 target sites be inserted into the genome to expand the number of available target sites that can be stochastically edited. After cells grow or differentiate and are harvested, routine DNA sequencing, scRNA-seq, or fluorescence in situ hybridization (FISH) can be used to read the resultant edits within the target sites (Frieda et al. 2017; Askary et al. 2020; Chow et al. 2021) and compare them among different cells within the biological system (e.g., organoid, tissue, tumor, or whole model organism) to infer a cell lineage tree. Despite its benefits over pulse-chase and optical forms of lineage tracing, this technology has important limitations. Cas9-based editing relies on double-stranded DNA breaks, which, if not properly repaired, can potentially compromise cellular health or promote carcinogenesis (Carroll 2018; Haapaniemi et al. 2018). In addition, due to multiple DSBs occuring at once, larger portions of the barcode can be unintentionally deleted, resulting in loss of lineage information.
Prime editing-based methods of lineage tracing, such as DNA Typewriter and peCHYRON (prime editing Cell HistorY Recording by Ordered iNsertion), overcome many key limitations (Loveless et al. 2021a; Choi et al. 2022) (Figure 2A). In each of these methods, a pegRNA targets a synthetic locus inserted into the genome that has been optimized for controlled rather than stochastic DNA editing. The synthetic target sequence is designed to require that the first portion of DNA must be edited before any subsequent edits can occur, thus ensuring a unidirectional editing process that can be read out at the end of an experiment in which cells have accumulated edits in a sequential fashion. DNA Typewriter and peCHYRON have significant advantages over CRISPR/Cas9-based lineage tracing: 1) no double-stranded DNA breaks are generated, such that edited DNA at the locus is not at risk of subsequent deletion, and 2) the order in which edits were introduced is explicitly captured due to unidirectional editing. Although the efficiency of these more controllable prime editing-based methods have been limited by low editing rates of barcode insertions longer than a few base pairs, researchers are rapidly identifying means to improve prime editing’s capacity and efficiency (Koeppel et al. 2023; Mathis et al. 2023; Yu et al. 2023; Zhao et al. 2023).
Figure 2: Prime editing-based lineage tracing records the order of DNA edits and events linked to edits.

(a) Prime editing-based lineage tracing can record the order in which edits were introduced by using a synthetic DNA target introduced into the cell’s genome. DNA Typewriter uses a tandem array of identical prime editing target sites in which all but the first in the series are truncated to prevent editing. The prime editing guide RNA (pegRNA) edits the first target site with high specificity; the resulting edit disrupts the first target site (preventing subsequent editing) and completes (unlocks) the second target site so that it is now recognized by the pegRNA; similar editing and unlocking of the subsequent target sites occurs such that cells accumulate defined DNA edits in a sequentially ordered fashion within the barcode. (b) Prime editing-based recording methods chronicle the temporal order of biological events by linking production of a pegRNA to a specific signal or event (e.g., Wnt/β-catenin signal activity, phage infection, light exposure, or other signals of interest); pegRNAs are placed under transcriptional control of a signal/event-responsive promoter and encode a specific edit to the barcode when a biological event of interest is experienced by the cell. Using DNA Typewriter, biological events are thus sequentially recorded into the editing target, allowing for reconstruction of event order based on the final edited sequence of the barcode.
Base editing methods have also been used for lineage tracing. The adenine base editor (ABE) was developed by fusing a Cas9 nickase to a tRNA adenosine deaminase to catalyze the conversion of A•T to G•C at the target site. In a similar approach, Cas9 nickase was fused to a single-stranded DNA-specific cytidine deaminase, which can catalyze the conversion of cytosine to uracil for C•G to A•T editing (Gaudelli et al. 2017; Huang et al. 2021). More advanced base editing methods such as DOMINO (DNA-based Ordered Memory and Iteration Network Operator), ZOMBIE (Zombie is Optical Measurement of Barcodes by In situ Expression), and CAMERA (CRISPR-mediated Analog Multi-Event Recording Apparatus) have been applied to control the order of DNA editing events to facilitate lineage tracing (Tang and Liu 2018; Farzadfard et al. 2019; Askary et al. 2020). However, because these methods are reliant upon predefined target sites and make only single-nucleotide changes, they generally do not produce the stochastic edits and barcode diversity required for more extensive and complex lineage tree reconstruction.
Applications of several of the methods described above have aimed not just to record lineage through stochastic cellular barcoding, but to chronicle biological activities such as signal transduction or key protein interactions by linking the biological event to insertion of a predefined DNA barcode or edit (Figure 2B). For example, recent methods such as ENGRAM (ENhancer-driven Genomic Recording of transcriptional Activity in Multiplex) and P3 (Protein-Protein Proximity) editing can couple expression of a pegRNA with the activation of certain signaling cascades or upon specific protein-protein interactions, respectively (Chen et al. 2021b; Choi et al. 2023). Activation of Wnt/β-catenin signaling, which has long been known to play a key role in the development of normal hair follicles and follicular tumors (Chan et al. 1999; Huelsken et al. 2001; Matos et al. 2020; Hile and Harms 2021), was successfully recorded by DNA editing through both the prime editing-based ENGRAM system and the CRISPR/Cas9-based MEMOIR (Memory by Engineered Mutagenesis with Optical In situ Readout) system (Frieda et al. 2017; Chen et al. 2021b). CAMERA base editing has been used to record other biological events such as exposure to visible light and infection by a phage (Tang and Liu 2018). These genomic editing tools offer the potential to link any transcriptionally-active event of interest to the production of a specific gRNA, which acts as a cellular historian to edit the DNA barcode to record the event. If coupled with unidirectional editing methods such as DNA Typewriter or peCHYRON, signal-responsive DNA edits can record not just whether a biological event occurred, but can chronicle the order in which events linked to distinct pegRNAs occurred in a cell (Figure 2B). These capabilities represent a powerful new dimension for lineage tracing research, which could be applied to understand the complex sequence of biological signals that direct the morphogenesis of the skin and its appendages (Fuchs 2016; Sun et al. 2020; Zhang and Chen 2023) and how these go awry in models of disease or carcinogenesis.
In this section, we introduced the diverse genome editing-based lineage tracing technologies that generate barcode diversity in vivo, allowing for high resolution lineage tracing from a monoclonal population of cells. For more details about these methods, we direct the reader to several excellent reviews (VanHorn and Morris 2021; Sankaran et al. 2022; Yao et al. 2022). In the following sections, we focus on existing and potential applications of CRISPR/Cas9-based lineage and molecular recording technologies.
Lineage Tracing in Organoids, Organs, and Model Organisms
Starting with John Sulston’s monumental achievement in 1983 of visually tracking and recording the lineage of each individual cell in the model organism Caenorhabditis elegans (Sulston et al. 1983), developmental biologists have aimed to generate comparable complete maps of the development of more complicated biological systems, including human organoids and whole organs like the brain (McKenna et al. 2016; Raj et al. 2018; Chan et al. 2019; He et al. 2022). Over the last decade, great strides in single-cell indexing methods and high-throughput DNA sequencing technologies have allowed researchers to develop atlases that catalog developmental gene expression programs during organismal development at single-cell resolution (Svensson et al. 2018; Aldridge and Teichmann 2020). Lineage tracing technologies have the potential to complement these efforts by allowing scientists to generate lineage trees for tracking the diverse states that pluripotent cells traverse to reach their ultimate fate in a mature tissue in models that are not amenable to continuous visual tracking as done by Sulston. Although still in their nascent phase, these technologies have so far shed light on how healthy development progresses and promise to illuminate how congenital disorders and malformations may result from disrupted developmental lineages.
