Abstract
Background:
Lipedema is a disease typically affecting women with a symmetrical, painful fat distribution disorder, which is hypothesized to be caused by impaired adipogenesis, inflammation, and extracellular matrix remodeling, leading to fibrosis and the development of edema in lipedema subcutaneous adipose tissue. The pathogenesis and molecular processes leading to lipedema have not yet been clarified.
Methods:
A whole transcriptome analysis of subcutaneous tissue of lipedema stages I (n = 12), II (n = 9), and III (n = 8) compared with hypertrophied subcutaneous tissue (n = 4) was performed. Further data about hormonal substitution and body morphology were collected. The study is registered at ClinicalTrials.gov (NCT05861583).
Results:
We identified several differentially expressed genes involved in mechanisms leading to the development of lipedema. Some genes, such as PRKG2, MEDAG, CSF1R, BICC1, ERBB4, and ACP5, are involved in adipogenesis, regulating the development of mature adipocytes from mesenchymal stem cells. Other genes, such as MAFB, C1Q, C2, CD68, CD209, CD163, CD84, BCAT1, and TREM2, are predicted to be involved in lipid accumulation, hypertrophy, and the inflammation process. Further genes such as SHTN1, SCN7A, and SCL12A2 are predicted to be involved in the regulation and transmission of pain.
Conclusions:
In summary, the pathogenesis and development of lipedema might be caused by alterations in adipogenesis, inflammation, and extracellular matrix remodeling, leading to fibrosis and the formation of edema resulting in this painful disease. These processes differ from hypertrophied adipose tissue and may therefore play a main role in the formation of lipedema.
Takeaways
Question: How does lipedema develop and what are the pathomechanisms that lead to this disease?
Findings: Via whole transcriptome analysis of subcutaneous fatty tissue of patients with lipedema compared to hypertrophied fatty tissue, we identified 137 differentially expressed genes involved in mechanisms, leading to the development of lipedema. The pathogenesis of lipedema might be caused by alterations in the gene expression of mechanisms such as adipogenesis, inflammation, and extracellular matrix remodeling, leading to fibrosis and the formation of edema.
Meaning: Lipedema appears to be a multifactorial disease with altered adipogenesis, inflammatory processes, fibrosis, and also nociception.
INTRODUCTION
Lipedema is an underdiagnosed disease affecting female patients showing a symmetrical subcutaneous fat distribution disorder of the upper and lower extremities.1 The patients have a limited quality of life caused by painful subcutaneous tissue, the development of edema, and the increased formation of hematoma.2 The increase in lipedema-associated subcutaneous adipose tissue is nearly unaffected by dietary or athletic interventions as well as physical devices. It is often misunderstood as obesity or lymphedema, which may lead to wrong therapeutic options being chosen.3,4 Diagnosis criteria for lipedema include only clinical symptoms and morphology.5 Lipedema is a painful accumulation of fatty tissue in specific regions such as the hips, thighs, lower legs, and/or arms.6 It can be subdivided into different stages as described by Kruppa et al.7
Lipedema is a chronic disease mostly starting with puberty.8 Some patients notice a progression boost after hormonal changes caused by pregnancy or menopause,8,9 wherefore hormonal changes are hypothesized to lead to an impetus of this chronic disease.10 A genetical predisposition as a main factor leading to lipedema is also discussed.11 Obesity is also a driving force for the progress of the disease.3,8
Standard therapies such as compression therapy by wearing compression garments and manual lymphatic drainage can temporarily reduce pain intensity.12 A long-term effect can be reached by tumescent liposuction.13–15 Liposuction slows the disease’s progress by reducing side effects from later staged symptoms, but it cannot cure lipedema.16,17
In conclusion, the etiopathogenesis of lipedema is rarely understood, the diagnostic tools are not highly specific, and the therapeutic options are limited. Therefore, this study is focusing on the differentiation of subcutaneous fatty tissue leading to this disease. We identified a specific RNA-expression pattern of lipedema-associated adipose tissue. Specific genes could be identified to play a role in proliferation and differentiation as well as inflammation of lipedema-associated tissue. Additionally, a questionnaire was performed about hormonal substitution and body morphology (see Fig. 1).
Fig. 1.
Graphical abstract. Biopsies from subcutaneous fat tissue of the thigh from patients with lipedema with different stages (study group) or hypertrophied fat tissue (control group) were obtained intraoperatively. The samples were processed and analyzed using NGS sequencing. The RNA expression differences between lipedema and the control group were then compared and the up- and down-regulated genes were shown in heatmaps. The altered gene expression pattern in lipedema patients provides information about which processes might differ. These include changes in adipogenesis, inflammation, and fibrosis of the subcutaneous tissue. This illustration was created by using Bioicons (https://bioicons.com/): sequence_histogram is licensed under CC0 by Marcel Tisch; nudeotide-t/a/c/g-0 is licensed under CC0 by Emmett Leddin; C-DNA is licensed under CC-BY 4.0 Unported by DBCLS, and the icon is modified in size; lllumina_miseq is licensed under CC-BY 4.0 Unported by DBCLS; tweezers_noS is licensed under CC-BY 4.0 Unported by DBCLS; microtube-dosed is licensed under CC-BY 4.0 Unported by DBLS, and the icon is modified by overlay; obese-female is licensed under CC-BY 3.0 Unported by Servier and modified by detail clipping; fat tissue is licensed under CC-BY 3.0 Unported by Servier.
