Abstract
The potential contribution of a plasmid DNA construct to vaccine-elicited protective immunity was explored in the simian immunodeficiency virus (SIV)/macaque model of AIDS. Making use of soluble major histocompatibility class I/peptide tetramers and peptide-specific killing assays to monitor CD8+ T-lymphocyte responses to a dominant SIV Gag epitope in genetically selected rhesus monkeys, a codon-optimized SIV gag DNA vaccine construct was shown to elicit a high-frequency SIV-specific cytotoxic T-lymphocyte (CTL) response. This CTL response was demonstrable in both peripheral blood and lymph node lymphocytes. Following an intravenous challenge with the highly pathogenic viral isolate SIVsm E660, these vaccinated monkeys developed a secondary CTL response that arose with more rapid kinetics and reached a higher frequency than did the postchallenge CTL response in control plasmid-vaccinated monkeys. While peak plasma SIV RNA levels were comparable in the experimentally and control-vaccinated monkeys during the period of primary infection, the gag plasmid DNA-vaccinated monkeys demonstrated better containment of viral replication by 50 days following SIV challenge. These findings indicate that a plasmid DNA vaccine can elicit SIV-specific CTL responses in rhesus monkeys, and this vaccine-elicited immunity can facilitate the generation of secondary CTL responses and control of viral replication following a pathogenic SIV challenge. These observations suggest that plasmid DNA may prove a useful component of a human immunodeficiency virus type 1 vaccine.
There is an emerging consensus that an effective human immunodeficiency virus type 1 (HIV-1) vaccine must elicit both virus-specific neutralizing antibody and cytotoxic T-lymphocyte (CTL) responses (21). The strategies that will be needed to generate these different types of immune responses are likely to be very different. Finding an approach for inducing an antibody response that is capable of neutralizing a diversity of primary patient HIV-1 isolates has proven elusive (17). However, HIV-1-specific CTL can be generated in rodents and nonhuman primates using a number of vaccine modalities, including live recombinant vectors (10–12, 35, 36), peptides (3, 9, 13, 28, 34, 40), and plasmid DNA (2, 4–6, 22, 26, 38, 39). It will be important to characterize systematically the CTL responses elicited by these various vaccine approaches to determine the magnitude and durability of the responses, as well as the contribution that such responses can make in containing HIV-1 replication. Such data will be needed to select the optimal vaccine approaches to carry forward into human testing.
DNA vaccination, the direct injection of plasmid DNA expressing genes encoding the proteins of pathogens, has proven to be highly immunogenic (7, 24, 27, 31, 32, 43, 44). Studies are currently ongoing to examine the potential utility of plasmid DNA as a component of an AIDS vaccine (2, 4–6, 26, 38, 39). Numerous reports have demonstrated that DNA vaccines can elicit HIV-1 and simian immunodeficiency virus (SIV)-specific CTL, T-helper cells, and antibodies in mice and rhesus monkeys (19, 25, 37, 39, 45–48). Several trials of DNA vaccines, either alone or in combination with other vaccine modalities, have shown promising results in nonhuman primates (2, 19, 22, 25, 32, 38). Because they are relatively stable, are inexpensive to produce, and do not carry with them the pathogenic risks inherent in immunization with live vectors, DNA vaccines represent an attractive solution to the HIV-1 pandemic.
The SIV/macaque model is useful in accessing HIV-1 vaccine strategies (20). SIV isolates are lentiviruses that are similar in nucleotide sequence to HIV-1. Moreover, certain SIV isolates induce an AIDS-like disease in Asian macaque species. The power of this animal model for exploring the efficacy of HIV-1 vaccine strategies has been enhanced by the definition of a dominant SIV Gag CTL epitope in rhesus monkeys expressing the major histocompatibility complex (MHC) class I allele Mamu-A*01 (1, 29). The immune response to this epitope can be monitored in infected and vaccinated animals using both functional CTL assays and flow cytometric analysis of tetrameric MHC class I/peptide-binding CD8+ T lymphocytes (8, 15–17, 35).
This study was initiated to evaluate a plasmid DNA vaccine-elicited SIV Gag-specific CTL response in rhesus monkeys and to determine the contribution of that restricted immune response to containing a highly pathogenic primary SIV infection.
MATERIALS AND METHODS
Animals.
All animals were maintained in accordance with the guidelines of the Committee on Animals for the Harvard Medical School and Guide for the Care and Use of Laboratory Animals (6a).
Selection of Mamu-A*01+ rhesus monkeys.
Rhesus monkeys were screened for the presence of the Mamu-A*01 allele using a PCR-based technique as previously described (15–17). EDTA-preserved whole blood from rhesus monkeys was subjected to Ficoll diatrizoate density gradient centrifugation to isolate leukocytes, and the washed cell pellets were resuspended in 200 μl of phosphate-buffered saline (PBS). DNA extraction was then carried out using a QIAmp blood kit (Qiagen Inc., Chatsworth, Calif.). PCR was performed on 200 to 500 μg of extracted DNA using allele-specific primers in a 50-μl reaction consisting of 60 mM Tris (pH 8.5), 2 mM MgCl2, 15 mM ammonium sulfate, 2 mM deoxynucleoside triphosphates (0.5 mM each), and 5 μl of Taq polymerase. Primers A*01/F (5′-GAC AGC GAC GCC GCG AGC CAA-3′) and A*01/R (5′-GCT GCA GCG TCT CCT TCC CC-3′) were used at a final concentration of 800 nM each. Two additional primers specific for a conserved MHC class II sequence (based on the macaque homologue of HLA-DRB3) were included in the reaction as internal positive controls. Primers 5′ MDRB (5′-GCC TCG AGT GTC CCC CCA GCA CGT TTC-3′) and 3′ MDRB (5′-GCA AGC TTT CAC CTC GCC GCT G-3′) were used at a final concentration of 680 nM each. PCR was carried out using a Perkin-Elmer GeneAmp System 9600 thermocycler (Perkin-Elmer Inc., Norwalk, Conn.). Samples were denatured at 96°C for 2 min, followed by 5 cycles of 25 s at 96°C and 60 s at 72°C, followed by 21 cycles of 25 s at 96°C, 50 s at 67°C, and 45 s at 72°C, followed by 4 cycles of 25 s at 96°C, 60 s at 55°C, and 80 s at 72°C. PCR products were analyzed by 1% agarose gel electrophoresis. Ten microliters of each PCR product was loaded per lane.
Potential Mamu-A*01+ animals were identified by the presence of two bands, a 685-bp amplified product and a 260-bp band. DNA sequence analysis was then performed on all potential positive samples to confirm nucleotide sequence identity with the published Mamu-A*01 prototype sequence (14). Prior to sequencing, amplified DNA was treated with 1 U of shrimp alkaline phosphatase and 10 U of exonuclease I per reaction for 15 min at 37°C followed by 15 min at 80°C. The sequencing templates were then purified using a QIAquick PCR purification kit (Qiagen). For each template, 70 ng of DNA was used for DNA sequencing together with 5 pmol of primer. Four PCR primers were used for sequencing: A*01/F and A*01/R, whose sequences are shown above, and B/1+ (5′-CTG CGC GGC TAC TAC AAC CA-3′) and G/1+ (5′-ATG TAA TCC TTG CCG TCG TA-3′). Sequencing was carried out at a central core sequencing facility on an ABI-373 stretch DNA sequencing machine, using ABI AmpliTaq FS dye terminator chemistry (Perkin-Elmer). All animals used in this study were genotypically Mamu-A*01+, based on the above screening as well as functional CTL assay.
Immunization of rhesus monkeys.
Monkeys were immunized with 5 mg of plasmid DNA, 2.5 mg in each quadricep muscle by intramuscular inoculation. Either an SIV gag-expressing plasmid (n = 5) or an empty control vector (n = 4) was inoculated. Injections were performed using a Biojector needle-free injection system and the supplied Biojector no. 3 syringe as instructed by the manufacturer (Bioject, Portland, Oreg.).
Recombinant vaccine constructs.
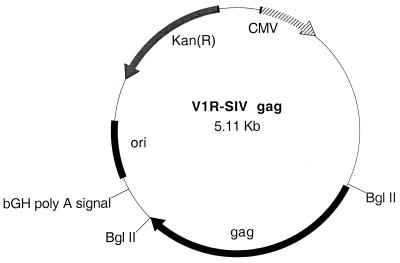
The vaccination plasmid, V1R-SIV gag (Fig. 1), is a pUC-based vector that utilizes the human cytomegalovirus immediate-early promoter with intron A and bovine growth hormone transcription terminator/polyadenylation signal as expression regulatory elements and expresses full-length SIV gag (30). The SIV gag open reading frame is homologous to that of SIVmac239 and was synthesized using optimal codons for human gene expression (18).
FIG. 1.
SIV gag DNA expression vector V1R-SIV gag. Kan(R), kanamycin resistance gene; CMV, cytomegalovirus; ori, origin of replication; bGH, bovine growth hormone.
Western blotting.
Human RD cells in duplicate 25-cm2 tissue culture flasks were transfected with either mock preparation (PBS) or 10 μg of V1R-SIV gag DNA construct using a calcium phosphate-based transfection kit (Pharmacia Biotech, Piscataway, N.J.). Twenty-four hours later, the cells were washed with PBS three times and lysed in 0.5 ml of lysis buffer (50 mM Tris, 5 mM EDTA, 150 mM NaCl, 0.5% Nonidet P-40, 1 mM phenylmethylsulfonyl fluoride, aprotinin [0.01 mg/ml; Sigma Chemical Co., St. Louis, Mo.]) at 4°C for 30 min. The aliquots of cell lysate (20 μl) were mixed with an equal volume of 2× sample buffer (Novex, San Diego, Calif.) and heated at 100°C for 5 min. The proteins were then resolved by electrophoresis on 4 to 20% gradient sodium dodecyl sulfate (SDS)-polyacrylamide gels (Novex). Following electrophoresis, proteins were transferred onto a polyvinylidene difluoride membrane (Millipore) under a constant current of 200 mA. The membrane was blocked with BLOTTO solution (5% nonfat dry milk, 2% fetal calf serum, 0.2% Tween 20, and 10 mM Tris-Cl [pH 8.0] in PBS) overnight at room temperature and then incubated with anti-p27 monoclonal antibody (1:1,000 dilution; ABI, Columbia, Md.) for 2 h at room temperature. After washing three times with BLOTTO solution, the membrane was incubated with horseradish peroxidase-conjugated goat anti-mouse antibody for 1 h. The membrane was further washed with BLOTTO solution and developed with an ECL (enhanced chemiluminescence reagent kit (Amersham, Arlington Park, Ill.) for luminescent detection.
