ABSTRACT
A comprehensive pangenomic approach was employed to analyze the genomes of 75 type II methylotrophs spanning various genera. Our investigation revealed 256 exact core gene families shared by all 75 organisms, emphasizing their crucial role in the survival and adaptability of these organisms. Additionally, we predicted the functionality of 12 hypothetical proteins. The analysis unveiled a diverse array of genes associated with key metabolic pathways, including methane, serine, glyoxylate, and ethylmalonyl-CoA (EMC) metabolic pathways. While all selected organisms possessed essential genes for the serine pathway, Methylooceanibacter marginalis lacked serine hydroxymethyltransferase (SHMT), and Methylobacterium variabile exhibited both isozymes of SHMT, suggesting its potential to utilize a broader range of carbon sources. Notably, Methylobrevis sp. displayed a unique serine-glyoxylate transaminase isozyme not found in other organisms. Only nine organisms featured anaplerotic enzymes (isocitrate lyase and malate synthase) for the glyoxylate pathway, with the rest following the EMC pathway. Methylovirgula sp. 4MZ18 stood out by acquiring genes from both glyoxylate and EMC pathways, and Methylocapsa sp. S129 featured an A-form malate synthase, unlike the G-form found in the remaining organisms. Our findings also revealed distinct phylogenetic relationships and clustering patterns among type II methylotrophs, leading to the proposal of a separate genus for Methylovirgula sp. 4M-Z18 and Methylocapsa sp. S129. This pangenomic study unveils remarkable metabolic diversity, unique gene characteristics, and distinct clustering patterns of type II methylotrophs, providing valuable insights for future carbon sequestration and biotechnological applications.
IMPORTANCE
Methylotrophs have played a significant role in methane-based product production for many years. However, a comprehensive investigation into the diverse genetic architectures across different genera of methylotrophs has been lacking. This study fills this knowledge gap by enhancing our understanding of core hypothetical proteins and unique enzymes involved in methane oxidation, serine, glyoxylate, and ethylmalonyl-CoA pathways. These findings provide a valuable reference for researchers working with other methylotrophic species. Furthermore, this study not only unveils distinctive gene characteristics and phylogenetic relationships but also suggests a reclassification for Methylovirgula sp. 4M-Z18 and Methylocapsa sp. S129 into separate genera due to their unique attributes within their respective genus. Leveraging the synergies among various methylotrophic organisms, the scientific community can potentially optimize metabolite production, increasing the yield of desired end products and overall productivity.
KEYWORDS: hypothetical proteins, isozymes, methane, persistent, PPanGOLLiN, serine pathway
INTRODUCTION
The field of genomics has predominantly focused on utilizing reference genomes as guiding maps, offering insights into the genetic makeup of a “typical” individual within each species (1). However, a single reference genome imposes limitations and fails to capture the full extent of genetic variation within a species. To address this issue, researchers have begun creating and utilizing pangenomes, which represent collections of all genomic DNA sequences identified across individuals within a species (2). The increase in the number of sequenced genomes has led to the recognition of graph-based pangenomes as a platform for studying diversity in a population or species, ranging from point mutations to large chromosomal rearrangements (3). Pangenomes can be represented as directed graphs, capturing structural and single variants (4). Despite their benefits, incorporating graph genomes into research practice is challenging due to the need for new tools, data structures, and formats as well as the difficulty in integrating them with existing software and databases (5). However, pangenomes have enabled notable discoveries that would have been difficult or impossible with traditional reference genomes and are evident from the identification of 51% gene families from the dispensable genomes of Glycine soja in 2014 (6).
Methanotrophs, a subset of methylotrophs, are crucial in regulating the global carbon cycle by converting methane into carbon dioxide (7). They are classified into three categories: type I and type X (γ-proteobacteria), type II (α-proteobacteria), and Verrucomicrobia (8). Type I and type II methanotrophs have distinct metabolic capabilities owing to differences in their intracellular structures, carbon assimilation pathways, and other metabolic features (9). Unlike type I methanotrophs, type II lacks intracytoplasmic membranes, which house methane monooxygenases (MMOs) (10). As a result, type II methanotrophs have cytoplasmic and transmembrane MMOs (11). Additionally, type I methanotrophs utilize the ribulose monophosphate pathway for carbon assimilation, while type II methanotrophs utilize the serine pathway (10, 12). This pathway enables type II methanotrophs to co-incorporate methane and CO2, giving them more versatility in carbon assimilation (13). Moreover, type II methylotrophs have the ability to fix atmospheric nitrogen, making them suitable for growth in nitrogen-limited environments (14). They also have a high acetyl-CoA flux, making them a potential microbial cell-factory platform for methane-derived biomanufacturing (13). Furthermore, through the application of pangenomic analysis, Oshkin et al. were able to distinguish two closely related type II methanotrophic genera, Methylosinus and Methylocystis. Their study revealed a diverse range of enzymes involved in methane oxidation and dinitrogen fixation, as well as genomic determinants for cell motility and photosynthesis within the accessory genome of these methanotrophic bacteria (15). Overall, the distinct metabolic features of type II methylotrophs make them a fascinating subject of study and a potential tool for methane biomanufacturing.
The primary objective of this study is to comprehensively explore the genetic diversity and evolutionary patterns of type II methylotrophs (Fig. 1). To achieve this, we employed a cutting-edge pangenomic approach to analyze the complete genetic content of type II methylotrophs in various environments. Through this comprehensive analysis, we aim to identify the unique genetic traits and pathways that underlie the evolutionary processes of type II methylotrophs. The findings of this study will have significant implications for developing biotechnological applications and strategies for mitigating methane emissions, thereby providing a valuable contribution to the field of environmental microbiology.
Fig 1.
Schematic overview of the study determined to analyze the genetic pattern in type II methylotrophs using pangenome.
MATERIALS AND METHODS
Genome mining and selection
The genera and species of type II methylotrophs available in the National Center for Biotechnology Information (NCBI) Refseq and GenBank assembly database were compiled into a Microsoft Excel (v365) file (16, 17). The genomic fasta (.fna) files of these organisms were obtained from the NCBI Refseq and GenBank using an automated Python program that utilized file transfer protocol. The completeness of the selected genomes was assessed using Benchmarking Universal Single-Copy Orthologs (BUSCO) analysis, which calculated the percentage of completeness (C), fragmented (F), and missing genes (M) in each genome (18). Assembly statistics such as scaffold N50 and L50, contig N50 and L50, coverage, total gap length, and spanned gaps were also collected for each genome to ensure quality. Genome assemblies with at least 94% completeness, less than 1,000 total gap length, and less than 5 spanned gaps were selected for pangenome analysis.
Genome annotation
To remove the biasness and variations in gene prediction and annotations, all the genomes used in pangenome analysis were annotated using Prokka v1.8 with ab initio algorithm (Prodigal v2.6.3) (19). To ensure accurate gene annotations, genus-specific parameters were employed. This approach allows the annotation algorithm to consider the same gene and codon patterns identified within the particular genus under study. Prokka employs an integration pipeline that incorporates various databases to enhance the annotation process. The databases used BLASTp, UniProt, RefSeq, Rfam, and TIGRFAMs, which provide valuable information for the identification and characterization of proteins. SignalP is used to identify signal peptides, aiding in the prediction of protein localization and its function. The prediction of transfer RNA (tRNA) and ribosomal RNA (rRNA) was performed using Aragorn (v1.2.41) and Barrnap v0.9, respectively. To identify CRISPR sequences, the software minced v.4.2 was utilized.
Pangenome analysis
The pangenome analysis was performed using graph-based analysis tool PPanGGOLiN (20). PPanGGOLiN is a statistical and graphical model-based tool that can construct pangenomes for large sets of prokaryotic genomes. The input files consisted of PROKKA-annotated genomic FASTA (.fna) and GTF files, with their coding regions classified into homologous gene families. The PPanGGOLiN pipeline was uploaded onto Google Colab for pangenome analysis. The tool utilizes information on protein-coding genes and their genomic context to build a graph, where each node represents a gene family, and each edge indicates genetic contiguity between families that are neighbors in the genomes. This approach has the advantage of being able to handle fragmented genome assemblies and maintain linkages between gene families even when genome assembly gaps exist. Orthologous gene clustering is performed using a sequence similarity cutoff of 50% and sequence coverage cutoff of 80%. This ensures that genes with sufficient similarity and coverage are clustered together as orthologs. The output file contained the corresponding gene families from the pangenome, along with partition and quality parameters, such as percent identity (pident), expectation value (e-value), and bit score for each input gene. PPanGGOLiN classified the genes into three categories: persistent, cloud, and shell partitions. The persistent gene families refer to the conserved gene families present across the entire bacterial genomes under study. The genes present in an intermediate number of genomes are labeled as shell, whereas the unique genes specific to a particular species are termed as cloud genes (21, 22).
Physical protein-protein interaction and enrichment
To analyze the physical protein-protein interactions (PPIs) and subsequent gene ontology (GO) interactions between the persistent genes, the STRING (Search Tool for Interacting Genes Retrieval) database was utilized, which is a precomputed global resource for predicting functional associations between proteins (23). To enrich the PPI network, the persistent gene set was taken as input into the STRING database. The enriched genes with a P-value less than 1.6 × 10−16 were visualized using Cytoscape (version 3.9.1), a PPI visualization software (24). The Cytoscape plugin, StringApp, was used to perform pathway enrichment analysis and import PPI networks from the STRING database to Cytoscape. Methylosinus trichosporium OB3b was selected as the organism, and a confidence score cutoff of 0.40 was used to find the potential interactions between the genes. To unveil the relationships between the GO terms related to the input gene set, enrichment interactions were performed (25). The most enriched GO terms were screened based on the false discovery rate, which was less than 1.0 × 10−6. In the GO-enrichment network, the nodes corresponded to the GO terms, and the edges depicted the interactions. A heatmap was generated to show the relevance of the enriched GO terms (each node) in a particular molecular function (MF)/biological process (BP)/cellular component (CC).
The pangenome graph file generated by the PPanGGOLiN pipeline was visualized using Gephi software (version 0.9.2), enabling graph visualization and manipulation (26). The nodes and edges representing the exact core, which are shared by all the organisms, were selected and filtered. Statistical analysis was performed, encompassing metrics such as average degree, average weighted degree, network diameter, modularity, average clustering coefficient, and average path length. These analyses provide insights into the structure and characteristics of the pangenome network. To enhance visual comprehension, the nodes were ranked based on the number of sequences, and a color ranking was assigned to represent the modularity class. Initially, the “Noverlap” layout was employed for arranging the nodes, followed by manual grouping of nodes based on their biological functions. Functional categories for gene families were obtained from the UniProt database (27). These categories aid in understanding the functional diversity within the pangenome network. The final network image was downloaded as a vector image with default presets and subsequently edited using Inkscape software (version 1.2). The editing process involved marking the node clusters with their respective functional categories, thereby improving the clarity and interpretability of the network visualization.
Gene ontology
The output generated by PPanGGOLiN resulted in a comprehensive matrix file containing gene families and their associated gene IDs specific to type II methylotrophs. Obtaining UniProtKB information for the exact core data was crucial; however, each gene family had multiple UniProtKB matches (28). Since each gene family represented a cluster of genes from at least the specified number of organisms, or more, it was expected to have multiple ID matches. Therefore, mapping the exact core gene families with gene ID data was necessary to obtain the complete gene list. The original genome annotation file in .gff format, obtained from Prokka, included the UniProtKB ID for each gene. To extract the UniProtKB information for each gene ID, a Python script was utilized, which extracted the relevant data from the Prokka annotation file. The extracted information was then consolidated into a single Excel (.xlsx) file. This resulting output, containing the UniProtKB IDs for the entire exact core gene list, was subsequently fed into the “Retrieve/ID mapping” tool provided by UniProt. This tool facilitated the retrieval of gene ontology information corresponding to the UniProtKB IDs, enriching the data set with valuable functional annotations associated with the genes. The GO terms associated with the gene families were retrieved and grouped, enabling the determination of the total gene family count and gene count for each specific GO category. To assign scores to each GO term, the percentage of genes within that category was calculated in relation to the total mapped genes. To visualize the distribution of GO terms across different categories such as BP, MF, and CC, plots were created using ggplot2 in Python v3.10.8.
KEGG pathway analysis
Pathway analysis and functional categorization of gene families in this study were performed using the KEGG Mapper search tool (29). The Enzyme Commission numbers were used as references to search pathway IDs for the input enzyme data. To achieve functional categorization of gene families in the shell and cloud partitions, KEGG BlastKOALA (KEGG Orthology and Links Annotation) and GhostKOALA were utilized, respectively (30). The FASTA amino acid sequence file for both partitions was uploaded to the KEGG server, with the “genus_prokaryotes” and KEGG genes database selected for the search.
