Abstract
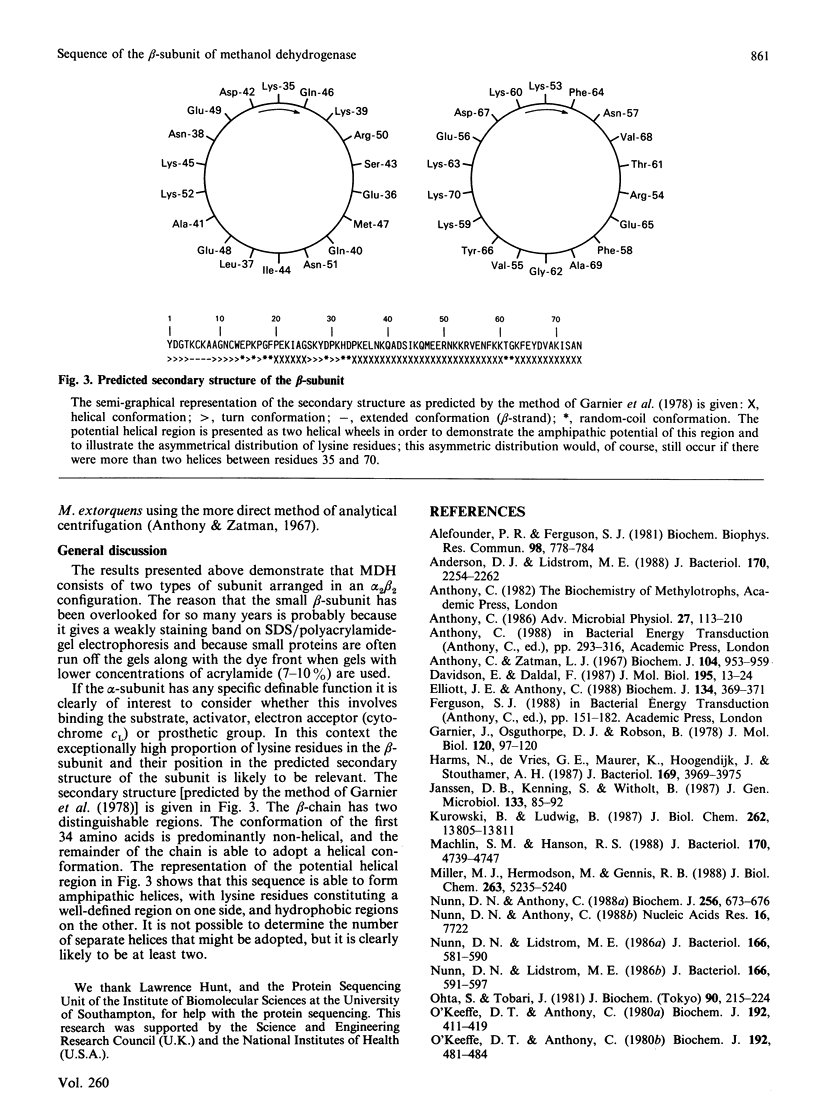
The nucleotide and deduced amino acid sequence of a novel small (beta) subunit of methanol dehydrogenase of Methylobacterium extorquens AM1 (previously Pseudomonas AM1) has been determined. Work with the whole protein has shown that is has an alpha 2 beta 2 configuration.
