Abstract
Gfi1 is a transcriptional repressor that plays a critical role in hematopoiesis. The repressive activity of Gfi1 is mediated mainly by its SNAG domain that interacts with and thereby recruits the histone demethylase LSD1 to its target genes. An important function of Gfi1 is to protect hematopoietic cells against stress-induced apoptosis, which has been attributed to its participation in the posttranscriptional modifications of p53 protein, leading to suppression of p53 activity. In this study, we show that Gfi1 upregulated the expression of Hemgn, a nuclear protein, through a 16-bp promoter region spanning from +47 to +63 bp relative to the transcription start site (TSS), which was dependent on its interaction with LSD1. We further demonstrate that Gfi1, Ikaros, and PU.1 are bound to this 16-bp region. However, while Ikaros activated Hemgn and collaborated with Gfi1 to augment Hemgn expression, it was not required for Gfi1-mediated Hemgn upregulation. In contrast, PU.1 repressed Hemgn and inhibited Hemgn upregulation by Gfi1. Notably, PU.1 knockdown and deficiency, while augmenting Hemgn expression, abolished Hemgn upregulation by Gfi1. PU.1 (Spi-1) is repressed by Gfi1. We show here that PU.1 repression by Gfi1 preceded and correlated well with Hemgn upregulation. Thus, our data strongly suggest that Gfi1 upregulates Hemgn by repressing PU.1. In addition, we demonstrate that Hemgn upregulation contributed to the anti-apoptotic activity of Gfi1 in a p53-independent manner.
Keywords: apoptosis, cell death, histone demethylase, transcription, transcription repressor, Gfi1, PU.1, Ikaros, Hemgn
Growth factor independence 1 (Gfi1) is a zinc-finger transcriptional repressor critically involved in hematopoiesis (1). Gfi1 is required for maintaining the quiescence and self-renewal capacity of hematopoietic stem cells (HSCs) and controls the development of HSCs into lymphoid and myeloid cells (2, 3, 4). Gfi1 deficiency significantly reduces the population of common lymphoid progenitors (CLP) and impairs T- and B-cell differentiation (5, 6). In the myeloid lineage, Gfi1 promotes neutrophil development at the expense of monocyte formation. Targeted deletion of Gfi1 in mice blocks neutrophil development leading to severe neutropenia accompanied by an expansion of atypical monocytes (7, 8, 9). As a transcriptional repressor, Gfi1 has been shown to repress genes that are key regulators of hematopoiesis, primarily by recruiting the histone demethylase LSD1 to the promoters of its target genes through its SNAG domain (10, 11).
Gfi1 has an important role in protecting hematopoietic cells against apoptosis. Overexpression of Gfi1 in T and B cells has been shown to abolish G1 cell cycle arrest and apoptosis induced by growth factor withdrawal (12, 13, 14). Targeted deletion of Gfi1 increases apoptosis in T-cell precursors and peripheral mature T-cells (15, 16). HSCs, T lymphoid, and myeloid precursors from Gfi1−/− mice are hypersensitive to stress-induced apoptosis (4). Gfi1 exerts its anti-apoptosis activity in part via its involvement in post-translational modifications of the C-terminal domain of the p53 protein, which inhibits p53 activity (17). We and others have shown that Gfi1 inhibited apoptosis through p53-independnt mechanisms (18, 19). Indeed, it has been shown recently that Gfi1 may inhibit apoptosis through modulation of sphingolipid metabolism by repressing SGPP1 in multiple myeloma (MM) cells and by regulating PRMT1-dependent methylation of proteins involved in DNA repair such as MRE11 and 53BP1, which is necessary for their function (20, 21).