Putative annotation to hypothetical proteins
The online webserver MOTIFSearch (https://www.genome.jp/tools/motif/) was employed to determine the motifs present in each hypothetical protein belonging to the exact core. This webserver offers a powerful platform for motif analysis by leveraging various databases and pattern repositories. This webserver leverages information from various sources including Pfam, NCBI-CDD (TIGRFAM, COG, and SMART), and PROSITE pattern to identify motifs within the amino acid sequences of the proteins. To ensure reliable results and minimize the false discovery rate, a statistical analysis was conducted using a low e-value threshold. The cutoff value of 1 × 10−5 was chosen as the criterion for significance. This threshold helps identify motifs with a high level of confidence, enhancing the accuracy and reliability of the motif predictions. Therefore, the putative functionality of the proteins was determined through a motif-based search, which helped assign potential functional annotations based on identified motifs. To ensure enhanced reliability and accuracy in determining the functionality of the identified genes, individual FASTA files were subjected to ProteInfer, a deep network for protein functional inference. By utilizing this server, we obtained GO terms along with associated confidence scores ranging from 0 to 1, providing a more comprehensive understanding of the functional annotations (31).
Phylogenetic analysis
The PPanGGOLiN workflow utilized a command-line tool to generate alignment files (.aln) encompassing the entire core gene families. These alignment files were subsequently converted to FASTA format. The resulting FASTA alignment files were concatenated, and further evolutionary analyses were carried out using MEGA v11. To infer the evolutionary history, the neighbor-joining method was employed. The Jones-Taylor-Thornton (JTT) matrix-based method was utilized to compute the evolutionary distances, which are measured in the number of amino acid substitutions per site for the final concatenated sequence file. To enhance the reliability of the analysis, the method was bootstrapped with 1,000 replications. In addition to the phylogenetic tree, pairwise distances and an overall mean distance analysis were conducted. Ambiguous positions were eliminated for each sequence pair using pairwise deletion. The resulting tree was drawn to scale, with branch lengths corresponding to the evolutionary distances employed in inferring the phylogenetic relationships. Finally, the tree was exported to an offline tree editing software FigTree v1.4.4 for final quality processing, providing a visually appealing representation of the evolutionary relationships (32).
Mapping of the genes based on pathways
A significant body of scientific literature exists that elucidates the genetic foundations of type II methylotrophs. To compile a comprehensive list of crucial pathway genes commonly found in type II methylotrophs, a meticulous manual search was conducted using PubMed and PubMed Central literature databases. In order to obtain the amino acid sequences corresponding to the identified genes, the UniProt Retrieve/ID mapping tool was employed, and the sequences were downloaded in the FASTA format. This FASTA file was subsequently uploaded to Google Collab (PPanGGOLiN), where the “align” command was utilized to align the gene set with the type II methylotrophs pangenome (33). The output data encompassed the matched gene families from the pangenome, along with partition and quality parameters such as percentage identity (pident), expectation value (e-value), and bitscore for each input gene. The term “pident” denotes the percentage of amino acid sequence positions that have identical residues, while “bitscore” is a statistical measure derived from a raw alignment score that assesses sequence similarity independently of sequence length and database size. “E-value” indicates the likelihood of sequence similarity occurring by chance. To generate a presence/absence matrix for the crucial genes, the corresponding gene family matrix data were retrieved. This matrix data were then utilized to create a heatmap using MS Excel v365.
RESULTS
Genome mining and genome screening for pangenomic studies
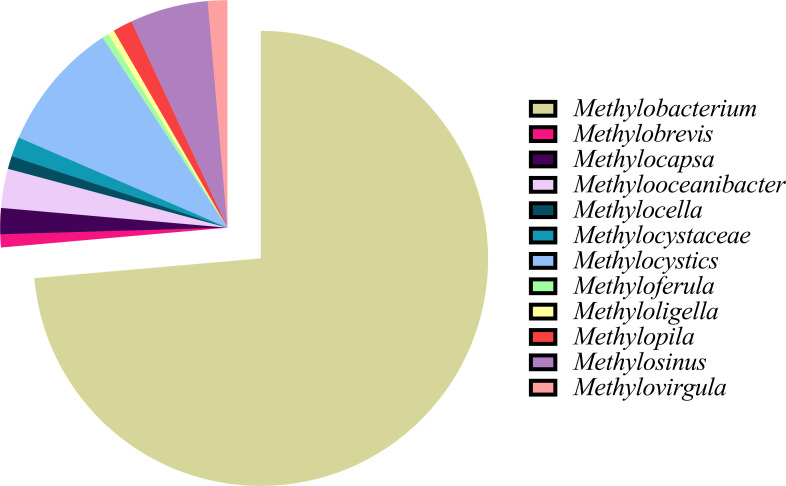
We utilized the NCBI RefSeq and GenBank databases and collected 216 type II methylotrophs of various genera (34, 35). Figure 2 visually represents the frequency distribution of these organisms within their respective genera, offering valuable insights into their taxonomic distribution. The distribution analysis reveals a diverse array of 11 distinct genera that have been identified thus far within the type II methylotrophs. Notably, among these genera, there exists an unclassified genus that has been designated with the family name Methylocystaceae. The predominant genus within the type II methylotrophs is Methylobacterium, comprising a significant portion (73.6%) of the community.
Fig 2.
Taxonomic distribution of type II methylotrophs within respective genera.
In addition to curating the data set, we performed BUSCO analysis to assess the completeness of the genomes. The visualization in Fig. S1 provides a clear depiction of the screening process, highlighting the assessment of each genome’s completeness, identification of missing fragments, and evaluation of duplication levels. A genome with less than 94% completeness was classified as a poor-quality genome for this study. This criterion has been established to ensure the inclusion of at least two organisms from each genus, allowing for a representative sampling across the taxonomic groups. By evaluating each genome against this critical criterion, we ensured that only genomes meeting the required standards were selected for subsequent analysis. From the initial collection of 216 organisms, the BUSCO analysis successfully screened and retained 75 organisms that met the predefined criteria (≥94%) for genome completeness.
The complete data set comprising 75 genomes, including information on genome size, GC content, genome coverage, and isolation site, has been compiled and presented in Table S1. Notably, the range of genome sizes varied significantly, with the smallest genome size observed in Methylooceanibacter marginalis (2,997,425 bp), while the largest genome size belonged to Methylobacterium nodulans (8,839,022 bp). The GC content across the genomes ranged from 58.3% to 73.0%, with an average value of 67.5%. Among the 75 genomes, 44 genomes exhibited a GC% equal to or greater than the average, suggesting a predominance of GC-rich genomes within the type II methylotroph community. Regarding genome coverage, 42 genomes had coverage values exceeding 100×, with the highest coverage reaching 1,680×. Conversely, 26 genomes displayed coverage below 100×, with the minimum coverage value noted at 12× for a particular genome. It is important to mention that sequence coverage data were unavailable for seven genomes in the assembly statistical report. Moreover, genomes with substantial total gap length and spanned gap values were excluded from the current study. All 75 organisms were found to inhabit mesophilic environments, with temperatures ranging from 25°C to 32°C. This temperature range represents the typical habitat for these type II methylotrophs.
Pangenome analysis and retrieval of precise core gene sets
A total of 256 precise core gene families were identified, comprising 20,188 genes distributed across 75 genomes. To establish a mapping between the 256 exact core gene families and their corresponding PROKKA gene IDs, a combination of Microsoft Excel and Python code was employed. Subsequently, the UniProt information associated with each PROKKA gene ID was retrieved from a consolidated Excel file, containing PROKKA annotations for all organisms examined in the study. These mappings allowed us to gather data on 241 distinct UniProtKBs, including their respective gene family affiliations and GO terms. It is noteworthy that each of the exact core gene families was represented in the final list of UniProtKB entries. For a comprehensive overview of the information pertaining to each specific UniProtKB, we refer readers to Table S2, which depicts an Excel file downloaded from UniProt, encompassing all relevant details associated with the respective UniProtKB entries. The functionality of the hypothetical proteins identified in this study has thoroughly been examined and discussed in subsequent sections.
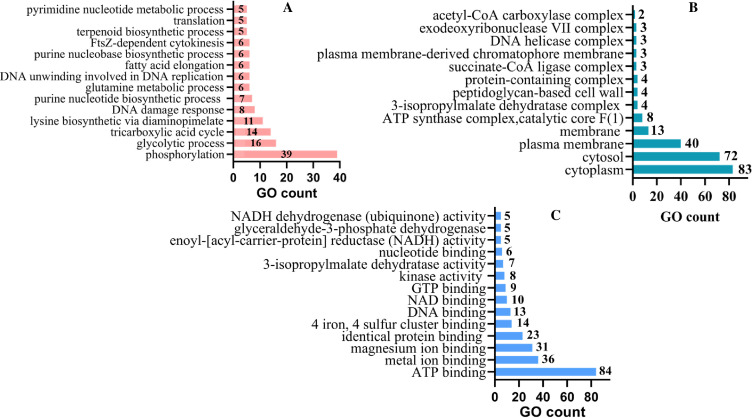
The distribution of gene ontology terms across biological processes, cellular components, and molecular functions is depicted in Fig. 3A through C, respectively. In the BP category, a total of 146 unique GO terms were observed, and those with frequencies exceeding 5 are highlighted in the plot. For the CC category, a total of 47 unique GO terms were identified, and the plot focuses on CC terms with frequencies greater than 1. In the MF category, a total of 98 unique GO terms were discovered, with the plot highlighting MF terms that occur more frequently than 5. The highest number of enriched genes (#39) in BP belongs to phosphorylation process, whereas in CC, cytoplasmic and periplasmic genes were observed to be 83 and 40, respectively. ATP binding and metal ion binding carried a significant number of genes (#84 and #36, respectively). Therefore, the analysis of GO terms related to BP, MF, and CC yielded insightful results that highlighted the intricate interconnections between these domains. The visualization in Fig. S2 underscores the harmonious coordination and cooperation among the GO terms (from 256 core gene families), forming a complex network that drives cellular activities. The interlinkage observed among the GO terms motivates the rationale behind categorizing the 256 exact core genes into 31 distinct categories, as presented in Table S3.
Fig 3.
The comprehensive analysis of GO term frequencies within the UniProtKBs data set. This analysis provides valuable insights into the prevalence and distribution of GO terms across three categories: (A) BP, (B) CC, and (C) MF.
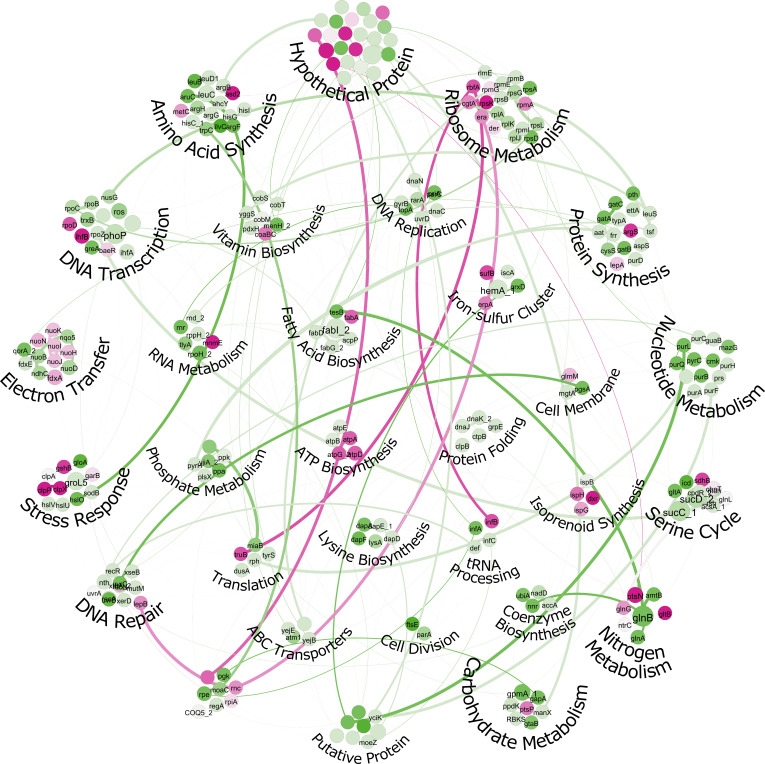
The physical protein-protein interactions of the 256 core genes are represented in Fig. 4. To gain further insights into these interactions, a statistical analysis using Gephi was conducted, and the results are provided as Fig. S3. The analysis revealed some interesting findings regarding the connectivity of these proteins. Specifically, the average degree of connectivity among the proteins was calculated to be 1.6, indicating that, on average, each protein interacts with approximately 1.6 other proteins in the network. Additionally, the average weighted degree, which takes into account the strength of these interactions, was found to be 31.1.
Fig 4.