Vaccine preparations.
Inoculated cultures of terrific broth (1.2% Bacto Tryptone, 2.4% Bacto Yeast Extract, 0.4% glycerol) supplemented with 15 mM KH2PO4, 65 mM K2HPO4, and 50 μg of kanamycin per ml were grown overnight with shaking at 37°C. Minipreparations of plasmids were made using a PERFECTprep plasmid DNA isolation kit (5 Prime-3 Prime, Inc., Boulder, Colo.). Maxipreparations of vaccine plasmids were made by standard alkaline lysis followed by double CsCl gradient banding. A 1-liter overnight culture was centrifuged for 15 min at 7,500 rpm and resuspended in 14 ml of lysozyme buffer with lysozyme (0.9% glucose, 10 mM EDTA, 25 mM Tris-Cl [pH 8], 4 mg of lysozyme per ml). The suspension was then lysed with 28 ml of 0.2 M NaOH–1% SDS, neutralized with 21 ml of sodium acetate 3 M (pH 4.8), and centrifuged for 20 min at 17,000 rpm in a Sorvall centrifuge (Sorvall, Braintree, Mass.). The supernatant was removed, centrifuged again for 20 min at 17,000 rpm, filtered, and ethanol precipitated with 3 volumes of 95% ethanol. Following a 30-min incubation at −20°C and centrifugation at 6,500 rpm for 30 min in a Sorvall centrifuge, the supernatants were discarded, and the ethanol-precipitated pellets were air dried and resuspended in 11.5 ml of TSE buffer (10 mM Tris-Cl [pH 8], 1 mM EDTA, 10 mM NaCl). Optical-grade CsCl (13.2 g; Life Technologies) and 2 ml of ethidium bromide (10 mg/ml) were added, and the solution was ultracentrifuged at 60,000 rpm overnight at 20°C. The CsCl-banded DNA was removed and then spun on a second CsCl gradient. Following double CsCl banding, the ethidium was extracted five times using CsCl-saturated isopropanol, and the DNA was precipitated using 3 volumes of ethanol. The DNA was washed with 70% ethanol, resuspended in TE buffer (10 mM Tris-Cl [pH 8], 1 mM EDTA), extracted with phenol-chloroform, extracted with chloroform, reprecipitated with ethanol, washed with 70% ethanol, and then resuspended in sterile TE buffer. The DNA was quantitated and adjusted to a concentration of 5 mg/ml with sterile isotonic saline.
Functional cytotoxicity assay.
Peripheral blood leukocytes (PBL) from Mamu-A*01+ rhesus monkeys were cultured with the optimal SIV peptide p11C (CTPYDINQM; 10 μg/ml) and maintained at a density of 2 × 106 cells/ml. On day 3 of culture, the medium was supplemented with recombinant human interleukin-2 (IL-2; 20 U/ml; provided by Hoffmann-La Roche), and cultures were maintained for an additional 7 days. Cells were then centrifuged over Ficoll-Hypaque and assessed as effectors in a standard 51Cr release assay at an effector-to-target ratio ranging from 20:1 to 0.6:1. Target cells were Mamu-A*01+ B-lymphoblastoid cell line (B-LCL) pulsed during overnight 51Cr labeling with 10 μg of either p11C or the control peptide p11B (ALSEGCTPYDIN) per ml. All wells were assayed in triplicate. Plates were incubated for 5 h in a humidified incubator at 37°C. Specific release was calculated as [(experimental release) − (spontaneous release)/(maximum release) − (spontaneous release)] × 100.
pCTL frequency determination by limiting dilution analysis (LDA).
Freshly isolated PBL were plated in twofold dilutions from 32,000 to 250 cells per well in 24-well replicates. The p11C-specific CTL were then expanded by the addition of 2 × 105 p11C-pulsed, irradiated, Mamu-A*01+ B-LCL. Microcultures were fed on days 5 and 10 by the addition of culture medium supplemented with recombinant IL-2 (10 U/ml). After 14 days of culture, the effector cells were assessed for cytotoxicity against 51Cr-labeled p11C- and control peptide p11B-pulsed target cells. Individual wells screened against p11C-pulsed targets that demonstrated counts 3 standard deviations higher than p11B-pulsed target cells were scored positive for p11C-specific CTL activity. By plotting the fraction of negative wells against the number of cells added per well, the p11C-specific precursor CTL (pCTL) frequency was extrapolated from the cell concentration at which 37% of the wells would be considered negative.
Mamu-A*01/peptide tetramer complex formation and staining of peptide-specific CD8αβ+ T lymphocytes.
Mamu-A*01/p11C tetramer complexes were prepared as previously described (15). Phycoerythrin (PE)-labeled streptavidin (Prozyme, San Leondro, Calif.) was mixed stepwise with biotinylated Mamu-A*01/peptide complexes at a molar ratio of 1:4 to produce the tetrameric complexes. All antibodies used in this study were directly coupled to fluorescein isothiocyanate (FITC), phycoerythrin-Texas red (ECD), or allophycocyanin (APC). The following monoclonal antibodies were used: anti-CD8α (Leu2a)-FITC (Becton Dickinson), anti-CD8αβ-ECD (Coulter), and anti-CD3-APC (FN18; kindly provided by D. M. Neville, Jr., National Institutes of Health, Bethesda, Md.).
The PE-coupled tetrameric Mamu-A*01/p11C complexes were used in combination with anti-CD8α-FITC, anti-CD8αβ-ECD, and anti-CD3-APC to stain 100 μl of fresh blood or 2 × 105 lymphocytes isolated by Ficoll-Hypaque density gradient centrifugation following in vitro peptide stimulation as previously described (15). The phenotyping of p11C-specific CD8+ T cells was done by staining 100 μl of whole blood with 1 μg of 488-Neutravadin (Molecular Probes, Eugene, Oreg.)-coupled tetrameric Mamu-A*01/p11C complexes in combination with anti-CD8αβ-ECD and anti-CD3-APC along with one of the following: anti-CD11a-PE, anti-CD28-PE, anti-CD45RA-PE, anti-CD49d-PE, anti-CD62L-PE, anti-CD95-PE, or anti-HLA-DR-PE.
Whole-blood samples were lysed using a Coulter Immunoprep reagent system and a Q-prep workstation (Beckman Coulter Inc.). Ten thousand gated events were collected, and samples were analyzed on a Coulter EPICS Elite ESP. The total number of Mamu-A*01/p11C tetramer-binding cells per microliter of whole blood was calculated as the product of (% Mamu-A*01/p11C tetramer binding) × (total white blood cell count per microliter of whole blood) × (% lymphocytes) × (% CD8+ cells). Total white blood cell counts and percent lymphocytes were determined by complete blood count analysis on 500 μl of EDTA-anticoagulated blood using an automated hematology analyzer (T540; Coulter Corporation).
Data analysis was performed using the EPICS Elite software (version 4.02; Beckman Coulter). Data presentation was performed by using WinMDI software version 2.7 (Joseph Trotter, La Jolla, Calif.) and Microsoft PowerPoint 97 (Microsoft, Redmond, Wash.).
SIVsm E660 challenge.
All animals were challenge intravenously with 50 50% monkey infective doses (MID50) of SIVsm E660 (kindly provided by V. Hirsch, National Institutes of Health).
In situ hybridization.
A 35S-labeled, single-stranded, antisense RNA probe (Lofstrand Laboratories, Gaithersburg, Md.) was used to detect viral mRNA. The hybridization was done on frozen sections as previously described (42). The sections were examined with a microscope equipped with epiluminescent illumination (Axiophot; Carl Ziess Inc., Jena, Germany). Cells were considered positive for viral gene expression if the grain count was more than six times higher than the background count.
RESULTS
V1R-SIV gag DNA vaccine expression of the gag gene.
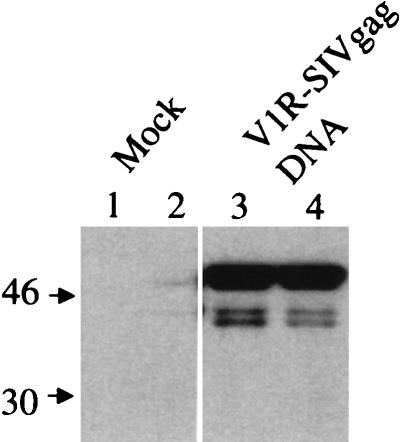
To examine protein expression from the V1R-SIV gag DNA vaccine (Fig. 1), we analyzed SIV Gag protein expression by transient transfection of human RD cells followed by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) of whole-cell lysates (Fig. 2). Figure 2 shows that V1R-SIV gag DNA-transfected cells express the SIV Gag protein to higher levels (lanes 3 and 4) than cells transiently transfected with the empty control vector (lanes 1 and 2).
FIG. 2.
In vitro expression of the V1R-SIV gag DNA vaccine construct in transiently transfected human RD cells. Human RD cells were transfected with either mock preparation (PBS; lanes 1 and 2) or 10 μg of V1R-SIV gag DNA construct (lanes 3 and 4) using calcium phosphate. Twenty-four hours later, the cells were washed and lysed, and the proteins were then resolved by SDS-PAGE. Sizes are indicated in kilodaltons.
SIV Gag-specific peripheral blood CD8+ T-lymphocyte responses after DNA immunization.
To evaluate the immunogenicity of a plasmid SIV gag DNA vaccine, Mamu-A*01+ rhesus monkeys were immunized by intramuscular inoculation with 5 mg of either an SIVmac 239 gag-expressing plasmid or a control empty plasmid. The immunization schedule included inoculations at 0, 1, 2, and 6 months. These monkeys were then monitored for the development of SIV Gag-specific CD8+ T-lymphocyte responses. PBL were assessed in whole blood without in vitro stimulation for Mamu-A*01/p11C tetramer binding to CD8+ T cells. In addition, CD8+ T lymphocytes from these animals were assessed for Mamu-A*01/p11C tetramer binding as well as for functional p11C-specific lytic activity after in vitro p11C stimulation. Finally, p11C-specific CTL precursor frequencies were determined by functional LDA. By week 4 following the first plasmid DNA immunization, four of the five SIV gag DNA-immunized monkeys had detectable Gag epitope-specific functional CTL activity. The Gag epitope-specific CTL responses in these four animals increased following subsequent DNA immunizations (Table 1). The control plasmid-immunized monkeys remained negative for Gag epitope-specific functional CTL activity throughout the course of the experiment.
