Abstract
Abnormal interaction between granulosa cells and oocytes causes disordered development of ovarian follicles. However, the interactions between oocytes and cumulus granulosa cells (CGs), oocytes and mural granulosa cells (MGs), and CGs and MGs remain to be fully explored. Using single-cell RNA-sequencing (scRNA-seq), we determined the transcriptional profiles of oocytes, CGs and MGs in antral follicles. Analysis of scRNA-seq data revealed that CGs may regulate follicular development through the BMP15-KITL-KIT-PI3K-ARF6 pathway with elevated expression of luteinizing hormone receptor (LHR). Because internalization of the LHR is regulated by Arf6, we constructed LHRN316S mice by CRISPR/Cas9 to further explore mechanisms of follicular development and novel treatment strategies for female infertility. Ovaries of LHRN316S mice exhibited reduced numbers of corpora lutea and ovulation. The LHRN316S mice had a reduced rate of oocyte maturation in vitro and decreased serum progesterone levels. Mating LHRN316S female mice with ICR wild type male mice revealed that the infertility rate of LHRN316S mice was 21.4% (3/14). Litter sizes from LHRN316S mice were smaller than those from control wild type female mice. The oocytes from LHRN316S mice had an increased rate of maturation in vitro after progesterone administration in vitro. Furthermore, progesterone treated LHRN316S mice produced offspring numbers per litter equivalent to WT mice. These findings provide key insights into cellular interactions in ovarian follicles and provide important clues for infertility treatment.
Graphical Abstract
Supplementary Information
The online version contains supplementary material available at 10.1007/s12539-024-00646-7.
Keywords: Oocytes, Granulosa cells, Follicle, LHRN316S, Single cell RNA-seq, Progesterone
Introduction
In mammalian reproduction, the ovarian follicle comprises a central oocyte with surrounding granulosa cells (GCs) and theca cells [1, 2]. The GCs regulate oocyte cortex dynamics and germinal vesicle (GV) position, anchor and spindle during oocyte maturation [3–5]. Bidirectional communication between oocytes and GCs contributes to oocyte development [1, 2, 6, 7], and abnormal interaction of this system causes disordered follicle development [8–10]. The GCs from antral follicles include both cumulus granulosa cells (CGs) and mural granulosa cells (MGs). MGs are located in the outer layer of GCs in antral follicles, and CGs closely encircle the oocyte. The two types of GCs have different responsiveness to extracellular signals, and different roles and fates in follicular development [11–14]. The MGs produce estrogen, which triggers the pituitary to secrete luteinizing hormone (LH), resulting in the promotion of oocyte maturation, cumulus expansion and ovulation. The CGs promote oocyte growth and maturation [15–17]. In a previous study, interactive regulation between oocytes and GCs during follicular development was shown at the transcriptional level via RNA-sequencing (RNA-seq) analysis [18]. However, the interactions between oocytes and CGs, oocytes and MGs, as well as CGs and MGs remain to be fully explored. Single cell RNA-seq (scRNA-seq) of the antral follicle would provide a targeted means to analyze these cellular interactions.
ADP-ribosylation factor 6 (Arf6) is a member of the ARF family of small GTPases that regulates multiple cellular events, cycling between active GTP and inactive GDP-bound forms [19]. Cavenagh et al. reported high expression levels of Arf6 in the human ovary [20]. Furthermore, it was reported that Arf6 was necessary for oocyte maturation [21]. Interestingly, Arf6 regulates the internalization of the luteinizing hormone receptor (LHR) and other G protein-coupled receptors (GPCRs) [22]. Previous studies showed that disruption of Arf6 in mice was fatal to embryos [23]. Therefore, Arf6 knockout mice do not provide a feasible model to study the effects of Arf6 on antral follicular development.
The LHR is essential for ovulation, progesterone production [24] and fertility [25]. In the ovary, LHR is present in theca cells, GCs, stromal cells and luteinized cells. Mutations and polymorphisms of LHR have been shown to cause infertility, subfertility and poor outcomes of fertility treatment [26]. In humans, the LHRN312S (rs2293275) variant is one of the most frequently studied polymorphic sites [27], LHCGR N312 may also be helpful for predicting reproductive outcomes in ART [28–30]. However, the underlying mechanism of the action of LHRN312S impacting female fertility is unknown, and there is no clear strategy for the treatment of female infertility caused by LHRN312S. In mice, the LHRN316S site corresponds to the LHRN312S site in humans and may provide a valuable model to study the potential mechanisms of LHRN316S actions on female fertility.
The current study detected the transcriptional profiles of oocytes, CGs and MGs in antral follicles by scRNA-seq and analyzed the interactions between these profiles. Then, we generated LHRN316S mice by CRISPR/Cas9 technology to explore the mechanisms of follicular development and a novel treatment strategy for female infertility.
Experiments
Animals
All procedures regarding to the animal experiment were approved by the Institutional Animal Care and Use Committee of Shanghai, and were conducted in line with the National Research Council Guide for Care and Use of Laboratory Animals. C57BL/6J and ICR mice were supplied by the company of Shanghai SLAC laboratory animal, China.
Isolation of Single Cells from Individual Antral Follicles
Female C57BL/6J mice at 6 weeks old were anesthetized and euthanized according to the experimental guidelines of animals in Shanghai Jiao Tong University. The ovaries were harvested and put into pre-cold phosphate-buffered saline (PBS). After that, mice ovaries were washed with pre-cold PBS and separated into several pieces under a stereomicroscope using a 29 G1/2 needle attached to an insulin syringe. Then, individual antral follicles were picked up with a mouth pipette and washed with cold PBS at least three times to remove any contaminating small cells. For collection of single cells, an individual antral follicle was transferred into a drop of PBS and gently punctured by a 29 G1/2 needle. Clumps of MGs and COCs were transferred into different drops of PBS. After several washes in PBS, MGs were scattered and a single cell was selected under a manipulation system. The COC was washed thoroughly in PBS with 1% hyaluronidase to separate CGs and the oocyte. A single oocyte or single CG was selected with a mouth pipette or manipulation system. Two oocytes were obtained from other follicles of similar size in repeated experiments.
cDNA Preparation and Library Preparation for Next Generation Sequencing
An individual cell was selected as described above and transferred into commercialized lysis buffer. The cDNA of single cell was reverse-transcribed with cell lysis using a SMARTer® Ultra™ Low Input RNA for Sequencing kit (Clontech Laboratories, Inc., Mountain View, CA) following the manuals. Briefly, RNA with polyA was reverse transcribed to cDNA using enzyme and primers for reverse transcription in the kit mentioned above. cDNA was amplified with 18 PCR cycles and purified step by step following the manuals. The amount and quality of purified cDNA libraries were determined by Agilent 2100. Fity nanogram tagmentation cDNA was prepared using Covaris S2 and a commercialized kit named NEBNext® Ultra™ for Illumina platform. The tagmentation cDNA was ligated with adapters after terminus repairing and dA tailing. Then, the cDNA with adapters was amplified with 12–15 PCR cycles. Paired-end sequencing was performed on a HiSeq 2500.
Read Mapping and Quantification of Gene Expression Levels
Raw data of each sample in FASTQ files were first processed into clean data by filtering the low-quality reads, reads with adapter and ploy-N. The quality of clean data was assessed based on the sequencing quality score of 20, of 30 and GC content. All the clean data was qualified with high quality and was used for following analysis.
Reference genome of mice and the annotation files were downloaded from the genomic libraries. Bowtie2 with version of v2.2.5 was used for building a reference genomic index. TopHat v2.0.14 was used for aligning the paired-end clean reads to the reference genome. The number of read mapping to each gene was counted by HTSeq v0.6.1. Then, the FPKM of each gene was determined in all the samples. A flow chart for the data analysis was shown in Fig. S1. A list of the package and software to analyze the data was shown in Supplementary Table S1.
Weighted Gene Co-expression Network Analysis (WGCNA)
Genes expressed at FPKM value ≥ 0.1 in any of the samples were used for constructing a signed network. Briefly, the correlation between the expression values of gene i and the eigengene of module q was defined as MMq(i) = cor[x(i), Eq], where x(i) is the expression profile of gene i and Eq is the eigengene of module q [31]. Modules were clustered using the method of dynamic hybrid cut.