The interconnections among gene families visualized using Gephi. Colors indicate the degree of association between gene families based on modularity statistical scores. Green nodes represent the strongest associations, while pink nodes indicate weaker connections
A significant portion of conserved gene families identified in this study is associated with fundamental cellular processes, including DNA replication, transcription, and translation. These conserved gene families are regarded as indispensable components across various environmental conditions. Notably, we observed conservation of genes related to nitrogen metabolism, which aligns with previous reports highlighting the nitrogen-fixing capabilities of type II methylotrophs (14). Additionally, the presence of conserved genes involved in isoprenoid synthesis suggests potential roles in secondary metabolite production. However, it is important to note that conservation of these genes does not necessarily imply the complete functionality of the corresponding metabolic pathways across all 75 organisms. Intriguingly, our analysis revealed the presence of 22 conserved hypothetical gene families across all the 75 organisms, the putative functions of which are discussed in detail in the subsequent section.
Functional annotation of exact core hypothetical genes
The functional annotation of exact core hypothetical genes represents a crucial step in unraveling the functional potential and understanding the significance of these genes within the broader context of the studied organisms or biological systems. Out of the 256 gene families in the exact core, a subset of 15 gene families remained unassigned in terms of their function. Table 1 presents the annotation results of hypothetical proteins, which might contribute valuable addition to the existing gene repertoire of methylotrophs. Table S4 provides a comprehensive overview of the detailed FASTA sequences and corresponding scores associated with the annotation of the hypothetical proteins.
TABLE 1.
Annotation of hypothetical proteins
| Gene familya | Annotation | GO function | Confidenceb |
|---|---|---|---|
| Methylobacterium_jeotgali_CDS_4118 | rpmE, 50S ribosomal protein L31 | Translation | Yes |
| Methylooceanibacter_caenitepidi_CDS_2654 | yhbS, N-acetyltransferase (GNAT) | Organonitrogen compound biosynthetic process | No |
| Methylobacterium_phyllostachyos_CDS_4295 | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex | Organophosphate metabolic process | Yes |
| Methylocella_tundrae_CDS_3415 | EC:3.4.11.9: Xaa-Pro aminopeptidase | Metalloaminopeptidase activity | Yes |
| Methylocystis_hirsuta_CDS_2285 | MoxR-like ATPase | Organonitrogen compound catabolic process/metalloendopeptidase activity | No |
| Methylooceanibacter_superfactus_CDS_2697 | ATPase family | Cell cycle process | No |
| Methylovirgula_sp._HY1_CDS_2003 | Glutamine amidotransferase | Pyrophosphatase activity | Yes |
| Methylobacterium_adhaesivum_CDS_1753 | NAc | Cytoplasm | No |
| Methylovirgula_sp._4M-Z18_CDS_4185 | RNA-binding domains | Lyase activity | No |
| Methylobacterium_terrae_CDS_3436 | NA | Phosphatidylinositol phosphorylation | No |
| Methylosinus_trichosporium_OB3b_CDS_4175 | Conserved protein YbaR | Enzyme inhibitor activity | No |
| Methylocystis_hirsuta_CDS_2870 | Lysine decarboxylase | Quinone metabolic process | Yes |
| Methylopila_sp._Yamaguchi_CDS_1376 | DNA polymerase III subunit | Binding | Yes |
| Methylooceanibacter_marginalis_CDS_2229 | Type III secretion system | Nitrogen compound metabolic process | Yes |
| Methylocystis_parvus_CDS_0309 | Polyhydroxyalkanoate synthesis repressor PhaR | rRNA processing/transcription factor binding | Yes |
| Methylooceanibacter_methanicus_CDS_0298 | Polyhydroxyalkanoate synthesis regulator protein | rRNA processing/transcription factor binding | Yes |
| Methylobacterium_brachythecii_CDS_2235 | Zn-finger domain | Cofactor binding | No |
| Methylobacterium_bullatum_CDS_2120 | ATPase, AAA+ | Transferase activity, transferring phosphorus-containing groups | No |
| Methylobrevis_albus_CDS_1240 | Haloacid dehalogenase | Catalytic activity | Yes |
| Methylocystis_parvus_CDS_1055 | SH3-like domain | Organonitrogen compound metabolic process | No |
| Methylobacterium_oxalidis_CDS_2397 | PtsH, phosphotransferase | Carbohydrate transport | Yes |
| Methylocapsa_sp._S129_CDS_5161 | Dipeptidyl aminopeptidase | Catalytic activity | Yes |
Information related to the gene families with regard to the FASTA sequences used in the study are provided in Table S4.
The confidence level is determined by the combined score derived from motif-based protein-level prediction from PROSITE and NCBI, as well as the GO process-based ProteInfer.
NA: not applicable.
Our efforts to unravel the functionality of the type II methylotrophs led to the discovery of 12 gene families. Through our investigations, we were able to shed light on the functions of these 12 predicted proteins. 50S ribosomal protein L31 is involved in the assembly and stabilization of the ribosomal subunits, facilitating the translation of mRNA into functional proteins (36). The NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, on the other hand, is part of the respiratory chain complex I, which is involved in energy production through oxidative phosphorylation (37). This subcomplex is responsible for the transfer of electrons from NADH to ubiquinone, a critical step in the electron transport chain. The Xaa-Pro aminopeptidase (EC:3.4.11.9) is involved in the cleavage of Xaa-Pro peptide bonds, contributing to protein degradation and turnover (38). Glutamine amidotransferase plays a key role in nitrogen metabolism by catalyzing the transfer of an amino group from glutamine to various acceptor molecules. Lysine decarboxylase participates in the decarboxylation of lysine, a process important for the synthesis of certain polyamines. The DNA polymerase III subunit is an essential component of the DNA replication machinery, contributing to accurate and efficient DNA synthesis (39). The type III secretion system enables the delivery of effector proteins from bacteria to host cells, playing a role in pathogenicity (40, 41). The polyhydroxyalkanoate (PHA) synthesis repressor PhaR regulates the synthesis of polyhydroxyalkanoates, which are storage compounds produced by bacteria. Additionally, the polyhydroxyalkanoate synthesis regulator protein controls the expression of genes involved in polyhydroxyalkanoate synthesis (42, 43). The haloacid dehalogenase enzyme is involved in the detoxification and degradation of halogenated organic compounds (44, 45). PtsH, also known as phosphotransferase, is a component of the sugar transport system, facilitating the phosphorylation of sugars for their uptake (46–48). Lastly, the dipeptidyl aminopeptidase enzyme participates in the cleavage of dipeptides, contributing to peptide metabolism.
Structure of shell and cloud
The observation of 3,433 gene families in the shell fraction and a staggering 69,732 in the cloud fraction highlights substantial evidence for horizontal gene transfer (HGT) occurring across the taxonomy, emphasizing the potential influence of genetic exchange in shaping the evolutionary dynamics of these methylotrophic organisms. Figure S4 presents the distribution of these genes across various taxonomic groups and cellular metabolic pathways. This section serves as a brief overview of the observed trends, acknowledging the presence of genes from other taxonomic lineages in the shell and cloud regions.
Type II methylotrophs, which belong to the α-proteobacteria, exhibit an interesting trend in their taxonomic distribution. As we progress from the persistent core genes (Fig. S4A) to the shell genes (Fig. S4B) and finally to the cloud genes (Fig. S4C), we observe a gradual decrease in the proportion of α-proteobacteria. This observation can be attributed to the presence of genes from other taxonomic lineages in the shell and cloud regions. It is likely that type II methylotrophs have acquired these genes through horizontal gene transfer from other bacteria, allowing them to adapt and thrive in diverse environments by harnessing the genetic repertoire of different organisms. Furthermore, an interesting observation in our analysis is the increasing fraction of genes associated with environmental signaling processes from persistent to cloud, accompanied by a decrease in the fraction of genes related to cellular metabolism. This trend suggests that cellular metabolism genes are highly conserved within the core genomes of type II methylotrophs, while the genes originating from other species are found in the cloud and shell regions, exhibiting greater variability and lack of conservation within the type II methanotrophic community.
Functional gene repertoire
Nitrogen-fixation capability
We have detected both molybdenum-iron (Mo-Fe) and vanadium-iron (V-Fe) nitrogenases across the diverse methylotrophic taxa investigated, spanning both terrestrial and atmospheric environments. Specifically, Mo-Fe nitrogenases were prevalent in 17 out of the 75 studied methylotrophic genera, including Methylocella spp., Methylocystis spp., Methyloferula spp., Methylosinus spp., Methylovirgula spp., and Methylocapsa spp. Concurrently, we identified the presence of the FixK transcriptional regulator, crucial for N2 fixation, in 17 organisms, encompassing Methylocapsa spp., Methyloferula spp., Methylopila spp., and Methylobacterium spp. Furthermore, among taxa possessing Mo-Fe nitrogenases, Methylocella spp., Methylosinus spp., and Methylovirgula spp. were observed to lack the FixK gene. However, Methylocapsa spp., Methylocystis bryophila, Methylocystis heyeri, and Methylocystis parvus were noteworthy for containing both Mo-Fe and V-Fe nitrogenases, with the presence of the VnfA transcriptional activator, indicative of specialized regulatory mechanisms in these organisms. Interestingly, our findings indicated that organisms harboring V-Fe nitrogenases lack the FixK gene.
Growth on C1 compounds
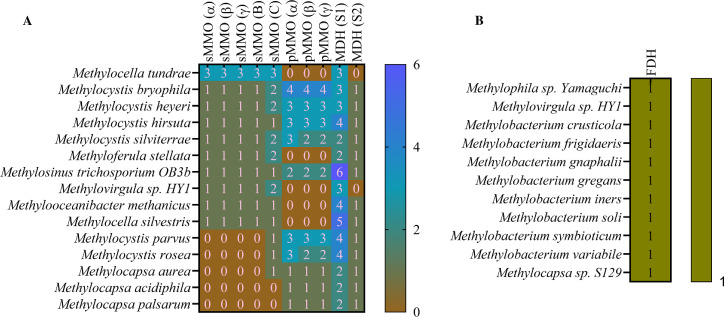
In our study, we investigated the presence of specific enzymes involved in methane oxidation, namely sMMO and pMMO, in all the 75 organisms. The 15 out of 75 organisms having MMOs are shown in Fig. 5A. Our findings revealed interesting patterns regarding the distribution of these enzymes across different organisms. Among the organisms examined, we identified 10 organisms that contain the sMMO hydrolase (α, β, and γ chains), while 10 organisms contain sMMO protein B (regulatory). Additionally, 13 organisms were found to possess sMMO protein C (reductase). Notably, the three additional organisms identified as Methylocystis parvus, Methylocystis rosea, and Methylocapsa aurea specifically harbored sMMO protein C. It is worth mentioning that in contrast to a previous study that reported two copies, our findings revealed that Methylocella tundrae exhibits three copies of all genes associated with sMMO, which are non-identical to each other. On the other hand, Methylocystis bryophila, Methylocystis hirsuta, Methylocystis silviterrae, and Methylosinus trichosporium contained two copies of protein C. Moreover, Methylocella tundrae, Methyloferula stellata, Methylovirgula sp. HY1, and Methylocella silvestris lacked pMMO proteins but possessed sMMO proteins. However, we noted that Methylocystis parvus, Methylocystis rosea, and Methylocapsa aurea, in addition to protein C, also contained all the pMMO genes. Interestingly, Methylocapsa aurea, Methylocapsa acidiphila, and Methylocapsa palsarum exclusively harbored pMMO genes, with one copy each. Notably, Methylocystis bryophila displayed four copies of pMMO genes.
Fig 5.
Presence-absence matrix (heatmap) illustrating the distribution of key genes involved in methane metabolism among the examined methanotrophic strains. Subfigure (A) displays the availability of genes associated with MMOs and MDH. Subfigure (B) focuses specifically on the presence or absence of formaldehyde dehydrogenase (FDH) genes.
Furthermore, we observed the presence of two subunits of methanol dehydrogenase (cytochrome c) in the organisms examined. Subunit 1 was detected in 73 organisms, with an average copy number of 3.16 per organisms. On the other hand, subunit II was found in 55 organisms, with an average copy number of 1.05 per organism. It is noteworthy that Methylovirgula sp. 4M-Z18 and Methylocapsa sp. S129 did not exhibit any methanol dehydrogenase (cytochrome c). However, it is interesting to highlight that Methylovirgula sp. 4M-Z18 possesses NAD-dependent methanol dehydrogenase, indicating an alternative pathway for methanol oxidation in this particular organism. Figure 5A illustrates the distribution of methanol dehydrogenase subunits among the organisms that possess either pMMO or sMMO. This subset of organisms was selected to highlight the presence of methanol dehydrogenase, which is a key enzyme involved in methanol utilization.