TABLE 1.
SIV Gag epitope p11C-specific CD8+ T-lymphocyte responses after plasmid DNA immunization
| Immunization | Monkey | Assay | Value at indicated wk after initial immunization
|
||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 4 | 6 | 8 | 9 | 10 | 12 | 16 | 24 | 25 | 26 | 28 | 29 | 30 | 31 | 32 | 34 | |||
| SIV gag | P091 | % Fresha | 0.0 | 0.0 | 0.3 | 0.1 | 0.1 | 0.2 | 0.2 | 0.5 | 0.3 | 0.9 | 0.6 | 0.1 | NDf | ND | ND | ND | ND |
| No. of cells/μl by tetramerb | ND | ND | 4 | ND | 1 | ND | ND | ND | 3 | 6 | 5 | ND | ND | ND | ND | ND | ND | ||
| No. of cells/μl by LDAc | 0 | 0.04 | 0.16 | 0.46 | 0.02 | 0.37 | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| % Peptide stimulatedd | 0.1 | 11.3 | 40.6 | 23.3 | 40.8 | 47.4 | 64.8 | 75.1 | 27.3 | 20.9 | 51.4 | 54.0 | ND | ND | ND | ND | ND | ||
| % Specific lysise | 0.0 | 46.5 | 46.3 | 48.4 | 51.0 | 47.1 | 73.6 | 82.2 | 64.0 | 46.4 | 58.3 | 62.5 | ND | ND | ND | ND | ND | ||
| N529 | % Fresh | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.1 | 0.0 | ND | ND | ND | ND | ND | |
| No. of cells/μl by tetramer | ND | ND | ND | ND | 0 | ND | ND | ND | 0 | 0 | 1 | 0 | ND | ND | ND | ND | ND | ||
| No. of cells/μl by LDA | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| % Peptide stimulated | 0.1 | 0.3 | 0.4 | 0.1 | 1.3 | 0.4 | 0.5 | ND | 0.2 | 0.2 | 0.4 | 0.7 | ND | ND | ND | ND | ND | ||
| % Specific lysis | 0.0 | 6.8 | 6.4 | 1.6 | 19.4 | 3.6 | 6.6 | ND | 6.1 | 11.6 | 13.0 | 18.2 | ND | ND | ND | ND | ND | ||
| T258 | % Fresh | 0.0 | 0.1 | 0.1 | 0.0 | 0.1 | 0.1 | 0.1 | 0.0 | 0.1 | 0.1 | 0.1 | 0.2 | 0.3 | 0.1 | 0.1 | 0.1 | 0.0 | |
| No. of cells/μl by tetramer | ND | ND | 1 | ND | 1 | ND | ND | ND | 0 | 2 | 2 | 4 | 5 | 2 | 1 | ND | 0 | ||
| No. of cells/μl by LDA | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| % Peptide stimulated | 0.1 | 11.5 | 7.9 | 7.5 | 13.3 | 21.1 | 21.5 | 15.1 | 8.6 | 6.8 | 13.1 | 17.4 | 20.0 | 19.7 | 14.2 | 3.4 | 3.6 | ||
| % Specific lysis | 0.0 | 22.4 | 32.8 | 28.2 | 50.3 | 38.7 | 43.5 | 56.6 | 51.6 | 39.2 | 47.1 | 69.2 | 68.3 | 80.6 | 65.2 | ND | 55.9 | ||
| 9298 | % Fresh | 0.0 | 0.0 | 0.1 | 0.0 | 0.0 | 0.2 | 0.1 | 0.1 | 0.1 | 0.1 | 0.3 | 0.0 | 0.2 | 0.1 | 0.2 | 0.2 | 0.1 | |
| No. of cells/μl by tetramer | ND | ND | 1 | ND | 0 | ND | ND | ND | 1 | 1 | 3 | 0 | 3 | 1 | 2 | 2 | 1 | ||
| No. of cells/μl by LDA | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| % Peptide stimulated | 1.1 | 25.0 | 32.5 | 18.4 | 26.9 | 34.1 | 26.2 | 19.3 | 19.7 | 21.7 | 32.0 | 25.9 | 30.1 | 26.1 | 12.8 | 14.8 | 16.7 | ||
| % Specific lysis | 7.5 | 26.5 | 47.7 | 27.8 | 43.5 | 28.2 | 48.1 | 31.4 | 61.0 | 37.8 | 41.3 | 52.1 | 55.1 | 50.5 | 47.5 | ND | 55.9 | ||
| 257 | % Fresh | 0.0 | 0.2 | 0.7 | ND | 0.1 | 0.3 | 0.1 | ND | 0.0 | 0.1 | 1.1 | ND | ND | ND | ND | ND | ND | |
| No. of cells/μl by tetramer | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| No. of cells/μl by LDA | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| % Peptide stimulated | ND | 36.0 | ND | ND | ND | 30.8 | 32.4 | ND | 29.3 | 13.8 | 15.3 | ND | ND | ND | ND | ND | ND | ||
| % Specific lysis | 0.0 | 55.0 | ND | ND | ND | 47.1 | 73.6 | ND | 52.0 | 53.9 | 45.3 | ND | ND | ND | ND | ND | ND | ||
| Control | P967 | % Fresh | ND | 0.0 | ND | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND |
| No. of cells/μl by tetramer | ND | ND | ND | ND | 0 | ND | ND | ND | 0 | 0 | 0 | ND | ND | ND | ND | ND | ND | ||
| No. of cells/μl by LDA | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| % Peptide stimulated | 0.3 | 0.0 | 0.0 | 6.6 | 0.3 | 0.2 | 0.2 | 3.0 | 0.8 | 1.8 | 0.2 | 0.2 | ND | ND | ND | ND | ND | ||
| % Specific lysis | 0.0 | 0.0 | 0.0 | 0.0 | 3.5 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 9.1 | 0.0 | ND | ND | ND | ND | ND | ||
| R468 | % Fresh | ND | 0.0 | ND | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND | |
| No. of cells/μl by tetramer | ND | ND | ND | ND | 0 | ND | ND | ND | 0 | 0 | 0 | ND | ND | ND | ND | ND | ND | ||
| No. of cells/μl by LDA | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| % Peptide stimulated | 0.2 | 0.0 | 0.0 | 0.0 | 0.3 | 0.2 | 0.2 | 2.7 | 0.7 | 0.5 | 0.4 | 0.2 | ND | ND | ND | ND | ND | ||
| % Specific lysis | 0.0 | 0.0 | 0.0 | 0.0 | 4.6 | 0.0 | 0.7 | 0.0 | 0.0 | 6.4 | 3.0 | 0.0 | ND | ND | ND | ND | ND | ||
| T720 | % Fresh | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | |
| No. of cells/μl by tetramer | ND | ND | 0 | ND | 0 | ND | ND | ND | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | ||
| No. of cells/μl by LDA | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| % Peptide stimulated | 0.3 | 0.1 | 0.1 | 7.6 | 4.1 | 2.3 | 1.0 | 1.3 | 1.0 | 1.1 | 1.5 | 0.1 | 0.1 | 1.0 | 0.9 | 0.3 | 0.4 | ||
| % Specific lysis | 11.0 | 0.0 | 0.0 | 0.0 | 4.5 | 0.0 | 1.7 | 0.0 | 11.8 | 19.5 | 0.0 | 8.0 | 1.2 | 7.6 | 6.1 | ND | 1.4 | ||
| V299 | % Fresh | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND | |
| No. of cells/μl by tetramer | ND | ND | 0 | ND | 0 | ND | ND | ND | 0 | 0 | 0 | 0 | ND | ND | ND | ND | ND | ||
| No. of cells/μl by LDA | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | ||
| % Peptide stimulated | 0.0 | 0.1 | 5.7 | 1.1 | 0.2 | 0.1 | 0.7 | 1.0 | 0.6 | 2.0 | 2.0 | 1.7 | ND | ND | ND | ND | ND | ||
| % Specific lysis | 3.9 | 0.0 | 5.4 | 0.0 | 5.7 | 0.0 | 0.0 | 0.8 | 13.6 | 6.9 | 5.1 | ND | ND | ND | ND | ND | ND | ||
CD8αβ+ PBL in whole blood were assessed for binding of the Mamu-A*01/p11C tetramer.
Total number of p11C-specific CD8αβ+ cells per microliter of whole blood as calculated by binding of the Mamu-A*01/p11C tetramer.
Total number of p11C-specific CD8αβ+ cells per microliter of whole blood as calculated by LDA.
PBL were cultured in vitro with p11C (10 μg/ml) for 12 days and then assessed for binding of the Mamu-A*01/p11C tetramer to CD8αβ+ T cells.
PBL were cultured in vitro with p11C (10 μg/ml) for 12 days and then assessed for p11C-specific lysis using a standard 51Cr release assay. Data represent percent p11C-specific lysis at an effector-to-target ratio of 20:1.
ND, not done.
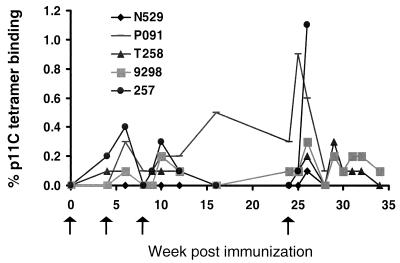
Mamu-A*01/p11C tetramer binding to unstimulated CD8αβ+ T cells in whole blood was also monitored in the monkeys (Table 1; Fig. 3). By week 6, 2 weeks after the second immunization, four of the five SIV gag DNA-vaccinated animals had detectable Mamu-A*01/p11C tetramer-binding CD8αβ+ T cells in whole blood, comprising 0.1 to 0.6% of all circulating CD8αβ+ T lymphocytes. Peak levels of Mamu-A*01/p11C tetramer-binding CD8αβ+ T cells in whole blood were seen 2 weeks after the fourth vaccination, following immunizations at week 24 (Fig. 4A). The control plasmid inoculated monkeys never developed Mamu-A*01/p11C tetramer-binding CD8+ T lymphocytes that could be detected in whole blood during the course of the experiment.