Gene co-expression networks, which could assess the intensity of gene interactions, were built according to the expression levels of genes (FPKM). The network was constructed based on the significant Pearson correlation pairs of genes [32]. Degree centrality, which determines relative importance of gene node, was defined as the number of links one node has to the others within a network [33, 34]. K-cores were used to find the very important subgraphs in the co-expression networks.
Gene Ontology Term and Pathway Enrichment Analysis
The main biological domain of the differential expression genes was analyzed by GO analysis, based on the Wey functional classification of NCBI [35]. The significant enrichment of GO terms was scored by hypergeometric test. False discovery rates were estimated using either Bonferonni or Benjamini–Hochberg procedures.
The significant biological functions of differentially expressed genes were analyzed by pathway enrichment analysis, based on the Fisher exact test. The P adjusted values were calculated by the algorithm of BH FDR [36]. The FDR values of all the reported pathway categories were less than 0.05.
Single-Cell Quantitative PCR
Single cell qPCR was applied for independent of the RNA-seq analyzed embryos, another four CGs, six MGs and five oocytes. Each single cell was snap frozen and stored at − 80 °C. HiScript II Q RT SuperMix for qPCR (+gDNA wiper) kit (Vazyme, China) was used for cell reverse transcription. Fast Start Universal SYBR Green Master Mix kits (Roche, Germany) was applied for primer-specific amplification with ABI PRISM 7500 system (Applied Biosystems, USA). The primers are shown in Supplementary Table S12.
Construction of LHRN316S Mutant Mice
Methods for generation of the point mutation in mice were described previously with modifications [37]. Briefly, Tris–EDTA buffer containing 50 ng/μg Cas9 mRNA, 50 ng/μL sgRNA1, 50 ng/μg sgRNA2 and 100 ng/μL ssODN donor was injected into the pronuclei of one-cell stage mouse embryos using an Eppendorf transfer Man NK2 micromanipulator. Injected embryos were transferred into KSOM medium and cultured overnight before transfer into pseudopregnant C57BL/6 female mice. Offspring tail-tip biopsies were subjected to PCR amplification using a primer pair specific for Lhr to amplify an Lhr fragment, and LHRN316S mutant mice were identified by genomic sequencing. C57BL/6 LHRN316S mice were mated with ICR wild-type mice to obtain C57BL/6 × ICR LHRN316S mice.
Hormone Assay
Blood was taken from the tail vein of mice. The levels of serum progesterone and serum estradiol were detected by the Ruijin Hospital, Shanghai Jiao Tong University School of Medicine (Shanghai, China).
Fertility Assessment and Progesterone Therapy
Adult LHRN316S female mice were injected intraperitoneally with 10 IU PMSG (Sansheng, China) and 10 IU human chorionic gonadotropin (Sansheng, China) 48 h apart. After approximately 14 h, metaphase II oocytes were collected from oviducts. Wild-type females with the same age as the LHRN316S mutant females were used as controls.
Mouse COCs isolated from 10 IU PMSG-primed mice were cultured in IVG medium: α-MEM supplemented with 5% FBS (Sigma), 150 μmol/L ascorbic acid, 1 × glutamax, 1 × penicillin/streptomycin, 100 μmol/L 2-mercaptoethanol, 55 μg/mL sodium pyruvate (NacalaiTesque), 0.1 IU/mL follicle-stimulating hormone (Follistim; MSD), 15 ng/mL BMP15 and 15 ng/mL growth differentiation factor 9 (R&D Systems), in the presence or absence of 10 μM progesterone. Maturation and cumulus expansion were detected at the end of 36 h of culture.
Six- to 12-week-old LHRN316S mutant female mice were selected to a continuous mating study. Wild-type female mice acted as control. Two female mice mated with one 8- to 10-week-old fertile wild-type male mouse. Then, the numbers of offspring per litter were recorded.
Fourteen LHRN316S female mice were mated with fertile WT males, and 14 WT females of the same age were mated with fertile WT males as a control group. Two months after mating, female mice not pregnant received progesterone treatment were injected subcutaneously with 50 mg/kg progesterone (progesterone treatment group). The mice that were still not pregnant were treated with a second dose of 50 mg/kg progesterone, and the number of post-generation was counted.
Histology
Mouse ovaries were fixed in 4% paraformaldehyde at 4 °C for 24 h. Then, the ovaries were encapsulated in paraffin and sectioned at 6 µm. Then, slides were stained with H&E. Images were obtained with microscope (Leica, DM2500) and digital camera (Leica, DFC550).
Statistics
The results were presented as mean ± standard error of the mean (SEM). Means were compared using two-tailed, unpaired Student’s t tests by SPSS software. P < 0.05 was considered as statistically significant. All experiments were repeated three times.
Results
Single-Cell Transcriptional Profiles of Oocytes, Cumulus Granulosa Cells and Mural Granulosa Cells in Antral Follicles
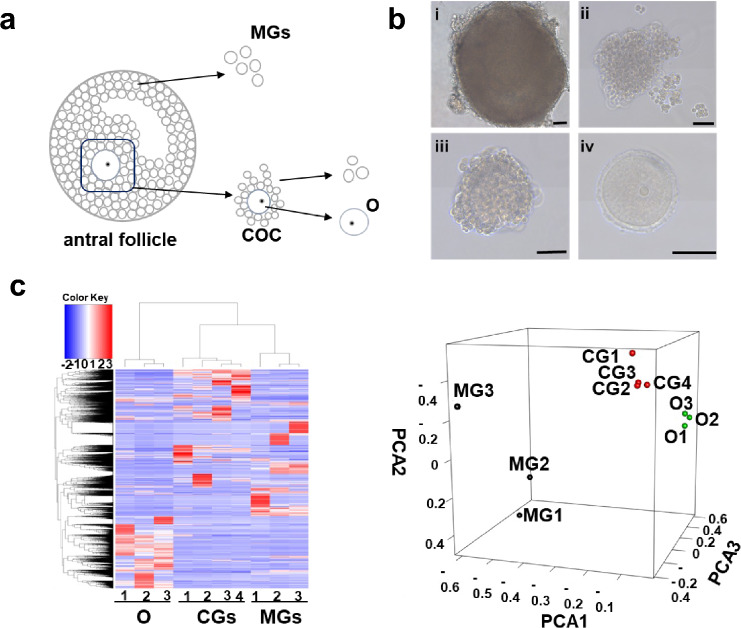
To investigate the interactions between oocytes and CGs, oocytes and MGs, and CGs and MGs, we detected cellular transcriptional profiles by scRNA-seq. Single cell types (oocytes, MGs and CGs) were isolated from antral follicles (Fig. 1a and b). Approximately 300 million sequencing reads derived from 10 single cells were mapped in this study. All genes expressed in these cells are listed in Supplementary Table S2. To ensure the reliability of scRNA-seq data, a Q-score higher than 30 (error rate < 0.1%) was acquired for each replication. To determine whether these gene expression profiles were correlated with different cell types, scRNA-seq data of all samples was analyzed by unsupervised hierarchical clustering. The cells that clustered together naturally were at the same cell type in all samples (Fig. 1c). These results were supported by principal-component analysis (Fig. 1d). Transcriptional profiles were obtained for oocytes, CGs and MGs in antral follicles.
Fig. 1.