MATERIALS AND METHODS
The study is an explorative study performing a transcriptome analysis of human biopsies of subcutaneous fatty tissue comparing the expression pattern of patients with lipedema [diagnosis of lipedema stage (St.) I, II, or III, staging criteria as described in Kruppa et al7], with hypertrophied healthy adipose tissue. To exclude effects caused by obesity, a subgroup analysis was performed corresponding to a body mass index (BMI) of greater than 25 and less than 34. Further, a questionnaire about hormonal substitution, body morphology [BMI, waist-to-height ratio (WHtR), waist-to-hip ratio (WHR)] was performed. (See appendix, Supplemental Digital Content 1, which displays the methods, http://links.lww.com/PRSGO/D612.) (See appendix, Supplemental Digital Content 2, which displays an overview of mean age, body measurements, and substitution of hormones, http://links.lww.com/PRSGO/D613.) (See appendix, Supplemental Digital Content 3, which displays hormonal substitution, http://links.lww.com/PRSGO/D614.) [See appendix, Supplemental Digital Content 4, which displays BMI, WHR, and WHtR. A, The comparison of the BMI between the control group (lipohypertrophy) and the patients with lipedema St. I–III. A significant difference between the control group and lipedema stage II (P = 0.0042) and stage III (P = 0.0003) could be found. B, The comparison of the WHR between the control group and the patients with lipedema St. I–III. A significant difference between the control group and lipedema St. II (P = 0.025) and St. III (P = 0.0112) could be found. C, The comparison of the WHtR between the control group and the patients with lipedema St. I–III. A significant difference between the control group and lipedema St. II (P = 0.0022) and St. III (P = 0.0001) could be detected. There is also a significant difference between lipedema St. I and II (P = 0.0246) as well as St. II and III (P = 0.0122). The figure shows the mean ± SD as well as the P value according to the significance level. The sample size is marked within the columns, http://links.lww.com/PRSGO/D615.] (See table, Supplemental Digital Content 5, which displays the comorbidities of the patients with lipedema whose tissue samples were sequenced. None of the test subjects had diabetes mellitus, and very few had hypercholesterolemia or hypotension. Some of the patients with lipedema had varicosis or hypothyroidism usually due to Hashimotos thyroiditis. The comorbidity of hypertension occurs particularly in St. III, http://links.lww.com/PRSGO/D616.)
Sampling and Treatment Protocol
Before liposuctioned native samples of subcutaneous fatty tissue of the thigh region were collected after local anesthesia, the skin was incised and the fatty tissue was sampled via forceps biopsy. Each sample consisted of nearly the same amount of fatty tissue. The tissue samples were incubated with RNA later (Sigma-Aldrich) at 4°C following the instruction protocol and were sent to Eurofins Genomics (Konstanz, Germany). The RNA was isolated from the tissue, and after quality control (RNA integrity number, rRNA ratio 28s/18s), a cDNA library was created via bead-based poly-A-selection. Next-generation sequencing (NGS) analysis (Illumina HiSeq 2500 technology and quality control using Illumina CASAVA software) was performed by Eurofins Genomics (Konstanz). The control group consisted of 4 patients. The patients with lipedema were stage-dependently divided into St. I (n = 12), St. II (n = 9), and St. III (n = 8). The raw data (fastq files) were downloaded from the servers and were further processed as described in NGS Analysis.
NGS Analysis
After download of the fastq files, sequences were mapped to the human reference genome (GRCh38) using the aligner hisat2 (version 2.1.0). After sorting the files for improved data processing using Samtools (version 1.9), featureCounts tables were generated using the GRCh38.110 annotation and the program featureCounts (version 2.0.3). The differential gene expression analysis of NGS data was performed in R using edgeR. Heatmaps were generated with Python (version 3.11.2) using bioinfokit (version 2.1.3).
Statistics of Clinical Data (Questionnaire)
Data are reported as mean ± SD. The statistical analysis was done with the t test calculator of GraphPad Dotmatics (https://www.graphpad.com/quickcalcs/ttest1.cfm) using an unpaired t test to determine the significance. Results were marked on their significance level as follows: P < 0.05 (*), P < 0.01 (**), P < 0.001 (***).
RESULTS
The average age was distributed homogeneously in all groups (Supplemental Digital Content 2, http://links.lww.com/PRSGO/D613).
Questionnaire
There is a significant difference between the control group and lipedema St. II (P = 0.0042) and St. III (P = 0.0003) looking at the mean values of BMI. There is a significant difference in WHR between the control group and lipedema St. II (P = 0.025) and St. III (P = 0.0122). Regarding WHtR, there is a significant difference between the control group and lipedema St. II (P = 0.0022) and St. III (P = 0.0001). Further, a significant difference between lipedema St. I and II (P = 0.0246) as well as St. II and III (P = 0.0122) could be detected.
Considering the substitution of hormonal contraceptives, patients with lipedema St. III showed the longest phase of substitution of hormones with an average of 15.41 (±8.49 SD) years. The difference to the control group is statistically significant (P = 0.0463) (Supplemental Digital Content 2–5, http://links.lww.com/PRSGO/D613, http://links.lww.com/PRSGO/D614, http://links.lww.com/PRSGO/D615, http://links.lww.com/PRSGO/D616).
Transcriptome Analysis
The analyzed samples have an RNA integrity number of 5–10 and have an average library length of 420–507 base pairs. Compared to the control group, 1 gene is significantly downregulated in lipedema St. I, but no significant upregulated gene could be found; in lipedema St. II, 43 upregulated genes as well as 14 downregulated genes could be identified; and in lipedema St. III, the expression level of 70 genes is upregulated and that of 9 genes is downregulated. The gene codes, names, fold changes, and P values are listed in Supplemental Digital Content 6–8. (See appendix, Supplemental Digital Content 6, which displays heatmaps of differentially expressed genes in lipedema, http://links.lww.com/PRSGO/D617.) (See appendix, Supplemental Digital Content 7, which displays NGS analyses and gene information about differentially expressed genes in lipedema, http://links.lww.com/PRSGO/D618.) (See table, Supplemental Digital Content 8, which displays differentially expressed genes in lipedema, http://links.lww.com/PRSGO/D619.)