The first demonstration that Cas9 editing could be used to generate sufficient barcode diversity to allow in vivo lineage tracing was termed GESTALT (Genome Editing of Synthetic Target Arrays for Lineage Tracing) and was applied to zebrafish (Figure 3). The authors generated transgenic zebrafish harboring arrays of CRISPR/Cas9 target sites and injected Cas9 pre-loaded with guide RNAs into zygotes (McKenna et al. 2016). Due to the injection of the editing machinery at the zygote stage and saturation of the genomic target sites, most editing took place during early stages of development. Interestingly, the authors found that blood lineages in particular were bottlenecked during development, with 98% of all blood in the adult zebrafish attributable to just 5 early ancestors. In other tissues, over half of the cells in each organ were represented by less than 7 tissue-specific lineages, indicating that the majority of each organ arose from just a handful of founder cells. These dominant lineages were generally organ-specific, although not exclusively so; for example, the same edit was found in >10% of both cardiomyocytes and intestinal bulb cells, indicating these divergent cell types shared a common progenitor. To determine the ancestral origins of distinct brain regions during neurogenesis, an improved version of GESTALT coupled to single-cell transcriptional profiling was applied to the zebrafish brain with a heat shock-inducible Cas9 to permit temporal control of DNA editing (Raj et al. 2018). This study found that though most cells in the forebrain, midbrain, and hindbrain arose from distinct lineages, some lineages spanned multiple brain regions, suggesting that edits either accumulated early in multi-potent progenitors or that some later lineages retain the ability to contribute to more than one brain region. Cutaneous biology researchers have posed similar questions aiming to understand how common progenitor cells contribute to the development of the epidermis and its appendages (hair follicles, sebaceous glands, sweat glands, and sensory structures like touch domes) as well as how malignant clones arise from various skin compartments to form the wide variety of cutaneous tumor types (Clayton et al. 2007; Arwert et al. 2012; Alcolea and Jones 2014; Ge et al. 2017; Andersen et al. 2019). DNA editing-based lineage tracing tools promise to make asking such questions more tractable.
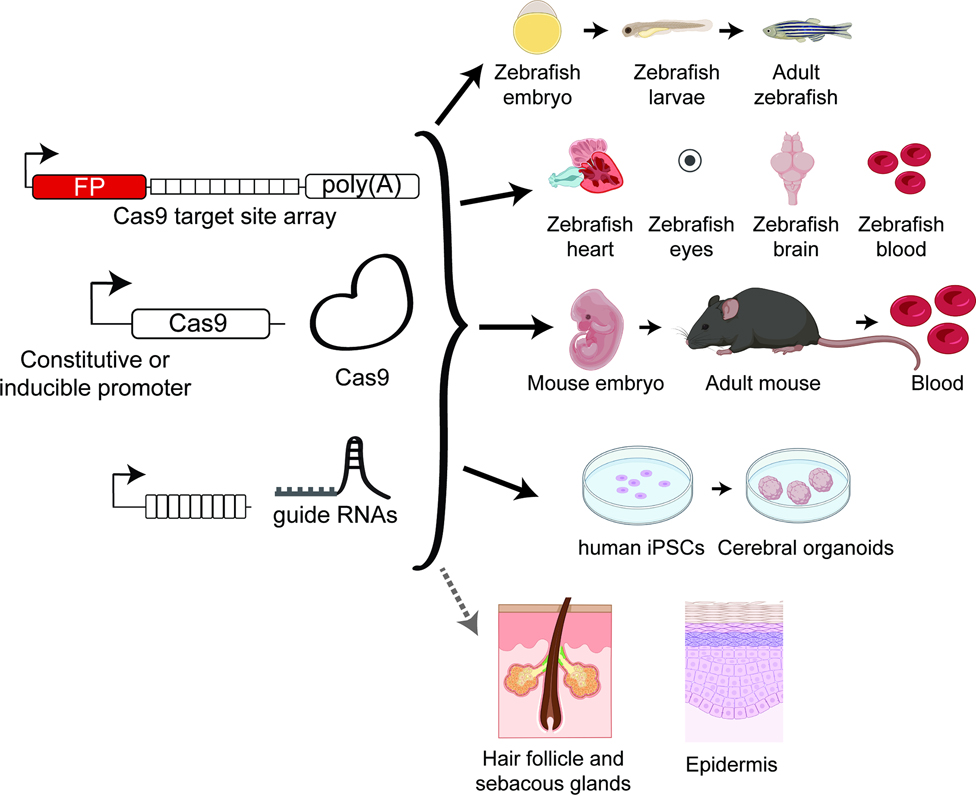
Figure 3: Cas9-based lineage tracing can map the fate of progenitor cells during the development of organoids, organs, and whole organisms.
Cas9-based methods dynamically generate diverse barcodes over many cellular generations and have been applied to map the development of whole zebrafish and its organs, mouse embryogenesis and hematopoiesis, and human induced pluripotent stem cell (iPSC)-derived cerebroids. An array of Cas9 targets (often linked to a transcribed fluorescent protein [FP]) and gRNAs are delivered into stem or progenitor cells; subsequent editing generates the barcode diversity needed for complex lineage tracing; importantly, the Cas9 editor, itself, can either be constitutively or inducibly expressed (e.g., under tetracycline- or heat shock-inducible promoters) to allow temporal control over when editing occurs during development. Such technology could be applied to animal or organoid models of skin to comprehensively map the developmental plan of the epidermis and its appendages.
Mammalian development has more recently been probed using Cas9-based barcoding methods. Mouse gastrulation was studied using a method similar to GESTALT by injecting a target array, Cas9, and multiple gRNAs into murine oocytes before they developed into embryos (Chan et al. 2019). Importantly, the researchers found that cells having a very similar state at the time of harvesting, as measured by scRNAseq, did not always share the same developmental origin; for example, while scRNAseq classified a cell population as embryonic endoderm, the lineage of a subset of these cells was clearly traced to an extra-embryonic progenitor population that likely migrated during development and later adopted an endodermal phenotype. This study underscores the enhanced precision of developmental mapping afforded by combining lineage tracing with transcriptomic profiling. Lineage recording has also been used successfully to trace blood cell expansion and maturation in adult mice by introducing barcodes via retroviral delivery into hematopoietic stem cells then tracking their progeny over time by reading out a static barcode (Gerrits et al. 2010; Lu et al. 2011; Sun et al. 2014b; Chan et al. 2019). Another study added an array of loxP sites for Cre recombinase to randomly recombine to create a unique cellular barcode in the 3’ untranslated region (UTR) of tdTomato (Pei et al. 2020). Reading the barcodes alongside the transcriptome allowed researchers to stratify adult blood cell populations by their progenitors. While sufficient for cataloging cell fates within the hematopoietic system, the limited diversity of static barcodes restricted the resolution at which lineage relationships could be determined and hematopoietic stem cells had to be re-engineered for each experiment. To overcome these limitations, researchers generated a more versatile mouse model for joint lineage tracing and transcriptomic profiling using a framework similar to GESTALT. This model showed that following bone marrow ablation, only a few hematopoietic stem cell clones generated the majority of progeny during hematopoiesis in adult mice (Bowling et al. 2020). While these investigators benchmarked Cas9-based lineage tracing methods in mouse hematopoietic lines, similar methods could be used to trace the fate of stem cells in the skin and determine whether a similar clonal bottleneck occurs during skin development, tissue regeneration after wounding, or tumorigenesis.
Lineage tracing tools can also be used to analyze organoid systems that model human tissue and are more easily manipulated ex vivo. In a study of human induced pluripotent stem cell (iPSC)-derived brain organoids that applied spatial RNA-seq in tandem with lineage tracing, researchers developed a method termed iTracer (He et al. 2022). This system provides two channels of barcoding to expand lineage tracing capability: the construct used for DNA editing contains a U6 promoter-driven gRNA as well as a fluorophore gene (to allow positive cell sorting) with a random barcode in its 3’ UTR. The fluorophore gene itself contains the gRNA target within its 3’ coding region, but it is not edited until expression of Cas9, which is controlled by a doxycycline-inducible promoter. Thus, gene editing could be induced after initiating cerebroid formation by adding doxycycline to the medium at various days during the protocol to generate edits. The researchers then used a spatial transcriptomics platform, Visium, to recover cell lineage and spatial relationships in tandem (Ståhl et al. 2016). Analysis of the permanent CRISPR-induced edits and unique barcodes revealed that each barcode was clustered based on location within the organoid, demonstrating that lineages were spatially constrained early in tissue differentiation. Investigators seeking to understand how stem cells residing in the bulge area of the hair follicle contribute to wound repair, for example, could use such a system to map the precise lineage, transcriptome, and the ultimate physical destination of cells in the healed tissue (Sada et al. 2016; Aragona et al. 2017; Joost et al. 2018; Phan et al. 2021).
Developmental biologists have more recently appreciated the importance of epigenetic modifications as a driver of differentiation (Meissner 2010; Cantone and Fisher 2013). Thus, profiling a cell’s lineage and transcriptome as well as epigenetic modifications would provide a more comprehensive roadmap of developmental programs (Lu et al. 2020). Toward this goal, a group recently developed a mouse model called DARLIN (Cas9-TdT CRISPR Array Repair LINeage tracing) (Li et al. 2023). Similar to GESTALT, this system generated barcode diversity for lineage tracing by inserting an array of Cas9 gRNAs and targets into safe harbor loci in a mouse line. Cell lineage, gene expression, and epigenetic modifications were simultaneously profiled by splitting single cells into cytoplasmic and nuclear fractions; the cytoplasmic fraction was used to capture all mRNAs including lineage-specific edited transcripts while the nuclear fraction was labeled with GpC methyltransferase to mark open chromatin domains. Using bisulfite DNA sequencing, the researchers were then able to determine the open versus closed chromatin status at genomic loci and compare these among clones.