Transcriptional profiles of single cells from the three types of cells examined in an antral follicle. a Schematic diagram of isolating individual cell types from a single antral follicle. b Morphology of a freshly isolated mouse antral follicle (i), mural granulosa cell (MG) (ii), cumulus-oocyte-complex (COC) (iii), and oocyte (O) (iv). c Unsupervised clustering of the transcriptome of all the samples. d Principal component analysis (PCA) of single cell expression patterns from the three type cells. Bar 50 μm
Analysis of the Signals for the Communication Between Oocytes and Granulosa Cells
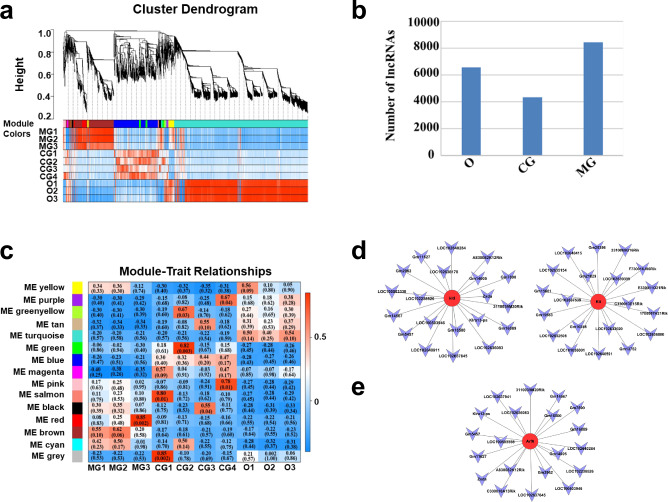
To study the regulation of oocytes by GCs, scRNA-seq data (fragments per kilobase per million mapped fragments, FPKM ≥ 0.1) were subjected to weighted correlation network analysis (WGCNA), and 15 gene-network modules were established (Fig. 2a and b, and Supplementary Table S3). Gene Ontology (GO) analysis of these modules revealed several key biological processes related to oocyte maturation. These biological processes were mainly enriched in the brown and turquoise modules shown in Supplementary Tables S4 and S5. Genes that functioned in the processes of oocyte maturation and follicle development were preferentially selected from these two modules. Approximately 300 selected genes were subjected to the construction of a co-expression network (Fig. S2 and Supplementary Table S6). In the analysis of the gene co-expression network, the expression of differential genes reflects the intergenic synergistic relationships. To identify the tightness of the co-expressed relationships among these genes, K-core values of genes were calculated (Supplementary Table S11) to focus on genes with K-core values ≥ 17 (Top2).
Fig. 2.
Gene-network modules established by weighted correlation network analysis (WGCNA), and oocyte polarity established gene-related long non-coding RNA (lncRNA). a WGCNA dendrogram indicating the expression of different gene modules in all 10 single-cell samples. b Module trait relationship followed by P values in parentheses between modules and different samples. c The number of lncRNAs detected. d and e LncRNAs correlated with genes Kitl, Kit and Arf6. Genes with different colored cycles mean different ranks in the network (ranks from higher to lower are red, green, yellow, dark blue and light blue). CG cumulus granulosa cells, O oocytes, MG mural granulosa cells
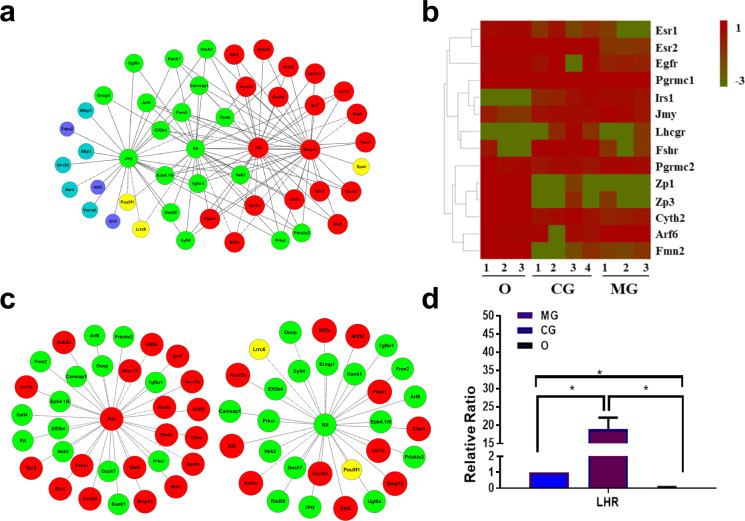
The Kitl-Kit, which is a canonical signal pathway involved in the communication between oocytes and GCs [38–43], was identified within this co-expression network (Fig. 3a). Nineteen genes that co-expressed with Kitl are shown in Fig. 3b. Upstream of this pathway, Bmp15 was co-expressed with Kitl (Fig. 3b). It has been reported that bone morphogenetic protein 15 (Bmp15) is secreted from oocytes and regulates the expression of Kitl in CGs [1]. Thirty genes were co-expressed with Kit (Fig. 3b). All the genes co-expressed with Kit and Kitl were represented in a network shown in Fig. 3a. Among these co-expressed genes, Arf6 was selected as a candidate that functioned downstream of the Kitl-Kit signal pathway.
Fig. 3.
Gene-network and gene expression differences. a Genes co-expressed with Kit ligand (Kitl) and Kit in the network. b Genes co-expressed with Kitl in the network. Genes co-expressed with Kit in the network. c The gene expression of related genes in the three types of cells. d The validation of Lhr expression by qRT-PCR. O-oocytes, CG-cumulus granulosa cells, MG-mural granulosa cells. *P < 0.05, n.s. no significant difference
Arf6 was highly expressed in the GV oocyte samples (average FPKM = 13.63, Supplementary Table S2 shows FPKM values of all genes). It was reported that cytohesin 2 activates Arf6 in a phosphoinositide 3-kinase (PI3K)-dependent manner during pre-adipocyte migration [44]. Interestingly, cytohesin 2 was also highly expressed in oocytes in this study (average FPKM = 18.88, Supplementary Table S2). It has been reported that the intra-oocyte PI3K pathway was activated by KITL-KIT in growing oocytes, inducing multiple effects, such as promoting the secretion of BMP15 [40, 45]. Therefore, we predicted that GCs may regulate oocyte maturation and follicular development through the BMP15-KITL-KIT-PI3K-ARF6 pathway.
Identification of Long Non-Coding RNAs Correlated with Genes Involved in Oocyte Maturation Based on WGCNA
Long non-coding RNAs (IncRNAs) are typically over 200 nucleotides in length and involved in diverse biological processes. The total numbers of lncRNAs detected in oocytes, CGs and MGs were 6545, 4357 and 8444, respectively (Fig. 2c). To identify possible lncRNAs involved in follicular development, module membership assignment and the weight of each gene from brown were calculated in WGCNA (Supplementary Tables S7 and S8). The top 20 lncRNAs correlated with Bmp15, Kitl, Kit and Arf6 were identified, as shown in Figs. 2d, e and S3, and Supplementary Tables S9 and S10. These lncRNAs may regulate follicular development through molecular members involved in the BMP15-KITL-KIT-PI3K-ARF6 pathway.
Generation of LHRN316S Mice
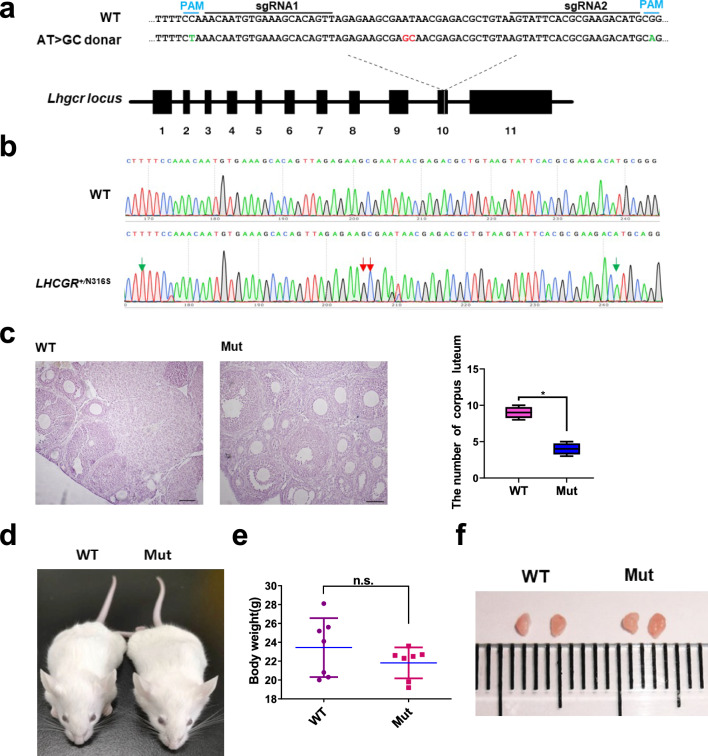
It is known that Arf6 is necessary for antral follicular development [46], but the Arf6 knockout mouse exhibits embryonic lethality [23]. It was reported that Arf6 regulates the internalization of LHR and other GPCRs [22]. Moreover, LHR directly activates the Arf-nucleotide binding site opener protein, which in turn activates Arf6 by promoting GTP/GDP exchange [47]. Consistent with the literature, the present study found that Arf6 was highly expressed in the three kinds of cells examined (Fig. 3c) and the Lhr was highly expressed in CGs (Figs. 3d and S4). These findings suggested that the Lhr may act downstream of Arf6 in follicular development. In humans, the N312S mutation of LHR has been shown to potentially inhibit oocyte maturation [27, 48]. This mutation corresponds to the murine LHR mutation at N316S (LHR N316S). To research the role of LHRN316S in follicular development in mice, we generated the LHRN316S mouse model using CRISPR/Cas9 technology (Fig. 4a, blue: silence mutation; red: N to S mutation). Genomic sequencing was used to confirm the successful generation of LHRN316S mice, and heterozygous LHRN316S mice were selected for further study (Figs. 4b and S5).