FIG. 3.
SIV Gag p11C tetramer binding to peripheral blood CD8+ lymphocytes of SIV gag DNA-immunized Mamu-A*-01+ rhesus monkeys. Arrows indicate the weeks at which the monkeys were immunized. Percent p11C tetramer binding represents Mamu-A*01/p11C tetramer-binding CD8αβ+ T cells in unstimulated whole blood samples at each time point tested.
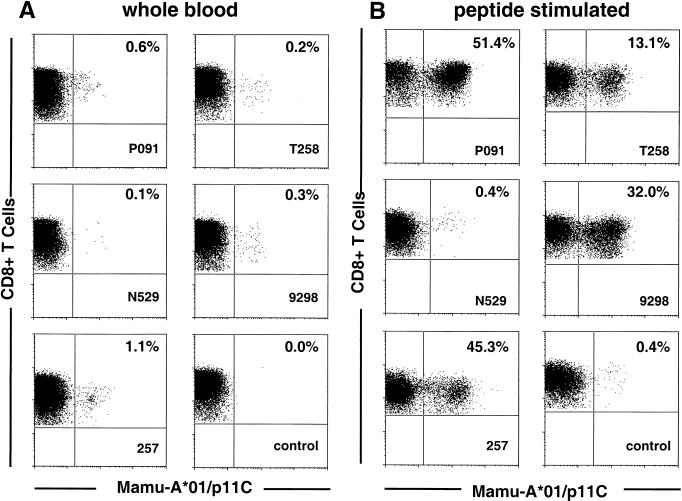
FIG. 4.
SIV Gag p11C tetramer binding-CD8αβ+ T cells in whole blood of SIV gag DNA-immunized Mamu-A*-01+ rhesus monkeys 2 weeks after the fourth immunization. Cells were gated on CD8αβ+ CD3+ lymphocytes. (A) Staining of unstimulated whole blood; (B) staining of lymphocytes after in vitro expansion with p11C.
PBL from the vaccinated monkeys were also stimulated in vitro with the Gag peptide p11C in IL-2-containing medium. These cells were assessed for Mamu-A*01/p11C tetramer binding and functional Gag epitope-specific CTL activity. The peptide-stimulated PBL from four of the five gag DNA-vaccinated animals demonstrated persistent high levels of Mamu-A*01/p11C tetramer-binding CD8+ T cells (Table 1; Fig. 4B). Moreover, after in vitro peptide stimulation, the levels of Mamu-A*01/p11C tetramer-binding CD8+ T cells in these monkeys mirrored the levels of functional Gag epitope-specific CTL activity (Table 1; Fig. 5). The tetramer-binding CD8+ T lymphocytes in the PBL of the single monkey that developed only a low level of tetramer-binding cells in whole blood, N529 (Fig. 4A), failed to expand to a significant level (Fig. 4B). None of the control DNA-immunized animals developed T-lymphocyte populations that demonstrated significant levels of Mamu-A*01/p11C tetramer-binding CD8αβ+ T cells after in vitro peptide stimulation. Using LDA, we were able to detect vaccine-induced p11C-specific CD8+ pCTL in only one of the four gag DNA-vaccinated monkeys tested, animal P091 (Table 1). As has been shown by others, the p11C-specific pCTL frequency calculated by LDA was typically 10- to 100-fold lower than the number of p11C-specific CD8+ CTL measured by direct tetramer staining in fresh blood.
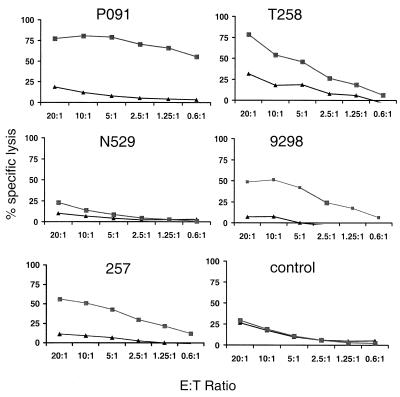
FIG. 5.
Plasmid DNA vaccine-induced Gag epitope-specific functional CTL activity in PBL of vaccinated monkeys 2 weeks after the fourth immunization. PBL of the vaccinated monkeys were activated in vitro with p11C in IL-2-containing medium and assessed for lysis of p11C-pulsed (■) and control peptide p11B-pulsed (▴) B-LCL target cells.
DNA vaccine-induced SIV Gag-specific CD8+ T-lymphocyte responses in lymph nodes.
Since HIV-1 replication occurs predominantly in lymph nodes of infected individuals, it is important that an HIV-1 vaccine elicit virus-specific CD8+ CTL that are resident in secondary lymphoid organs. To determine whether DNA vaccination elicited such a response, inguinal lymph nodes were obtained from the immunized animals 1 week after the third immunization and analyzed for SIV Gag epitope-specific CD8+ T lymphocytes. Lymph node cell suspensions from two of the five plasmid SIV gag DNA-vaccinated animals (P091 and 9298) showed detectable levels of Mamu-A*01/p11C tetramer-binding CD8αβ+ T cells (Table 2). After p11C stimulation, lymph node cells from these same two animals, as well as a third vaccinated animal (T258), showed high levels of Mamu-A*01/p11C tetramer binding as well as functional p11C-specific lytic activity. No SIV Gag epitope-specific CD8+ T lymphocytes were detected in the lymph nodes of the control plasmid DNA-immunized monkeys.
TABLE 2.
Lymph node T lymphocytes from SIV gag plasmid DNA-vaccinated monkeys demonstrate Gag epitope-specific CTL responses 10 weeks following the initial immunization
| Immunization | Monkey | % CD8αβ tetramer binding
|
% Specific lysisc | |
|---|---|---|---|---|
| Fresha | Peptide stimulatedb | |||
| SIV gag | P091 | 0.1 | 17.2 | 65.7 |
| N529 | 0.0 | 1.0 | 13.0 | |
| T258 | 0.0 | 12.7 | 35.7 | |
| 9298 | 0.1 | 28.1 | 36.5 | |
| 257 | NDd | ND | ND | |
| Control | P967 | 0.0 | 1.3 | 4.1 |
| R468 | 0.0 | 0.2 | 6.5 | |
| T720 | 0.0 | 0.2 | 3.2 | |
| V299 | 0.0 | 3.3 | 2.1 | |
Lymphocytes from freshly obtained lymph nodes were directly assessed for binding of the Mamu-A*01/p11C tetramer to CD8αβ+ T cells.
Lymph node cells were cultured in vitro with p11C (10 μg/ml) for 12 days and then assessed for binding of the Mamu-A*01/p11C tetramer to CD8αβ+ T cells.
Lymph node cells were cultured in vitro with p11C (10 μg/ml) for 12 days and then assessed for p11C-specific lysis using a standard 51Cr release assay. Data represent percent specific lysis at an effector-to-target ratio of 5:1.
ND, not done.
Secondary CTL responses in DNA-vaccinated monkeys after SIV challenge.
We then assessed the protection afforded these monkeys by plasmid DNA vaccine-elicited immunity against SIV challenge as well as the durability of this protection. A subset of the animals were challenged 4 weeks after the final DNA immunization (SIV gag DNA-vaccinated animals P091 and N529; control DNA-vaccinated animals R468, P967, and V299); the remaining animals were challenged 9 weeks after the final DNA immunization. All animals were challenged with an intravenous inoculation of 50 MID50 of the highly pathogenic SIVsm E660. The challenged monkeys were monitored for the development of SIV Gag epitope-specific CTL responses by Mamu-A*01/p11C tetramer binding in whole blood, as well as by tetramer binding and functional CTL activity after in vitro p11C stimulation of PBL. All SIV gag plasmid DNA-vaccinated animals, regardless of the time interval between the final immunization and viral challenge, developed strong secondary CTL responses to the challenge virus. The SIV gag DNA-vaccinated monkeys developed higher levels of Mamu-A*01/p11C tetramer-binding cells in whole blood, and these cells appeared earlier after virus challenge than did the control DNA-vaccinated monkeys (Table 3). After the initial burst of virus replication, the levels of circulating Mamu-A*01/p11C tetramer-binding CD8αβ+ T cells in the SIV gag DNA-vaccinated monkeys and those of the control DNA-vaccinated monkeys did not differ significantly (Fig. 6).
TABLE 3.