The differentially expressed genes, which are predicted to be involved in adipogenesis, obese adipose tissue remodeling, lipotoxicity, and inflammation, are displayed in a heatmap (Supplemental Digital Content 6, http://links.lww.com/PRSGO/D617). A total of 24 genes were identified that are upregulated in both St. II and St. III. A slight dynamic in the form of an increase or decrease in the expression level with increasing stage could be observed (Supplemental Digital Content 8, http://links.lww.com/PRSGO/D619).
BMI-corrected Subgroup Analysis
The patients in the control group (n = 3) and the lipedema St. III group (n = 3) were selected depending on their BMI. The BMI of both groups is homogenous with an average BMI of 28.07 (±4.48 SD) in the control group and an average BMI of 27.1 (±1.02 SD) in St. I, 29.6 (±1.23 SD) in St. II, and 31.4 (±2.73 SD) in St. II. There is no significant difference in BMI between control and study groups [(P (St. I) = 0.733; P (St. II) = 0.606; P (St. III) = 0.3374]. A total of 7 genes are differentially expressed in St. II and 5 genes are significantly upregulated in St. III in a BMI-corrected manner in lipedema (Supplemental Digital Content 6 and 7, http://links.lww.com/PRSGO/D617, http://links.lww.com/PRSGO/D618).
DISCUSSION
The average age of the control group and the lipedema groups is distributed homogeneously, wherefore an age-dependent regulation can be excluded. The results of the transcriptome analysis include many characterized and uncharacterized transcripts (Supplemental Digital Content 7, http://links.lww.com/PRSGO/D618), and this discussion focuses on characterized transcripts. As hypothesized, lipedema might be hereditary,4,18 and a hormone-dependent dysregulation via estrogen is also discussed. Our data show a significant difference in the duration of contraceptive hormone substitution between the control group and St. III. Lipedema-associated fatty tissue shows a higher distribution density of estrogen receptors and stimulates itself by releasing estrogen as a paracrine hormone.10 The effect of progesterone is also present in adipose tissue by increased expression of progesterone receptors in subcutaneous tissue.19 In female rats, it could be shown that progesterone especially in addition to estrogen substitution is leading to adiposity.20 As sexual hormones are also accumulated in adipose tissue, the effect of estrogen and progesterone might be potentiated.21,22 We identified some genes that are regulated in a hormone-dependent manner and whose expression is dysregulated in lipedema. Our data indicate that ACTG2, MEDAG, and NEGR1 are differentially expressed in lipedema. ACTG2 is known to be involved in the formation of adipose tissue in correlation to the hormone level.23 NGER1 is known to regulate central nervous activity in different brain areas under feeding conditions,24 and a dysregulation in neurons as well as subcutaneous tissue might lead to obesity.25 In mouse models, a gender-specific alteration of NEGR1 expression with an obese phenotype in mice could be found.26 MEDAG is a mesenteric estrogen–dependent adipogenesis factor, which is highly regulated via estrogen and is an important factor in adipogenesis.27
Differentially Expressed Genes Involved in Adipogenesis
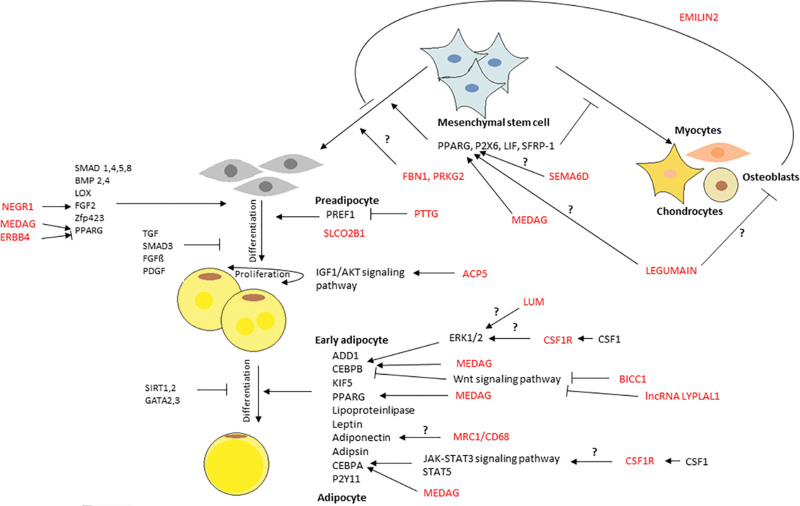
Adipocytes are formed from mesenchymal stem cells (see Fig. 2). The pool of mesenchymal stem cells is also used for the formation of muscle tissue, osteoblasts, and chondrocytes. Undifferentiated mesenchymal stem cells rise to preadipocytes that later become early adipocytes. The first step is driven by Peroxisome proliferator-activated receptor gamma (PPARγ) as main regulator.28 Some of our upregulated genes in lipedema such as SEMA6D, MEDAG, and LEGUMAIN are predicted to regulate PPARγ.27,29,30. We identified two additional genes—FBN1 and PRKG2—upregulated in lipedema that are also predicted to lead to the development of preadipocytes via an unknown pathway.31–33 The formation of preadipocytes might lead to a forced adipogenesis, and therefore, to a leakage of osteoblasts, chondrocytes, and myocytes. The reversal strategy might be the upregulation of EMILIN2 confirmed by our data.34,35 In patients with lipedema, a valgus osteoarthritis in the knee and a skew foot formation is observed.36 The reason for this has always been seen in the massive burden on the fatty tissue. Supporting our data, it should also be considered that depletion of the mesenchymal stem cell pool and a shift in balance toward adipogenesis may result in a loss of bone strength, a reduced cartilage mass, and muscle weakness.
Fig. 2.