The presence of formaldehyde dehydrogenase (FDH), which facilitates the direct oxidation of formaldehyde to formic acid, was observed in only 10 organisms (Fig. 5B). Interestingly, none of the examined organisms possessed either of sMMO or pMMO genes, except for Methylovirgula sp. HY1. This suggests that in methanotrophs, the direct oxidation of formaldehyde to formic acid may not be the primary pathway, and alternative assimilation pathways are likely in place, via the serine pathway. However, these 10 organisms that do possess FDH have the ability to utilize formaldehyde as a carbon source and can efficiently convert it to carbon dioxide. The metabolic flux in these organisms is directed toward carbon dioxide production, indicating the importance of formaldehyde metabolism in their carbon assimilation strategies. It is worth noting that among the 11 organisms, Methylocapsa sp. S129 stands out as it contains a non-identical form of formaldehyde dehydrogenase compared to the other nine organisms.
In summary, our study identified a total of 15 organisms (out of 75 organisms) capable of methane oxidation, classifying them as methanotrophs. Among these organisms, five contained only sMMO, five contained only pMMO, and five contained both types of enzymes. Notably, the Methylocapsa genus exclusively possessed pMMO, while Methylocella and Methyloferula solely exhibited sMMO. The distribution of sMMO copies among the 10 organisms was found to be 12, whereas the number of pMMO copies totaled 22. This observation suggests that organisms possessing pMMO may exhibit higher methane oxidation rates compared to those with sMMO. The detection of methanol dehydrogenase subunit 1 in the examined organisms suggests their ability to utilize methanol as a carbon source, except for Methylocapsa sp. S129. This finding implies that these organisms have the enzymatic machinery necessary for methanol oxidation, a characteristic trait of methanotrophs. However, it is intriguing to note that Methylovirgula sp. 4M-Z18 possesses NAD-dependent methanol dehydrogenase, indicating an alternative pathway for methanol oxidation. The exact nature and mechanism of methanol oxidation in Methylovirgula sp. 4M-Z18 warrant further investigation to elucidate the specific enzymatic reactions involved in this process.
Serine pathway
In our study, we made an intriguing observation regarding the presence of the key enzyme serine hydroxymethyltransferase (SHMT) in the examined organisms. We found that SHMT was present in 74 organisms, with an average copy number of 1.31 per organism, highlighting its prevalence and importance in methylotrophic metabolism. However, it is worth noting that Methylooceanibacter marginalis was an exception to this pattern, as it did not possess SHMT. An interesting finding emerged when examining Methylobacterium variabile, as it exhibited both isozymes of SHMT. This unique characteristic suggests that Methylobacterium variabile has the potential to utilize a broader range of carbon sources, including succinate, methane, and methanol. In our study, we made an intriguing discovery concerning the presence of serine-glyoxylate transaminase (SGT) in all 75 examined organisms. As far as our current knowledge extends, the literature does not thoroughly explore the isozymes of SGT. Nevertheless, in the course of our inquiry, we observed three distinct isozymes of this enzyme across various organisms. Of notable interest is the revelation concerning Methylobrevis sp., which harbors a distinctive SGT isozyme absent in all other studied organisms. Another notable observation is that Methylobrevis albus shares a common isozyme with 70 other organisms, indicating a conserved functional role for this particular isozyme among a significant subset of the organisms. Additionally, there are 11 more organisms that share the most common isozyme but also possess a non-identical isozyme, suggesting additional metabolic diversity within this group.
Our investigation revealed the ubiquitous presence of hydroxypyruvate reductase (HPR) in all 75 examined organisms but with 22 different isoforms. Several organisms, including Methylocapsa sp. S129, Methylopila sp. Yamaguchi, Methylobacterium jeotgali, and Methylobacterium crusticola, exhibited a distinct and entirely different type of the HPR enzyme, indicating the presence of unique functional adaptations in their glyoxylate and hydroxypyruvate metabolic pathways. Furthermore, a putative hydroxypyruvate reductase was observed in 73 organisms, with an average copy number of 1.14 per organism. Notably, Methylobacterium ajmalii displayed the highest number of copies of this enzyme, indicating a potentially significant role in the metabolic activities of this particular organism. Further investigations on the enzymes, glycerate 2-kinase, phosphoglycerate mutase, phosphopyruvate hydratase, phosphoenolpyruvate carboxylase, malate dehydrogenase, and malyl-CoA lyase, showed that these enzymes are indeed present in all the 75 organisms but were observed in the shell and cloud region due to their variations in isoforms.
Glyoxylate cycle and ethylmalonyl-CoA pathway
During our analysis of the isocitrate lyase enzyme, we made interesting observations regarding its presence among the methane-oxidizing organisms. Out of the 15 organisms capable of oxidizing methane, only 6 organisms, namely Methyloferula stellata, Methylovirgula sp. HY1, Methylocapsa silvestris, Methylocapsa aurea, and Methylocapsa palsarum, were found to possess the isocitrate lyase enzyme. This indicates that these organisms utilize the glyoxylate pathway for carbon assimilation. On the other hand, the remaining nine organisms may follow ethylmalonyl-CoA (EMC) pathway instead. Interestingly, among the methylotrophic organisms, Methylobacterium gossipiicola, Methylovirgula ligni, Methylovirgula sp. 4MZ18, and Methylocapsa acidiphila were found to possess isocitrate lyase, indicating their potential utilization of the glyoxylate pathway for carbon assimilation. Another key enzyme in the glyoxylate pathway is malate synthase. We further investigated the presence of malate synthase among these organisms. We observed that malate synthase was present in 31 organisms, either in the G-form or the A-form. Notably, all the species mentioned above that possess isocitrate lyase were also found to contain malate synthase, except for Methylobacterium gossipiicola, suggesting the presence of the glyoxylate pathway in the six methanotrophic strains. Among these organisms, Methylocapsa sp. S129 was the only one to possess the A-form of malate synthase, while the rest had the G-form. Interestingly, Methylovirgula sp. HY1 and Methyloferula stellata not only possessed the common G-form of malate synthase but also exhibited distinct G-forms in their genomes.
Based on our expectation that nine organisms possess the glyoxylate pathway, we hypothesized that the remaining 66 organisms would likely follow the ethylmalonyl-CoA pathway. To explore this further, we focused on the central coenzyme, crotonyl-CoA carboxylase/reductase. Interestingly, our analysis revealed the presence of crotonyl-CoA carboxylase/reductase in 67 organisms, with an average copy number of 1.03. This finding aligned with our expectation that the majority of the organisms would utilize the EMC pathway. It was intriguing to note that the methylotrophic strain Methylovirgula sp. 4MZ18 possessed both the enzymes associated with the glyoxylate pathway and the EMC pathway. Another enzyme of interest within the EMC pathway is methylsuccinyl-CoA dehydrogenase. We observed the presence of this enzyme in all 67 organisms, consistent with the presence of crotonyl-CoA carboxylase/reductase. However, among these organisms, 33 exhibited a distinct isozyme copy of this enzyme. Notably, Methylobacterium gossipiicola was found to contain both crotonyl-CoA carboxylase/reductase and methylsuccinyl-CoA dehydrogenase, indicating its adherence to the EMC pathway, despite the presence of isocitrate lyase within its genome.
Phylogenetic analysis
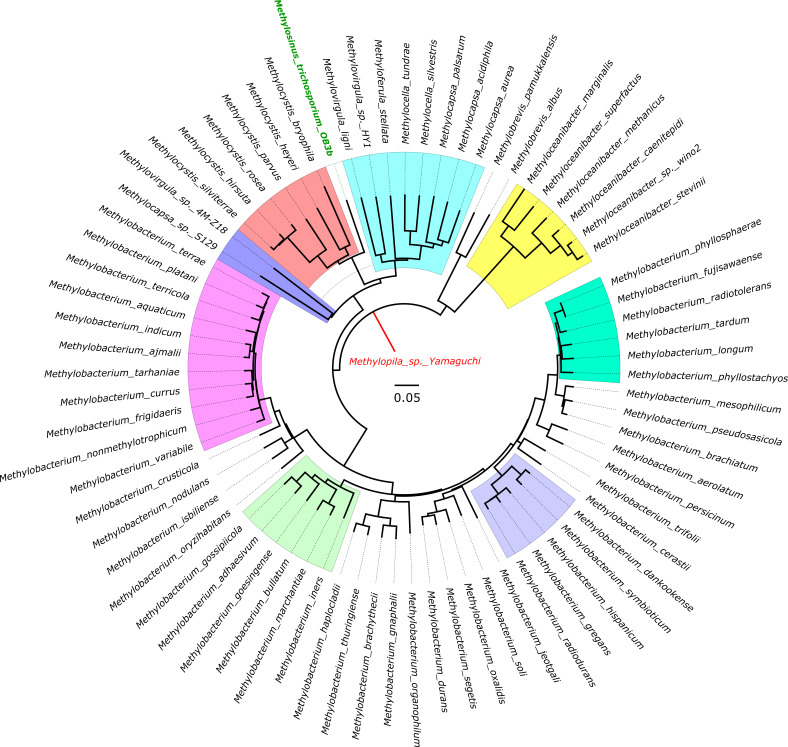
The phylogenetic tree analysis revealed intriguing patterns of classification among the organisms. Two major divisions were observed (shown in Fig. 6), with one comprising the Methylobacterium genus and the other consisting of the remaining genera. The genetic patterns observed in relation to the enzymes shed light on the unique characteristics of Methylobrevis compared to other methylotrophs during formaldehyde assimilation. Methylobrevis exhibited distinct isozymes or isoforms of enzymes present in other organisms, setting it apart from the common methanotrophic clade. Notably, Methylocapsa sp. S129 displayed a distinct genomic architecture and was separated from other Methylocapsa species. This separation was evident through the absence of methanol dehydrogenase in this particular methylotroph. Despite this distinction, Methylocapsa sp. S129 showed high similarity with Methylovirgula sp. 4M-Z18. It is worth mentioning that Methylovirgula sp. 4M-Z18 was also separated from other Methylovirgula species. Given the interconnectivity and lack of association with other organisms, we propose the establishment of a new separate genus for Methylocapsa sp. S129 and Methylovirgula sp. 4M-Z18.
Fig 6.
Dendrogram illustrating the hierarchical relationships among 75 type II methylotrophs. Branch lengths scaled at a ratio of 1 cm = 0.05, ensuring precise clustering of evolutionarily related methylotrophs. The outgroup is highlighted in red letters, while the methanotroph of interest is denoted by green letters. Noteworthy clads are highlighted clusters of closely related methylotrophs.
A cluster comprising the Methylocapsa, Methyloferula, Methylocella, and Methylovirgula genera was observed (highlighted in bluish green). Additionally, the model organism Methylosinus trichosporium OB3b formed a common cluster (highlighted in orange) with the Methylocystis genus. It is evident that the organisms within the bluish green cluster and the orange cluster are primarily methanotrophs, with the exception of Methylovirgula ligni. The Methylooceanibacter genus displayed a distinct cluster without association to other organisms, indicating that all species within this genus are closely related. However, an exception occurred with Methylooceanibacter methanicus, which exhibited the ability to oxidize methane. A cluster highlighted in pink contained Methylobacterium species with no significant distance between the genera. It is worth noting that Methylobacterium ajmalii, despite being reported for its methane-oxidizing capacity, did not have any mapped genes within the pangenome analysis. This suggests the need for a comprehensive search within the genome, as a distinct isoform of MMOs may be present. Similar considerations apply to its sister clade, Methylobacterium indicum.
While our focus primarily revolved around major pathways such as methane oxidation, the serine pathway, glyoxylate pathway, and EMC pathway, it is essential to give due attention to other pathway genes. Understanding the capabilities of these type II methylotrophs to grow on various C1-C6 compounds requires a comprehensive investigation of additional pathways. The observed variations among these organisms highlight the diversity and adaptability within this group of microorganisms, emphasizing the need for further exploration and characterization of their unique metabolic capabilities.
DISCUSSION
Exact core genes
This investigation underscores the significance of conserved gene families in driving essential cellular processes such as DNA replication, transcription, and translation, which are indispensable for cellular survival across diverse environmental conditions. These processes are tightly interconnected, as evidenced by the strong interconnections revealed by GO terms, in terms of BP, MF, and CC. For instance, phosphorylation, a critical post-translational modification, is intricately linked to key metabolic pathways like glycolysis and the tricarboxylic acid cycle (TCA), pivotal for energy production (49, 50). Chistoserdova et al. documented the occurrence of complete TCA cycles, along with a full serine cycle, in a very few type II methylotrophs, notably including Methylobacterium extorquens and Granulibacter bethesdensis (51). However, in addition to this, M. trichosporium OB3b possesses a closed and reversible TCA cycle capable of channeling TCA intermediates toward pyruvate, acetyl CoA, and manoyl-CoA (52). Nonetheless, the interconnectedness of the EMC pathway, serine cycle, and TCA cycle confers an advantageous control mechanism over carbon flux within the TCA cycle (53). Similarly, the DNA damage response pathway, crucial for maintaining genomic integrity, intertwines with DNA replication and topological changes (54, 55). Additionally, biosynthetic processes such as nucleotide biosynthesis, including purine and pyrimidine pathways, are interconnected, playing pivotal roles in DNA and RNA synthesis (56, 57). Furthermore, the investigation highlights the interplay between various biosynthetic pathways, such as lysine biosynthesis and fatty acid elongation, elucidating the intricate relationship between amino acid metabolism and lipid synthesis (58–60). Processes like FtsZ-dependent cytokinesis underscore the importance of cellular division, while terpenoid biosynthesis demonstrates the interconnectedness between metabolic processes and antioxidant defense mechanisms (61, 62).