Fig. 4.
The generation of luteinizing hormone receptor (LHR)N316S mice. a Schematic view of the strategy used to generate the N316S point mutation (LHRN316S). Base pair substitutions for N316S are labeled in red. Silent mutations to prevent cleavage of the precisely mutated alleles are labeled in green. b Sanger sequencing chromatogram of genomic DNA from a wildtype (WT) mouse and the LHRN316S F0 founder. Red arrows indicate the base pair substitutions for the N316S point mutation. Green arrows indicate the silent mutations. c Morphological comparison of ovaries from LHRN316S and control female mice (left). The number of corpora lutea in different groups (right). Bar 100 μm. d The WT and LHRN316S C57BL/6 × ICR mice 8 weeks after birth. e The mean body weight of LHRN316S female mice was indistinguishable from that of controls (female littermates of LHRN316S female mice). f Images of whole ovaries from 8-week-old LHRN316S and control mice. *P < 0.05, n.s.-no significant difference
LHRN316S Causes Subfertility in Female Mice
In present study, the number of corpora lutea was significantly lower in the LHRN316S group compared with that in the control group (P < 0.05, Fig. 4c). However, there were no differences in body weight (Fig. 4d and e), ovarian morphology (Fig. 4f) and the ovary-to-body weight ratio (Fig. 5a) of 6-week-old LHRN316S vs control mice (P > 0.05). To investigate fertility, 6- to 10-week-old LHRN316S and control female mice were mated naturally and the number of offspring per litter was recorded. Although most LHRN316S females (11/14) produced offspring after mating with fertile males, the number of offspring per litter was lower than the control group (P < 0.05), indicating subfertility (Fig. 5b). Moreover, the number of ovulated oocytes induced by pregnant mare serum gonadotropin (PMSG) and human chorionic gonadotropin was lower in LHRN316S compared with that in control females (Fig. 5c). These results suggested that the LHRN316S female mice were subfertile.
Fig. 5.
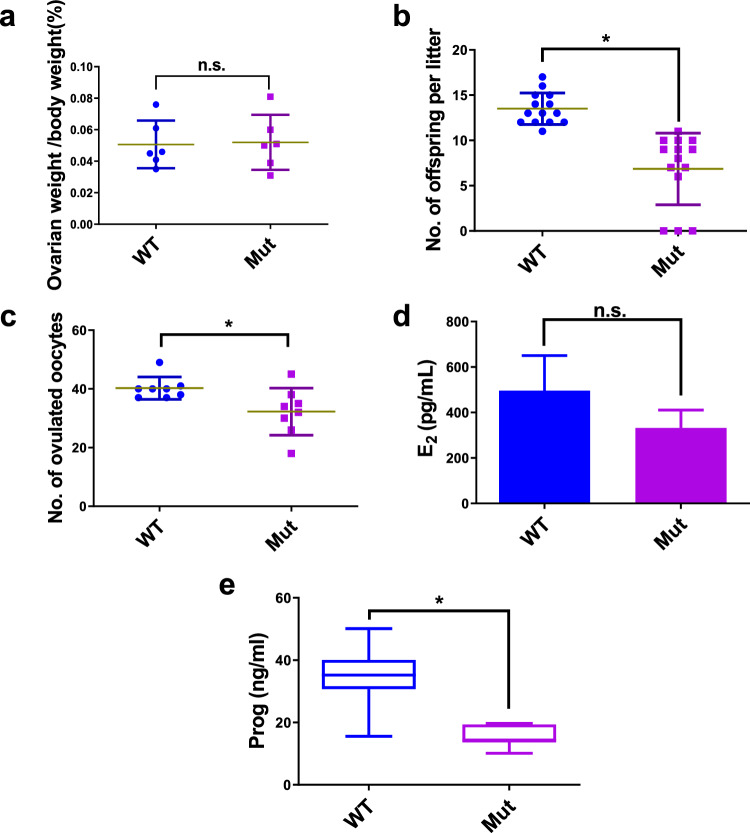
Low fertility of luteinizing hormone receptor (LHR)N316S adult female mice. a The relative ovarian weight of LHRN316S mice was indistinguishable from wild type (WT) mice. b Litter sizes from C57BL/6 × ICR (n = 14) LHRN316S female mice mated with ICR WT male mice were smaller than those from C57BL/6 × ICR (n = 14) WT female mice mated with ICR WT male mice. c The mean number of ovulated oocytes per female mouse after hormonal stimulation. d Estradiol (E2) levels in WT and LHRN316S mice. e Progesterone (Prog) homone levels in WT and LHRN316S mice. LHRN316S and WT adult female mice were used at the same ages. Data shown as mean ± SEM, n ≥ 6 mice per group. *P < 0.05, n.s.-no significant differences
Because the LHRN316S female mice were subfertile, potential hormonal changes caused by LHRN316S were examined in 6-week-old female mice. Compared with the findings in wild type (WT) mice, there were no significant differences in serum estradiol levels (Fig. 5d), but significantly lower serum progesterone levels in 6-week-old LHRN316S mice (Fig. 5e), suggesting that the reduced oocyte maturation caused by LHRN316S may be due to decreased progesterone.
Progesterone Improves Oocyte Maturation and Reproduction of LHRN316S Mice
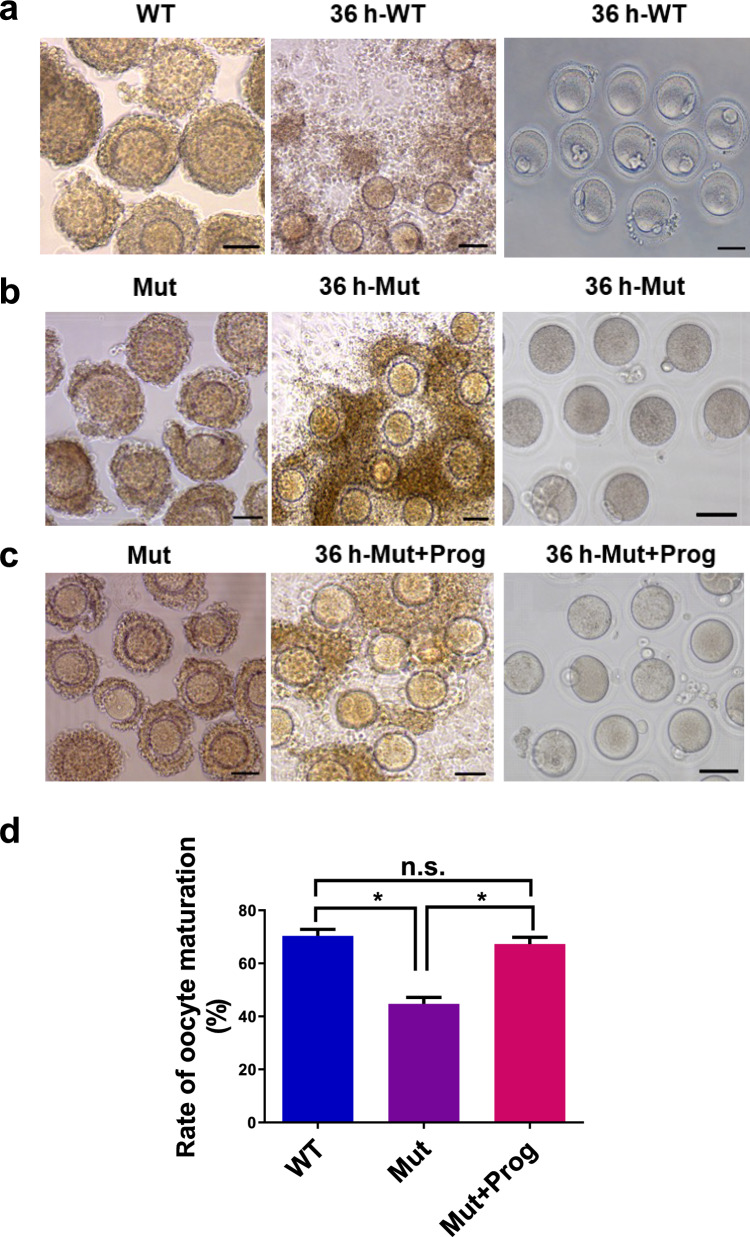
The present study examined progesterone actions on oocyte maturation in mice using in vitro and in vivo models. After cumulus-oocyte complexes (COCs) isolated from PMSG-primed mice were cultured for in vitro maturation (IVM) for 36 h, the number of mature oocytes was lower from LHRN316S mice (44%) than from WT mice (70%, P < 0.05) (Fig. 6a, b and d). The oocyte maturation rate of progesterone-treated COCs from LHRN316S mice was higher (67%) than untreated COCs from LHRN316S mice (44%, P < 0.05) and was equivalent to the COCs from WT mice (P > 0.05) (Fig. 6c and d). These results suggested that progesterone can significantly promote oocyte maturation in vitro.