Hemgn, also known as embryonic development-associated gene (EDAG) in humans, is a nuclear protein that is primarily expressed in HSCs and early progenitor cells (22, 23). HSCs from Hemgn−/− mice are functionally impaired, displaying defective engraftment activity in competitive repopulation assay that is associated with increased apoptosis and enhanced IFN-γ response (24). Conversely, Hemgn overexpression in human and mouse hematopoietic progenitor cells enhances their proliferation, survival, and self-renewal, which promotes their expansion in mice (25, 26, 27, 28, 29). When overexpressed, Hemgn also favors myelopoiesis over lymphopoiesis in mice (30). Hemgn is transcriptionally activated by HOXB4 and GATA1, and its expression is induced in response to DNA damage (24, 31, 32, 33). Significantly, overexpression of Hemgn has been observed in acute myeloid leukemia (AML) and acute lymphoblastic leukemia (ALL) and may confer resistance to chemotherapy (26, 27, 28, 29).
In this study, we show that Gfi1 upregulated Hemgn expression through the 16-bp region (+47/+63 bp relative to TSS) in the Hemgn promoter, which was dependent on its interaction with LSD1. We further demonstrate that Gfi1, Ikaros, and PU.1 are bound to this 16-bp region. Notably, Ikaros activated Hemgn, but was not required for Hemgn upregulation by Gfi1. In contrast, PU.1 repressed Hemgn, and its knockdown or deficiency abolished Hemgn upregulation. Gfi1 has been shown to repress PU.1 (34). We show that PU.1 repression by Gfi1 preceded and correlated well with Hemgn upregulation. Together, these data strongly suggest that Gfi1 upregulates Hemgn by repressing PU.1. In addition, we demonstrate that Hemgn upregulation contributed to the anti-apoptotic activity of Gfi1 in a p53-independent manner.
Results
Gfi1 upregulates Hemgn expression
Although Gfi1 is known as a transcriptional repressor, we were interested to know whether Gfi1 may also activate the expression of certain genes. Gene chip microarray was employed to assess the gene expression profile in the murine pro-B BaF3 cells transduced with a doxycycline (Dox)-inducible lentiviral expression construct encoding Gfi1 (BaF/Gfi1) (18). We observed the upregulation of several genes, notably including Hemgn, in BaF/Gfi1 cells in which Gfi1 expression was induced by the addition of Dox. Upregulation of Hemgn upon Dox induction was confirmed subsequently in BaF/Gfi1 cells by quantitative real-time RT-PCR (qRT-PCR) and Western blot analysis (Fig.1, A and B). We next examined whether Gfi1 upregulated Hemgn in murine myeloid 32D and human Burkitt's lymphoma Ramos cells transduced with the Dox-inducible Gfi1-expressing lentiviral construct (18, 35). As shown in Fig. 1, A and B, Hemgn mRNA and protein levels were significantly increased in Dox-treated 32D/Gfi1 and Ramos/Gfi1 cells. In contrast, Hemgn mRNA expression was significantly reduced in the Lin- bone marrow (BM) cells from Gfi1−/− mice as compared to Gfi1+/+ BM cells (Fig. 1C). Hemgn expression has been shown to be induced by DNA damage (24, 36). We further examined whether Gfi1 influenced Hemgn expression in response to DNA damage. BaF/Gfi1 and Ramos/Gfi1 cells were first treated with Dox to induce Gfi1 expression, followed by treatment with doxorubicin (Doxo) to induce DNA damage. As shown in Fig. 1D, Hemgn expression was augmented in response to Doxo treatment in both cell lines and treatment with a combination of Dox and Doxo led to further increases in Hemgn expression. Collectively, these results established that Gfi1 upregulates Hemgn expression in mouse and human hematopoietic cells. As Ramos cells lack functional p53, the results obtained in Ramos cells also indicated that Hemgn upregulation by Gfi1 and DNA damage was independent of p53.
Figure 1.