Tracing the Origins and Drivers of Cancer
Identifying the cells of origin in tumors is essential to delineate the early mechanisms driving carcinogenesis. Moreover, understanding the states that a cell must traverse during malignant transformation and its primary driver signals could allow pharmacologic targeting of malignant stem cells to prevent cancer initiation or recurrence, including in the skin (Sánchez-Danés et al. 2016). Applying the dynamic barcoding methods outlined above, researchers have engineered tumor cells ex vivo, introduced them into model systems, and subsequently harvested the primary tumor or sites of metastases for lineage tracing (Aalam et al. 2023). Metastases are of particular interest to clinicians since treatment effectiveness and survival are minimal for metastatic disease. Moreover, 2–5% of cancers are diagnosed as metastases of unknown primary tumor (Varadhachary 2007), which reduces clinicians’ ability to choose the most effective treatment for the underlying cancer. Using lineage tracing methods to directly link metastasized cells to ancestral cells in cancer models, we can enhance our understanding of tumor origin, progression, and spread (Figure 4A).
Figure 4: Lineage tracing in cancer models can reveal key oncogenic steps and drivers of metastatic behavior.
(a) Lineage tracing allows investigators to track a cancer progenitor cell through its clonal expansion, epithelial-to-mesenchymal transition (EMT), and leaving the site of origin through the vasculature to metastasize, seed, and grow in distant sites. The resultant barcode sequence at secondary metastatic sites can identify the clone of origin and can be coupled with transcriptomic analysis to understand the critical drivers of its metastatic behavior, which could dictate the therapeutic strategy. (b) Cancer cells can be engineered to express the machinery for DNA editing-based lineage tracing studies; this approach has revealed the origins and drivers of metastases in mouse xenograft models of lung and pancreatic carcinomas and could be adapted to study metastatic behavior in melanoma or other invasive skin cancers.
Researchers focused on a wide variety of cancers have applied Cas9-based lineage tracing methods to tumor xenograft models in order to probe metastatic origins (Quinn et al. 2021; Simeonov et al. 2021; Yang et al. 2022). Aggressive human KRAS-mutant lung adenocarcinoma lines were engineered with synthetic arrays of Cas9 sites and gRNAs along with luciferase as an optical signal to locate where the cancer had metastasized (Quinn et al. 2021) (Figure 4B). The results showed that 30,000 profiled metastatic lung cancer cells had nearly as many unique lineage states and that individual cell clades had varying degrees of metastatic behavior. Some clades of related cells remained entirely in their original location, while others seeded several metastatic progeny into one other tissue, and some seeded many metastatic cells across multiple tissues. Lineage tracing allowed identification of clades that were then transcriptionally compared to identify genes directly or inversely correlated with metastatic potential. To validate these findings, candidate driver or suppressor genes were either up- or down-regulated to directly test their effect on invasiveness of multiple different cell types. This study demonstrates the power of coupling lineage tracing with transcriptional profiling to simultaneously trace a tumors’ origin and delineate key controllers of its metastatic potential.
Similar lineage tracing was performed in a mouse pancreatic cancer transplant model using a variant of GESTALT with doxycycline-inducible Cas9 to initiate lineage recording in a desired temporal window (Simeonov et al. 2021). This allowed the investigators to define the origin of rare metastatic cells and identify bottlenecks during tumor implantation, as only a small minority of cells successfully engrafted. Another mouse model of lung adenocarcinoma used mouse embryonic stem cells engineered with a similar Cas9-based lineage tracing system that also allowed Cre-inducible activation of oncogenic KRAS and p53 mutation (Yang et al. 2022). Their study produced a high-resolution lineage map of lung cancer progression and found that the most aggressive metastatic clones were enriched in mesenchymal genes, showing a direct correlation between metastatic potential and progression along the epithelial-to-mesenchymal (EMT) transition (Figure 4B) (Kalluri and Weinberg 2009).
Leveraging DNA sequence-based lineage tracing in the skin
In sum, CRISPR/Cas9-based lineage tracing has been leveraged to study development at both early and late timepoints and cancer metastases in diverse model systems and human organoids. While many of these innovative technologies were pioneered in extra-cutaneous systems, we propose they can be readily applied to many unanswered questions in cutaneous biology and dermatology to better understand and treat genodermatoses, congenital nevi, and vascular malformations as examples. Applying DNA sequencing-based lineage tracing methods to models used to study cutaneous biology including mice, human organotypic cultures (Simpson et al. 2010; Smits et al. 2017), and more recently described iPSC-derived appendage-bearing organoids (Lee et al. 2020) would allow us to understand how the epidermis and its appendages continuously renew the skin, hair, and nails and how its stem cells respond to external insults and injuries (Figure 1C and Figure 3). While advanced lineage tracing tools have been used up to now to understand the development of internal organs like the brain, the external location of the skin and the potential for xenografting human stem cell-derived tissues or organoids onto mouse skin further amplifies the experimental possibilities. By combining lineage tracing tools with intravital imaging (Obeidy et al. 2018) investigators can elucidate both the genetic lineage and spatiotemporal fate of progenitor cells during human skin morphogenesis and regeneration or scarring after wounding or other damage like radiation. Recent studies have applied scRNA-seq to skin organoid systems both ex vivo and as in vivo xenografts (Lee et al. 2020) in which one can both define cell types and infer lineage by computationally derived RNA velocities (Stabell et al. 2023). However, the accuracy of such methods can be greatly enhanced when supplemented with and verified by direct lineage tracing using the dynamic barcoding methods described above (He et al. 2022).
In dermatology, the study of melanoma could benefit greatly from the application of advanced lineage tracing methods multiplexed with transcriptomics to define the subsets of this cancer with invasive behavior that pose true risk to patients to help determine the need for staging and the aggressiveness of therapy. Lineage tracing methods could also help map the origins of melanoma, which may arise de novo or from existing nevi or lentigines, and to define the early fate-determining signals that dictate outcomes ranging from atypical melanocytic lesions that may remain in situ indefinitely to rapidly progressive nodular tumors that readily metastasize (Figure 4A–B). Melanocytic stem cells have been tracked using a fluorescent label and a tyrosinase marker, which allowed investigators to localize invasive tumor cells over time in a mouse model of melanoma driven by BRAF V600E mutation and ultraviolet radiation (Moon et al. 2017). Applying more advanced CRISPR/Cas9-based lineage tracing methods would allow investigators to elucidate more details about the ancestral origin of melanoma in this and other in vivo models in addition to mapping the epigenetic and transcriptional landscape of melanocyte stem cells that primes certain cancer progenitors for malignant transformation.
While many keratinocyte carcinomas can be treated with local surgical methods or topical chemotherapy, these are the most numerous human cancers by far and a subset of squamous cell carcinomas (and rarely basal cell carcinomas, BCC) do invade and metastasize. However, it is unknown if the cell of origin or lineage differ between more or less aggressive keratinocyte tumor subtypes, for example, superficial spreading BCC, which is easily treated, versus basosquamous carcinoma, which exhibits much more invasive behavior (Chiang et al. 2019). Even less is known about the origins and signaling drivers of more aggressive tumors arising from the skin such as sebaceous carcinoma and Merkel cell carcinoma, which have very high mortality rates once they have spread beyond the skin. Applying lineage tracing methods to murine (Lowry et al. 2016; Verhaegen et al. 2022) and/or xenograft models (Patel et al. 2012; Fang et al. 2020) of these aggressive skin cancer types could provide important advances to understand their cellular origins, define the signaling pathways that drive their aggressive behavior, and identify preventive strategies and effective treatments. Building on recent evidence showing specific chromatin modifiers exert a major influence on keratinocyte carcinomas in the skin (Ko et al. 2022), tracing cell lineage alongside epigenetic modifications could be applied to mouse models of skin carcinogenesis to understand how epigenetic factors influence cancer stem cells (Li et al. 2023).