Fig. 6.
Effect of progesterone on cumulus expansion and oocyte maturation. Cumulus oocyte complexes (COCs) were isolated from mice (48 h after pregnant mare serum gonadotropin stimulation) by puncturing large antral follicles and then cultured in IVM medium supplemented with or without 10 μmol/L progesterone for 36 h. a COCs from wild type (WT) mice were incubated in IVM medium. b COCs of luteinizing hormone receptor (LHR)N316S mice were incubated in IVM medium. c COCs from LHRN316S mice were incubated in IVM medium with 10 μmol/L progesterone. d The oocyte maturation rate of oocytes from LHRN316S mice incubated in IVM medium with or without progesterone. LHRN316S and WT female mice were used at the same ages. Bar 50 μm. *P < 0.05, n.s.-no significant differences
To examine whether progesterone can improve oocyte maturation in vivo, 14 LHRN316S and 14 WT female mice were mated with WT males. All WT mice became pregnant. However, three LHRN316S mice did not become pregnant, and the infertility rate of mutant mice was 21.4% (3/14). Female LHRN316S mice treated with progesterone in vivo produced offspring with normal litter sizes (Table 1).
Table 1.
Progesterone treatment of infertility mice
| Mice treatment | Total number of females | Pregnant mice | Proportion of pregnant mice (%) | Pups/mother | |
|---|---|---|---|---|---|
| LHCGR+/N316S mutation mice | None | 14 | 11 | 78 | 9 ± 1.9 |
| First progesterone treatmenta | 3 | 2 | 67 | 11 ± 0.5 | |
| Secondary progesterone treatmentb | 1 | 1 | 100 | 12 ± 0 | |
| Wild type mice | None | 14 | 14 | 100 | 14 ± 1.6 |
aInfertility mice in the None group receive the first progesterone treatment
bInfertility mice in the “First progesterone treatment” group receive the second progesterone treatment
Discussion
In this study, scRNA-seq was applied to investigate the transcriptional profiles of oocytes, CGs and MGs, and then the interactions between these cell types in antral follicles. The scRNA-seq data revealed that the Kitl-Kit pathway was shown in the network. The Kitl-Kit pathway is a canonical signal pathway involved in the development of follicles [40, 42]. KIT is a multi-domain transmembrane tyrosine kinase expressed in multiple cell types, such as germ cells [49]. KITL, the ligand of KIT, is expressed in GCs and can promote oocyte growth by binding to the KIT receptor. It was reported that the interaction of KIT and KITL can activate the PI3K pathway by causing receptor dimerization and autophosphorylation [50].
The current study found that Bmp15, which is upstream of the Kitl-Kit signaling pathway, was highly expressed in oocytes. Bmp15 was the first reported ovarian determining gene and was used as an indicator of the oocytes ability to sustain folliculogensis [51]. Bmp15 can promote the expression of Kitl [52, 53] and the proliferation of GCs [54]. Another study found that reduced Bmp15 expression led to decreased Kitl expression in oocytes and GCs [55]. Decreased expression of Bmp15 and Kitl in follicles may increase follicle atresia and decrease ovarian follicle reserve [55]. Thus, we predicted that the BMP15-KITL-KIT-PI3K pathway was significant for oocyte development regulated by GCs.
The Kitl expressed by GCs binds to the Kit receptor and activates many downstream molecules in oocytes. Factors downstream of the BMP15-KITL-KIT-PI3K pathway include Arf6, which the present study showed was highly expressed in GV oocytes. Therefore, Arf6 may be one of the key factors regulating follicular development, consistent with findings from previous studies [21, 56, 57]. Past work reported that Arf6 is activated by cytohesin 2 in a PI3K-dependent manner during preadipocyte migration [44]. Interestingly, the present study found that cytohesin 2 was also highly expressed in GV oocytes. Past research showed that the activity of Arf6 could be regulated by PI3K in other cell types [44]. The intra-oocyte PI3K pathway can be activated by KITL-KIT in growing oocytes [36, 40]. During follicle development, oocyte and granulosa cells coordinate intensively for the progress of follicle maturation. According to our RNA-seq data and literature, Bmp15 is almost exclusively expressed in oocytes. Bmp15 could trigger the expression kitl in granulosa cells. Then, secreted KITL binds to its receptor KIT which was expressed on oocytes and granulosa cells. After this, Arf6 was activated in both oocytes and granulosa cells. In oocytes, BMP15-KITL-KIT-PI3K-ARF6 was supposed to regulate the cytoskeleton arrangement and meiosis. In granulosa cells, activated Arf6 triggers the LHR signals which promote the growth and secreting the nursing materials for follicle growth. Together, the maturation of oocytes and granulosa cells is coordinated via BMP15-KITL-KIT-PI3K-ARF6 pathway during follicle development.
The present study also found that lncRNAs were highly expressed in mouse follicles, and correlated with Bmp15, Kitl, Kit and Arf6. This result indicates that follicular development may be regulated by lncRNAs through factors involved in the BMP15-KITL-KIT-PI3K-ARF6 pathway.
In the present study, Arf6 was highly expressed in oocytes, CGs and MGs, and Lhr was highly expressed in CGs. It is well known that BMP15-KITL-KIT pathway and Lhr play important roles in oocytes maturation. However, whether oocyte secreted BMP15 could regulate the activity of Lhr on granulosa is unclear. Through single cell RNA-seq, we revealed Arf6 is a downstream effector of BMP15-KITL-KIT pathway. Moreover, Arf6 could regulate the internalization and signaling of LHR on granulosa [22, 58]. Therefore, this study demonstrated the links between BMP15-KITL-KIT and ARF6-LHR. To our knowledge, it is the first time to depict the important roles of BMP15-KITL-KIT/ARF6-LHR pathway in the communication between oocytes and granulosa cells during the maturation of follicles. In humans, the LHRN312S mutation may inhibit oocyte maturation [27]. We established the LHRN316S mutant mouse model by CRISPR/Cas9 to evaluate the impact of this LHR mutation in follicular development. For humans, it was reported that the proportion of individuals heterozygous for the LHRN312S mutation was higher than those homozygous for asparagine (N/N) or serine (S/S) at this site [59]. Therefore, we chose heterozygous LHRN316S mice for further study.