Gfi1 upregulates Hemgn expression.A and B, cells as indicated were left untreated (Ctr) or treated with Dox (1 μg/ml) for 24 h, followed by evaluation of Hemgn mRNA and protein expression by qRT-PCR (A) and Western blot analysis (B). C, Hemgn mRNA levels were analyzed in Lin- BM cells from Gfi1+/+ and Gfi1−/− mice. D, BaF/Gfi1 (left panel) and Ramos/Gfi1 (right panel) cells were left untreated (Ctr) or treated with Dox for 6 h and then incubated with or without Doxo for 16 h prior to evaluation of Hemgn mRNA levels. Data are presented as mean ± SD (N = 3). Statistically significant differences in Hemgn expression levels: ∗∗p < 0.01; ∗∗∗p < 0.001.
LSD1 interaction is required for Gfi1-mediated Hemgn upregulation
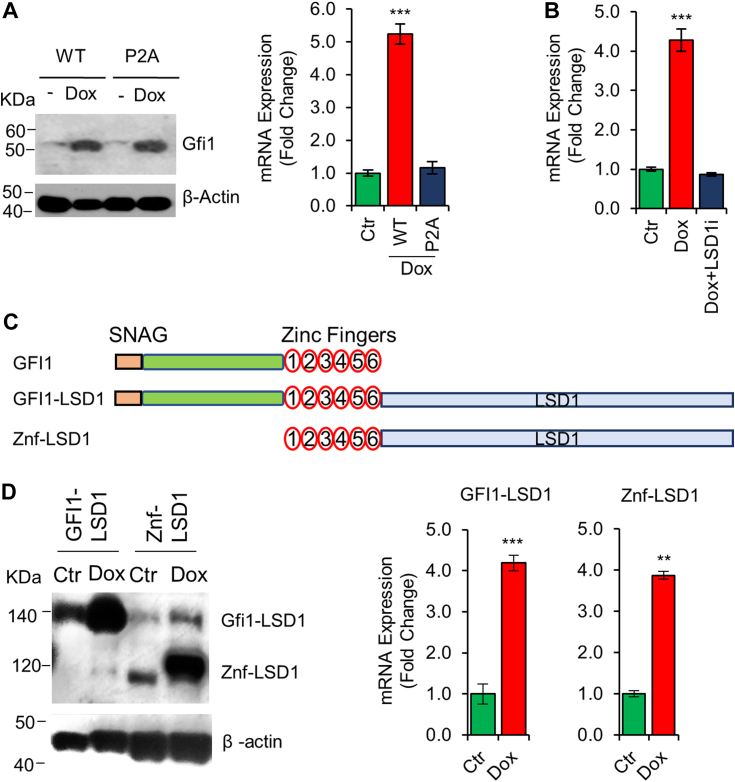
It has been shown that the SNAG domain of Gfi1 is required for Gfi1-mediated transcriptional repression by interacting with LSD1. A P2A point mutation in the SNAG domain impedes Gfi1 interaction with LSD1, rendering it inactive as a transcriptional repressor (1, 37). We examined the impact of the P2A mutation on Gfi1-mediated Hemgn upregulation in BaF3 cells transduced with the Dox-inducible Gfi1 P2A mutant (Fig. 2A). Gfi1-mediated Hemgn upregulation was abolished by the P2A mutation, suggesting that Hemgn upregulation by Gfi1 relied on its capability to recruit LSD1 via the SNAG domain. We further investigated the effect of an LSD1 inhibitor GSK2879552 (LSD1i), which inhibits LSD1 histone demethylase activity, on Gfi1-mediated Hemgn upregulation. As shown in Fig. 2B, LSD1i treatment effectively suppressed Hemgn upregulation by Gfi1 in BaF/Gfi1 cells.
Figure 2.