Dermal fibroblasts are known to play key roles in epidermal differentiation, appendage development, and wound repair, but also drive fibrotic pathologies such as keloids and scleroderma and can even fuel inflammatory disorders (Leask et al. 2004; Driskell et al. 2013). The marked heterogeneity of skin fibroblasts populations has been characterized both by Cre-based lineage tracing as well as single-cell RNA sequencing-based pseudotime trajectory analysis (Driskell et al. 2013; Guerrero-Juarez et al. 2019). While these novel studies identified crucial divergences in fibroblast lineage, such as distinct fibroblast populations responsible for a first and second wave of wound healing, an understanding of the precise signals that drive fibroblasts into different lineages remain elusive. Using the methods described herein, investigators have the tools needed to map lineage hierarchies of fibroblasts throughout cutaneous development, during regeneration versus scar formation after wounding, and within models of fibrotic (or non-fibrotic) disorders. Understanding fibroblast fate, function, and pathogenic behavior could uncover much-needed treatments for sclerosing disorders and advance long-standing goals like hair follicle regeneration and scarless wound healing (Mascharak et al. 2021).
Summary
DNA editing-based methods have rapidly expanded researchers’ ability to interrogate cell lineage relationships required to reconstruct complex morphogenetic programs. Newer dynamic barcoding strategies allow investigators to generate detailed maps of cell lineage, which can now be overlaid with single-cell gene expression and epigenetic information. Even greater diversity of DNA barcodes afforded by Cas9-based editing or arrays of truncated prime editing targets that must be edited in sequence has provided additional bandwidth to construct more expansive lineage trees to map cellular origins and relationships in higher-order biological systems like organoids, organs, or even whole organisms. Moreover, methods such as peCHYRON and DNA Typewriter provide a temporal directionality to barcoding, which allows more accurate mapping of lineages by reading sequential edits inserted over time. Finally, linking sequential DNA editing to specific biological events, investigators can translate the edited DNA template into a historical record of signals that a cell has received over the course of development, tissue repair, or carcinogenesis. While use of these more advanced DNA editing tools has been limited in cutaneous biology, their application to the fields of hematology, immunology, and neurobiology has yielded deep insights into the developmental trajectories of stem and progenitor cells and the origins and metastatic potential of cancerous cells derived from their respective organ systems. By introducing cutaneous biologists to the remarkable toolkit of DNA editors capable of advanced lineage tracing and cataloging of biological events, we hope to have expanded the notion of what is possible by applying these cutting-edge techniques to the study of skin biology. Together, these remarkable technologies in DNA editing offer cutaneous biologists the opportunity to re-imagine our capacity to create a blueprint of the morphogenetic programs that support skin development and regeneration and to understand how these go awry in cutaneous malformations, diseases, and malignancies and how they might be reversed. Perhaps most excitingly, prime editing-based molecular recording tools can now, using the nucleotide alphabet, chronicle cellular responses to external signaling cues or injury. These advances in DNA sequence-based recording should allow investigators to optimize stem cell mobilization and differentiation protocols and aspire to build fully functional skin grafts and appendages, bringing real-world regenerative medicine within reach in the field of dermatology.
Practical considerations
Each specific method of lineage tracing offers both benefits and limitations (Table 1). For example, optical lineage tracing via fluorescence offers optimal spatial resolution, but the time-consuming nature of live microscopy can be limiting, while CRISPR/Cas9 edits can accumulate over many cellular generations and be interpreted retrospectively. While prime editor-based tracing affords sequential barcoding to delineate complex lineages, it requires delivery of multiple large transgenes that are not needed for RNA sequencing-based inferred lineages. Below we describe additional practical considerations to help users appropriately plan and optimize their use of DNA editing-based lineage tracing strategies.
Table 1. Lineage Tracing Methods.
| Method | Advantages | Disadvantages |
|---|---|---|
| Fluorescence-based optical lineage tracing | High temporal and spatial resolution Can be coupled with fluorescent biosensors to capture physiologic signals along with lineage Readily applied to the skin surface using two-photon microscopy |
Low throughput (number of cells traced is limited by the visual window and available fluorophores) Data collection is limited to the labeled cell population |
| RNA-sequencing- based lineage inference | Provides a broad description of cell state via transcriptomics Analysis can be performed using many widely available software tools No delivery of fluorescent markers or complicated editing machinery |
Data is limited to the time-point/s chosen for RNA collection Inferred lineages have considerable uncertainty |
| CRISPR/Cas9-based lineage tracing | Can be coupled with RNA-seq to provides a broad description of cell state via transcriptomics Continuous DNA editing allows dynamic barcoding to construct more complicated lineages |
DNA barcodes can be inadvertently deleted Double-stranded DNA breaks may have deleterious effects Order of DNA edits is not explicitly captured |
| Prime editor-based lineage tracing | Can be coupled with RNA-seq to provides a broad description of cell state via transcriptomics Continuous DNA editing allows dynamic barcoding to construct more complicated lineages Can engineer the barcode to be unidirectional to prevent loss of edits Can be used to record the temporal order of edits or biological signals No double-stranded DNA breaks |
Editing rate is generally low Analysis software is nascent and not widely available Delivery of large editing machinery into cells can be challenging |
High copy number delivery methods
To generate high-resolution lineage maps, it is best to integrate high copy numbers (termed multiplicity of infection [MOI]) of lineage tracing components into cells prior to expansion and differentiation. High copy numbers of components are desirable to provide more DNA editing targets per cell and higher expression of the editor protein to enhance the rate of successful editing. Together, these factors provide more potential barcode diversity, which enhances the ability to distinguish subclades for high-resolution lineage tracing.
Transduction is the most widely used method to establish cell lines with high-copy numbers of editing components. Optimization of MOI is feasible for viral packaging vectors such as lentivirus and adenovirus, however, their packaging capacity of ~9 kilobases and ~4.7 kilobases, respectively, limits their ability to carry large transgenes required for more complex editing platforms (Dong et al. 1996; Kumar et al. 2001). Recent advances in optimizing smaller or split prime editors have enhanced viral delivery of lineage-tracing cargo (Davis et al. 2023; Doman et al. 2023).
Transposases are also utilized for high-copy number transgene delivery, but can integrate into non-desirable sites, potentially disrupting native genes that encode critical proteins or tumor suppressors. Piggybac and sleeping beauty transposases preferentially integrate at specific genomic sequences and the cells in which a deleterious integration event occurs are generally diluted out through reduced fitness over several generations. The upper bound for transposon cargo is >100 kilobases, making them suitable for large transgene integration (Rostovskaya et al. 2012; Li et al. 2013). Larger transgene capacity allows editing components to be integrated alongside selection markers, which can be used to increase the copy number of integrants by selecting cells with the highest puromycin resistance or sorting those with highest fluorescence intensity (Chardon et al. 2023). However, the reversible nature of DNA transposition makes it possible for later excision of the editing machinery, which can preclude further editing.
Timing and saturation of editing activity
The lineage tracing methods outlined in this review can be restrained by using inducible Cas9 expression to control the precise timing of DNA editing. This is important because sustained editing by Cas9 or its derived editing systems can lead to “saturation” of DNA target sites. For example, in the original description of GESTALT, the authors found the majority of editing happened very early in zebrafish development due to exhaustion of DNA target sites (McKenna et al. 2016). To address this, investigators can employ systems that initiate Cas9 editing only during discrete temporal windows to reduce the risk of target site saturation and to strategically time editing to occur during periods of particular interest in the model system. Under the Tet-ON system (Das et al. 2016), application of a tetracycline class drug like doxycycline leads to specific activation of a transcription factor that drives expression of Cas9. This system has been utilized in lineage tracing systems ranging from cerebral organoids to full mouse development from embryonic stem cells (He et al. 2022; Li et al. 2023). While the Tet-ON platform has been frequently used to control Cas9-based lineage tracing experiments, other tunable protein induction and degradation methods have been developed for mammalian systems, which could permit more precise control over editing enzymes to match the biology under study (Mullick et al. 2006; Kallunki et al. 2019; Yesbolatova et al. 2020).