The numbers of corpora lutea and ovulated oocytes were lower in LHRN316S mice than in control mice, although there were no differences in body weight and ovary-to-body weight ratios between the two groups. Furthermore, LHRN316S females exhibited an infertility rate of 21.4% when mated with WT males. In addition, pregnant LHRN316S mice ultimately produced a reduced number of offspring per litter compared with control mice. The number of oocytes from hormone-induced ovulation was also significantly lower for LHRN316S compared with that for control mice. These results indicated that the LHRN316S caused infertility or subfertility in mice. Similar to our findings, previous studies reported that it failed to ovulate when LHR was abnormal in females [25].
The levels of progesterone in LHRN316S mice were significantly lower than those in control mice. Past research showed that production of progesterone by GCs during the maturation of COCs was affected by LH and follicle-stimulating hormone [60]. In porcine oocytes, progesterone decreased connexin in MGs by a progesterone receptor (PR)-mediated pathway, leading to the recovery of meiosis [61]. Therefore, it is reasonable to speculate that LHRN316S may affect oocyte maturation and fertility by reducing progesterone levels.
The role of progesterone in the actions of the LHRN316S mutant was investigated in vivo and in vitro. In vitro, using isolated COCs from PMSG-stimulated mice, the number of oocytes reaching metaphase II was significantly reduced for LHRN316S mice compared with that for control mice. Interestingly, the oocyte maturation rate was significantly higher for progesterone-treated versus untreated cultured COCs from LHRN316S mice, and was similar to that of control COCs. Then, we explored the in vivo impact of progesterone on the fertility of LHRN316S mice. Initially infertile LHRN316S females (identified after natural mating with WT males) all generated offspring after progesterone treatment.
Thus, these results of both in vivo and in vitro models illustrated that LHRN316S affects the fertility of mice by reducing the level of progesterone. Consistent with our conclusion, reduced progesterone during the pre-ovulatory follicle stage was reported to lower pregnancy rates [62]. Conversely, a rising plasma progesterone concentration had a significant positive correlation with ovulation [63]. Simon et al. reported that progesterone was significant to the outcome of IVF [64]. However, there are also some opposite actions reported for progesterone. For example, progesterone inhibited follicle growth and the proliferation of GCs [65]. Higher progesterone levels were found in cows with low antral follicle counts in comparison with cows exhibiting high antral follicle counts [66]. Increased progesterone levels induced ovulation failure in crossbred Holstein heifers [67]. Progesterone alone did not have a positive effect on follicular growth [68]. Considering these diverse conclusions, we predict that the different effects of progesterone on follicular development and fertility may reflect the different actions of progesterone during distinct stages of ovary function.
Conclusions
In conclusion, our current study detected the transcriptional profiles of oocytes, CGs and MGs by scRNA-seq and investigated the interactions between these cellular profiles. The results showed that oocytes could be regulated by CGs through the BMP15-KITL-KIT-PI3K-ARF6 signaling pathway. We generated LHRN316S mutant mice by CRISPR/Cas9 technology, and revealed that LHRN316S led to decreased progesterone levels, which reduced oocyte maturation and resulted in infertility or subfertility. These results also suggested that progesterone therapy may be an effective means for the clinical treatment of female infertility or subfertility.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 Fig. S1. A flow chart for the data analysis (PDF 133 KB)
Supplementary file2 Fig. S2. Co-expression network of genes related to the regulation of oocyte development (PDF 1542 KB)
Supplementary file3 Fig. S3. Long non-coding RNAs (lncRNAs) correlated with genes involved in oocyte development (PDF 223 KB)
Supplementary file4 Fig. S4. Single cell Lhr and Arf6 expression were validated by qRT-PCR. Mann-Whitney test was used to test the differences between oocytes (O), mural granulosa cells (MG) and cumulus granulosa cells (CG). *: P < 0.05, n.s.: no significant differences (PDF 51 KB)
Supplementary file5 Fig. S5. Sanger sequencing chromatogram of genomic DNA samples from wildtype (WT) and LHRN316S mice (PDF 49 KB)
Author Contributions
C.Z., Y.Q.N. and C.L.M. performed Conceptualization, Investigation, Experiments, Data analysis and wrote the manuscript. B.F.X. provided specimens. G.G.T. performed bioinformatics analysis. X.Y.L. contributed to writing—review, editing. W.W.C., A.J.Z. and D.L.L. contributed to conceptualization, investigation, supervision and provided specimens. J.W. contributed to conceptualization supervision, writing—review and editing, project administration, funding acquisition.
Funding
This work was supported by the Shanghai Jiao Tong University 2030 Initiative, Key Research and Development Program of Ningxia Hui Autonomous Region (2022BFH02004), the interdisciplinary program of Shanghai Jiao Tong University (ZH2018ZDA31).
Data Availability Statement
The datasets generated and analysed during the current study are available in the NCBI Gene Expression Omnibus database [GSE241721]. https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE241721].
Declarations
Conflict of interest
There is no disclosure of potential conflicts of interest.
Research involving animals
All procedures regarding to the animal experiment were approved by the Experimental Animal Ethics and Use Committee of Shanghai Jiao Tong University (Number: 201703004) and were conducted in line with the National Research Council Guide for Care and Use of Laboratory Animals. The ethical statement file was shown in Supplementary Material.
Footnotes
Chen Zhang, Yongqiang Nie, Bufang Xu and Chunlan Mu have contributed equally.
Contributor Information
Weiwei Cheng, Email: 18017316001@163.com.
Aijun Zhang, Email: zhaj1268@163.com.
Dali Li, Email: dlli@bio.ecnu.edu.cn.
Ji Wu, Email: jiwu@sjtu.edu.cn.
References
- 1.Kidder GM, Vanderhyden BC (2010) Bidirectional communication between oocytes and follicle cells: ensuring oocyte developmental competence. Can J Physiol Pharmacol 88:399–413. 10.1139/y10-009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Dupont J, Scaramuzzi RJ (2016) Insulin signalling and glucose transport in the ovary and ovarian function during the ovarian cycle. Biochem J 473:1483–1501. 10.1042/BCJ20160124 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Brunet S, Maro B (2007) Germinal vesicle position and meiotic maturation in mouse oocyte. Reproduction 133:1069–1072. 10.1530/REP-07-0036 [DOI] [PubMed] [Google Scholar]
- 4.Barrett SL, Albertini DF (2010) Cumulus cell contact during oocyte maturation in mice regulates meiotic spindle positioning and enhances developmental competence. J Assist Reprod Genet 27:29–39. 10.1007/s10815-009-9376-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Gittens JE, Barr KJ, Vanderhyden BC et al (2005) Interplay between paracrine signaling and gap junctional communication in ovarian follicles. J Cell Sci 118:113–122. 10.1242/jcs.01587 [DOI] [PubMed] [Google Scholar]
- 6.Pelland AM, Corbett HE, Baltz JM (2009) Amino acid transport mechanisms in mouse oocytes during growth and meiotic maturation. Biol Reprod 81:1041–1054. 10.1095/biolreprod.109.079046 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Su YQ, Sugiura K, Wigglesworth K et al (2008) Oocyte regulation of metabolic cooperativity between mouse cumulus cells and oocytes: BMP15 and GDF9 control cholesterol biosynthesis in cumulus cells. Development 135:111–121. 10.1242/dev.009068 [DOI] [PubMed] [Google Scholar]