Potential regulation of adipogenesis in lipedema. The signaling pathway of adipogenesis displays the difference of mesenchymal stem cells to mature adipocytes via known signaling cascades. Known regulating factors of adipogenesis are marked in black. Factors that might be involved in the development of lipedema based on our data analysis were shown in red and their activating or inhibiting effects were labeled accordingly.
In the second step, the preadipocytes differentiate into early adipocytes triggered by signaling molecules such as FGF2, PREF-1, and PPARγ. PPARγ and PREF-1 are proadipogenic factors, whereas the role of FGF2 might be ambivalent.37 NEGR1 is inducing FGF2, and MEDAG is inducing PPARγ leading to differentiation.27,38,39 PREF-1 is negatively regulated by PTTG,40 which can be a reversal strategy to slow down adipogenesis, or depending on the protein level, the loss of PPTG needs to be compensated by upregulation of the RNA expression. Both NEGR1 and MEDAG are upregulated in lipedema. The known marker gene for preadipocytes SLCO2B141 is also enriched in our data, suggesting that preadipocytes and differentiation might play an important role in lipedema. The stimulation of proliferation takes place via the IGF/AKT signaling pathway via ACP5.42 In lipedema, ACP5 is increased leading to a faster proliferation rate compared to hypertrophied fatty tissue. Early adipocytes differentiate and mature into late adipocytes, stimulated by factors such as CEPBA, CEBPB, PPARγ, and adiponectin. MEDAG is directly activating CEBPB and PPARγ,27 whereas CEBPB is inhibited via Wnt signaling pathway. BICC1 and the lncRNA LYPLAL1 are leading to an inhibition of the Wnt signaling pathway, whereby the inhibition of adipocytes differentiation disappears.43,44 The proadipogenic factors BICC1 and lncRNA LYPLAL1 are both upregulated in lipedema. CEBPB is also activated via the extracellular signal-regulated kinase signaling pathway by CSF1.45 We identified an increased expression of the CSF1R in lipedema-associated fatty tissue, which might cause an increased sensitivity to CSF1 in adipocytes finally leading to an activation of CEBPB. In accordance with our data, Nankam et al46 identified the soluble CSF1 as marker of lipedema in peripheral blood. CSF1 is also known to play a role in the activation of the JAK-STAT3 and STAT5 pathways, which are both involved in adipogenesis via CEBPA.47,48 We further identified MRC1 and CD68, which are both upregulated in lipedema. The MRC1/CD68 ratio is known to correlate with the adiponectin level and both proteins are predicted to regulate adipogenesis and lipogenesis via different pathways.49
Cell Growth, Lipotoxicity, Inflammation, and Pain
Mature adipocytes are growing and become hypertrophic by increasing their lipid storage via lipid accumulation in lipid droplets.50,51 We identified several genes that are predicted to be involved in these processes: MAFB, F13A1, and NTRK3.52–54 The hypertrophy leads to hypoxia of the subcutaneous tissue, which on the one hand leads to the release of proangiogenic factors, but on the other hand also causes apoptosis and necrosis. The proangiogenic and prolymphogenic factors OLFLM3 and CNN1 are leading to angiogenesis in obese fatty tissue,55,56 and both are differentially expressed in lipedema. Therefore, tissue destruction increases, and a chronic inflammatory process of fatty tissue develops: in St. II and III, most inflammatory factors are increasingly expressed (see Fig. 3), leading to a destruction of the extracellular matrix (ECM). Three factors LUM, EMILIN2, and FBN1 are upregulated in lipedema and are components of the proteoglycan–microfibril scaffold of the ECM, which becomes degraded by ECM processing enzymes, resulting in a massive increase of mechanical stress.57 The tissue thus fibrosed fires up again the inflammation process, leading to swollen and painful tissue. More than 90% of patients with lipedema describe a neuropathic pain, which might be caused by sensory neurons’ activity intensified by this chronic inflammation.58 We identified 2 neuronal ion channels SLC12A2 and SCN7A, and 1 neuronal factor SHTN1, which are differentially expressed in lipedema and which might be involved in the origin and regulation of pain.59–61
Fig. 3.
Potential pathological processes of lipedema. The process of fatty tissue lipedema might be caused by changes in cell size, impaired lipid accumulation, lymphangiogenesis and angiogenesis, inflammation and ECM remodeling, and fibrosis. These processes are fired by hypoxia, vascular dysfunction, and immune response, which contributes to the development of a chronic inflammatory process. This ultimately leads to changes in the ECM resulting in fibrosed tissue with increased pain. Genes that are involved in the development of lipedema via these pathological processes based on our data analysis are shown in red.
Obesity as a Symptom of Lipedema
Obesity seems to be an intertwined comorbidity of lipedema, particularly with progress in increased stage. There is a difference in weight between the control group and the patients with lipedema caused by the increase of lipedema-associated fatty tissue. Considering the BMI, the WHR, and the WHtR, the WHR and the WHtR appear to represent a possibility to measure fat distribution rather than overweight, whereas BMI is the best method to describe the obesity effect (Supplemental Digital Content 4, http://links.lww.com/PRSGO/D615).
BMI-corrected Subgroup Analysis
The BMI-corrected subgroup analysis showed that some genes are exclusively associated with lipedema. ERBB4 is downregulated in lipedema and is predicted to be involved in the adipogenesis. ERBB4 might be a direct regulator of the differentiation of preadipocytes via the extracellular signal-regulated kinase/PPARγ pathway.62 It is known that ERBB4 is downregulated in adipose tissue. Further ERBB4 is a proangiogenic factor that is strongly expressed in healthy adipose tissue but downregulated in hypertrophied and hypoxic adipose tissue. The hypoxia might be caused by the reduced formation of blood vessels due to the lack of proangiogenic growth factor ERBB4.63
We identified BCAT1 upregulated in lipedema, which is predicted to be upregulated with weight gain, whereas downregulated with weight loss.64,65 There are different hypotheses about the effect of BCAT1: the cytosolic BCAT1 initiates the formation of branched chained amino acids, which are predicted to be involved in metabolic malfunction of different diseases such as obesity and diabetes, but the tissue-specific mechanism is actually unknown.66 In BCAT1 knockout mice, increased energy expenditure was found in a wide variety of tissues.67 It is postulated that BCAT1 is a proangiogenic factor and proliferation factor, leading to cell growth and tissue growth in tumors68 or might be involved in inflammation processes.69 The role of BCAT1 in adipose tissue is unclear, but it seems to be a key regulator of lipedema. BCAT is upregulated in lipedema St. II and III and even independently of obesity.