Library generation
The majority of lineage tracing methods described in this review are reliant upon high-fidelity DNA sequencing in order to accurately “read” the target or TAPE in which editing or barcoding has occurred. Moreover, optimal primer design is necessary to sufficiently amplify the edited loci from genomic DNA with few off-target products. To optimize the protocol for sequencing the DNA “tape” (where edits have been recorded), the region can be amplified via quantitative PCR to capture amplified sites before PCR saturation, and then non-quantitative thermocyclers can be used once an optimal PCR protocol has been determined. For primer design, Primer-BLAST is an excellent resource to generate primers specific to the target with minimal off-target amplifications, as well as primer designing tools available on softwares such as Snapgene and Benchling (Sofi et al. 2022). The primers should include sequences specific to the region of interest as well as adaptors for a second round of PCR to index the sample. Gel electrophoresis via agarose gel or Tapestation devices can be used to validate the correct size of the amplicon. The desired library can be isolated via gel extraction or size selected via purification with magnetic beads. We suggest cleaning and checking library size after each PCR step during initial protocol optimization to ensure reliable results.
Data analysis
Lineage tracing methods that utilize Cas9 to generate random, non-ordered edits are analyzed via phylogenetic analysis methods, which can be mathematically complex. Data from these assays are compared to the phylogenetic analysis of accumulated mutations in the genome to determine phylogenetic maps of cell ancestry and lineage (similar analysis is used to generate evolutionary trees) (Pavlopoulos et al. 2010). This data can be analyzed with assistance from software available on Github developed in several of the methods described in this review (McKenna et al. 2016; Raj et al. 2018; Bowling et al. 2020; Yang et al. 2022). The Cassiopeia pipeline for single-cell lineage tracing data may be useful as well (Jones et al. 2020).
Because the TAPE in DNA typewriter and peCHYRON is sequentially edited and the sequence of the TAPE is highly repetitive, the editing data produced from these experiments must be analyzed differently than those produced with GESTALT or related methods. The scripts used in the original DNA Typewriter and peCHYRON papers are similarly available on Github and Figshare (Loveless et al. 2021a; Choi et al. 2022). Familiarity with Regular Expressions (RegEx) is also advised for analysis of this type of sequencing data.
ACKNOWLEDGEMENTS
JFN is supported by the Institute for Stem Cell and Regenerative Medicine Fellowship fund. CLS is supported by National Institutes of Health grant K08 AR075846.
Footnotes
CONFLICT OF INTEREST
The University of Washington has filed a patent application for several technologies related to molecular recording, including DNA Typewriter, ENGRAM and P3 editing, on which J.S. is listed as inventor. J.S. is on the scientific advisory board, a consultant, and/or a co-founder of Adaptive Biotechnologies, Cajal Neuroscience, Camp4 Therapeutics, Guardant Health, Maze Therapeutics, Pacific Biosciences, Phase Genomics, Prime Medicine, Scale Biosciences, and Sixth Street Capital.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- Aalam SMM, Nguyen LV, Ritting ML, Kannan N. Clonal tracking in cancer and metastasis. Cancer Metastasis Rev [Internet]. 2023. Nov 1; Available from: 10.1007/s10555-023-10149-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alcolea MP, Jones PH. Tracking cells in their native habitat: lineage tracing in epithelial neoplasia. Nat Rev Cancer. 2013. Mar;13(3):161–71. [DOI] [PubMed] [Google Scholar]
- Alcolea MP, Jones PH. Lineage analysis of epidermal stem cells. Cold Spring Harb Perspect Med. 2014. Jan 1;4(1):a015206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aldridge S, Teichmann SA. Single cell transcriptomics comes of age. Nat Commun. 2020. Aug 27;11(1):4307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersen MS, Hannezo E, Ulyanchenko S, Estrach S, Antoku Y, Pisano S, et al. Tracing the cellular dynamics of sebaceous gland development in normal and perturbed states. Nat Cell Biol. 2019. Aug;21(8):924–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anzalone AV, Randolph PB, Davis JR, Sousa AA, Koblan LW, Levy JM, et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature. 2019. Dec;576(7785):149–57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aragona M, Dekoninck S, Rulands S, Lenglez S, Mascré G, Simons BD, et al. Defining stem cell dynamics and migration during wound healing in mouse skin epidermis. Nat Commun. 2017. Mar 1;8:14684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arwert EN, Hoste E, Watt FM. Epithelial stem cells, wound healing and cancer. Nat Rev Cancer. 2012. Feb 24;12(3):170–80. [DOI] [PubMed] [Google Scholar]
- Askary A, Sanchez-Guardado L, Linton JM, Chadly DM, Budde MW, Cai L, et al. In situ readout of DNA barcodes and single base edits facilitated by in vitro transcription. Nat Biotechnol. 2020. Jan;38(1):66–75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhang HEC, Ruddy DA, Krishnamurthy Radhakrishna V, Caushi JX, Zhao R, Hims MM, et al. Studying clonal dynamics in response to cancer therapy using high-complexity barcoding. Nat Med. 2015. May;21(5):440–8. [DOI] [PubMed] [Google Scholar]
- Bowling S, Sritharan D, Osorio FG, Nguyen M, Cheung P, Rodriguez-Fraticelli A, et al. An Engineered CRISPR-Cas9 Mouse Line for Simultaneous Readout of Lineage Histories and Gene Expression Profiles in Single Cells. Cell. 2020. Jun 11;181(6):1410–22.e27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calderon D, Blecher-Gonen R, Huang X, Secchia S, Kentro J, Daza RM, et al. The continuum of Drosophila embryonic development at single-cell resolution. Science. 2022. Aug 5;377(6606):eabn5800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cantone I, Fisher AG. Epigenetic programming and reprogramming during development. Nat Struct Mol Biol. 2013. Mar;20(3):282–9. [DOI] [PubMed] [Google Scholar]
- Carroll D p53 Throws CRISPR a Curve. Trends Pharmacol Sci. 2018. Sep;39(9):783–4. [DOI] [PubMed] [Google Scholar]
- Chan EF, Gat U, McNiff JM, Fuchs E. A common human skin tumour is caused by activating mutations in beta-catenin. Nat Genet. 1999. Apr;21(4):410–3. [DOI] [PubMed] [Google Scholar]
- Chan MM, Smith ZD, Grosswendt S, Kretzmer H, Norman TM, Adamson B, et al. Molecular recording of mammalian embryogenesis. Nature. 2019. Jun;570(7759):77–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chardon FM, McDiarmid TA, Page NF, Martin B, Domcke S, Regalado SG, et al. Multiplex, single-cell CRISPRa screening for cell type specific regulatory elements. bioRxiv [Internet]. 2023. Mar 28; Available from: 10.1101/2023.03.28.534017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen PJ, Hussmann JA, Yan J, Knipping F, Ravisankar P, Chen PF, et al. Enhanced prime editing systems by manipulating cellular determinants of editing outcomes. Cell. 2021a. Oct 28;184(22):5635–52.e29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen R, Zhu Z, Ji S, Geng Z, Hou Q, Sun X, et al. Sweat gland regeneration: Current strategies and future opportunities. Biomaterials. 2020. Oct;255:120201. [DOI] [PubMed] [Google Scholar]
- Chen W, Choi J, Nathans JF, Agarwal V, Martin B, Nichols E, et al. Multiplex genomic recording of enhancer and signal transduction activity in mammalian cells [Internet]. bioRxiv. 2021b. [cited 2023 Sep 27]. p. 2021.11.05.467434. Available from: https://www.biorxiv.org/content/10.1101/2021.11.05.467434v1
- Chiang A, Tan CZ, Kuonen F, Hodgkinson LM, Chiang F, Cho RJ, et al. Genetic Mutations Underlying Phenotypic Plasticity in Basosquamous Carcinoma. J Invest Dermatol. 2019. Nov;139(11):2263–71.e5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi J, Chen W, Liao H, Li X, Shendure J. A molecular proximity sensor based on an engineered, dual-component guide RNA. bioRxiv [Internet]. 2023. Aug 14; Available from: 10.1101/2023.08.14.553235 [DOI] [Google Scholar]