- 8.Fan HY, Liu Z, Cahill N et al (2008) Targeted disruption of Pten in ovarian granulosa cells enhances ovulation and extends the life span of luteal cells. Mol Endocrinol 22:2128–2140. 10.1210/me.2008-0095 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kawamura K, Cheng Y, Suzuki N et al (2013) Hippo signaling disruption and Akt stimulation of ovarian follicles for infertility treatment. Proc Natl Acad Sci USA 110:17474–17479. 10.1073/pnas.1312830110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nagashima T, Kim J, Li Q et al (2011) Connective tissue growth factor is required for normal follicle development and ovulation. Mol Endocrinol 25:1740–1759. 10.1210/me.2011-1045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wigglesworth K, Lee KB, Emori C et al (2015) Transcriptomic diversification of developing cumulus and mural granulosa cells in mouse ovarian follicles. Biol Reprod 92:23. 10.1095/biolreprod.114.121756 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chen J, Torcia S, Xie F et al (2013) Somatic cells regulate maternal mRNA translation and developmental competence of mouse oocytes. Nat Cell Biol 15:1415–1423. 10.1038/ncb2873 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hummitzsch K, Anderson RA, Wilhelm D et al (2015) Stem cells, progenitor cells, and lineage decisions in the ovary. Endocr Rev 36:65–91. 10.1210/er.2014-1079 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Diaz FJ, Wigglesworth K, Eppig JJ (2007) Oocytes determine cumulus cell lineage in mouse ovarian follicles. J Cell Sci 120:1330–1340. 10.1242/jcs.000968 [DOI] [PubMed] [Google Scholar]
- 15.Eppig JJ (2001) Oocyte control of ovarian follicular development and function in mammals. Reproduction 122:829–838. 10.1530/rep.0.1220829 [DOI] [PubMed] [Google Scholar]
- 16.Hussein TS, Thompson JG, Gilchrist RB (2006) Oocyte-secreted factors enhance oocyte developmental competence. Dev Biol 296:514–521. 10.1016/j.ydbio.2006.06.026 [DOI] [PubMed] [Google Scholar]
- 17.Su YQ, Wu X, O’Brien MJ et al (2004) Synergistic roles of BMP15 and GDF9 in the development and function of the oocyte-cumulus cell complex in mice: genetic evidence for an oocyte-granulosa cell regulatory loop. Dev Biol 276:64–73. 10.1016/j.ydbio.2004.08.020 [DOI] [PubMed] [Google Scholar]
- 18.Zhang Y, Yan Z, Qin Q et al (2018) Transcriptome landscape of human folliculogenesis reveals oocyte and granulosa cell interactions. Mol Cell 72(1021–1034):e4. 10.1016/j.molcel.2018.10.029 [DOI] [PubMed] [Google Scholar]
- 19.Donaldson JG, Jackson CL (2011) ARF family G proteins and their regulators: roles in membrane transport, development and disease. Nat Rev Mol Cell Biol 12:362–375. 10.1038/nrm3117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Cavenagh MM, Whitney JA, Carroll K et al (1996) Intracellular distribution of Arf proteins in mammalian cells. Arf6 is uniquely localized to the plasma membrane. J Biol Chem 271:21767–21774. 10.1074/jbc.271.36.21767 [DOI] [PubMed] [Google Scholar]
- 21.Duan X, Zhang HL, Pan MH et al (2018) Vesicular transport protein Arf6 modulates cytoskeleton dynamics for polar body extrusion in mouse oocyte meiosis. Biochim Biophys Acta Mol Cell Res 1865:455–462. 10.1016/j.bbamcr.2017.11.016 [DOI] [PubMed] [Google Scholar]
- 22.Claing A (2004) Regulation of G protein-coupled receptor endocytosis by ARF6 GTP-binding proteins. Biochem Cell Biol 82:610–617. 10.1139/o04-113 [DOI] [PubMed] [Google Scholar]
- 23.Suzuki T, Kanai Y, Hara T et al (2006) Crucial role of the small GTPase ARF6 in hepatic cord formation during liver development. Mol Cell Biol 26:6149–6156. 10.1128/MCB.00298-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Breen SM, Andric N, Ping T et al (2013) Ovulation involves the luteinizing hormone-dependent activation of G(q/11) in granulosa cells. Mol Endocrinol 27:1483–1491. 10.1210/me.2013-1130 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ascoli M, Fanelli F, Segaloff DL (2002) The lutropin/choriogonadotropin receptor, a 2002 perspective. Endocr Rev 23:141–174. 10.1210/edrv.23.2.0462 [DOI] [PubMed] [Google Scholar]
- 26.Themmen AP (2005) An update of the pathophysiology of human gonadotrophin subunit and receptor gene mutations and polymorphisms. Reproduction 130:263–274. 10.1530/rep.1.00663 [DOI] [PubMed] [Google Scholar]
- 27.Lindgren I, Nenonen H, Henic E et al (2019) Gonadotropin receptor variants are linked to cumulative live birth rate after in vitro fertilization. J Assist Reprod Genet 36:29–38. 10.1007/s10815-018-1318-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nagulapalli A, Vembu R, Pandurangi M et al (2023) Luteinizing hormone/human chorionic gonadotropin receptor N312S single-nucleotide polymorphism and its impact on clinical and reproductive outcomes in assisted reproductive technology: a prospective cohort study. Cureus 15:e47217. 10.7759/cureus.47217 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Pirtea P, de Ziegler D, Marin D et al (2022) Gonadotropin receptor polymorphisms (FSHR N680S and LHCGR N312S) are not predictive of clinical outcome and live birth in assisted reproductive technology. Fertil Steril 118:494–503. 10.1016/j.fertnstert.2022.06.011 [DOI] [PubMed] [Google Scholar]
- 30.Jin H, Yang H, Zheng J et al (2023) Post-trigger luteinizing hormone concentration to positively predict oocyte yield in the antagonist protocol and its association with genetic variants of LHCGR. J Ovarian Res 16:189. 10.1186/s13048-023-01271-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Xue ZG, Huang K, Cai CC et al (2013) Genetic programs in human and mouse early embryos revealed by single-cell RNA sequencing. Nature 500:593–597. 10.1038/nature12364 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Pujana MA, Han JDJ, Starita LM et al (2007) Network modeling links breast cancer susceptibility and centrosome dysfunction. Nat Genet 39:1338–1349. 10.1038/ng.2007.2 [DOI] [PubMed] [Google Scholar]
- 33.Prieto C, Risueno A, Fontanillo C et al (2008) Human gene coexpression landscape: confident network derived from tissue transcriptomic profiles. PLoS ONE 3:e3911. 10.1371/journal.pone.0003911 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Barabasi AL, Oltvai ZN (2004) Network biology: understanding the cell’s functional organization. Nat Rev Genet 5:101–113. 10.1038/nrg1272 [DOI] [PubMed] [Google Scholar]
- 35.Ashburner M, Ball CA, Blake JA et al (2000) Gene ontology: tool for the unification of biology. Nat Genet 25:25–29. 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Dupont J, Reverchon M, Cloix L et al (2012) Involvement of adipokines, AMPK, PI3K and the PPAR signaling pathways in ovarian follicle development and cancer. Int J Dev Biol 56:959–967. 10.1387/ijdb.120134jd [DOI] [PubMed] [Google Scholar]
- 37.Shao YJ, Guan YT, Wang LR et al (2014) CRISPR/Cas-mediated genome editing in the rat via direct injection of one-cell embryos. Nat Protoc 9:2493–2512. 10.1038/nprot.2014.171 [DOI] [PubMed] [Google Scholar]
- 38.Packer AI, Hsu YC, Besmer P et al (1994) The ligand of the c-kit receptor promotes oocyte growth. Dev Biol 161:194–205. 10.1006/dbio.1994.1020 [DOI] [PubMed] [Google Scholar]
- 39.Ye Y, Kawamura K, Sasaki M et al (2009) Kit ligand promotes first polar body extrusion of mouse preovulatory oocytes. Reprod Biol Endocrinol 7:26. 10.1186/1477-7827-7-26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Hsueh AJ (2014) Fertility: the role of mTOR signaling and KIT ligand. Curr Biol 24:R1040–R1042. 10.1016/j.cub.2014.09.033 [DOI] [PubMed] [Google Scholar]
- 41.Jagarlamudi K, Rajkovic A (2012) Oogenesis: transcriptional regulators and mouse models. Mol Cell Endocrinol 356:31–39. 10.1016/j.mce.2011.07.049 [DOI] [PubMed] [Google Scholar]
- 42.Jones RL, Pepling ME (2013) KIT signaling regulates primordial follicle formation in the neonatal mouse ovary. Dev Biol 382:186–197. 10.1016/j.ydbio.2013.06.030 [DOI] [PubMed] [Google Scholar]