As named above, Lumican (LUM) is an ECM protein regulating adhesion of cells to the microfibril scaffold and linking interfibrillar molecules.70 LUM is also responsible to regulate cell growth, proliferation, autophagy, and apoptosis of different cells71 and might also be associated with the recruitment of neutrophils to inflamed tissue and wound healing. It is predicted that LUM regulates the Erk1/2 pathway, which is an important pathway in differentiation of adipocytes. In cell culture of adipocytes, it could be shown that higher levels of LUM lead to a forced adipogenesis.72 Known from cancer research, it is also discussed that LUM might be an antiangiogenic factor leading to a rarefication of blood vessels.73 LUM is an important factor for the integrity of the ECM. In the mouse model, it was found that overexpression of LUM leads to fibrosis and inflammation with a sex-specific alteration in LUM expression in adipose tissue of male and female mice.74
Several inflammation markers, such as CD84, CD163, and CD209, are upregulated in lipedema St. II and III, independent of the obesity effect. CD84 might be a membrane receptor of macrophages, which is predicted to be a key regulator of proinflammatory processes in adipose tissue.75,76 CD209 is known to be overexpressed in the adipose tissue of women supposed to inhibit adiponectin expression and upregulate leptin expression.77 CD163 and CD209 are both markers of anti-inflammatory M2 macrophages,78 which are regulated by different adipokines and cytokines as well as local hypoxia. Adipose tissue–specific macrophages might also be involved in regulating the development of adipose tissue and adipogenesis.79 Kruppa et al80 postulated also the importance of M2 macrophages in inflammation and the formation of fibrosis in lipedema confirming our data. CD163-positive macrophages are also predicted to be regulators of lipedema-associated activation of adipocyte differentiation from stem cells and increasing the lipid accumulation. Further, M2 macrophages are involved in the CD163-mediated hemoglobin scavenging mechanisms.81 This mechanism might lead to the release of anti-inflammatory cytokines. CD163 and hemoglobin are increased in the acute phase of inflammation and this mechanism is regarded to be a rescue mechanism to counteract the inflammation.82 This mechanism might be very important in lipedema-associated inflammation because we also identified hemoglobin subunit γ as well as an uncharacterized protein of the globin family downregulated in lipedema. This downregulation might be caused by compensating the oversupply of hemoglobin in acute inflammation of lipedema-associated fatty tissue.
The assumptions refer exclusively to the whole transcriptome analysis and must be validated by further investigations analyzing the tissue and cell specific protein levels and protein interaction mechanisms.
CONCLUSIONS
Lipedema seems to be a multifactorial disease with a dysregulation of cell differentiation, adipogenesis, and proliferation, leading to mechanical and hypoxia-induced cellular alterations. The pathological processes lead to inflammation, resulting in a cellular and extracellular remodeling with the expression of a lipedema-specific expression pattern of regulatory genes. This study elucidates aspects of the pathophysiology of lipedema and therefore gives a perspective for new therapeutic options in the future.
DISCLOSURES
The authors have no financial interest to declare in relation to the content of this article. The study was sponsored by the University Hospital Schleswig-Holstein, Luebeck, Germany. The patient counseling and sample collection of this study were supported by Hanse Clinic, Luebeck, Germany.
ACKNOWLEDGMENT
The authors acknowledge the financial support by Land Schleswig-Holstein within the funding program Open Access Publikationsfonds.
DECLARATION OF HELSINKI
This study was performed following the Declaration of Helsinki.
ETHICAL APPROVAL
The protocol was authorized by the institutional ethics committee (ethic vote, registration number 22-220) and is registered at ClinicalTrials.gov (NCT05861583).
Supplementary Material
Footnotes
Published online 8 November 2024.
Disclosure statements are at the end of this article, following the correspondence information.
Related Digital Media are available in the full-text version of the article on www.PRSGlobalOpen.com.
Clinical Relevance Statement: This study elucidates aspects of the pathophysiology of lipedema and therefore offers targets for new therapeutic options.
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary materials. Derived data supporting the findings of this study are available on request.