Turning to molecular function counts, the analysis reveals crucial binding events, including ATP and GTP binding, which are central to cellular energy transfer processes (63). Metal ion binding, particularly magnesium ion binding, underscores the importance of metal ions as cofactors in enzymatic reactions and for maintaining protein structural integrity (64–66). Moreover, protein-protein interactions, exemplified by identical protein binding and 4 iron-4 sulfur cluster binding, are vital for protein complex formation and electron transfer processes (67–69). Enzymatic activities like NAD binding and glyceraldehyde-3-phosphate dehydrogenase activity are pivotal for redox reactions and glycolysis, respectively, highlighting the diverse roles of proteins in cellular function and metabolism (70). Enzymes, such as NADH dehydrogenase (ubiquinone) activity and enoyl-[acyl-carrier-protein] reductase (NADH) activity, rely on NADH as a cofactor to catalyze specific reactions (71).
In addition, the study uncovered 22 hypothetical genes that were conserved among all 75 methylotrophs examined. Notably, during the annotation process, 2 enzymes stood out prominently: PhaR and haloacid dehalogenase, within the subset of 12 gene families. The Akira et al. study demonstrated that the presence of PhaR in an organism correlates with the production of short-chain polyhydroxyalkanoates (43). In our investigation, the presence of PhaR indicates that type II methylotrophs possess the capability to produce short-chain PHAs. Again, methylotrophs are recognized for their effectiveness in bioremediating halogenated compounds, underscoring the significance of the presence of haloacid dehalogenase (72).
Functional gene repertoire in shell and cloud
Nitrogen-fixation capability
Type II methylotrophs exhibit a remarkable trait of expressing nitrogenase (encoded by nifH) to harness atmospheric N2 as a nitrogen source, a pivotal adaptation in their ecological niche (14, 73). Auman et al. investigated this trait by assessing type II NifH amplification and acetylene reduction activity, confirming N2-fixing capability in Methylocystis spp. and Methylosinus spp. (14). Type II methanotrophs, exemplified by Methylosinus and Methylocella, are believed to play a significant role in N2 fixation within forest soil ecosystems and rice plants (74). Alongside them, diazotrophs like Rhizobium, Bradyrhizobium, and Mesorhizobium are renowned for their N2-fixing abilities and are commonly found in association with methylotrophs in rice paddies (75). While Mo-Fe nitrogenases are ubiquitous among diazotrophs, V-Fe nitrogenases are confined to a select group of prokaryotes, notably prevalent in Methylocystis strains isolated from wetlands (76). Notably, Mo-Fe nitrogenase is favored in Mo-rich conditions, whereas V-Fe nitrogenase predominates in Mo-deficient environments, with Oshkin et al. proposing this diversification as an adaptation to nutrient limitation in wetlands (15, 77). At standard temperatures (e.g., 30°C), V-Fe nitrogenase exhibits lower specific activity compared to its Mo-Fe counterpart, but at colder temperatures (e.g., 5°C), V-Fe nitrogenase surpasses Mo-Fe nitrogenase in activity (78). Moreover, V-Fe nitrogenase displays versatility by catalyzing both CO and N2 reductions, suggesting a potential interplay between carbon and nitrogen cycles in wetlands. Recent findings indicate the presence of V-Fe nitrogenase in Methylocystis bryophila S285 and Methylospira mobilis Shm1, although classical Mo-Fe nitrogenase genes are commonly found across methanotrophic genomes (79, 80).
We identified the presence of both V-Fe and Mo-Fe nitrogenases in Methylocystis bryophila, Methylocystis heyeri, and Methylocystis parvus, underscoring their significance in N2 fixation, which can be finely tuned simply by adjusting the growth medium. Furthermore, our investigation reveals the presence of the fixK gene, a transcription factor belonging to the Crp family, which governs N2 fixation, exhibiting both positive and negative regulation, mirroring findings in Rhizobium meliloti. In our study, we detected this gene in 15 methylotrophs, including prominent methanotroph strains—Methylocapsa, Methylocella, Methyloferula, and Methylopila. Of particular interest, strains associated with V-Fe nitrogenases—namely, Methylocystis bryophila, Methylocystis heyeri, and Methylocystis parvus—harbor the VnfA transcriptional activator as their N2-fixation regulatory protein, indicating a specialized adaptation within this subset of methanotrophs.
Growth on C1 compounds
The carbon assimilation pathways of aerobic methylotrophs that are capable of growing on C1 compounds as their source of carbon exhibit distinct mechanisms with regard to its enzyme characteristics (presence/absence of genes) (81). Methylotrophs typically rely on more commonly available C1 compounds such as methane, methanol, formaldehyde, or formic acid (82). Methanotrophs, a subset of methylotrophs, utilize methane as their primary substrate, which is oxidized by MMOs (sMMO or pMMO) to produce methanol (81). Among the type II methanotroph strains, Methylocapsa aurea exclusively possesses the pMMO form, while some strains exhibit either both forms of MMO (Methylosinus trichosporium OB3b) or solely sMMO (Methylocella and Methyloferula) (83, 84). In the case of Methylocella tundrae, it contains two similar but not identical sMMO operons, showcasing a 96% nucleotide identity between the mmoXYBZDCRG operons and an 88%–99% amino acid identity between the corresponding polypeptides (85). Although many methanotrophs harbor multiple copies of the pmoCAB operon, the existence of additional copies of the sMMO genes is quite not general (86). While, in our study, we identified three copies of the sMMO subunits (XYZ) within Methylocella tundrae, along with proteins B and C, deviating from the expected two copies.
The further oxidation of methanol in methylotrophs involves the action of methanol dehydrogenase, which converts methanol to formaldehyde (87). The calcium-dependent MDH is encoded by the Mxa operon, with its larger and smaller subunits denoted as mxaF and mxaI, respectively (88). This form is widespread among methanotrophs and is activated by the cytochrome c electron acceptor. In contrast, the lanthanide-dependent MDH, xoxF, dominates in acidophilic or acid-tolerant methylotrophs due to lanthanides’ stronger Lewis acid properties compared to calcium (89). This enhances the electrophilic nature of active carbons in PQQ, facilitating electron removal from methanol. In our study, among methanotrophs, Methylovirgula sp. 4M-Z18 exhibited NAD-dependent MDH, which donates electrons under both aerobic and anaerobic conditions, distinguishing it from cytochrome c-dependent MDH. Notably, this trait is commonly associated with gram-positive bacteria. The reports highlighted that Methylosinus trichosporium OB3b contains both xoxF and mxaF, a finding corroborated in our study (90). Additionally, we observed Methylocella spp. and Methyloferula spp. under the same category, which have not been reported previously.
The subsequent assimilation of formaldehyde in type II methylotrophs primarily relies on the serine cycle. This cycle allows the incorporation of formaldehyde into cellular metabolism for the synthesis of essential biomolecules (91). To complete the oxidation process, formaldehyde is further oxidized to formic acid by the action of formaldehyde dehydrogenase (92). Hence, methylotrophs derive the energy necessary for their growth by oxidizing C1 substrates through specific dehydrogenases. Unlike other organisms, they do not depend on a complete Krebs TCA cycle for their energy generation.
Serine pathway
The serine cycle is different from the other pathways in having carboxylic and amino acids as intermediates instead of the usual carbohydrates. In the initial step of the serine pathway, formaldehyde undergoes a reaction with glycine to produce L-serine. This crucial transformation is facilitated by the enzyme glycine/serine hydroxymethyltransferase, which is classified as EC 2.1.2.1. SHMT utilizes tetrahydrofolate as a cofactor in this reaction. When formaldehyde binds to SHMT, it generates a complex called 5,10-methylenetetrahydrofolate. During the enzymatic process, the formaldehyde moiety from 5,10-methylenetetrahydrofolate is transferred to glycine, leading to the formation of L-serine. In a study conducted by O’Connor et al., it was demonstrated that facultative methylotrophs possess multiple isozymes of SHMT (93). Interestingly, the presence and activity of these isozymes were found to vary depending on the carbon source used for growth. Specifically, one isozyme was found to be predominant when the organism was grown using methane or methanol, while the other isozyme dominated when succinate was utilized as the sole carbon source. This study identified multiple isozymes of SHMT, directly linked to the organism’s capacity for multi-substrate uptake.
In the subsequent step of the pathway, L-serine undergoes transamination with glyoxylate, utilizing the enzyme serine-glyoxylate transaminase (EC 2.6.1.45). This reaction leads to the production of 3-hydroxypyruvate and glycine. The glycine generated can be recycled and serve as a substrate for SHMT. Meanwhile, hydroxypyruvate is reduced to D-glycerate by the enzyme hydroxypyruvate reductase (EC 1.1.1.81). The resulting D-glycerate is further phosphorylated by glycerate 2-kinase (EC 2.7.1.165) to yield 2-phospho-D-glycerate. At this stage, Samanta et al. showed the bifurcation of the pathway in M. trichosporium OB3b, which is consistent for a few other type II methylotrophs (especially Methylocystis sp.) (94–96). A portion of the 2-phospho-D-glycerate is converted by phosphoglycerate mutase (2,3-diphosphoglycerate dependent) (EC 5.4.2.11) into 3-phospho-D-glycerate. The remainder of the 2-phospho-D-glycerate is transformed by phosphopyruvate hydratase (EC 4.2.1.11) into phosphoenolpyruvate. Phosphoenolpyruvate carboxylase (EC 4.1.1.31) subsequently facilitates the fixation of carbon dioxide, converting phosphoenolpyruvate into oxaloacetate. Oxaloacetate is then reduced to (S)-malate by the enzyme malate dehydrogenase (EC 1.1.1.37). A reaction catalyzed by malate-CoA ligase (EC 6.2.1.9) forms malyl coenzyme A, which is further cleaved by malyl-CoA lyase (EC 4.1.3.24) into acetyl-CoA and glyoxylate (97). The presence of the last two enzymes (EC 6.2.1.9 and EC 4.1.3.24), as well as EC 1.1.1.81 and EC 2.7.1.165, in methylotrophs signifies the existence of the serine pathway.
Glyoxylate and EMC pathway
The fate of acetyl-CoA in an organism depends on the presence or absence of the enzyme isocitrate lyase (EC 4.1.3.1), which serves as a key enzyme in the glyoxylate cycle. Indeed, some methylotrophs do have the key enzyme (isocitrate lyase) of that pathway, and they assimilate C1 compounds by what is known as the icl+ serine cycle. If the organism possesses this enzyme, acetyl-CoA is converted to glyoxylate through the glyoxylate cycle, a modified version of TCA cycle (98). The glyoxylate cycle requires two key enzymes, namely isocitrate lyase and malate synthase, in addition to certain TCA cycle enzymes (99). These enzymes are often referred to as anaplerotic enzymes since they play a crucial role in replenishing the intermediates of the TCA cycle. As a result, this pathway is commonly known as the glyoxylate bypass (100). However, it had been shown previously that there is no isocitrate lyase present during methylotrophic growth for majority of methylotrophs. However, in the absence of isocitrate lyase, acetyl-CoA is processed through the EMC pathway (98).
Within EMC pathway, a C4 compound known as acetoacetyl-CoA, derived from two acetyl-CoA molecules, undergoes a series of transformations to yield a C5 compound called 2-methylfumaryl-CoA (101). This conversion involves the hydration of 2-methylfumaryl-CoA to produce (2R,3S)-β-methylmalyl-CoA. The latter compound is then cleaved into glyoxylate and propanoyl-CoA. By condensing glyoxylate with another molecule of acetyl-CoA, (S)-malate is formed (102). Simultaneously, propionyl-CoA undergoes carboxylation via a dedicated pathway, leading to the production of succinate (103). The central enzyme in this pathway is crotonyl-CoA carboxylase/reductase, which exhibits the remarkable ability to carboxylate and reduce the four-carbon compound crotonyl-CoA simultaneously (104). This enzymatic reaction results in the formation of a five-carbon compound known as (2S)-ethylmalonyl-CoA. In certain methylotrophs, an enzyme called ethylmalonyl-CoA mutase plays a pivotal role in the conversion of acetyl-CoA to glyoxylate (105). This unique enzyme belongs to a distinct category of coenzyme B12-dependent acyl-CoA mutases. Alongside, propionyl-CoA carboxylase is also involved in this process (106). Furthermore, the ethylmalonyl-CoA pathway for acetate assimilation is finalized by the action of methylsuccinyl-CoA dehydrogenase (107).