- 43.Morohaku K, Tanimoto R, Sasaki K et al (2016) Complete in vitro generation of fertile oocytes from mouse primordial germ cells. Proc Natl Acad Sci USA 113:9021–9026. 10.1073/pnas.1603817113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Davies JC, Tamaddon-Jahromi S, Jannoo R et al (2014) Cytohesin 2/ARF6 regulates preadipocyte migration through the activation of ERK1/2. Biochem Pharmacol 92:651–660. 10.1016/j.bcp.2014.09.023 [DOI] [PubMed] [Google Scholar]
- 45.Lees-Murdock DJ, Lau HT, Castrillon DH et al (2008) DNA methyltransferase loading, but not de novo methylation, is an oocyte-autonomous process stimulated by SCF signalling. Dev Biol 321:238–250. 10.1016/j.ydbio.2008.06.024 [DOI] [PubMed] [Google Scholar]
- 46.Salvador LM, Mukherjee S, Kahn RA et al (2001) Activation of the luteinizing hormone/choriogonadotropin hormone receptor promotes ADP ribosylation factor 6 activation in porcine ovarian follicular membranes. J Biol Chem 276:33773–33781. 10.1074/jbc.M101498200 [DOI] [PubMed] [Google Scholar]
- 47.Mukherjee S, Gurevich VV, Jones JC et al (2000) The ADP ribosylation factor nucleotide exchange factor ARNO promotes beta-arrestin release necessary for luteinizing hormone/choriogonadotropin receptor desensitization. Proc Natl Acad Sci USA 97:5901–5906. 10.1073/pnas.100127097 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Lindgren I, Baath M, Uvebrant K et al (2016) Combined assessment of polymorphisms in the LHCGR and FSHR genes predict chance of pregnancy after in vitro fertilization. Hum Reprod 31:672–683. 10.1093/humrep/dev342 [DOI] [PubMed] [Google Scholar]
- 49.Comazzetto S, Murphy MM, Berto S et al (2019) Restricted hematopoietic progenitors and erythropoiesis require SCF from leptin receptor+ niche cells in the bone marrow. Cell Stem Cell 24(477–486):e6. 10.1016/j.stem.2018.11.022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lennartsson J, Ronnstrand L (2012) Stem cell factor receptor/c-Kit: from basic science to clinical implications. Physiol Rev 92:1619–1649. 10.1152/physrev.00046.2011 [DOI] [PubMed] [Google Scholar]
- 51.Liu JC, Yan ZH, Li B et al (2021) Di (2-ethylhexyl) phthalate impairs primordial follicle assembly by increasing PDE3A expression in oocytes. Environ Pollut 270:116088. 10.1016/j.envpol.2020.116088 [DOI] [PubMed] [Google Scholar]
- 52.Thomas FH, Vanderhyden BC (2006) Oocyte-granulosa cell interactions during mouse follicular development: regulation of kit ligand expression and its role in oocyte growth. Reprod Biol Endocrinol 4:19. 10.1186/1477-7827-4-19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Otsuka F, Shimasaki S (2002) A negative feedback system between oocyte bone morphogenetic protein 15 and granulosa cell kit ligand: its role in regulating granulosa cell mitosis. Proc Natl Acad Sci USA 99:8060–8065. 10.1073/pnas.122066899 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Otsuka F, Yao Z, Lee T et al (2000) Bone morphogenetic protein-15. Identification of target cells and biological functions. J Biol Chem 275:39523–39528. 10.1074/jbc.M007428200 [DOI] [PubMed] [Google Scholar]
- 55.Celik S, Ozkavukcu S, Celik-Ozenci C (2020) Altered expression of activator proteins that control follicle reserve after ovarian tissue cryopreservation/transplantation and primordial follicle loss prevention by rapamycin. J Assist Reprod Genet 37:2119–2136. 10.1007/s10815-020-01875-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Sun SC, Sun QY, Kim NH (2011) JMY is required for asymmetric division and cytokinesis in mouse oocytes. Mol Hum Reprod 17:296–304. 10.1093/molehr/gar006 [DOI] [PubMed] [Google Scholar]
- 57.Li H, Guo F, Rubinstein B et al (2008) Actin-driven chromosomal motility leads to symmetry breaking in mammalian meiotic oocytes. Nat Cell Biol 10:1301–1308. 10.1038/ncb1788 [DOI] [PubMed] [Google Scholar]
- 58.Kanamarlapudi V, Thompson A, Kelly E et al (2012) ARF6 activated by the LHCG receptor through the cytohesin family of guanine nucleotide exchange factors mediates the receptor internalization and signaling. J Biol Chem 287:20443–20455. 10.1074/jbc.M112.362087 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Rivron NC, Frias-Aldeguer J, Vrij EJ et al (2018) Blastocyst-like structures generated solely from stem cells. Nature 557:106–111. 10.1038/s41586-018-0051-0 [DOI] [PubMed] [Google Scholar]
- 60.Chian RC, Ao A, Clarke HJ et al (1999) Production of steroids from human cumulus cells treated with different concentrations of gonadotropins during culture in vitro. Fertil Steril 71:61–66. 10.1016/s0015-0282(98)00416-6 [DOI] [PubMed] [Google Scholar]
- 61.Shimada M, Terada T (2002) FSH and LH induce progesterone production and progesterone receptor synthesis in cumulus cells: a requirement for meiotic resumption in porcine oocytes. Mol Hum Reprod 8:612–618. 10.1093/molehr/8.7.612 [DOI] [PubMed] [Google Scholar]
- 62.Cavalieri J (2019) Absence of a corpus luteum and relatively lesser concentrations of progesterone during the period of pre-ovulatory follicle emergence results in lesser pregnancy rates in Bos indicus cattle. Anim Reprod Sci 204:39–49. 10.1016/j.anireprosci.2019.03.003 [DOI] [PubMed] [Google Scholar]
- 63.Gaur M, Purohit GN (2019) Follicular dynamics and colour Doppler vascularity evaluations of follicles and corpus luteum in relation to plasma progesterone during the oestrous cycle of Surti buffaloes. Reprod Domest Anim 54:585–594. 10.1111/rda.13400 [DOI] [PubMed] [Google Scholar]
- 64.Simon C, Branet L, Moreau J et al (2019) Association between progesterone to number of mature oocytes index and live birth in GnRH antagonist protocols. Reprod Biomed Online 38:901–907. 10.1016/j.rbmo.2019.01.009 [DOI] [PubMed] [Google Scholar]
- 65.Long H, Yu W, Yu S et al (2021) Progesterone affects clinic oocyte yields by coordinating with follicle stimulating hormone via PI3K/AKT and MAPK pathways. J Adv Res 33:189–199. 10.1016/j.jare.2021.02.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Bonato DV, Ferreira EB, Gomes DN et al (2022) Follicular dynamics, luteal characteristics, and progesterone concentrations in synchronized lactating Holstein cows with high and low antral follicle counts. Theriogenology 179:223–229. 10.1016/j.theriogenology.2021.12.006 [DOI] [PubMed] [Google Scholar]
- 67.Kornmatitsuk B, Kornmatitsuk S (2021) Circulating progesterone concentrations and preovulatory follicle diameters affecting ovulatory response in crossbred dairy heifers, following a 7-day progesterone-based synchronization protocol. Trop Anim Health Prod 53:102. 10.1007/s11250-020-02494-1 [DOI] [PubMed] [Google Scholar]
- 68.Paulino L, Barroso PAA, Silva AWB et al (2020) Effects of epidermal growth factor and progesterone on development, ultrastructure and gene expression of bovine secondary follicles cultured in vitro. Theriogenology 142:284–290. 10.1016/j.theriogenology.2019.10.031 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary file1 Fig. S1. A flow chart for the data analysis (PDF 133 KB)
Supplementary file2 Fig. S2. Co-expression network of genes related to the regulation of oocyte development (PDF 1542 KB)
Supplementary file3 Fig. S3. Long non-coding RNAs (lncRNAs) correlated with genes involved in oocyte development (PDF 223 KB)
Supplementary file4 Fig. S4. Single cell Lhr and Arf6 expression were validated by qRT-PCR. Mann-Whitney test was used to test the differences between oocytes (O), mural granulosa cells (MG) and cumulus granulosa cells (CG). *: P < 0.05, n.s.: no significant differences (PDF 51 KB)
Supplementary file5 Fig. S5. Sanger sequencing chromatogram of genomic DNA samples from wildtype (WT) and LHRN316S mice (PDF 49 KB)
Data Availability Statement
The datasets generated and analysed during the current study are available in the NCBI Gene Expression Omnibus database [GSE241721]. https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE241721].