REFERENCES
- 1.Herpertz U. Das Lipödem [Lipedema]. Z Lymphol. 1995;19:1–11. [Article in German.] [PubMed] [Google Scholar]
- 2.Reich-Schupke S, Schmeller W, Brauer WJ, et al. S1-Leitlinie Lipodem. J Dtsch Dermatol Ges. 2017;15:758–768. [DOI] [PubMed] [Google Scholar]
- 3.Meier-Vollrath I, Schmeller W. [Lipoedema—current status, new perspectives]. J Dtsch Dermatol Ges. 2004;2:181–186. [DOI] [PubMed] [Google Scholar]
- 4.Child AH, Gordon KD, Sharpe P, et al. Lipedema: an inherited condition. Am J Med Genet A. 2010;152A:970–976. [DOI] [PubMed] [Google Scholar]
- 5.Shavit E, Wollina U, Alavi A. Lipoedema is not lymphoedema: a review of current literature. Int Wound J. 2018;15:921–928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sandhofer M, Hanke CW, Habbema L, et al. Prevention of progression of lipedema with liposuction using tumescent local anesthesia: results of an International Consensus Conference. Dermatol Surg. 2020;46:220–228. [DOI] [PubMed] [Google Scholar]
- 7.Kruppa P, Georgiou I, Biermann N, et al. Lipedema-pathogenesis, diagnosis, and treatment options. Dtsch Arztebl Int. 2020;117:396–403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Fife CE, Maus EA, Carter MJ. Lipedema: a frequently misdiagnosed and misunderstood fatty deposition syndrome. Adv Skin Wound Care. 2010;23:81–92; quiz 93. [DOI] [PubMed] [Google Scholar]
- 9.Langendoen SI, Habbema L, Nijsten TE, et al. Lipoedema: from clinical presentation to therapy. A review of the literature. Br J Dermatol. 2009;161:980–986. [DOI] [PubMed] [Google Scholar]
- 10.Katzer K, Hill JL, McIver KB, et al. Lipedema and the potential role of estrogen in excessive adipose tissue accumulation. Int J Mol Sci . 2021;22:11720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Schook CC, Mulliken JB, Fishman SJ, et al. Differential diagnosis of lower extremity enlargement in pediatric patients referred with a diagnosis of lymphedema. Plast Reconstr Surg. 2011;127:1571–1581. [DOI] [PubMed] [Google Scholar]
- 12.Szolnoky G, Varga E, Varga M, et al. Lymphedema treatment decreases pain intensity in lipedema. Lymphology. 2011;44:178–182. [PubMed] [Google Scholar]
- 13.Baumgartner A, Hueppe M, Schmeller W. Long-term benefit of liposuction in patients with lipoedema: a follow-up study after an average of 4 and 8 years. Br J Dermatol. 2016;174:1061–1067. [DOI] [PubMed] [Google Scholar]
- 14.Schmeller W, Hueppe M, Meier-Vollrath I. Tumescent liposuction in lipoedema yields good long-term results. Br J Dermatol. 2012;166:161–168. [DOI] [PubMed] [Google Scholar]
- 15.Schmeller W, Meier-Vollrath I. Tumescent liposuction: a new and successful therapy for lipedema. J Cutan Med Surg. 2006;10:7–10. [DOI] [PubMed] [Google Scholar]
- 16.Kirstein F, Hamatschek M, Knors H, et al. Patient-reported outcomes of liposuction for lipedema treatment. Healthcare (Basel). 2023;11:2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Dadras M, Mallinger PJ, Corterier CC, et al. Liposuction in the treatment of lipedema: a longitudinal study. Arch Plast Surg. 2017;44:324–331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Paolacci S, Precone V, Acquaviva F, et al. ; GeneOb Project. Genetics of lipedema: new perspectives on genetic research and molecular diagnoses. Eur Rev Med Pharmacol Sci. 2019;23:5581–5594. [DOI] [PubMed] [Google Scholar]
- 19.O’Brien SN, Welter BH, Mantzke KA, et al. Identification of progesterone receptor in human subcutaneous adipose tissue. J Clin Endocrinol Metab. 1998;83:509–513. [DOI] [PubMed] [Google Scholar]
- 20.Gray JM, Wade GN. Food intake, body weight, and adiposity in female rats: actions and interactions of progestins and antiestrogens. Am J Physiol. 1981;240:E474–E481. [DOI] [PubMed] [Google Scholar]
- 21.Van Pelt RE, Gozansky WS, Hickner RC, et al. Acute modulation of adipose tissue lipolysis by intravenous estrogens. Obesity (Silver Spring). 2006;14:2163–2172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gavin KM, Cooper EE, Hickner RC. Estrogen receptor protein content is different in abdominal than gluteal subcutaneous adipose tissue of overweight-to-obese premenopausal women. Metabolism. 2013;62:1180–1188. [DOI] [PubMed] [Google Scholar]
- 23.Corton M, Botella-Carretero JI, Benguria A, et al. Differential gene expression profile in omental adipose tissue in women with polycystic ovary syndrome. J Clin Endocrinol Metab. 2007;92:328–337. [DOI] [PubMed] [Google Scholar]
- 24.Boender AJ, van Rozen AJ, Adan RA. Nutritional state affects the expression of the obesity-associated genes Etv5, Faim2, Fto, and Negr1. Obesity (Silver Spring). 2012;20:2420–2425. [DOI] [PubMed] [Google Scholar]
- 25.Walley AJ, Jacobson P, Falchi M, et al. Differential coexpression analysis of obesity-associated networks in human subcutaneous adipose tissue. Int J Obes (Lond). 2012;36:137–147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kaare M, Mikheim K, Lillevali K, et al. High-fat diet induces pre-diabetes and distinct sex-specific metabolic alterations in Negr1-deficient mice. Biomedicines. 2021;9:1148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhang H, Chen X, Sairam MR. Novel genes of visceral adiposity: identification of mouse and human mesenteric estrogen-dependent adipose (MEDA)-4 gene and its adipogenic function. Endocrinology. 2012;153:2665–2676. [DOI] [PubMed] [Google Scholar]
- 28.Ali AT, Hochfeld WE, Myburgh R, et al. Adipocyte and adipogenesis. Eur J Cell Biol. 2013;92:229–236. [DOI] [PubMed] [Google Scholar]
- 29.Kang S, Nakanishi Y, Kioi Y, et al. Semaphorin 6D reverse signaling controls macrophage lipid metabolism and anti-inflammatory polarization. Nat Immunol. 2018;19:561–570. [DOI] [PubMed] [Google Scholar]