SIV Gag epitope p11C-specific CD8+ T-lymphocyte responses of vaccinated animals after intravenous challenge with SIVsm E660
| Immunization | Monkey | Assay | Value at indicated day after challenge
|
||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 3 | 7 | 10 | 12 | 14 | 17 | 21 | 24 | 28 | 35 | 42 | 49 | 63 | 77 | |||
| SIV gag | P091 | % Fresha | 0.1 | 0.1 | 0.0 | 10.5 | 57.3 | 56.9 | 37.7 | 25.3 | NDe | 14.4 | 10.4 | 5.7 | 3.2 | 6.8 | 5.5 |
| No. of cells/μlb | 1 | 1 | 0 | 306 | 2,843 | 822 | 856 | 760 | ND | 241 | 156 | 65 | 39 | 68 | 46 | ||
| % Peptide stimulatedc | 54.0 | 48.1 | 62.2 | 20.7 | 34.4 | 76.0 | 59.5 | 84.5 | ND | 87.0 | 70.3 | 75.8 | 83.3 | 83.3 | 90.8 | ||
| % Specific lysisd | 62.5 | 62.3 | 80.6 | ND | ND | 73.5 | 79.8 | 87.1 | ND | 78.7 | ND | 98.1 | 81.5 | 75.8 | 59.0 | ||
| N529 | % Fresh | 0.0 | 0.0 | 0.0 | 1.0 | 9.1 | 11.2 | 4.5 | 3.7 | ND | 1.4 | 3.0 | 3.2 | 3.1 | 2.2 | 1.8 | |
| No. of cells/μl | 0 | 0 | 0 | 8 | 78 | 91 | 52 | 58 | ND | 9 | 38 | 21 | 36 | ND | 21 | ||
| % Peptide stimulated | 0.7 | 0.3 | 0.9 | 9.4 | 12.6 | 5.5 | 6.2 | 15.3 | ND | 5.1 | 2.7 | 5.1 | 12.2 | 28.7 | 25.9 | ||
| % Specific lysis | 18.2 | 7.4 | 6.0 | ND | 33.3 | 18.7 | 45.4 | 68.5 | ND | 34.3 | ND | 60.2 | 52.7 | 62.1 | 53.6 | ||
| T258 | % Fresh | 0.1 | 0.0 | 0.1 | 0.1 | 0.1 | 0.1 | 0.2 | 2.0 | 3.8 | 3.0 | 2.1 | 1.1 | 0.9 | 0.6 | 0.8 | |
| No. of cells/μl | 1 | 0 | 2 | 1 | 1 | 2 | 3 | 25 | 76 | 61 | 35 | 20 | 11 | 6 | 11 | ||
| % Peptide stimulated | 3.6 | 2.4 | 11.9 | 6.7 | 4.2 | 11.7 | 7.0 | 13.2 | 15.8 | 25.6 | 8.4 | 25.7 | 18.8 | 20.5 | 13.9 | ||
| % Specific lysis | 23.5 | ND | 60.8 | 39.7 | 21.2 | 53.3 | 50.5 | 54.4 | 71.0 | 55.9 | 50.1 | 41.4 | 45.4 | 42.3 | 38.1 | ||
| 9298 | % Fresh | 0.1 | 0.2 | 9.1 | 17.1 | 13.0 | 9.0 | 8.5 | 5.4 | 5.6 | 4.6 | 3.8 | 2.5 | 2.4 | 2.1 | 1.7 | |
| No. of cells/μl | 1 | 1 | 62 | 210 | 176 | 146 | 85 | 66 | 60 | 50 | 13 | 22 | 20 | 15 | 12 | ||
| % Peptide stimulated | 16.7 | 3.2 | 22.5 | 27.1 | 18.4 | 40.4 | 46.0 | 13.8 | 27.9 | 37.6 | 0.0 | 40.1 | 26.0 | 13.6 | 26.6 | ||
| % Specific lysis | 55.9 | ND | 56.0 | 65.5 | 41.8 | 71.0 | 77.8 | 51.3 | 77.7 | 52.6 | 58.0 | 45.8 | 37.1 | 13.1 | 42.3 | ||
| Control | P967 | % Fresh | 0.0 | 0.0 | 0.0 | 0.0 | 0.4 | 1.8 | 0.9 | 0.7 | ND | 0.6 | 0.9 | 1.9 | 1.4 | 3.3 | 1.7 |
| No. of cells/μl | 0 | 0 | 0 | 0 | 2 | 27 | 9 | 8 | ND | 5 | 8 | 10 | 10 | 31 | 17 | ||
| % Peptide stimulated | 0.2 | 0.2 | 0.3 | 0.2 | 45.7 | 37.5 | 60.2 | 63.9 | ND | 51.5 | 4.5 | 17.9 | 14.9 | 35.7 | 46.8 | ||
| % Specific lysis | 0.0 | 3.4 | 0.0 | 0.0 | 46.0 | 30.1 | 61.6 | 67.2 | ND | 48.4 | ND | ND | 26.5 | 39.6 | 40.4 | ||
| R468 | % Fresh | 0.0 | 0.0 | 0.0 | 0.0 | 0.6 | 3.3 | 1.3 | 0.6 | ND | 0.6 | 0.4 | 0.2 | 0.2 | 0.4 | —f | |
| No. of cells/μl | 0 | 0 | 0 | 0 | 11 | 103 | 23 | 11 | ND | 8 | 4 | 2 | 2 | 5 | — | ||
| % Peptide stimulated | 0.2 | 0.1 | 0.2 | 0.4 | 10.1 | 36.1 | 45.9 | 46.6 | ND | 18.9 | 3.7 | 15.7 | 6.3 | 14.3 | — | ||
| % Specific lysis | 0.0 | 2.5 | 2.5 | 0.0 | 36.4 | 49.9 | 70.1 | 80.4 | ND | 40.2 | ND | 62.0 | 33.5 | 56.0 | — | ||
| T720 | % Fresh | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.1 | 1.6 | 1.8 | 1.3 | 0.8 | 0.5 | 0.5 | 0.7 | 0.9 | 0.8 | |
| No. of cells/μl | 0 | 0 | 0 | 0 | 0 | 1 | 38 | 32 | 34 | 18 | 10 | 6 | 6 | 9 | 3 | ||
| % Peptide stimulated | 0.4 | 0.8 | 0.2 | 0.3 | 0.1 | 2.9 | 26.9 | 17.9 | 28.8 | 21.0 | 1.6 | 16.7 | 8.0 | 4.2 | 6.3 | ||
| % Specific lysis | 1.4 | ND | 4.7 | 0.0 | 2.1 | 14.6 | 67.1 | 54.4 | 77.0 | 48.4 | 15.0 | 39.0 | 23.5 | 1.5 | 7.7 | ||
| V299 | % Fresh | 0.0 | 0.0 | 0.0 | 0.1 | 0.0 | 0.3 | 1.2 | 1.0 | ND | 0.7 | 0.7 | 1.0 | 1.7 | 1.4 | 1.7 | |
| No. of cells/μl | 0 | 0 | 0 | 0 | 0 | 2 | 9 | 8 | ND | 2 | 4 | 5 | 14 | 10 | 13 | ||
| % Peptide stimulated | 1.7 | 0.3 | 4.8 | 0.5 | 2.6 | 52.6 | 41.6 | 38.9 | ND | 5.6 | 31.8 | 21.3 | 29.4 | 17.4 | 20.5 | ||
| % Specific lysis | 13.8 | ND | 7.2 | 0.7 | 4.2 | 72.8 | 74.6 | 62.0 | ND | ND | 47.6 | 52.0 | 61.0 | 54.3 | 55.4 | ||
| J8N | % Fresh | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.2 | 4.3 | 1.1 | 2.5 | 2.2 | 2.9 | 3.9 | 6.1 | 5.6 | ND | |
| No. of cells/μl | 0 | ND | 0 | 0 | 0 | 0 | 19 | 3 | 9 | 9 | 10 | 19 | 27 | 25 | ND | ||
| % Peptide stimulated | 0.6 | 0.5 | 0.3 | 0.5 | 0.4 | 0.7 | 4.0 | 9.3 | 4.7 | 2.5 | ND | 13.8 | 3.7 | 2.8 | 0.0 | ||
| % Specific lysis | 3.7 | ND | 4.7 | 0.5 | 5.6 | ND | ND | 24.6 | ND | 3.4 | 6.7 | 19.7 | 0.7 | 3.5 | 2.8 | ||
CD8αβ+PBL in whole blood were assessed for binding of the Mamu-A*01/p11C tetramer.
Total number of CD8αβ+ cells binding the Mamu-A*01/p11C tetramer per microliter of whole blood.
PBL were cultured in vitro with p11C (10 μg/ml) for 12 days and then assessed for binding of the Mamu-A*01/p11C tetramer to CD8αβ+ T cells.
PBL were cultured in vitro with p11C (10 μg/ml) for 12 days and then assessed for p11C-specific lysis using a standard 51Cr release assay. Data represent percent p11C-specific lysis at an effector-to-target ratio of 10:1.
ND, not done.
Animal was euthanized.
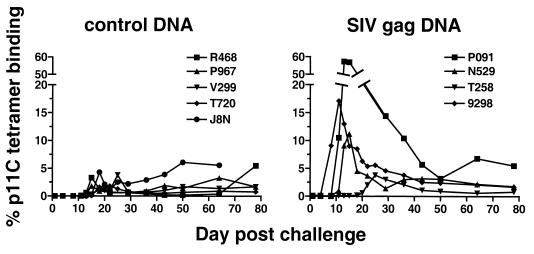
FIG. 6.
SIV Gag p11C tetramer binding CD8αβ+ T cells in whole blood after SIVsm E660 challenge in control DNA- and SIV gag DNA-immunized monkeys. Percent p11C tetramer binding represents Mamu-A*01/p11C tetramer-binding CD8αβ+ T cells in unstimulated whole blood samples at each time point tested.
The phenotypes of the Mamu-A*01/p11C tetramer-binding and nonbinding CD8αβ+ T cells in the vaccinated monkeys after virus challenge were analyzed by four-color flow cytometry. CD8αβ+ T cells were analyzed for binding of the Mamu-A*01/p11C 01 tetrameric complex and CD11a, CD28, CD45RA, and MHC class II DR expression (data not shown). The phenotypes of the Mamu-A*01/p11C tetramer-binding and nonbinding CD8αβ+ T cells were similar. Moreover, the phenotypic characteristics were similar to those of Mamu-A*01/p11C tetramer-binding and nonbinding CD8αβ+ T cells seen in monkeys chronically infected with SIVmac (15). The Mamu-A*01/p11C tetramer-binding T cells showed a high level of CD11a expression, a relatively low level of CD45RA expression, and heterogeneous CD28 and MHC class II expression.
DNA-vaccinated monkeys develop secondary CTL responses in lymph nodes after challenge.
Inguinal lymph nodes were obtained from the monkeys at the time of peak viral replication on day 15 postchallenge, as well as 28 and 42 days postchallenge. Lymphocytes from these lymph nodes were analyzed for secondary Gag epitope-specific CTL responses by tetramer staining and functional assays. At the time of peak virus replication during primary infection, as was seen in the peripheral blood, the SIV gag DNA-vaccinated animals developed higher levels of Mamu-A*01/p11C tetramer-binding CD8αβ+ T cells than did the control-vaccinated animals as measured in freshly isolated lymph nodes cells (Table 4). This difference between the experimentally and control-vaccinated groups of monkeys was not apparent in the Gag peptide-stimulated lymphocytes when assessed for tetramer binding or functional CTL activity. As was the case in the peripheral blood, the differences in the levels of SIV Gag epitope-specific CD8+ T-cell responses in the lymph nodes of the SIV gag DNA-vaccinated animals compared to the control DNA-vaccinated animals were not apparent in the specimens obtained 28 and 42 days after challenge.