Full text
PDF




Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alefounder P. R., Ferguson S. J. A periplasmic location for methanol dehydrogenase from Paracoccus denitrificans: implications for proton pumping by cytochrome aa3. Biochem Biophys Res Commun. 1981 Feb 12;98(3):778–784. doi: 10.1016/0006-291x(81)91179-7. [DOI] [PubMed] [Google Scholar]
- Anderson D. J., Lidstrom M. E. The moxFG region encodes four polypeptides in the methanol-oxidizing bacterium Methylobacterium sp. strain AM1. J Bacteriol. 1988 May;170(5):2254–2262. doi: 10.1128/jb.170.5.2254-2262.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anthony C. Bacterial oxidation of methane and methanol. Adv Microb Physiol. 1986;27:113–210. doi: 10.1016/s0065-2911(08)60305-7. [DOI] [PubMed] [Google Scholar]
- Anthony C., Zatman L. J. The microbial oxidation of methanol. Purification and properties of the alcohol dehydrogenase of Pseudomonas sp. M27. Biochem J. 1967 Sep;104(3):953–959. doi: 10.1042/bj1040953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davidson E., Daldal F. Primary structure of the bc1 complex of Rhodopseudomonas capsulata. Nucleotide sequence of the pet operon encoding the Rieske cytochrome b, and cytochrome c1 apoproteins. J Mol Biol. 1987 May 5;195(1):13–24. doi: 10.1016/0022-2836(87)90323-8. [DOI] [PubMed] [Google Scholar]
- Garnier J., Osguthorpe D. J., Robson B. Analysis of the accuracy and implications of simple methods for predicting the secondary structure of globular proteins. J Mol Biol. 1978 Mar 25;120(1):97–120. doi: 10.1016/0022-2836(78)90297-8. [DOI] [PubMed] [Google Scholar]
- Harms N., de Vries G. E., Maurer K., Hoogendijk J., Stouthamer A. H. Isolation and nucleotide sequence of the methanol dehydrogenase structural gene from Paracoccus denitrificans. J Bacteriol. 1987 Sep;169(9):3969–3975. doi: 10.1128/jb.169.9.3969-3975.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurowski B., Ludwig B. The genes of the Paracoccus denitrificans bc1 complex. Nucleotide sequence and homologies between bacterial and mitochondrial subunits. J Biol Chem. 1987 Oct 5;262(28):13805–13811. [PubMed] [Google Scholar]
- Machlin S. M., Hanson R. S. Nucleotide sequence and transcriptional start site of the Methylobacterium organophilum XX methanol dehydrogenase structural gene. J Bacteriol. 1988 Oct;170(10):4739–4747. doi: 10.1128/jb.170.10.4739-4747.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller M. J., Hermodson M., Gennis R. B. The active form of the cytochrome d terminal oxidase complex of Escherichia coli is a heterodimer containing one copy of each of the two subunits. J Biol Chem. 1988 Apr 15;263(11):5235–5240. [PubMed] [Google Scholar]
- Nunn D. N., Anthony C. The nucleotide sequence and deduced amino acid sequence of the cytochrome cL gene of Methylobacterium extorquens AM1, a novel class of c-type cytochrome. Biochem J. 1988 Dec 1;256(2):673–676. doi: 10.1042/bj2560673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunn D. N., Anthony C. The nucleotide sequence and deduced amino acid sequence of the genes for cytochrome cL and a hypothetical second subunit of the methanol dehydrogenase of Methylobacterium AM1. Nucleic Acids Res. 1988 Aug 11;16(15):7722–7722. doi: 10.1093/nar/16.15.7722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunn D. N., Lidstrom M. E. Isolation and complementation analysis of 10 methanol oxidation mutant classes and identification of the methanol dehydrogenase structural gene of Methylobacterium sp. strain AM1. J Bacteriol. 1986 May;166(2):581–590. doi: 10.1128/jb.166.2.581-590.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunn D. N., Lidstrom M. E. Phenotypic characterization of 10 methanol oxidation mutant classes in Methylobacterium sp. strain AM1. J Bacteriol. 1986 May;166(2):591–597. doi: 10.1128/jb.166.2.591-597.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Keeffe D. T., Anthony C. The two cytochromes c in the facultative methylotroph Pseudomonas am1. Biochem J. 1980 Nov 15;192(2):411–419. doi: 10.1042/bj1920411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta S., Tobari J. Two cytochromes c of Methylomonas J. J Biochem. 1981 Jul;90(1):215–224. doi: 10.1093/oxfordjournals.jbchem.a133452. [DOI] [PubMed] [Google Scholar]
- Raftery M. A., Hunkapiller M. W., Strader C. D., Hood L. E. Acetylcholine receptor: complex of homologous subunits. Science. 1980 Jun 27;208(4451):1454–1456. doi: 10.1126/science.7384786. [DOI] [PubMed] [Google Scholar]
- Steinrücke P., Steffens G. C., Panskus G., Buse G., Ludwig B. Subunit II of cytochrome c oxidase from Paracoccus denitrificans. DNA sequence, gene expression and the protein. Eur J Biochem. 1987 Sep 15;167(3):431–439. doi: 10.1111/j.1432-1033.1987.tb13356.x. [DOI] [PubMed] [Google Scholar]
- Woese C. R. Bacterial evolution. Microbiol Rev. 1987 Jun;51(2):221–271. doi: 10.1128/mr.51.2.221-271.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]