- Choi J, Chen W, Minkina A, Chardon FM, Suiter CC, Regalado SG, et al. A time-resolved, multi-symbol molecular recorder via sequential genome editing. Nature. 2022. Aug;608(7921):98–107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chow KHK, Budde MW, Granados AA, Cabrera M, Yoon S, Cho S, et al. Imaging cell lineage with a synthetic digital recording system. Science [Internet]. 2021. Apr 9;372(6538). Available from: 10.1126/science.abb3099 [DOI] [PubMed] [Google Scholar]
- Clayton E, Doupé DP, Klein AM, Winton DJ, Simons BD, Jones PH. A single type of progenitor cell maintains normal epidermis. Nature. 2007. Mar 8;446(7132):185–9. [DOI] [PubMed] [Google Scholar]
- Cockburn K, Annusver K, Gonzalez DG, Ganesan S, May DP, Mesa KR, et al. Gradual differentiation uncoupled from cell cycle exit generates heterogeneity in the epidermal stem cell layer. Nat Cell Biol. 2022. Dec;24(12):1692–700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cotsarelis G, Sun TT, Lavker RM. Label-retaining cells reside in the bulge area of pilosebaceous unit: implications for follicular stem cells, hair cycle, and skin carcinogenesis. Cell. 1990. Jun 29;61(7):1329–37. [DOI] [PubMed] [Google Scholar]
- Das AT, Tenenbaum L, Berkhout B. Tet-On Systems For Doxycycline-inducible Gene Expression. Curr Gene Ther. 2016;16(3):156–67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis JR, Banskota S, Levy JM, Newby GA, Wang X, Anzalone AV, et al. Efficient prime editing in mouse brain, liver and heart with dual AAVs. Nat Biotechnol [Internet]. 2023. May 4; Available from: 10.1038/s41587-023-01758-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doman JL, Pandey S, Neugebauer ME, An M, Davis JR, Randolph PB, et al. Phage-assisted evolution and protein engineering yield compact, efficient prime editors. Cell. 2023. Aug 31;186(18):3983–4002.e26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong JY, Fan PD, Frizzell RA. Quantitative analysis of the packaging capacity of recombinant adeno-associated virus. Hum Gene Ther. 1996. Nov 10;7(17):2101–12. [DOI] [PubMed] [Google Scholar]
- Doudna JA, Charpentier E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science. 2014. Nov 28;346(6213):1258096. [DOI] [PubMed] [Google Scholar]
- Driskell RR, Lichtenberger BM, Hoste E, Kretzschmar K, Simons BD, Charalambous M, et al. Distinct fibroblast lineages determine dermal architecture in skin development and repair. Nature. 2013. Dec 12;504(7479):277–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Efremova M, Vento-Tormo M, Teichmann SA, Vento-Tormo R. CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat Protoc. 2020. Apr;15(4):1484–506. [DOI] [PubMed] [Google Scholar]
- Fang B, Kannan A, Zhao S, Nguyen QH, Ejadi S, Yamamoto M, et al. Inhibition of PI3K by copanlisib exerts potent antitumor effects on Merkel cell carcinoma cell lines and mouse xenografts. Sci Rep. 2020. Jun 1;10(1):8867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farzadfard F, Gharaei N, Higashikuni Y, Jung G, Cao J, Lu TK. Single-Nucleotide-Resolution Computing and Memory in Living Cells. Mol Cell. 2019. Aug 22;75(4):769–80.e4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fleck JS, Jansen SMJ, Wollny D, Zenk F, Seimiya M, Jain A, et al. Inferring and perturbing cell fate regulomes in human brain organoids. Nature. 2023. Sep;621(7978):365–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frangoul H, Altshuler D, Cappellini MD, Chen YS, Domm J, Eustace BK, et al. CRISPR-Cas9 Gene Editing for Sickle Cell Disease and β-Thalassemia. N Engl J Med. 2021. Jan 21;384(3):252–60. [DOI] [PubMed] [Google Scholar]
- Frieda KL, Linton JM, Hormoz S, Choi J, Chow KHK, Singer ZS, et al. Synthetic recording and in situ readout of lineage information in single cells. Nature. 2017. Jan 5;541(7635):107–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuchs E Epithelial Skin Biology: Three Decades of Developmental Biology, a Hundred Questions Answered and a Thousand New Ones to Address. Curr Top Dev Biol. 2016. Feb 8;116:357–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaudelli NM, Komor AC, Rees HA, Packer MS, Badran AH, Bryson DI, et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature. 2017. Nov 23;551(7681):464–71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gay D, Kwon O, Zhang Z, Spata M, Plikus MV, Holler PD, et al. Fgf9 from dermal γδ T cells induces hair follicle neogenesis after wounding. Nat Med. 2013. Jul;19(7):916–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerrits A, Dykstra B, Kalmykowa OJ, Klauke K, Verovskaya E, Broekhuis MJC, et al. Cellular barcoding tool for clonal analysis in the hematopoietic system. Blood. 2010. Apr 1;115(13):2610–8. [DOI] [PubMed] [Google Scholar]
- Ge Y, Gomez NC, Adam RC, Nikolova M, Yang H, Verma A, et al. Stem Cell Lineage Infidelity Drives Wound Repair and Cancer. Cell. 2017. May 4;169(4):636–50.e14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guerrero-Juarez CF, Dedhia PH, Jin S, Ruiz-Vega R, Ma D, Liu Y, et al. Single-cell analysis reveals fibroblast heterogeneity and myeloid-derived adipocyte progenitors in murine skin wounds. Nat Commun. 2019. Feb 8;10(1):650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guitart JR Jr, Johnson JL, Chien WW. Research Techniques Made Simple: The Application of CRISPR-Cas9 and Genome Editing in Investigative Dermatology. J Invest Dermatol. 2016. Sep;136(9):e87–93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haapaniemi E, Botla S, Persson J, Schmierer B, Taipale J. CRISPR-Cas9 genome editing induces a p53-mediated DNA damage response. Nat Med. 2018. Jul;24(7):927–30. [DOI] [PubMed] [Google Scholar]
- Haghverdi L, Büttner M, Wolf FA, Buettner F, Theis FJ. Diffusion pseudotime robustly reconstructs lineage branching. Nat Methods. 2016. Oct;13(10):845–8. [DOI] [PubMed] [Google Scholar]
- Hampel S, Chung P, McKellar CE, Hall D, Looger LL, Simpson JH. Drosophila Brainbow: a recombinase-based fluorescence labeling technique to subdivide neural expression patterns. Nat Methods. 2011. Mar;8(3):253–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He L, Li Y, Li Y, Pu W, Huang X, Tian X, et al. Enhancing the precision of genetic lineage tracing using dual recombinases. Nat Med. 2017. Dec;23(12):1488–98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herring CA, Chen B, McKinley ET, Lau KS. Single-Cell Computational Strategies for Lineage Reconstruction in Tissue Systems. Cell Mol Gastroenterol Hepatol. 2018. Feb 13;5(4):539–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heumos L, Schaar AC, Lance C, Litinetskaya A, Drost F, Zappia L, et al. Best practices for single-cell analysis across modalities. Nat Rev Genet. 2023. Aug;24(8):550–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He Z, Maynard A, Jain A, Gerber T, Petri R, Lin HC, et al. Lineage recording in human cerebral organoids. Nat Methods. 2022. Jan;19(1):90–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hile G, Harms PW. Update on Molecular Genetic Alterations of Cutaneous Adnexal Neoplasms. Surg Pathol Clin. 2021. Jun;14(2):251–72. [DOI] [PubMed] [Google Scholar]
- Hsu YC, Fuchs E. Building and Maintaining the Skin. Cold Spring Harb Perspect Biol [Internet]. 2022. Jul 1;14(7). Available from: 10.1101/cshperspect.a040840 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang TP, Newby GA, Liu DR. Precision genome editing using cytosine and adenine base editors in mammalian cells. Nat Protoc. 2021. Feb;16(2):1089–128. [DOI] [PubMed] [Google Scholar]
- Huelsken J, Vogel R, Erdmann B, Cotsarelis G, Birchmeier W. beta-Catenin controls hair follicle morphogenesis and stem cell differentiation in the skin. Cell. 2001. May 18;105(4):533–45. [DOI] [PubMed] [Google Scholar]