TABLE 4.
Lymph node T lymphocytes from SIV gag plasmid DNA-vaccinated monkeys demonstrate secondary Gag epitope-specific CTL responses after SIVsm E660 challenge
| Immunization | Monkey | Assay | Value at indicated day postchallenge
|
||
|---|---|---|---|---|---|
| 14 | 28 | 42 | |||
| SIV gag | P091 | % Fresha | 17.5 | 7.3 | 2.6 |
| % Peptide stimulatedb | 28.0 | 20.2 | NDd | ||
| % Specific Lysisc | 55.0 | 64.9 | ND | ||
| N529 | % Fresh | 9.3 | 4.3 | 4.2 | |
| % Peptide stimulated | 12.3 | 4.2 | ND | ||
| % Specific lysis | 59.0 | 26.1 | ND | ||
| T258 | % Fresh | 0.1 | 0.6 | 1.1 | |
| % Peptide stimulated | 2.6 | 8.8 | 6.3 | ||
| % Specific lysis | 21.2 | 36.7 | 20.7 | ||
| 9298 | % Fresh | 6.5 | 4.4 | 4.1 | |
| % Peptide stimulated | 9.4 | 7.2 | 3.9 | ||
| % Specific lysis | 45.5 | 16.5 | 9.7 | ||
| Control | P967 | % Fresh | 0.8 | 0.5 | 0.8 |
| % Peptide stimulated | 50.6 | 17.1 | ND | ||
| % Specific lysis | 56.7 | 41.8 | ND | ||
| R468 | % Fresh | 2.2 | 0.4 | 0.2 | |
| % Peptide stimulated | 64.1 | 6.5 | ND | ||
| % Specific lysis | 65.8 | 27.5 | ND | ||
| T720 | % Fresh | 0.2 | 1.1 | 1.7 | |
| % Peptide stimulated | 0.8 | 10.8 | 7.3 | ||
| % Specific lysis | 0.0 | 25.1 | 14.4 | ||
| V299 | % Fresh | 0.7 | ND | ND | |
| % Peptide stimulated | 31.0 | ND | ND | ||
| % Specific lysis | 63.8 | ND | ND | ||
| J8N | % Fresh | 0.7 | 2.7 | 5.6 | |
| % Peptide stimulated | 3.3 | 8.6 | 2.3 | ||
| % Specific lysis | ND | ND | 0.3 | ||
Lymphocytes from freshly obtained lymph nodes were directly assessed for binding of the Mamu-A*01/p11C tetramer to CD8αβ+ T cells.
Lymph node cells were cultured in vitro with p11C peptide (10 μg/ml) for 12 days and then assessed for binding of the Mamu-A*01/p11C tetramer to CD8αβ+ T cells.
Lymph node cells were cultured in vitro with p11C peptide (10 μg/mL) for 12 days and then assessed for p11C-specific lysis using a standard 51Cr release assay. Data represent percent p11C-specific lysis at an effector-to-target ratio of 10:1.
ND, not done.
Secondary CTL responses do not alter the level of virus replication in lymph nodes immediately after challenge.
Inguinal lymph nodes harvested on days 15 and 42 after viral challenge were analyzed for viral RNA levels by in situ hybridization. As shown in Table 5, no differences were apparent between the gag DNA- and control DNA-vaccinated monkeys in the SIV RNA detected in these lymph nodes. Therefore, the potent secondary CTL responses seen in the SIV gag DNA-vaccinated animals failed to affect the level of SIV replication in lymph nodes during the first weeks following infection.
TABLE 5.
In situ hybridization studies of SIV RNA in lymph node sections from plasmid DNA-vaccinated monkeys after SIVsm E660 virus challenge
| Immunization | Monkey | No. of RNA-positive cells/mm3
|
|
|---|---|---|---|
| Day 15 | Day 42 | ||
| SIV gag | P091 | NDa | 2.7 |
| N529 | 3.9 | 0.3 | |
| T258 | 0.8 | 0.5 | |
| 9298 | 0.7 | 0.1 | |
| Control | R468 | 0.7 | 0.9 |
| P967 | ND | 0.0 | |
| T720 | 2.0 | 0.4 | |
| J8N | 0.2 | 0.5 | |
ND, not done.
Peripheral blood CD4+ T lymphocyte counts after challenge.
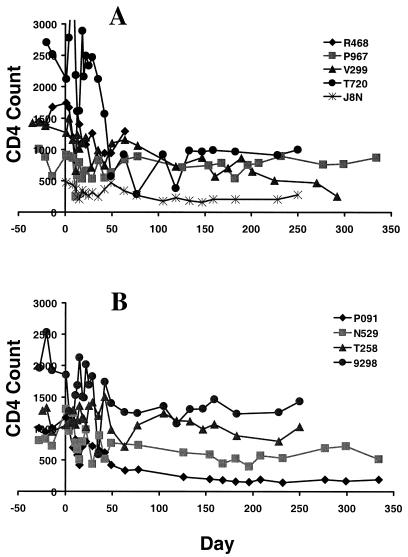
Clinical sequelae of infection were assessed in the weeks immediately after challenge by monitoring peripheral blood CD4+ T-lymphocyte counts in the SIV gag DNA- and control DNA-vaccinated monkeys (Fig. 7). By day 50 after challenge, two of the control (V299 and T720) and one of the experimentally vaccinated (P091) monkeys demonstrated a significant loss of circulating CD4+ T lymphocytes. Thus, there was no apparent effect of vaccination on CD4+ T-lymphocyte preservation in the early weeks following viral challenge. However, one control DNA-vaccinated animal (R468), the monkey with the highest level of virus replication, had to be euthanized on day 64 postchallenge with clinical signs of SIV disease (weight loss, lethargy, decreasing platelet counts, and anemia). All of the animals after day 50 have maintained their peripheral blood CD4+ T-lymphocyte counts and have remained healthy.
FIG. 7.
Peripheral blood CD4+ T-lymphocyte counts after intravenous SIVsm E660 challenge in Mamu-A*01+ rhesus monkeys immunized with the control (A) or the SIV gag (B) DNA vaccine.
SIV gag DNA-vaccinated monkeys have lower viral loads at set point after viral challenge than control DNA-vaccinated monkeys.
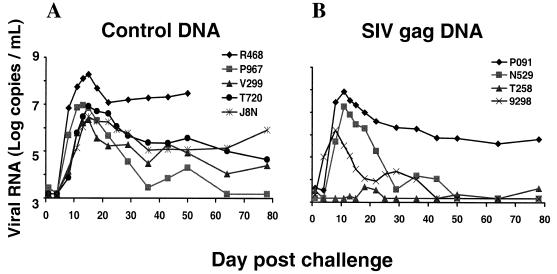
The best predictor of eventual clinical course in SIVsm E660-infected macaques is plasma viral load at set point, 8 weeks after infection (23). We therefore assessed plasma viral RNA levels through day 80 of infection in these challenged monkeys. The control DNA-vaccinated animals all had high levels of virus replication which peaked at day 15; these levels stabilized at set point, between 104.3 and 107.5 viral RNA copies per ml (Fig. 8). The SIV gag DNA-vaccinated animals had comparable levels of virus replication during primary infection, peaking at day 15. However, in contrast to the control DNA-vaccinated monkeys, the SIV gag DNA-vaccinated animals were better able to control the viral infection, with three of four vaccinated animals having undetectable plasma levels of virus (<1,500 viral RNA copies/ml) at day 50 postchallenge (Fig. 8). Day 50 plasma samples from the three SIV gag-vaccinated animals with undetectable levels of viral RNA were then reanalyzed using a more sensitive assay (41). Two of the three animals (T258 and 9298) maintained undetectable levels of virus (<300 viral RNA copies/ml). The other animal (N529) showed a low level of viral replication (6,000 viral RNA copies/ml).
FIG. 8.
Plasma viral load analysis after intravenous SIVsm E660 challenge in Mamu-A*01+ rhesus monkeys immunized with the control (A) or the SIV gag (B) DNA vaccine.
DISCUSSION
While a number of recent studies in nonhuman primates have suggested that plasmid DNA immunization may prove useful as a component of an HIV vaccine, caveats associated with each of these studies have made it difficult to generalize from these findings with any sense of certainty. Although plasmid DNA immunization protected chimpanzees from infection with HIV-1 (5), the poor replicative ability and lack of pathogenicity in chimpanzees of the SF2 challenge isolate used in that study have raised questions as to the significance of that observation. We have previously reported that plasmid DNA priming and recombinant envelope protein boosting protected macaques from infection with a chimeric SIV/HIV (SHIV) challenge (22). However, the lack of pathogenicity in macaques of the SHIV isolate used in that study makes it difficult to conclude anything from that observation concerning the eventual utility of such a vaccine approach in humans. Finally, a DNA priming-recombinant poxvirus boosting approach has recently shown some protective efficacy in macaques against a pathogenic SHIV challenge (33). In this study, however, a prior nonpathogenic SHIV challenge in the same monkeys obscures the contribution of the immunizations on the protection that was seen. The present study was designed to assess gag DNA immunization alone in eliciting Gag-specific CTL and protection against a highly pathogenic SIV challenge.
It has been difficult to generalize from HIV vaccine studies in nonhuman primates not only because of the diversity of immunization regimens employed but also because of differences in the challenge viruses used. In the various studies that have been reported, the plasmid DNA constructs used have differed in construction and in the methods by which they have been delivered as immunogens. It is clear now that the immunogenicity of a plasmid DNA vaccine in a higher primate is tied to the promoters that are employed in their construction, whether or not codon optimization has been used in their construction, the quality and quantity of DNA used in each inoculation, the route by which the plasmid DNA is administered, and how the plasmid is formulated. The results of the present experiments indicate that when at least some of these variables are optimized, a plasmid DNA vaccine can elicit a CTL response of high enough frequency to be detected in unstimulated whole blood using the soluble MHC class I tetramer technology.