- 30.Jafari A, Qanie D, Andersen TL, et al. Legumain regulates differentiation fate of human bone marrow stromal cells and is altered in postmenopausal osteoporosis. Stem Cell Rep. 2017;8:373–386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Davis MR, Arner E, Duffy CR, et al. Expression of FBN1 during adipogenesis: relevance to the lipodystrophy phenotype in Marfan syndrome and related conditions. Mol Genet Metab. 2016;119:174–185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Diaz-Gonzalez F, Wadhwa S, Rodriguez-Zabala M, et al. Biallelic cGMP-dependent type II protein kinase gene (PRKG2) variants cause a novel acromesomelic dysplasia. J Med Genet. 2022;59:28–38. [DOI] [PubMed] [Google Scholar]
- 33.Yi X, Wu P, Liu J, et al. Candidate kinases for adipogenesis and osteoblastogenesis from human bone marrow mesenchymal stem cells. Mol Omics. 2021;17:790–795. [DOI] [PubMed] [Google Scholar]
- 34.Schiavinato A, Keene DR, Wohl AP, et al. Targeting of EMILIN-1 and EMILIN-2 to fibrillin microfibrils facilitates their incorporation into the extracellular matrix. J Invest Dermatol. 2016;136:1150–1160. [DOI] [PubMed] [Google Scholar]
- 35.Da Ros F, Persano L, Bizzotto D, et al. Emilin-2 is a component of bone marrow extracellular matrix regulating mesenchymal stem cell differentiation and hematopoietic progenitors. Stem Cell Res Ther. 2022;13:2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Buso G, Depairon M, Tomson D, et al. Lipedema: a call to action! Obesity (Silver Spring). 2019;27:1567–1576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kim S, Ahn C, Bong N, et al. Biphasic effects of FGF2 on adipogenesis. PLoS One. 2015;10:e0120073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Yoo A, Joo Y, Cheon Y, et al. Neuronal growth regulator 1 promotes adipocyte lipid trafficking via interaction with CD36. J Lipid Res. 2022;63:100221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Boender AJ, van Gestel MA, Garner KM, et al. The obesity-associated gene Negr1 regulates aspects of energy balance in rat hypothalamic areas. Physiol Rep. 2014;2:e12083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Espina AG, Mendez-Vidal C, Moreno-Mateos MA, et al. Induction of Dlk1 by PTTG1 inhibits adipocyte differentiation and correlates with malignant transformation. Mol Biol Cell. 2009;20:3353–3362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Karakus E, Schmid A, Leiting S, et al. Role of the steroid sulfate uptake transporter soat (Slc10a6) in adipose tissue and 3T3-L1 adipocytes. Front Mol Biosci. 2022;9:863912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lang P, Patlaka C, Andersson G. Tartrate-resistant acid phosphatase type 5/ACP5 promotes cell cycle entry of 3T3-L1 preadipocytes by increasing IGF-1/Akt signaling. FEBS Lett. 2021;595:2616–2627. [DOI] [PubMed] [Google Scholar]
- 43.Yang Y, Fan J, Xu H, et al. Long noncoding RNA LYPLAL1-AS1 regulates adipogenic differentiation of human mesenchymal stem cells by targeting desmoplakin and inhibiting the Wnt/beta-catenin pathway. Cell Death Discov. 2021;7:105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Kraus MR, Clauin S, Pfister Y, et al. Two mutations in human BICC1 resulting in Wnt pathway hyperactivity associated with cystic renal dysplasia. Hum Mutat. 2012;33:86–90. [DOI] [PubMed] [Google Scholar]
- 45.Nandipati KC, Subramanian S, Agrawal DK. Protein kinases: mechanisms and downstream targets in inflammation-mediated obesity and insulin resistance. Mol Cell Biochem. 2017;426:27–45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Nankam PAN, Cornely M, Kloting N, et al. Is subcutaneous adipose tissue expansion in people living with lipedema healthier and reflected by circulating parameters? Front Endocrinol (Lausanne). 2022;13:1000094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Guo L, Bertola DR, Takanohashi A, et al. Bi-allelic CSF1R mutations cause skeletal dysplasia of dysosteosclerosis-pyle disease spectrum and degenerative encephalopathy with brain malformation. Am J Hum Genet. 2019;104:925–935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wei S, Nandi S, Chitu V, et al. Functional overlap but differential expression of CSF-1 and IL-34 in their CSF-1 receptor-mediated regulation of myeloid cells. J Leukoc Biol. 2010;88:495–505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Moreno-Navarrete JM, Ortega F, Gomez-Serrano M, et al. The MRC1/CD68 ratio is positively associated with adipose tissue lipogenesis and with muscle mitochondrial gene expression in humans. PLoS One. 2013;8:e70810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Jo J, Gavrilova O, Pack S, et al. Hypertrophy and/or hyperplasia: dynamics of adipose tissue growth. PLoS Comput Biol. 2009;5:e1000324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Bischof J, Salzmann M, Streubel MK, et al. Clearing the outer mitochondrial membrane from harmful proteins via lipid droplets. Cell Death Discov. 2017;3:17016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kaartinen MT, Arora M, Heinonen S, et al. Transglutaminases and obesity in humans: association of F13A1 to adipocyte hypertrophy and adipose tissue immune response. Int J Mol Sci . 2020;21:8289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Dai C, Kayton NS, Shostak A, et al. Stress-impaired transcription factor expression and insulin secretion in transplanted human islets. J Clin Invest. 2016;126:1857–1870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Bove M, Monto F, Guillem-Llobat P, et al. NT3/TrkC pathway modulates the expression of UCP-1 and adipocyte size in human and rodent adipose tissue. Front Endocrinol (Lausanne). 2021;12:630097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Stalin J, Imhof BA, Coquoz O, et al. Targeting OLFML3 in colorectal cancer suppresses tumor growth and angiogenesis, and increases the efficacy of anti-PD1 based immunotherapy. Cancers (Basel). 2021;13:4625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Yang Y, Li S, Sun X, et al. CCN1 secreted by human adipose-derived stem cells enhances wound healing and promotes angiogenesis through activating the AKT signalling pathway. Int Wound J. 2023;20:1667–1677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Mariman EC, Wang P. Adipocyte extracellular matrix composition, dynamics and role in obesity. Cell Mol Life Sci. 2010;67:1277–1292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Chakraborty A, Crescenzi R, Usman TA, et al. Indications of peripheral pain, dermal hypersensitivity, and neurogenic inflammation in patients with lipedema. Int J Mol Sci . 2022;23:10313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Laird JM, Garcia-Nicas E, Delpire EJ, et al. Presynaptic inhibition and spinal pain processing in mice: a possible role of the NKCC1 cation-chloride co-transporter in hyperalgesia. Neurosci Lett. 2004;361:200–203. [DOI] [PubMed] [Google Scholar]