Phylogenetic analysis
The genus Methylobacterium encompasses a diverse group of pink-pigmented facultatively methylotrophic bacteria (108). Methylobacterium demonstrate the ability to utilize one-carbon compounds such as formate, formaldehyde, and methanol as their sole carbon and energy source (109). Furthermore, they exhibit versatility in utilizing multi-carbon growth substrates. They have evolved to thrive in a wide range of environmental conditions, including extremes of temperature, salinity, drought, and pH, as well as acidic and alkaline habitats (110). These bacteria are commonly found in agroecosystems and can be isolated from various parts of plants (111). Methylobacterium play a crucial role in providing essential nutrients (nitrogen- and phosphorus-associated compounds) to plants during stressful periods, thereby enhancing their tolerance to abiotic stress (112). Multiple reports have shed light on the genetic and environmental diversity within the type II methylotrophs, revealing intriguing characteristics (113). One example is the Methylobrevis genus, which consists of two known species: Methylobrevis albus and Methylobrevis pamukkalensis. Of the two, Methylobrevis pamukkalensis stands out as a facultative halotolerant methylotroph (114). The classification of Methylobacterium within the type II methylotrophy and its role as a methanotroph have been the subject of extensive debate (115). The type species of Methylobacterium, e.g., Methylobacterium organophilum, was described for the first time as a facultative methane-utilizing bacterium (116). Notably, Bijlani et al. conducted a study demonstrating the growth of Methylobacterium ajmalli using both methane and methanol (117). Consequently, Methylobacterium species are often harnessed for metabolite production due to their capacity to grow on methane and its oxidation by-products, such as methanol and formaldehyde (118). However, the methanotrophy capacity of the remaining Methylobacterium species is yet to be fully elucidated. The genetic architecture of these bacteria explored through the phylogenetic analysis revealed a separate group from methanotrophs.
Hence, this study delves into the adaptability of methylotrophs—from their diverse substrate preferences to their distinct genetic architecture—for thriving in various environmental conditions and contributing to bioremediation efforts. In the future, a comprehensive analysis of individual isozymes involved in methane oxidation and the serine pathway across diverse methylotrophs will offer insights into variations in catalytic structure, enzyme-substrate conformations, and reaction mechanisms.
Conclusion
Our pangenomic approach has provided valuable insights into the metabolic genetic distributions across the 75 type II methylotrophic strains. We identified 256 exact core gene families, encompassing essential housekeeping genes associated with central dogma processes, with the exception of 22 hypothetical proteins whose functions were elucidated. Notably, among the 12 identified hypothetical proteins, 1 was involved in translation, 1 related to nitrogen metabolism, and 2 with catalytic activity (haloacid dehalogenase and dipeptidyl aminopeptidase), showcasing their significance.
Our analysis of the shell and cloud regions revealed a trend of HGT, indicating the acquisition of a larger fraction of genes in the cloud region from other taxonomic communities. This highlights the adaptive strategy of type II methylotrophs to thrive in versatile environments by incorporating genetic material from diverse sources. Furthermore, our study unraveled a diverse repertoire of genes associated with crucial metabolic pathways, including methane oxidation, serine pathway, glyoxylate pathway, and ethylmalonyl-CoA pathway. We made intriguing observations, such as Methylocella tundrae’s possession of three copies of sMMO components, distinguishing it from other methanotrophs. Additionally, Methylooceanibacter marginalis lacked SHMT, while Methylobacterium variabile exhibited both isozymes of SHMT, suggesting their potential to utilize a broader range of carbon sources.
Phylogenetic analysis and distinct clustering patterns among the type II methylotrophs led us to propose a separate or possibly new genus for Methylovirgula sp. 4M-Z18 and Methylocapsa sp. S129, underscoring their unique characteristics and evolutionary divergence within the methylotrophic community. By unraveling the genetic foundations of type II methylotrophs, their potential for sustainable solutions in various fields, including bioremediation, biofuel production, and carbon capture, can be unlocked. These findings contribute to the advancement of environmental and industrial biotechnology, offering promising avenues for harnessing the metabolic capabilities of these organisms in practical applications.
ACKNOWLEDGMENTS
The authors acknowledge the support by the Department of Chemical and Biological Engineering at the South Dakota School of Mines and Technology. We would like to thank Dr. Ram N. Singh from South Dakota School of Mines and Technology (Rapid City, SD, USA) for annotating the organisms using PROKKA.
This research was funded by the National Science Foundation in the form of the BuG ReMeDEE initiative (Award #1736255, #1849206, and #1920954).
Conceptualization, D.S., S.R., and R.K.S.; methodology, D.S. and S.R.; software, D.S., S.R., and P.S.; validation, D.S., S.R., and P.S.; formal analysis, D.S., S.R., P.S., and R.K.S.; investigation, D.S. and R.K.S.; resources, R.K.S.; data curation, D.S., S.R., and R.K.S.; writing—original draft preparation, D.S.; writing—review and editing, D.S., S.R., P.S., and R.K.S.; visualization, D.S. and S.R.; supervision, R.K.S.; project administration, R.K.S.; funding acquisition, R.K.S. All authors reviewed the manuscript.
Contributor Information
Rajesh K. Sani, Email: Rajesh.Sani@sdsmt.edu.
Katrine Whiteson, University of California Irvine, Irvine, California, USA.
DATA AVAILABILITY
The original contributions presented in the study are included in the article and supplemental material. The PPanGGOLiN algorithm is available for access at https://github.com/labgem/PPanGGOLiN.
SUPPLEMENTAL MATERIAL
The following material is available online at https://doi.org/10.1128/msystems.00248-24.
Comprehensive visualization of the screening process, depicting the evaluation of each genome's completeness percentage.
Visualization of the intricate interconnections and collaborative relationships among the GO terms.
Statistical analysis of the 256 nodes determined using Gephi.
Distribution of genes across taxonomic groups and cellular metabolic pathways.
Legends to supplemental figures and tables.
The complete data set comprising 75 genomes, including information on genome size, GC content, genome coverage, and isolation site.
Detailed UniProt entries and descriptions of 256 exact core gene families.
Categorizing the 256 exact core genes into 31 distinct categories with their annotation, number of organisms, and number of sequences.
A comprehensive overview of the detailed FASTA sequence, and corresponding scores associated with the annotation of the hypothetical proteins.
ASM does not own the copyrights to Supplemental Material that may be linked to, or accessed through, an article. The authors have granted ASM a non-exclusive, world-wide license to publish the Supplemental Material files. Please contact the corresponding author directly for reuse.
REFERENCES
- 1. Sherman RM, Salzberg SL. 2020. Pan-genomics in the human genome era. Nat Rev Genet 21:243–254. doi: 10.1038/s41576-020-0210-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Whibley A, Kelley JL, Narum SR. 2021. The changing face of genome assemblies: guidance on achieving high‐quality reference genomes. Mol Ecol Resour 21:641–652. doi: 10.1111/1755-0998.13312 [DOI] [PubMed] [Google Scholar]
- 3. Sherman RM, Forman J, Antonescu V, Puiu D, Daya M, Rafaels N, Boorgula MP, Chavan S, Vergara C, Ortega VE, et al. 2019. Assembly of a pan-genome from deep sequencing of 910 humans of African descent. Nat Genet 51:30–35. doi: 10.1038/s41588-018-0273-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Hübner S. 2022. Are we there yet? Driving the road to evolutionary graph-pangenomics. Curr Opin Plant Biol 66:102195. doi: 10.1016/j.pbi.2022.102195 [DOI] [PubMed] [Google Scholar]
- 5. Ballouz S, Dobin A, Gillis JA. 2019. Is it time to change the reference genome? Genome Biol. 20:159. doi: 10.1186/s13059-019-1774-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Li Y, Zhou G, Ma J, Jiang W, Jin L, Zhang Z, Guo Y, Zhang J, Sui Y, Zheng L, et al. 2014. De novo assembly of soybean wild relatives for pan-genome analysis of diversity and agronomic traits. Nat Biotechnol 32:1045–1052. doi: 10.1038/nbt.2979 [DOI] [PubMed] [Google Scholar]
- 7. Samanta D, Govil T, Saxena P, Gadhamshetty V, Krumholz LR, Salem DR, Sani RK. 2022. Enhancement of methane catalysis rates in Methylosinus trichosporium OB3b. Biomolecules 12:560. doi: 10.3390/biom12040560 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Samanta D, Govil T, Salem DR, Krumholz LR, Gerlach R, Gadhamshetty V, Sani RK. 2019. Methane monooxygenases: their regulations and applications in biofuel production, p 187–206. In Microbes for sustainable development and bioremediation. CRC Press. [Google Scholar]
- 9. Trotsenko YA, Murrell JC. 2008. Metabolic aspects of aerobic obligate methanotrophy⋆. Adv Appl Microbiol 63:183–229. doi: 10.1016/S0065-2164(07)00005-6 [DOI] [PubMed] [Google Scholar]
- 10. Khider ML, Brautaset T, Irla M. 2021. Methane monooxygenases: central enzymes in methanotrophy with promising biotechnological applications. World J Microbiol Biotechnol 37:72. doi: 10.1007/s11274-021-03038-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Whiddon KT, Gudneppanavar R, Hammer TJ, West DA, Konopka MC. 2019. Fluorescence‐based analysis of the intracytoplasmic membranes of type I methanotrophs. Microb Biotechnol 12:1024–1033. doi: 10.1111/1751-7915.13458 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Khmelenina VN, But SY, Rozova ON, Trotsenko YA. 2019. Metabolic features of aerobic methanotrophs: news and views. Curr Issues Mol Biol 33:85–100. doi: 10.21775/cimb.033.085 [DOI] [PubMed] [Google Scholar]
- 13. Nguyen DTN, Lee OK, Nguyen TT, Lee EY. 2021. Type II methanotrophs: a promising microbial cell-factory platform for bioconversion of methane to chemicals. Biotechnol Adv 47:107700. doi: 10.1016/j.biotechadv.2021.107700 [DOI] [PubMed] [Google Scholar]
- 14. Auman AJ, Speake CC, Lidstrom ME. 2001. nifH sequences and nitrogen fixation in type I and type II methanotrophs . Appl Environ Microbiol 67:4009–4016. doi: 10.1128/AEM.67.9.4009-4016.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Oshkin IY, Miroshnikov KK, Grouzdev DS, Dedysh SN. 2020. Pan-genome-based analysis as a framework for demarcating two closely related methanotroph genera Methylocystis and Methylosinus. Microorganisms 8:768. doi: 10.3390/microorganisms8050768 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Pruitt KD, Tatusova T, Maglott DR. 2005. NCBI reference sequence (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 33:D501–D504. doi: 10.1093/nar/gki025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Benson DA, Karsch-Mizrachi I, Clark K, Lipman DJ, Ostell J, Sayers EW. 2012. GenBank. Nucleic Acids Res. 40:D48–D53. doi: 10.1093/nar/gkr1202 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM. 2015. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31:3210–3212. doi: 10.1093/bioinformatics/btv351 [DOI] [PubMed] [Google Scholar]
- 19. Seemann T. 2014. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. doi: 10.1093/bioinformatics/btu153 [DOI] [PubMed] [Google Scholar]