The central goal of this study was to assess CTL immediately following virus challenge to determine whether the CTL that were generated following infection arose earlier and reached a higher frequency in the experimentally vaccinated monkeys than in those animals that received the control vaccinations. It has been well documented that immunization with a number of effective vaccines does not elicit sterilizing immunity. Rather, vaccine-elicited immunity facilitates the generation of secondary immune responses that contain the spread of replicating virus during the first days following virus infection, limiting the pathogenicity of the virus. Priming for the induction of such a secondary immune response is certainly a realistic goal for a plasmid DNA immunogen. In fact, the present study clearly indicates that high-frequency secondary CTL responses are observed in the gag DNA-vaccinated monkeys following infection with SIV. Moreover, the statistically significant containment of SIV replication at the time of set point in the gag DNA-vaccinated monkeys suggests that those secondary CTL responses may play a useful role in containing SIV replication.
The present study includes a clinical evaluation of the SIV-challenged monkeys for more than 300 days following infection. It has been clearly demonstrated that viral set point postinfection in SIVsm E660-infected macaques predicts the clinical outcome of the infection (23). Thus a higher virus load by 8 weeks postinfection likely indicates a rapid clinical course of disease. Therefore, there is reason to suppose that the gag DNA-vaccinated monkeys will have a more benign clinical course than the control DNA-vaccinated monkeys. The control DNA-vaccinated monkey (R468) with the highest level of virus replication had to be euthanized on day 64 postchallenge with clinical signs of SIV disease. Interestingly, the one SIV gag DNA-vaccinated monkey that failed to control virus replication (P091), despite a measurable decline in CD4 counts, has remained healthy.
The use of the soluble MHC class I tetramer technology for monitoring vaccine-elicited CTL in these monkeys is of enormous value. This technical approach allows us to quantitate with ease and precision Gag epitope-specific CTL in lymph nodes and peripheral blood without relying on in vitro stimulation of the antigen-specific lymphocyte populations. It also allows us to compare the relative efficacy of DNA vaccination with that of other previously evaluated vaccine technologies. Interestingly, the immunongenicity of the gag DNA plasmid vaccine construct used in the present study does not appear to be quite as great as the immunogenicity of a recombinant MVA construct in rhesus monkeys (35). It will be important to assess the relative protection afforded by these different immunogens against SIV challenge. It is, however, likely that the immune responses elicited by this DNA immunogen might be augmented by formulating the DNA with a variety of novel adjuvants or delivering the DNA with cytokines. Finally, there is also reason to assume that combining DNA immunogens with live vectors may also further augment the vaccine-elicited CTL responses.
ACKNOWLEDGMENTS
This work was supported by National Institutes of Health grants CA50139 and AI85343.
REFERENCES
- 1.Allen T M, Sidney J, del Guercio M F, Glickman R L, Lensmeyer G L, Wiebe D A, DeMars R, Pauza C D, Johnson R P, Sette A, Watkins D I. Characterization of the peptide binding motif of a rhesus MHC class I molecule (Mamu-A*01) that binds an immunodominant CTL epitope from simian immunodeficiency virus. J Immunol. 1998;160:6062–6071. [PubMed] [Google Scholar]
- 2.Barouch D H, Santra S, Steenbeke T D, Zheng X X, Perry H C, Davies M E, Freed D C, Craiu A, Strom T B, Shiver J W, Letvin N L. Augmentation and suppression of immune responses to an HIV-1 DNA vaccine by plasmid cytokine/Ig administration. J Immunol. 1998;161:1875–1882. [PubMed] [Google Scholar]
- 3.Belyakov I M, Derby M A, Ahlers J D, Kelsall B L, Earl P, Moss B, Strober W, Berzofsky J A. Mucosal immunization with HIV-1 peptide vaccine induces mucosal and systemic cytotoxic T lymphocytes and protective immunity in mice against intrarectal recombinant HIV-vaccinia challenge. Proc Natl Acad Sci USA. 1998;95:1709–1714. doi: 10.1073/pnas.95.4.1709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Boyer J D, Chattergoon M, Shah A, Ginsberg R, MacGregor R R, Weiner D B. HIV-1 DNA based vaccine induces a CD8 mediated cross-clade CTL response. Dev Biol Stand. 1998;95:147–153. [PubMed] [Google Scholar]
- 5.Boyer J D, Ugen K E, Wang B, Agadjanyan M, Gilbert L, Bagarazzi M L, Chattergoon M, Frost P, Javadian A, Williams W V, Refaeli Y, Ciccarelli R B, McCallus D, Coney L, Weiner D B. Protection of chimpanzees from high-dose heterologous HIV-1 challenge by DNA vaccination. Nat Med. 1997;3:526–532. doi: 10.1038/nm0597-526. [DOI] [PubMed] [Google Scholar]
- 6.Boyer J D, Wang B, Ugen K E, Agadjanyan M, Javadian A, Frost P, Dang K, Carrano R A, Ciccarelli R, Coney L, Williams W V, Weiner D B. In vivo protective anti-HIV immune responses in non-human primates through DNA immunization. J Med Primatol. 1996;25:242–250. doi: 10.1111/j.1600-0684.1996.tb00022.x. [DOI] [PubMed] [Google Scholar]
- 6a.Committee on Care and Use of Laboratory Animals. Guide for the care and use of laboratory animals. Washington, D.C.: Institute of Laboratory Animal Resources, National Research Council; 1996. [Google Scholar]
- 7.Donnelly J J, Ulmer J B, Shiver J W, Liu M A. DNA vaccines. Annu Rev Immunol. 1997;15:617–648. doi: 10.1146/annurev.immunol.15.1.617. [DOI] [PubMed] [Google Scholar]
- 8.Egan M A, Kuroda M J, Voss G, Schmitz J E, Charini W A, Lord C I, Forman M A, Letvin N L. Use of major histocompatibility complex class I/peptide/β2M tetramers to quantitate CD8+ cytotoxic T lymphocytes specific for dominant and nondominant viral epitopes in simian-human immunodeficiency virus-infected rhesus monkeys. J Virol. 1999;73:5466–5472. doi: 10.1128/jvi.73.7.5466-5472.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gorse G J, Keefer M C, Belshe R B, Matthews T J, Forrest B D, Hsieh R H, Koff W C, Hanson C V, Dolin R, Weinhold K J, Frey S E, Ketter N, Fast P E. A dose-ranging study of a prototype synthetic HIV-1MN V3 branched peptide vaccine. The National Institute of Allergy and Infectious Diseases AIDS Vaccine Evaluation Group. J Infect Dis. 1996;173:330–339. doi: 10.1093/infdis/173.2.330. [DOI] [PubMed] [Google Scholar]
- 10.Hirsch V M, Fuerst T R, Sutter G, Carroll M W, Yang L C, Goldstein S, Piatak M, Jr, Elkins W R, Alvord W G, Montefiori D C, Moss B, Lifson J D. Patterns of viral replication correlate with outcome in simian immunodeficiency virus (SIV)-infected macaques: effect of prior immunization with a trivalent SIV vaccine in modified vaccinia virus Ankara. J Virol. 1996;70:3741–3752. doi: 10.1128/jvi.70.6.3741-3752.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hu S L, Abrams K, Barber G N, Moran P, Zarling J M, Langlois A J, Kuller L, Morton W R, Benveniste R E. Protection of macaques against SIV infection by subunit vaccines of SIV envelope glycoprotein gp160. Science. 1992;255:456–459. doi: 10.1126/science.1531159. [DOI] [PubMed] [Google Scholar]
- 12.Hu S L, Fultz P N, McClure H M, Eichberg J W, Thomas E K, Zarling J, Singhal M C, Kosowski S G, Swenson R B, Anderson D C, et al. Effect of immunization with a vaccinia-HIV env recombinant on HIV infection of chimpanzees. Nature. 1987;328:721–723. doi: 10.1038/328721a0. [DOI] [PubMed] [Google Scholar]
- 13.Kelleher A D, Emery S, Cunningham P, Duncombe C, Carr A, Golding H, Forde S, Hudson J, Roggensack M, Forrest B D, Cooper D A. Safety and immunogenicity of UBI HIV-1MN octameric V3 peptide vaccine administered by subcutaneous injection. AIDS Res Hum Retroviruses. 1997;13:29–32. doi: 10.1089/aid.1997.13.29. [DOI] [PubMed] [Google Scholar]
- 14.Knapp L A, Lehmann E, Piekarczyk M S, Urvater J A, Watkins D I. A high frequency of Mamu-A*01 in the rhesus macaque detected by polymerase chain reaction with sequence-specific primers and direct sequencing. Tissue Antigens. 1997;50:657–661. doi: 10.1111/j.1399-0039.1997.tb02927.x. [DOI] [PubMed] [Google Scholar]
- 15.Kuroda M J, Schmitz J E, Barouch D H, Craiu A, Allen T M, Sette A, Watkins D I, Forman M A, Letvin N L. Analysis of Gag-specific cytotoxic T lymphocytes in simian immunodeficiency virus-infected rhesus monkeys by cell staining with a tetrameric major histocompatibility complex class I-peptide complex. J Exp Med. 1998;187:1373–1381. doi: 10.1084/jem.187.9.1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kuroda M J, Schmitz J E, Charini W A, Nickerson C E, Lifton M A, Lord C I, Forman M A, Letvin N L. Emergence of CTL coincides with clearance of virus during primary simian immunodeficiency virus infection in rhesus monkeys. J Immunol. 1999;162:5127–5133. [PubMed] [Google Scholar]