- 60.Ke CB, He WS, Li CJ, et al. Enhanced SCN7A/Nax expression contributes to bone cancer pain by increasing excitability of neurons in dorsal root ganglion. Neuroscience. 2012;227:80–89. [DOI] [PubMed] [Google Scholar]
- 61.Zeidler M, Kummer KK, Schopf CL, et al. NOCICEPTRA: gene and microRNA signatures and their trajectories characterizing human iPSC-derived Nociceptor maturation. Adv Sci (Weinh). 2021;8:e2102354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Wang Y, Zhang J, Chu X, et al. MiR-146a-5p, targeting ErbB4, promotes 3T3-L1 preadipocyte differentiation through the ERK1/2/PPAR-gamma signaling pathway. Lipids Health Dis. 2022;21:54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Nugroho DB, Ikeda K, Barinda AJ, et al. Neuregulin-4 is an angiogenic factor that is critically involved in the maintenance of adipose tissue vasculature. Biochem Biophys Res Commun. 2018;503:378–384. [DOI] [PubMed] [Google Scholar]
- 64.Chen J, Meng Y, Zhou J, et al. Identifying candidate genes for type 2 diabetes mellitus and obesity through gene expression profiling in multiple tissues or cells. J Diabetes Res. 2013;2013:970435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Mardinoglu A, Kampf C, Asplund A, et al. Defining the human adipose tissue proteome to reveal metabolic alterations in obesity. J Proteome Res. 2014;13:5106–5119. [DOI] [PubMed] [Google Scholar]
- 66.Vanweert F, Schrauwen P, Phielix E. Role of branched-chain amino acid metabolism in the pathogenesis of obesity and type 2 diabetes-related metabolic disturbances BCAA metabolism in type 2 diabetes. Nutr Diabetes. 2022;12:35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Heinonen S, Jokinen R, Rissanen A, et al. White adipose tissue mitochondrial metabolism in health and in obesity. Obes Rev. 2020;21:e12958. [DOI] [PubMed] [Google Scholar]
- 68.Shu X, Zhan PP, Sun LX, et al. BCAT1 activates PI3K/AKT/mTOR pathway and contributes to the angiogenesis and tumorigenicity of gastric cancer. Front Cell Dev Biol. 2021;9:659260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Papathanassiu AE, Ko JH, Imprialou M, et al. BCAT1 controls metabolic reprogramming in activated human macrophages and is associated with inflammatory diseases. Nat Commun. 2017;8:16040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Guzman-Ruiz R, Tercero-Alcazar C, Rabanal-Ruiz Y, et al. Adipose tissue depot-specific intracellular and extracellular cues contributing to insulin resistance in obese individuals. FASEB J. 2020;34:7520–7539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Giatagana EM, Berdiaki A, Tsatsakis A, et al. Lumican in carcinogenesis-revisited. Biomolecules. 2021;11:1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Strieder-Barboza C, Flesher CG, Geletka LM, et al. Lumican modulates adipocyte function in obesity-associated type 2 diabetes. Adipocyte. 2022;11:665–675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Sharma B, Ramus MD, Kirkwood CT, et al. Lumican exhibits anti-angiogenic activity in a context specific manner. Cancer Microenviron. 2013;6:263–271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Wolff G, Taranko AE, Meln I, et al. Diet-dependent function of the extracellular matrix proteoglycan Lumican in obesity and glucose homeostasis. Mol Metab. 2019;19:97–106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Ramalho T, Pahlavani M, Kalupahana N, et al. Eicosapentaenoic acid regulates inflammatory pathways through modulation of transcripts and miRNA in adipose tissue of obese mice. Biomolecules. 2020;10:1292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Verschuren L, Kooistra T, Bernhagen J, et al. MIF deficiency reduces chronic inflammation in white adipose tissue and impairs the development of insulin resistance, glucose intolerance, and associated atherosclerotic disease. Circ Res. 2009;105:99–107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Kim J, Lee SK, Kim D, et al. Adipose tissue LECT2 expression is associated with obesity and insulin resistance in Korean women. Obesity (Silver Spring). 2022;30:1430–1441. [DOI] [PubMed] [Google Scholar]
- 78.Fujisaka S, Usui I, Kanatani Y, et al. Telmisartan improves insulin resistance and modulates adipose tissue macrophage polarization in high-fat-fed mice. Endocrinology. 2011;152:1789–1799. [DOI] [PubMed] [Google Scholar]
- 79.Fujisaka S. The role of adipose tissue M1/M2 macrophages in type 2 diabetes mellitus. Diabetol Int. 2021;12:74–79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Kruppa P, Gohlke S, Lapinski K, et al. Lipedema stage affects adipocyte hypertrophy, subcutaneous adipose tissue inflammation and interstitial fibrosis. Front Immunol. 2023;14:1223264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Wolf S, Rannikko JH, Virtakoivu R, et al. A distinct M2 macrophage infiltrate and transcriptomic profile decisively influence adipocyte differentiation in lipedema. Front Immunol. 2022;13:1004609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Kristiansen M, Graversen JH, Jacobsen C, et al. Identification of the haemoglobin scavenger receptor. Nature. 2001;409:198–201. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.