LSD1 is required for Gfi1-mediated Hemgn upregulation.A, BaF/Gfi1 and BaF/Gfi1P2A cells were left untreated or treated with Dox (1 μg/ml) for 24 h and examined for expression of Gfi1 proteins (left panel) and Hemgn mRNA (right panel). B, BaF/Gfi1 cells were treated with Dox overnight and then with LSD1i (1 μM) for 24 h prior to evaluation of Hemgn mRNA levels. C, schematic diagrams of Gfi1-LSD1 and Znf-LSD1 fusion proteins. D, BaF3 cells expressing the Dox-inducible fusion proteins as indicated were left untreated or treated with Dox for 24 h and then examined for expression of the fusion proteins using the anti-Flag antibody (left panel) and Hemgn mRNA (right panel). Data are shown as mean ± SD (n = 3). Statistical significances: ∗∗p < 0.01; ∗∗∗p < 0.001.
We sought to examine if LSD1 interaction alone is sufficient for Gfi1-mediated Hemgn upregulation. To address this, BaF3 cells were transduced with the Dox-inducible lentiviral constructs encoding two different fusion proteins consisting of LSD1 fused to either the full-length Gfi1 (Gfi1-LSD1) or only the 6 C-terminal zinc finger domains that are required for DNA binding (ZNF-LSD1) (Fig. 2C). The expression of the fusion proteins upon Dox induction was confirmed by Western blotting analysis (Fig. 2D). Notably, Gfi1-LSD1 and ZNF-LSD1 were equally capable of upregulating Hemgn expression upon induction of their expression with Dox. Together, these results indicated that LSD1 interaction is essential and sufficient for Gfi1-mediated upregulation of Hemgn.
A 16-bp sequence of Hemgn spanning from +47 to +63 bp is required for Gfi1-mediated upregulation
We next investigated whether Gfi1 activated the Hemgn promoter. Consistent with upregulation of Hemgn mRNA expression, Gfi1 activated the Hemgn promoter fragment spanning from −1972 to +63 bp in luciferase reporter assay (fragment I; Fig. 3A). Progressive truncation of the Hemgn promoter fragment from the 5′ end revealed that a fragment spanning from −70 to +63 bp (fragment VI) was sufficient for its activation by Gfi1 (Fig. 3B). However, Gfi1 failed to activate the Hemgn promoter fragments spanning from −88 to −34 bp and −51 to −1 bp that were placed upstream of the adenovirus minimal promoter MLP (major late promoter), suggesting that the region responsible for Gfi1-mediated activation lies downstream of TSS. Interestingly, the deletion of a 16-bp nucleotide sequence from the 3′ ends of fragments I and II abolished their activation by Gfi1. We then deleted the various regions downstream of the TSS in the Hemgn promoter fragment VI (−70/+63 bp) to more precisely map the region important for Gfi1-mediated activation. Like fragments I and II, deletion of the 16-bp sequence from the 3′ end abrogated its activation by Gfi1 although the 3′ region of 30 bp was required for optimal activation (fragment VIf). Together, these data demonstrated that the +47/+63-bp region of the Hemgn promoter is critical for its activation by Gfi1.
Figure 3.
The 16-bp region spanning from +47 to +63 bp of Hemgn is essential for Gfi1-mediated Hemgn upregulation.A, schematic diagrams of the various mouse Hemgn promoter fragments (upper panel) and the different deletion mutants derived from fragment VI (middle panel), and the nucleotide sequence of Hemgn promoter spanning from +33 to +63 bp and the mutations introduced in this region of fragment II. B, the reporter constructs containing the different promoter fragments were transfected into BaF/Gfi1 cells, followed by treatment with Dox (1 μg/ml) for 24 h. Promoter activation, shown as fold changes as compared to untreated cells, was determined. Data are shown as mean ± SD (n = 3). ∗∗p < 0.01; ∗∗∗p < 0.001.