- 17.Kuroda M J, Schmitz J E, Charini W A, Nickerson C E, Lord C I, Forman M A, Letvin N L. Comparative analysis of cytotoxic T lymphocytes in lymph nodes and peripheral blood of simian immunodeficiency virus-infected rhesus monkeys. J Virol. 1999;73:1573–1579. doi: 10.1128/jvi.73.2.1573-1579.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lathe R. Synthetic oligonucleotide probes deduced from amino acid sequence data. Theoretical and practical considerations. J Mol Biol. 1985;183:1–12. doi: 10.1016/0022-2836(85)90276-1. [DOI] [PubMed] [Google Scholar]
- 19.Lekutis C, Shiver J W, Liu M A, Letvin N L. HIV-1 env DNA vaccine administered to rhesus monkeys elicits MHC class II-restricted CD4+ T helper cells that secrete IFN-gamma and TNF-alpha. J Immunol. 1997;158:4471–4477. [PubMed] [Google Scholar]
- 20.Letvin N L. Animal models for the study of human immunodeficiency virus infections. Curr Opin Immunol. 1992;4:481–485. doi: 10.1016/s0952-7915(06)80043-9. [DOI] [PubMed] [Google Scholar]
- 21.Letvin N L. Progress in the development of an HIV-1 vaccine. Science. 1998;280:1875–1880. doi: 10.1126/science.280.5371.1875. [DOI] [PubMed] [Google Scholar]
- 22.Letvin N L, Montefiori D C, Yasutomi Y, Perry H C, Davies M E, Lekutis C, Alroy M, Freed D C, Lord C I, Handt L K, Liu M A, Shiver J W. Potent, protective anti-HIV immune responses generated by bimodal HIV envelope DNA plus protein vaccination. Proc Natl Acad Sci USA. 1997;94:9378–9383. doi: 10.1073/pnas.94.17.9378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lifson J D, Nowak M A, Goldstein S, Rossio J L, Kinter A, Vasquez G, Wiltrout T A, Brown C, Schneider D, Wahl L, Lloyd A L, Williams J, Elkins W R, Fauci A S, Hirsch V M. The extent of early viral replication is a critical determinant of the natural history of simian immunodeficiency virus infection. J Virol. 1997;71:9508–9514. doi: 10.1128/jvi.71.12.9508-9514.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Liu M A, Fu T M, Donnelly J J, Caulfield M J, Ulmer J B. DNA vaccines. Mechanisms for generation of immune responses. Adv Exp Med Biol. 1998;452:187–191. [PubMed] [Google Scholar]
- 25.Liu M A, Yasutomi Y, Davies M E, Perry H C, Freed D C, Letvin N L, Shiver J W. Vaccination of mice and nonhuman primates using HIV-gene-containing DNA. Antibiot Chemother. 1996;48:100–104. doi: 10.1159/000425163. [DOI] [PubMed] [Google Scholar]
- 26.Lu S, Arthos J, Montefiori D C, Yasutomi Y, Manson K, Mustafa F, Johnson E, Santoro J C, Wissink J, Mullins J I, Haynes J R, Letvin N L, Wyand M, Robinson H L. Simian immunodeficiency virus DNA vaccine trial in macaques. J Virol. 1996;70:3978–3991. doi: 10.1128/jvi.70.6.3978-3991.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.McDonnell W M, Askari F K. DNA vaccines. N Engl J Med. 1996;334:42–45. doi: 10.1056/NEJM199601043340110. [DOI] [PubMed] [Google Scholar]
- 28.Miller M D, Gould-Fogerite S, Shen L, Woods R M, Koenig S, Mannino R J, Letvin N L. Vaccination of rhesus monkeys with synthetic peptide in a fusogenic proteoliposome elicits simian immunodeficiency virus-specific CD8+ cytotoxic T lymphocytes. J Exp Med. 1992;176:1739–1744. doi: 10.1084/jem.176.6.1739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Miller M D, Yamamoto H, Hughes A L, Watkins D I, Letvin N L. Definition of an epitope and MHC class I molecule recognized by gag-specific cytotoxic T lymphocytes in SIVmac-infected rhesus monkeys. J Immunol. 1991;147:320–329. [PubMed] [Google Scholar]
- 30.Montgomery D L, Shiver J W, Leander K R, Perry H C, Friedman A, Martinez D, Ulmer J B, Donnelly J J, Liu M A. Heterologous and homologous protection against influenza A by DNA vaccination: optimization of DNA vectors. DNA Cell Biol. 1993;12:777–783. doi: 10.1089/dna.1993.12.777. [DOI] [PubMed] [Google Scholar]
- 31.Montgomery D L, Ulmer J B, Donnelly J J, Liu M A. DNA vaccines. Pharmacol Ther. 1997;74:195–205. doi: 10.1016/s0163-7258(97)82003-7. [DOI] [PubMed] [Google Scholar]
- 32.Robinson H L, Lu S, Feltquate D M, Torres C T, Richmond J, Boyle C M, Morin M J, Santoro J C, Webster R G, Montefiori D, Yasutomi Y, Letvin N L, Manson K, Wyand M, Haynes J R. DNA vaccines. AIDS Res Hum Retroviruses. 1996;12:455–457. doi: 10.1089/aid.1996.12.455. [DOI] [PubMed] [Google Scholar]
- 33.Robinson H L, Montefiori D C, Johnson R P, Manson K H, Kalish M L, Lifson J D, Rizvi T A, Lu S, Hu S L, Mazzara G P, Panicali D L, Herndon J G, Glickman R, Candido M A, Lydy S L, Wyand M S, McClure H M. Neutralizing antibody-independent containment of immunodeficiency virus challenges by DNA priming and recombinant pox virus booster immunizations. Nat Med. 1999;5:526–534. doi: 10.1038/8406. [DOI] [PubMed] [Google Scholar]
- 34.Sauzet J P, Moog C, Krivine A, Martinon F, Bossus M, Gras-Masse H, Tartar A, Guillet J G, Gomard E. Adjuvant is required when using Env lipopeptide construct to induce HIV type 1-specific neutralizing antibody responses in mice in vivo. AIDS Res Hum Retroviruses. 1998;14:901–909. doi: 10.1089/aid.1998.14.901. [DOI] [PubMed] [Google Scholar]
- 35.Seth A, Ourmanov I, Kuroda M J, Schmitz J E, Carroll M W, Wyatt L S, Moss B, Forman M A, Hirsch V M, Letvin N L. Recombinant modified vaccinia virus Ankara-simian immunodeficiency virus gag pol elicits cytotoxic T lymphocytes in rhesus monkeys detected by a major histocompatibility complex class I/peptide tetramer. Proc Natl Acad Sci USA. 1998;95:10112–10116. doi: 10.1073/pnas.95.17.10112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Shen L, Chen Z W, Miller M D, Stallard V, Mazzara G P, Panicali D L, Letvin N L. Recombinant virus vaccine-induced SIV-specific CD8+ cytotoxic T lymphocytes. Science. 1991;252:440–443. doi: 10.1126/science.1708168. [DOI] [PubMed] [Google Scholar]
- 37.Shiver J W, Davies M E, Perry H C, Freed D C, Liu M A. Humoral and cellular immunities elicited by HIV-1 vaccination. J Pharm Sci. 1996;85:1317–1324. doi: 10.1021/js9600991. [DOI] [PubMed] [Google Scholar]
- 38.Shiver J W, Davies M E, Yasutomi Y, Perry H C, Freed D C, Letvin N L, Liu M A. Anti-HIV env immunities elicited by nucleic acid vaccines. Vaccine. 1997;15:884–887. doi: 10.1016/s0264-410x(96)00251-4. [DOI] [PubMed] [Google Scholar]
- 39.Shiver J W, Perry H C, Davies M E, Freed D C, Liu M A. Cytotoxic T lymphocyte and helper T cell responses following HIV polynucleotide vaccination. Ann N Y Acad Sci. 1995;772:198–208. doi: 10.1111/j.1749-6632.1995.tb44745.x. [DOI] [PubMed] [Google Scholar]
- 40.Staats H F, Nichols W G, Palker T J. Mucosal immunity to HIV-1: systemic and vaginal antibody responses after intranasal immunization with the HIV-1 C4/V3 peptide T1SP10 MN(A) J Immunol. 1996;157:462–472. [PubMed] [Google Scholar]
- 41.Suryanarayana K, Wiltrout T A, Vasquez G M, Hirsch V M, Lifson J D. Plasma SIV RNA viral load determination by real-time quantification of product generation in reverse transcriptase-polymerase chain reaction. AIDS Res Hum Retroviruses. 1998;14:183–189. doi: 10.1089/aid.1998.14.183. [DOI] [PubMed] [Google Scholar]
- 42.Tenner-Racz K, Stellbrink H J, van Lunzen J, Schneider C, Jacobs J P, Raschdorff B, Grosschupff G, Steinman R M, Racz P. The unenlarged lymph nodes of HIV-1-infected, asymptomatic patients with high CD4 T cell counts are sites for virus replication and CD4 T cell proliferation. The impact of highly active antiretroviral therapy. J Exp Med. 1998;187:949–959. doi: 10.1084/jem.187.6.949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ulmer J B, Deck R R, Yawman A, Friedman A, Dewitt C, Martinez D, Montgomery D L, Donnelly J J, Liu M A. DNA vaccines for bacteria and viruses. Adv Exp Med Biol. 1996;397:49–53. doi: 10.1007/978-1-4899-1382-1_7. [DOI] [PubMed] [Google Scholar]
- 44.Ulmer J B, Donnelly J J, Liu M A. Toward the development of DNA vaccines. Curr Opin Biotechnol. 1996;7:653–658. doi: 10.1016/s0958-1669(96)80078-4. [DOI] [PubMed] [Google Scholar]
- 45.Wang B, Boyer J, Srikantan V, Coney L, Carrano R, Phan C, Merva M, Dang K, Agadjanan M, Gilbert L, et al. DNA inoculation induces neutralizing immune responses against human immunodeficiency virus type 1 in mice and nonhuman primates. DNA Cell Biol. 1993;12:799–805. doi: 10.1089/dna.1993.12.799. [DOI] [PubMed] [Google Scholar]
- 46.Wang B, Boyer J, Srikantan V, Ugen K, Gilbert L, Phan C, Dang K, Merva M, Agadjanyan M G, Newman M, et al. Induction of humoral and cellular immune responses to the human immunodeficiency virus type 1 virus in nonhuman primates by in vivo DNA inoculation. Virology. 1995;211:102–112. doi: 10.1006/viro.1995.1383. [DOI] [PubMed] [Google Scholar]
- 47.Wang B, Ugen K E, Srikantan V, Agadjanyan M G, Dang K, Refaeli Y, Sato A I, Boyer J, Williams W V, Weiner D B. Gene inoculation generates immune responses against human immunodeficiency virus type 1. Proc Natl Acad Sci USA. 1993;90:4156–4160. doi: 10.1073/pnas.90.9.4156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Yasutomi Y, Robinson H L, Lu S, Mustafa F, Lekutis C, Arthos J, Mullins J I, Voss G, Manson K, Wyand M, Letvin N L. Simian immunodeficiency virus-specific cytotoxic T-lymphocyte induction through DNA vaccination of rhesus monkeys. J Virol. 1996;70:678–681. doi: 10.1128/jvi.70.1.678-681.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]