- 20. Gautreau G, Bazin A, Gachet M, Planel R, Burlot L, Dubois M, Perrin A, Médigue C, Calteau A, Cruveiller S, Matias C, Ambroise C, Rocha EPC, Vallenet D. 2020. PPanGGOLiN: depicting microbial diversity via a partitioned pangenome graph. PLoS Comput Biol 16:e1007732. doi: 10.1371/journal.pcbi.1007732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Saxena P, Rauniyar S, Thakur P, Singh RN, Bomgni A, Alaba MO, Tripathi AK, Gnimpieba EZ, Lushbough C, Sani RK. 2023. Integration of text mining and biological network analysis: identification of essential genes in sulfate-reducing bacteria. Front Microbiol 14:1086021. doi: 10.3389/fmicb.2023.1086021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Thakur P, Alaba MO, Rauniyar S, Singh RN, Saxena P, Bomgni A, Gnimpieba EZ, Lushbough C, Goh KM, Sani RK. 2023. Text-mining to identify gene sets involved in biocorrosion by sulfate-reducing bacteria: a semi-automated workflow. Microorganisms 11:119. doi: 10.3390/microorganisms11010119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering C von. 2019. STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613. doi: 10.1093/nar/gky1131 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. 2003. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13:2498–2504. doi: 10.1101/gr.1239303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Gene Ontology Consortium . 2004. The gene ontology (GO) database and informatics resource. Nucleic Acids Res 32:D258–D261. doi: 10.1093/nar/gkh036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Bastian M. 2017. Gephi (version 0.9. 2)
- 27. UniProt Consortium . 2019. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47:D506–D515. doi: 10.1093/nar/gky1049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Boutet E, Lieberherr D, Tognolli M, Schneider M, Bairoch A. 2007. UniProtKB/Swiss-Prot, p 89–112. In Edwards D (ed), Plant bioinformatics: methods and protocols. Humana Press, Totowa, NJ. [Google Scholar]
- 29. Kanehisa M, Sato Y. 2020. KEGG mapper for inferring cellular functions from protein sequences. Protein Sci 29:28–35. doi: 10.1002/pro.3711 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Kanehisa M, Sato Y, Morishima K. 2016. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J Mol Biol 428:726–731. doi: 10.1016/j.jmb.2015.11.006 [DOI] [PubMed] [Google Scholar]
- 31. Sanderson T, Bileschi ML, Belanger D, Colwell LJ. 2023. ProteInfer: deep networks for protein functional inference. Elife 12:e80942. doi: 10.7554/eLife.80942 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Kumar S, Tamura K, Nei M. 1994. MEGA: molecular evolutionary genetics analysis software for microcomputers. Bioinformatics 10:189–191. doi: 10.1093/bioinformatics/10.2.189 [DOI] [PubMed] [Google Scholar]
- 33. Gautreau G, Bazin A, Gachet M, Planel R, Burlot L, Dubois M, Perrin A, Médigue C, Calteau A, Cruveiller S, Matias C, Ambroise C, Rocha EPC, Vallenet D. 2020. PPanGGOLiN: depicting microbial diversity via a partitioned pangenome graph. PLoS Comput Biol 16:e1007732. doi: 10.1371/journal.pcbi.1007732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Pruitt KD, Tatusova T, Maglott DR. 2007. NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 35:D61–D65. doi: 10.1093/nar/gkl842 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Benson DA, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW. 2009. GenBank. Nucleic Acids Res 37:D26–D31. doi: 10.1093/nar/gkn723 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Aseev LV, Koledinskaya LS, Boni IV. 2020. Autogenous regulation in vivo of the rpmE gene encoding ribosomal protein L31 (bL31), a key component of the protein–protein intersubunit bridge B1b. RNA 26:814–826. doi: 10.1261/rna.074237.119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Wong KS, Snider JD, Graham C, Greenblatt JF, Emili A, Babu M, Houry WA. 2014. The MoxR ATPase RavA and its cofactor ViaA interact with the NADH: ubiquinone oxidoreductase I in Escherichia coli. PLoS ONE 9:e85529. doi: 10.1371/journal.pone.0085529 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Yang M, Zheng J, Jia H, Song M. 2016. Functional characterization of X-prolyl aminopeptidase from Toxoplasma gondii. Parasitology 143:1443–1449. doi: 10.1017/S0031182016000986 [DOI] [PubMed] [Google Scholar]
- 39. Leipe DD, Aravind L, Koonin EV. 1999. Did DNA replication evolve twice independently? Nucleic Acids Res. 27:3389–3401. doi: 10.1093/nar/27.17.3389 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Norais C, Servant P, Bouthier-de-la-Tour C, Coureux P-D, Ithurbide S, Vannier F, Guerin PP, Dulberger CL, Satyshur KA, Keck JL, Armengaud J, Cox MM, Sommer S. 2013. The Deinococcus radiodurans DR1245 protein, a DdrB partner homologous to YbjN proteins and reminiscent of type III secretion system chaperones. PLoS ONE 8:e56558. doi: 10.1371/journal.pone.0056558 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Notti RQ, Stebbins CE. 2016. The structure and function of type III secretion systems, p 241–264. In Virulence mechanisms of bacterial pathogens [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Anderson AJ, Dawes EA. 1990. Occurrence, metabolism, metabolic role, and industrial uses of bacterial polyhydroxyalkanoates. Microbiol Rev 54:450–472. doi: 10.1128/mr.54.4.450-472.1990 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Maehara A, Taguchi S, Nishiyama T, Yamane T, Doi Y. 2002. A repressor protein, PhaR, regulates polyhydroxyalkanoate (PHA) synthesis via its direct interaction with PHA. J Bacteriol 184:3992–4002. doi: 10.1128/JB.184.14.3992-4002.2002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Kuznetsova E, Nocek B, Brown G, Makarova KS, Flick R, Wolf YI, Khusnutdinova A, Evdokimova E, Jin K, Tan K, Hanson AD, Hasnain G, Zallot R, de Crécy-Lagard V, Babu M, Savchenko A, Joachimiak A, Edwards AM, Koonin EV, Yakunin AF. 2015. Functional diversity of haloacid dehalogenase superfamily phosphatases from Saccharomyces cerevisiae: biochemical, structural, and evolutionary insights. J Biol Chem 290:18678–18698. doi: 10.1074/jbc.M115.657916 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Yusupova YR, Skripnikova VS, Kivero AD, Zakataeva NP. 2020. Expression and purification of the 5′-nucleotidase YitU from Bacillus species: its enzymatic properties and possible applications in biotechnology. Appl Microbiol Biotechnol 104:2957–2972. doi: 10.1007/s00253-020-10428-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Deutscher J, Francke C, Postma PW. 2006. How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria. Microbiol Mol Biol Rev 70:939–1031. doi: 10.1128/MMBR.00024-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Rodionova IA, Zhang Z, Mehla J, Goodacre N, Babu M, Emili A, Uetz P, Saier MH. 2017. The phosphocarrier protein HPR of the bacterial phosphotransferase system globally regulates energy metabolism by directly interacting with multiple enzymes in Escherichia coli. J Biol Chem 292:14250–14257. doi: 10.1074/jbc.M117.795294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Deutscher J, Reizer J, Fischer C, Galinier A, Saier MH, Steinmetz M. 1994. Loss of protein kinase-catalyzed phosphorylation of HPr, a phosphocarrier protein of the phosphotransferase system, by mutation of the ptsH gene confers catabolite repression resistance to several catabolic genes of Bacillus subtilis. J Bacteriol 176:3336–3344. doi: 10.1128/jb.176.11.3336-3344.1994 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Macek B, Forchhammer K, Hardouin J, Weber-Ban E, Grangeasse C, Mijakovic I. 2019. Protein post-translational modifications in bacteria. Nat Rev Microbiol 17:651–664. doi: 10.1038/s41579-019-0243-0 [DOI] [PubMed] [Google Scholar]
- 50. Fuhrmann JJ. 2021. Microbial metabolism, p 57–87. In Principles and applications of soil microbiology. Elsevier. [Google Scholar]
- 51. Chistoserdova L, Kalyuzhnaya MG, Lidstrom ME. 2009. The expanding world of methylotrophic metabolism. Annu Rev Microbiol 63:477–499. doi: 10.1146/annurev.micro.091208.073600 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Naizabekov S, Lee EY. 2020. Genome-scale metabolic model reconstruction and in silico investigations of methane metabolism in Methylosinus trichosporium OB3b. Microorganisms 8:437. doi: 10.3390/microorganisms8030437 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Nguyen DTN, Lee OK, Lim C, Lee J, Na J-G, Lee EY. 2020. Metabolic engineering of type II methanotroph, Methylosinus trichosporium OB3b, for production of 3-hydroxypropionic acid from methane via a malonyl-CoA reductase-dependent pathway. Metab Eng 59:142–150. doi: 10.1016/j.ymben.2020.02.002 [DOI] [PubMed] [Google Scholar]
- 54. Ljungman M. 2010. The DNA damage response—repair or despair? Environ Mol Mutagen 51:879–889. doi: 10.1002/em.20597 [DOI] [PubMed] [Google Scholar]
- 55. Mott ML, Berger JM. 2007. DNA replication initiation: mechanisms and regulation in bacteria. Nat Rev Microbiol 5:343–354. doi: 10.1038/nrmicro1640 [DOI] [PubMed] [Google Scholar]
- 56. Copley SD, Smith E, Morowitz HJ. 2007. The origin of the RNA world: co-evolution of genes and metabolism. Bioorg Chem 35:430–443. doi: 10.1016/j.bioorg.2007.08.001 [DOI] [PubMed] [Google Scholar]
- 57. Goncheva MI, Chin D, Heinrichs DE. 2022. Nucleotide biosynthesis: the base of bacterial pathogenesis. Trends Microbiol. 30:793–804. doi: 10.1016/j.tim.2021.12.007 [DOI] [PubMed] [Google Scholar]
- 58. Hirasawa T, Shimizu H. 2016. Recent advances in amino acid production by microbial cells. Curr Opin Biotechnol 42:133–146. doi: 10.1016/j.copbio.2016.04.017 [DOI] [PubMed] [Google Scholar]
- 59. Zhang Y-M, Rock CO. 2008. Membrane lipid homeostasis in bacteria. Nat Rev Microbiol 6:222–233. doi: 10.1038/nrmicro1839 [DOI] [PubMed] [Google Scholar]
- 60. Drackley, JK. "Lipid Metabolism." Farm animal metabolism and nutrition 1 (2000): 97-119. [Google Scholar]
- 61. Egan AJ, Vollmer W. 2013. The physiology of bacterial cell division. Ann N Y Acad Sci 1277:8–28. doi: 10.1111/j.1749-6632.2012.06818.x [DOI] [PubMed] [Google Scholar]
- 62. Banerjee A, Sharkey TD. 2014. Methylerythritol 4-phosphate (MEP) pathway metabolic regulation. Nat Prod Rep 31:1043–1055. doi: 10.1039/c3np70124g [DOI] [PubMed] [Google Scholar]
- 63. Bergman J. 1999. ATP: the perfect energy currency for the cell. Creation Research Society Quarterly 36:2–9. [Google Scholar]
- 64. Ardito F, Giuliani M, Perrone D, Troiano G, Lo Muzio L. 2017. The crucial role of protein phosphorylation in cell signaling and its use as targeted therapy. Int J Mol Med 40:271–280. doi: 10.3892/ijmm.2017.3036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Connolly T, Gilmore R. 1986. Formation of a functional ribosome-membrane junction during translocation requires the participation of a GTP-binding protein. J Cell Biol 103:2253–2261. doi: 10.1083/jcb.103.6.2253 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Hartwig A. 2001. Role of magnesium in genomic stability. Mutat Res 475:113–121. doi: 10.1016/s0027-5107(01)00074-4 [DOI] [PubMed] [Google Scholar]
- 67. Lill Roland. 2009. Function and biogenesis of iron–sulphur proteins. Nature 460:831–838. doi: 10.1038/nature08301 [DOI] [PubMed] [Google Scholar]
- 68. Marceau AH. 2012. Functions of single-strand DNA-binding proteins in DNA replication, recombination, and repair, p 1–21. In Keck JL (ed), Single-stranded DNA binding proteins: methods and protocols. Humana Press, Totowa, NJ. [DOI] [PubMed] [Google Scholar]
- 69. Vetter IR, Wittinghofer A. 1999. Nucleoside triphosphate-binding proteins: different scaffolds to achieve phosphoryl transfer. Quart Rev Biophys 32:1–56. doi: 10.1017/S0033583599003480 [DOI] [PubMed] [Google Scholar]
- 70. Yang J, Zhou R, Ma Z. 2019. Autophagy and energy metabolism, p 329–357. In Qin ZH (ed), Autophagy: biology and diseases. Springer Singapore, Singapore. [DOI] [PubMed] [Google Scholar]
- 71. Vick JE, Clomburg JM, Blankschien MD, Chou A, Kim S, Gonzalez R. 2015. Escherichia coli enoyl-acyl carrier protein reductase (FabI) supports efficient operation of a functional reversal of the β-oxidation cycle. Appl Environ Microbiol 81:1406–1416. doi: 10.1128/AEM.03521-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Giri DD, Singh SK, Giri A, Dwivedi H, Kumar A. 2021. Bioremediation potential of methylotrophic bacteria, p 199–207. In Microbe mediated remediation of environmental contaminants. Elsevier. [Google Scholar]