- Hu Y, Peng T, Gao L, Tan K. CytoTalk: De novo construction of signal transduction networks using single-cell transcriptomic data. Sci Adv [Internet]. 2021. Apr;7(16). Available from: 10.1126/sciadv.abf1356 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hwang B, Lee W, Yum SY, Jeon Y, Cho N, Jang G, et al. Lineage tracing using a Cas9-deaminase barcoding system targeting endogenous L1 elements. Nat Commun. 2019. Mar 15;10(1):1234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jindal K, Adil MT, Yamaguchi N, Yang X, Wang HC, Kamimoto K, et al. Single-cell lineage capture across genomic modalities with CellTag-multi reveals fate-specific gene regulatory changes. Nat Biotechnol [Internet]. 2023. Sep 25; Available from: 10.1038/s41587-023-01931-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin S, Guerrero-Juarez CF, Zhang L, Chang I, Ramos R, Kuan CH, et al. Inference and analysis of cell-cell communication using CellChat. Nat Commun. 2021. Feb 17;12(1):1088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones MG, Khodaverdian A, Quinn JJ, Chan MM, Hussmann JA, Wang R, et al. Inference of single-cell phylogenies from lineage tracing data using Cassiopeia. Genome Biol. 2020. Apr 14;21(1):92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joost S, Jacob T, Sun X, Annusver K, La Manno G, Sur I, et al. Single-Cell Transcriptomics of Traced Epidermal and Hair Follicle Stem Cells Reveals Rapid Adaptations during Wound Healing. Cell Rep. 2018. Oct 16;25(3):585–97.e7. [DOI] [PubMed] [Google Scholar]
- Kalhor R, Mali P, Church GM. Rapidly evolving homing CRISPR barcodes. Nat Methods. 2017. Feb;14(2):195–200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kallunki T, Barisic M, Jäättelä M, Liu B. How to Choose the Right Inducible Gene Expression System for Mammalian Studies? Cells [Internet]. 2019. Jul 30;8(8). Available from: 10.3390/cells8080796 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalluri R, Weinberg RA. The basics of epithelial-mesenchymal transition. J Clin Invest. 2009. Jun;119(6):1420–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ko EK, Anderson A, Zou J, Huang S, Cho S, Alawi F, et al. H3K36M provokes cellular plasticity to drive aberrant glandular formation and squamous carcinogenesis [Internet]. bioRxiv. 2022. [cited 2023 Dec 13]. p. 2022.08.19.504575. Available from: https://www.biorxiv.org/content/10.1101/2022.08.19.504575v1 [DOI] [PMC free article] [PubMed]
- Koeppel J, Weller J, Peets EM, Pallaseni A, Kuzmin I, Raudvere U, et al. Prediction of prime editing insertion efficiencies using sequence features and DNA repair determinants. Nat Biotechnol. 2023. Oct;41(10):1446–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature. 2016. May 19;533(7603):420–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kong W, Biddy BA, Kamimoto K, Amrute JM, Butka EG, Morris SA. CellTagging: combinatorial indexing to simultaneously map lineage and identity at single-cell resolution. Nat Protoc. 2020. Mar;15(3):750–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wetterstrand Kris A.. The cost of sequencing a human genome [Internet]. Genome.gov. NHGRI; 2019. [cited 2023 Dec 7]. Available from: https://www.genome.gov/about-genomics/fact-sheets/Sequencing-Human-Genome-cost [Google Scholar]
- Kumar M, Keller B, Makalou N, Sutton RE. Systematic determination of the packaging limit of lentiviral vectors. Hum Gene Ther. 2001. Oct 10;12(15):1893–905. [DOI] [PubMed] [Google Scholar]
- Leask A, Denton CP, Abraham DJ. Insights into the molecular mechanism of chronic fibrosis: the role of connective tissue growth factor in scleroderma. J Invest Dermatol. 2004. Jan;122(1):1–6. [DOI] [PubMed] [Google Scholar]
- Lee J, Rabbani CC, Gao H, Steinhart MR, Woodruff BM, Pflum ZE, et al. Hair-bearing human skin generated entirely from pluripotent stem cells. Nature. 2020. Jun;582(7812):399–404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li L, Bowling S, McGeary SE, Yu Q, Lemke B, Alcedo K, et al. A mouse model with high clonal barcode diversity for joint lineage, transcriptomic, and epigenomic profiling in single cells. Cell [Internet]. 2023. Oct 12; Available from: 10.1016/j.cell.2023.09.019 [DOI] [PubMed] [Google Scholar]
- Li R, Zhuang Y, Han M, Xu T, Wu X. piggyBac as a high-capacity transgenesis and gene-therapy vector in human cells and mice. Dis Model Mech. 2013. May;6(3):828–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Livet J, Weissman TA, Kang H, Draft RW, Lu J, Bennis RA, et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature. 2007. Nov 1;450(7166):56–62. [DOI] [PubMed] [Google Scholar]
- Loveless TB, Carlson CK, Hu VJ, Dentzel Helmy CA, Liang G, Ficht M, et al. Molecular recording of sequential cellular events into DNA [Internet]. bioRxiv. 2021a. [cited 2023 Sep 27]. p. 2021.11.05.467507. Available from: https://www.biorxiv.org/content/10.1101/2021.11.05.467507v1 [DOI] [PMC free article] [PubMed]
- Loveless TB, Grotts JH, Schechter MW, Forouzmand E, Carlson CK, Agahi BS, et al. Lineage tracing and analog recording in mammalian cells by single-site DNA writing. Nat Chem Biol. 2021b. Jun;17(6):739–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowry WE, Flores A, White AC. Exploiting Mouse Models to Study Ras-Induced Cutaneous Squamous Cell Carcinoma. J Invest Dermatol. 2016. Aug;136(8):1543–8. [DOI] [PubMed] [Google Scholar]
- Lun XK, Bodenmiller B. Profiling Cell Signaling Networks at Single-cell Resolution. Mol Cell Proteomics. 2020. May;19(5):744–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu R, Neff NF, Quake SR, Weissman IL. Tracking single hematopoietic stem cells in vivo using high-throughput sequencing in conjunction with viral genetic barcoding. Nat Biotechnol. 2011. Oct 2;29(10):928–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu Y, Chan YT, Tan HY, Li S, Wang N, Feng Y. Epigenetic regulation in human cancer: the potential role of epi-drug in cancer therapy. Mol Cancer. 2020. Apr 27;19(1):79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mascharak S, desJardins-Park HE, Davitt MF, Griffin M, Borrelli MR, Moore AL, et al. Preventing Engrailed-1 activation in fibroblasts yields wound regeneration without scarring. Science [Internet]. 2021. Apr 23;372(6540). Available from: 10.1126/science.aba2374 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masuyama N, Mori H, Yachie N. DNA barcodes evolve for high-resolution cell lineage tracing. Curr Opin Chem Biol. 2019. Oct;52:63–71. [DOI] [PubMed] [Google Scholar]
- Mathis N, Allam A, Kissling L, Marquart KF, Schmidheini L, Solari C, et al. Predicting prime editing efficiency and product purity by deep learning. Nat Biotechnol. 2023. Aug;41(8):1151–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matos I, Asare A, Levorse J, Ouspenskaia T, de la Cruz-Racelis J, Schuhmacher LN, et al. Progenitors oppositely polarize WNT activators and inhibitors to orchestrate tissue development. Elife [Internet]. 2020. Apr 20;9. Available from: 10.7554/eLife.54304 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKenna A, Findlay GM, Gagnon JA, Horwitz MS, Schier AF, Shendure J. Whole-organism lineage tracing by combinatorial and cumulative genome editing. Science. 2016. Jul 29;353(6298):aaf7907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meissner A Epigenetic modifications in pluripotent and differentiated cells. Nat Biotechnol. 2010. Oct;28(10):1079–88. [DOI] [PubMed] [Google Scholar]
- Moon H, Donahue LR, Choi E, Scumpia PO, Lowry WE, Grenier JK, et al. Melanocyte Stem Cell Activation and Translocation Initiate Cutaneous Melanoma in Response to UV Exposure. Cell Stem Cell. 2017. Nov 2;21(5):665–78.e6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullick A, Xu Y, Warren R, Koutroumanis M, Guilbault C, Broussau S, et al. The cumate gene-switch: a system for regulated expression in mammalian cells. BMC Biotechnol. 2006. Nov 3;6:43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Obeidy P, Tong PL, Weninger W. Research Techniques Made Simple: Two-Photon Intravital Imaging of the Skin. J Invest Dermatol. 2018. Apr;138(4):720–5. [DOI] [PubMed] [Google Scholar]
- Patel GK, Yee CL, Terunuma A, Telford WG, Voong N, Yuspa SH, et al. Identification and characterization of tumor-initiating cells in human primary cutaneous squamous cell carcinoma. J Invest Dermatol. 2012. Feb;132(2):401–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pavlopoulos GA, Soldatos TG, Barbosa-Silva A, Schneider R. A reference guide for tree analysis and visualization. BioData Min. 2010. Feb 22;3(1):1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pei W, Shang F, Wang X, Fanti AK, Greco A, Busch K, et al. Resolving Fates and Single-Cell Transcriptomes of Hematopoietic Stem Cell Clones by PolyloxExpress Barcoding. Cell Stem Cell. 2020. Sep 3;27(3):383–95.e8. [DOI] [PubMed] [Google Scholar]