- 73. Cui J, Zhang M, Chen L, Zhang S, Luo Y, Cao W, Zhao J, Wang L, Jia Z, Bao Z. 2022. Methanotrophs contribute to nitrogen fixation in emergent macrophytes. Front Microbiol 13:851424. doi: 10.3389/fmicb.2022.851424 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Cui J, Zhao J, Wang Z, Cao W, Zhang S, Liu J, Bao Z. 2020. Diversity of active root-associated methanotrophs of three emergent plants in a eutrophic wetland in northern China. AMB Express 10:48. doi: 10.1186/s13568-020-00984-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Bao Z, Watanabe A, Sasaki K, Okubo T, Tokida T, Liu D, Ikeda S, Imaizumi-Anraku H, Asakawa S, Sato T, Mitsui H, Minamisawa K. 2014. A rice gene for microbial symbiosis, Oryza sativa CCaMK, reduces CH4 flux in a paddy field with low nitrogen input. Appl Environ Microbiol 80:1995–2003. doi: 10.1128/AEM.03646-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Rubio LM, Ludden PW. 2005. Maturation of nitrogenase: a biochemical puzzle. J Bacteriol 187:405–414. doi: 10.1128/JB.187.2.405-414.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Lei S, Pulakat L, Gavini N. 2000. Activation of vanadium nitrogenase expression in Azotobacter vinelandii DJ54 revertant in the presence of molybdenum. FEBS Lett 482:149–153. doi: 10.1016/s0014-5793(00)02052-4 [DOI] [PubMed] [Google Scholar]
- 78. Hu Y, Lee CC, Ribbe MW. 2012. Vanadium nitrogenase: a two-hit wonder? Dalton Trans 41:1118–1127. doi: 10.1039/c1dt11535a [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Oshkin IY, Miroshnikov KK, Danilova OV, Hakobyan A, Liesack W, Dedysh SN. 2019. Thriving in wetlands: ecophysiology of the spiral-shaped methanotroph Methylospira mobilis as revealed by the complete genome sequence. Microorganisms 7:683. doi: 10.3390/microorganisms7120683 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Han D, Dedysh SN, Liesack W. 2018. Unusual genomic traits suggest Methylocystis bryophila S285 to be well adapted for life in peatlands. Genome Biol Evol 10:623–628. doi: 10.1093/gbe/evy025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Hanson RS, Hanson TE. 1996. Methanotrophic bacteria. Microbiol Rev 60:439–471. doi: 10.1128/mr.60.2.439-471.1996 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Wang Y, Fan L, Tuyishime P, Zheng P, Sun J. 2020. Synthetic methylotrophy: a practical solution for methanol-based biomanufacturing. Trends Biotechnol. 38:650–666. doi: 10.1016/j.tibtech.2019.12.013 [DOI] [PubMed] [Google Scholar]
- 83. Dunfield PF, Belova SE, Vorob’ev AV, Cornish SL, Dedysh SN. 2010. Methylocapsa aurea sp. nov., a facultative methanotroph possessing a particulate methane monooxygenase, and emended description of the genus Methylocapsa. Int J Syst Evol Microbiol. 60:2659–2664. doi: 10.1099/ijs.0.020149-0 [DOI] [PubMed] [Google Scholar]
- 84. DiSpirito AA, Semrau JD, Murrell JC, Gallagher WH, Dennison C, Vuilleumier S. 2016. Methanobactin and the link between copper and bacterial methane oxidation. Microbiol Mol Biol Rev 80:387–409. doi: 10.1128/MMBR.00058-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Farhan Ul Haque M, Xu H-J, Murrell JC, Crombie A. 2020. Facultative methanotrophs–diversity, genetics, molecular ecology and biotechnological potential: a mini-review. Microbiology 166:894–908. doi: 10.1099/mic.0.000977 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Villada JC, Duran MF, Lee PK. 2019. Genomic evidence for simultaneous optimization of transcription and translation through codon variants in the pmoCAB operon of type Ia methanotrophs. mSystems 4:e00342. doi: 10.1128/mSystems.00342-19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Keltjens JT, Pol A, Reimann J, Op den Camp HJM. 2014. PQQ-dependent methanol dehydrogenases: rare-earth elements make a difference. Appl Microbiol Biotechnol 98:6163–6183. doi: 10.1007/s00253-014-5766-8 [DOI] [PubMed] [Google Scholar]
- 88. Chu F, Lidstrom ME. 2016. XoxF acts as the predominant methanol dehydrogenase in the type I methanotroph Methylomicrobium buryatense. J Bacteriol 198:1317–1325. doi: 10.1128/JB.00959-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Pol A, Barends TRM, Dietl A, Khadem AF, Eygensteyn J, Jetten MSM, Op den Camp HJM. 2014. Rare earth metals are essential for methanotrophic life in volcanic mudpots. Environ Microbiol 16:255–264. doi: 10.1111/1462-2920.12249 [DOI] [PubMed] [Google Scholar]
- 90. Le T-K, Lee Y-J, Han GH, Yeom S-J. 2021. Methanol dehydrogenases as a key biocatalysts for synthetic methylotrophy. Front Bioeng Biotechnol 9:787791. doi: 10.3389/fbioe.2021.787791 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Anthony C. 2011. How half a century of research was required to understand bacterial growth on C1 and C2 compounds; the story of the serine cycle and the ethylmalonyl-CoA pathway. Sci Prog 94:109–137. doi: 10.3184/003685011X13044430633960 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Liesivuori J, Savolainen AH. 1991. Methanol and formic acid toxicity: biochemical mechanisms. Pharmacol Toxicol 69:157–163. doi: 10.1111/j.1600-0773.1991.tb01290.x [DOI] [PubMed] [Google Scholar]
- 93. O’Connor ML, Hanson RS. 1975. Serine transhydroxymethylase isoenzymes from a facultative methylotroph. J Bacteriol 124:985–996. doi: 10.1128/jb.124.2.985-996.1975 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Quayle JR. 1980. Microbial assimilation of C1 compounds. Biochem Soc Trans 8:1–10. doi: 10.1042/bst0080001 [DOI] [PubMed] [Google Scholar]
- 95. Samanta D, Singh RN, Goh KM, Sani RK. 2024. Integrating metabolomics and whole genome sequencing to elucidate the metabolic pathways in Methylosinus trichosporium OB3b. Syst Microbiol Biomanufacturing 4:661–674. doi: 10.1007/s43393-023-00214-y [DOI] [Google Scholar]
- 96. Caspi R. MetaCyc. SRI International. Available from: https://biocyc.org/META/NEW-IMAGE?type=PATHWAY&object=PWY-1622 [Google Scholar]
- 97. Barta TM, Hanson RS. 1993. Genetics of methane and methanol oxidation in Gram-negative methylotrophic bacteria. Antonie Van Leeuwenhoek 64:109–120. doi: 10.1007/BF00873021 [DOI] [PubMed] [Google Scholar]
- 98. Schneider K, Peyraud R, Kiefer P, Christen P, Delmotte N, Massou S, Portais JC, Vorholt JA. 2012. The ethylmalonyl-CoA pathway is used in place of the glyoxylate cycle by Methylobacterium extorquens AM1 during growth on acetate. J Biol Chem 287:757–766. doi: 10.1074/jbc.M111.305219 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99. Kunze M, Pracharoenwattana I, Smith SM, Hartig A. 2006. A central role for the peroxisomal membrane in glyoxylate cycle function. Biochim Biophys Acta 1763:1441–1452. doi: 10.1016/j.bbamcr.2006.09.009 [DOI] [PubMed] [Google Scholar]
- 100. Alber BE, Spanheimer R, Ebenau-Jehle C, Fuchs G. 2006. Study of an alternate glyoxylate cycle for acetate assimilation by Rhodobacter sphaeroides. Mol Microbiol 61:297–309. doi: 10.1111/j.1365-2958.2006.05238.x [DOI] [PubMed] [Google Scholar]
- 101. Ahn JW, Hong J, Kim KJ. 2023. Crystal structure of mesaconyl-CoA hydratase from Methylorubrum extorquens CM4. J Microbiol Biotechnol 33:485–492. doi: 10.4014/jmb.2212.12003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Khomyakova M, Bükmez Ö, Thomas LK, Erb TJ, Berg IA. 2011. A methylaspartate cycle in haloarchaea. Science 331:334–337. doi: 10.1126/science.1196544 [DOI] [PubMed] [Google Scholar]
- 103. Orlowska K, Fling RR, Nault R, Sink WJ, Schilmiller AL, Zacharewski T. 2022. Dioxin-elicited decrease in cobalamin redirects hepatic propionyl-CoA metabolism to the β–oxidation-like pathway resulting in acrylyl-CoA conjugate accumulation. J Biol Chem 298:102301. doi: 10.1016/j.jbc.2022.102301 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104. Blažič M, Kosec G, Baebler Š, Gruden K, Petković H. 2015. Roles of the crotonyl-CoA carboxylase/reductase homologues in acetate assimilation and biosynthesis of immunosuppressant FK506 in Streptomyces tsukubaensis. Microb Cell Fact 14:164. doi: 10.1186/s12934-015-0352-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105. Mai DHA, Nguyen TT, Lee EY. 2021. The ethylmalonyl-CoA pathway for methane-based biorefineries: a case study of using Methylosinus trichosporium OB3b, an alpha-proteobacterial methanotroph, for producing 2-hydroxyisobutyric acid and 1,3-butanediol from methane . Green Chem 23:7712–7723. doi: 10.1039/D1GC02866A [DOI] [Google Scholar]
- 106. Wongkittichote P, Ah Mew N, Chapman KA. 2017. Propionyl-CoA carboxylase - a review. Mol Genet Metab 122:145–152. doi: 10.1016/j.ymgme.2017.10.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107. Erb TJ, Fuchs G, Alber BE. 2009. (2S)-methylsuccinyl-CoA dehydrogenase closes the ethylmalonyl-CoA pathway for acetyl-CoA assimilation. Mol Microbiol 73:992–1008. doi: 10.1111/j.1365-2958.2009.06837.x [DOI] [PubMed] [Google Scholar]
- 108. Alessa O, Ogura Y, Fujitani Y, Takami H, Hayashi T, Sahin N, Tani A. 2021. Comprehensive comparative genomics and phenotyping of Methylobacterium species. Front Microbiol 12:740610. doi: 10.3389/fmicb.2021.740610 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109. Sun J, Steindler L, Thrash JC, Halsey KH, Smith DP, Carter AE, Landry ZC, Giovannoni SJ. 2011. One carbon metabolism in SAR11 pelagic marine bacteria. PLoS One 6:e23973. doi: 10.1371/journal.pone.0023973 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110. Verma P, Suman A. 2018. Wheat microbiomes: ecological significances, molecular diversity and potential bioresources for sustainable agriculture. EC Microbiol 14:641–665. [Google Scholar]
- 111. Verma P, Yadav AN, Khannam KS, Panjiar N, Kumar S, Saxena AK, Suman A. 2015. Assessment of genetic diversity and plant growth promoting attributes of psychrotolerant bacteria allied with wheat (Triticum aestivum) from the northern hills zone of India. Ann Microbiol 65:1885–1899. doi: 10.1007/s13213-014-1027-4 [DOI] [Google Scholar]
- 112. Tamosiune I, Baniulis D, Stanys V. 2017. Role of endophytic bacteria in stress tolerance of agricultural plants: diversity of microorganisms and molecular mechanisms, p 1–29. In Probiotics in agroecosystem [Google Scholar]
- 113. Dedysh SN. 2009. Exploring methanotroph diversity in acidic northern wetlands: molecular and cultivation-based studies. Microbiology 78:655–669. doi: 10.1134/S0026261709060010 [DOI] [Google Scholar]
- 114. Haoxin LV. 2018. Exploration of lanthanide-dependent methylotrophic bacteria [DOI] [PubMed] [Google Scholar]
- 115. Higgins IJ, Best DJ, Hammond RC, Scott . 1981. Methane-oxidizing microorganisms. Microbiol Rev 45:556–590. doi: 10.1128/mr.45.4.556-590.1981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116. Patt TE, Cole GC, Hanson RS. 1976. Methylobacterium, a new genus of facultatively methylotrophic bacteria. Int J Syst Bacteriol 26:226–229. doi: 10.1099/00207713-26-2-226 [DOI] [Google Scholar]
- 117. Bijlani S, Singh NK, Eedara VVR, Podile AR, Mason CE, Wang CCC, Venkateswaran K. 2021. Methylobacterium ajmalii sp. nov., isolated from the international space station. Front Microbiol 12:639396. doi: 10.3389/fmicb.2021.639396 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118. Vadivukkarasi P, Jayashree S, Seshadri S. 2018. Occurrence and ecological significance of Methylobacterium. Trop Ecol 59:575–587. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Comprehensive visualization of the screening process, depicting the evaluation of each genome's completeness percentage.
Visualization of the intricate interconnections and collaborative relationships among the GO terms.
Statistical analysis of the 256 nodes determined using Gephi.
Distribution of genes across taxonomic groups and cellular metabolic pathways.
Legends to supplemental figures and tables.
The complete data set comprising 75 genomes, including information on genome size, GC content, genome coverage, and isolation site.
Detailed UniProt entries and descriptions of 256 exact core gene families.
Categorizing the 256 exact core genes into 31 distinct categories with their annotation, number of organisms, and number of sequences.
A comprehensive overview of the detailed FASTA sequence, and corresponding scores associated with the annotation of the hypothetical proteins.
Data Availability Statement
The original contributions presented in the study are included in the article and supplemental material. The PPanGGOLiN algorithm is available for access at https://github.com/labgem/PPanGGOLiN.