- Perli SD, Cui CH, Lu TK. Continuous genetic recording with self-targeting CRISPR-Cas in human cells. Science [Internet]. 2016. Sep 9;353(6304). Available from: 10.1126/science.aag0511 [DOI] [PubMed] [Google Scholar]
- Phan QM, Sinha S, Biernaskie J, Driskell RR. Single-cell transcriptomic analysis of small and large wounds reveals the distinct spatial organization of regenerative fibroblasts. Exp Dermatol. 2021. Jan;30(1):92–101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pollen AA, Nowakowski TJ, Shuga J, Wang X, Leyrat AA, Lui JH, et al. Low-coverage single-cell mRNA sequencing reveals cellular heterogeneity and activated signaling pathways in developing cerebral cortex. Nat Biotechnol. 2014. Oct;32(10):1053–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Potten CS, Allen TD. Control of epidermal proliferative units (EPUs). An hypothesis based on the arrangement of neighbouring differentiated cells. Differentiation. 1975. Aug 11;3(1–3):161–5. [DOI] [PubMed] [Google Scholar]
- Quinn JJ, Jones MG, Okimoto RA, Nanjo S, Chan MM, Yosef N, et al. Single-cell lineages reveal the rates, routes, and drivers of metastasis in cancer xenografts. Science [Internet]. 2021. Feb 26;371(6532). Available from: 10.1126/science.abc1944 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raj B, Wagner DE, McKenna A, Pandey S, Klein AM, Shendure J, et al. Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nat Biotechnol. 2018. Jun;36(5):442–50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rompolas P, Deschene ER, Zito G, Gonzalez DG, Saotome I, Haberman AM, et al. Live imaging of stem cell and progeny behaviour in physiological hair-follicle regeneration. Nature. 2012. Jul 26;487(7408):496–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rostovskaya M, Fu J, Obst M, Baer I, Weidlich S, Wang H, et al. Transposon-mediated BAC transgenesis in human ES cells. Nucleic Acids Res. 2012. Oct;40(19):e150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sada A, Jacob F, Leung E, Wang S, White BS, Shalloway D, et al. Defining the cellular lineage hierarchy in the interfollicular epidermis of adult skin. Nat Cell Biol. 2016. Jun;18(6):619–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sánchez-Danés A, Hannezo E, Larsimont JC, Liagre M, Youssef KK, Simons BD, et al. Defining the clonal dynamics leading to mouse skin tumour initiation. Nature. 2016. Aug 18;536(7616):298–303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sankaran VG, Weissman JS, Zon LI. Cellular barcoding to decipher clonal dynamics in disease. Science. 2022. Oct 14;378(6616):eabm5874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt ST, Zimmerman SM, Wang J, Kim SK, Quake SR. Quantitative Analysis of Synthetic Cell Lineage Tracing Using Nuclease Barcoding. ACS Synth Biol. 2017. Jun 16;6(6):936–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simeonov KP, Byrns CN, Clark ML, Norgard RJ, Martin B, Stanger BZ, et al. Single-cell lineage tracing of metastatic cancer reveals selection of hybrid EMT states. Cancer Cell. 2021. Aug 9;39(8):1150–62.e9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simpson CL, Kojima SI, Getsios S. RNA interference in keratinocytes and an organotypic model of human epidermis. Methods Mol Biol. 2010;585:127–46. [DOI] [PubMed] [Google Scholar]
- Smits JPH, Niehues H, Rikken G, van Vlijmen-Willems IMJJ, van de Zande GWHJF, Zeeuwen PLJM, et al. Immortalized N/TERT keratinocytes as an alternative cell source in 3D human epidermal models. Sci Rep. 2017. Sep 19;7(1):11838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sofi MY, Shafi A, Masoodi KZ. Primer designing tools. In: Bioinformatics for Everyone. Elsevier; 2022. p. 127–39. [Google Scholar]
- Spanjaard B, Hu B, Mitic N, Olivares-Chauvet P, Janjuha S, Ninov N, et al. Simultaneous lineage tracing and cell-type identification using CRISPR-Cas9-induced genetic scars. Nat Biotechnol. 2018. Jun;36(5):469–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stabell AR, Lee GE, Jia Y, Wong KN, Wang S, Ling J, et al. Single-cell transcriptomics of human-skin-equivalent organoids. Cell Rep. 2023. May 30;42(5):112511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ståhl PL, Salmén F, Vickovic S, Lundmark A, Navarro JF, Magnusson J, et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science. 2016. Jul 1;353(6294):78–82. [DOI] [PubMed] [Google Scholar]
- Sulston JE, Schierenberg E, White JG, Thomson JN. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev Biol. 1983. Nov;100(1):64–119. [DOI] [PubMed] [Google Scholar]
- Sun BK, Siprashvili Z, Khavari PA. Advances in skin grafting and treatment of cutaneous wounds. Science. 2014a. Nov 21;346(6212):941–5. [DOI] [PubMed] [Google Scholar]
- Sun J, Ramos A, Chapman B, Johnnidis JB, Le L, Ho YJ, et al. Clonal dynamics of native haematopoiesis. Nature. 2014b. Oct 16;514(7522):322–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun X, Are A, Annusver K, Sivan U, Jacob T, Dalessandri T, et al. Coordinated hedgehog signaling induces new hair follicles in adult skin. Elife [Internet]. 2020. Mar 17;9. Available from: 10.7554/eLife.46756 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Svensson V, Vento-Tormo R, Teichmann SA. Exponential scaling of single-cell RNA-seq in the past decade. Nat Protoc. 2018. Apr;13(4):599–604. [DOI] [PubMed] [Google Scholar]
- Tang W, Liu DR. Rewritable multi-event analog recording in bacterial and mammalian cells. Science [Internet]. 2018. Apr 13;360(6385). Available from: 10.1126/science.aap8992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tumbar T, Guasch G, Greco V, Blanpain C, Lowry WE, Rendl M, et al. Defining the epithelial stem cell niche in skin. Science. 2004. Jan 16;303(5656):359–63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Urnov FD. Imagine CRISPR cures. Mol Ther. 2021. Nov 3;29(11):3103–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- VanHorn S, Morris SA. Next-Generation Lineage Tracing and Fate Mapping to Interrogate Development. Dev Cell. 2021. Jan 11;56(1):7–21. [DOI] [PubMed] [Google Scholar]
- Varadhachary GR. Carcinoma of unknown primary origin. Gastrointest Cancer Res. 2007. Nov;1(6):229–35. [PMC free article] [PubMed] [Google Scholar]
- Verhaegen ME, Harms PW, Van Goor JJ, Arche J, Patrick MT, Wilbert D, et al. Direct cellular reprogramming enables development of viral T antigen-driven Merkel cell carcinoma in mice. J Clin Invest [Internet]. 2022. Apr 1;132(7). Available from: 10.1172/JCI152069 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang T, Chen X, Wang K, Ju J, Yu X, Wang S, et al. Cre-loxP-mediated genetic lineage tracing: Unraveling cell fate and origin in the developing heart. Front Cardiovasc Med. 2023. Feb 27;10:1085629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang D, Jones MG, Naranjo S, Rideout WM 3rd, Min KHJ, Ho R, et al. Lineage tracing reveals the phylodynamics, plasticity, and paths of tumor evolution. Cell. 2022. May 26;185(11):1905–23.e25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yao M, Ren T, Pan Y, Xue X, Li R, Zhang L, et al. A New Generation of Lineage Tracing Dynamically Records Cell Fate Choices. Int J Mol Sci [Internet]. 2022. Apr 30;23(9). Available from: 10.3390/ijms23095021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yesbolatova A, Saito Y, Kitamoto N, Makino-Itou H, Ajima R, Nakano R, et al. The auxin-inducible degron 2 technology provides sharp degradation control in yeast, mammalian cells, and mice. Nat Commun. 2020. Nov 11;11(1):5701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu G, Kim HK, Park J, Kwak H, Cheong Y, Kim D, et al. Prediction of efficiencies for diverse prime editing systems in multiple cell types. Cell. 2023. May 11;186(10):2256–72.e23. [DOI] [PubMed] [Google Scholar]
- Zhang B, Chen T. Local and systemic mechanisms that control the hair follicle stem cell niche. Nat Rev Mol Cell Biol [Internet]. 2023. Oct 30; Available from: 10.1038/s41580-023-00662-3 [DOI] [PubMed] [Google Scholar]
- Zhao Z, Shang P, Mohanraju P, Geijsen N. Prime editing: advances and therapeutic applications. Trends Biotechnol. 2023. Aug;41(8):1000–12. [DOI] [PubMed] [Google Scholar]