Analysis of the 30-bp sequence in the 3′ region utilizing transcription factor binding prediction software (http://tfbind.hgc.jp/) identified potential binding sites for Gfi1, Ikaros, and PU.1. Notably, PU.1 shares its consensus binding motif with Ikaros (38). We introduced mutations in the Gfi1 and Ikaros/PU.1 binding site either alone or in combination in the Hemgn promoter fragment II (−567/+63 bp). Interestingly, mutation of either the Gfi1 or Ikaros/PU.1 binding site significantly diminished Gfi1-mediated activation whereas mutation of both binding sites completely abolished its activation by Gfi1, suggesting that Gfi1 may collaborate with Ikaros/PU.1 in the activation of the Hemgn promoter.
Gfi1, Ikaros, and PU.1 bind to the Hemgn core promoter
Chromatin immunoprecipitation (ChIP) assays were then performed to investigate whether Gfi1 and Ikaros bind to the Hemgn core promoter in BaF/Gfi1, BaF/Ik, and BaF/Gfi1/Ik cells, which expressed Gfi1, Ikaros, or both in response to Dox (Fig. 4B). As shown in Fig. 5A, both Gfi1 and Ikaros bound to the Hemgn core promoter, but not to the 3-kb upstream region. BaF3 cells expressed a very low level of endogenous Gfi1 that was barely detectable by Western blot analysis (data not shown). It appeared that overexpression of Gfi1 increased Ikaros binding to the Hemgn core promoter. On the other hand, Ikaros overexpression had no effect on Gfi1 binding to the Hemgn core promoter.
Figure 4.
Gfi1 and Ikaros bind to the Hemgn core promoter.A, cells as indicated were cultured with and without Dox for 24 h. ChIP assays were performed using antibodies against Gfi1 and Ikaros or IgG as a control, followed by PCR to amplify the Hemgn core promoter and the 3-kb upstream region. B, expression of Gfi1 and Ikaros in BaF/Ik and BaF/Gfi1/Ik cells. C, BaF3/Gfi1/Ik cells were transfected with the reporter plasmid containing fragment I or fragment VII (upper panel). The green and red arrows denote the forward and reverse primers, respectively, used to amplify the proximal (P1) and distal (P2) plasmid sequences. ChIP experiments were conducted as in A using cells transfected with fragment I (middle panel) and fragment VII (bottom panel). Data are shown as mean ± SD (n = 3). ∗∗p < 0.01; ∗∗∗p < 0.001.
Figure 5.
PU.1 binds to the Hemgn core promoter. BaF/Gfi1/PUER cells were transfected with the reporter plasmid containing fragment I or fragment VII, and cultured without or with Dox (1 μg/ml) and 4-HT (100 nM) for 24 h. ChIP experiments were carried out using antibodies against Gfi1, PU.1 or IgG. A, qPCR was conducted to amplify the Hemgn core promoter and the 3 kb upstream region in cells transfected with fragment VII (upper panel). Expression of PUER along with endogenous PU.1 and Gfi1 was examined (lower panel). B, the P1 and P2 plasmid sequences were amplified using cells transfected with fragment I (upper panel) or fragment VII (lower panel). Data are shown as mean ± SD (n = 3). ∗∗p < 0.01; ∗∗∗p < 0.001.
We further conducted plasmid-ChIP experiments to determine whether Gfi1 and Ikaros are specifically bound to the 16-bp (+47/+63 bp) region of the Hemgn promoter in BaF/GFI1/Ik cells. The cells were transfected with the reporter plasmid containing either fragment I (−1972/+63 bp) or fragment VII (−1972/+47 bp) lacking the 16 bp region (Fig. 3). After ChIP with antibodies against Gfi1 or Ikaros, the precipitated DNA was analyzed by PCR with primers that amplify the plasmid DNA sequences 215 bp and approximately 2.5 kb (as a negative control) downstream of the Hemgn promoter fragments (Fig. 4C). Notably, Gfi1 and Ikaros bound to the Hemgn promoter fragment I, but not to fragment VII or the plasmid sequence 2.5 kb downstream, demonstrating that they are both specifically bound to the 16-bp region of the Hemgn promoter.
To examine whether PU.1 bound to the 16-bp sequence of Hemgn promoter, we generated BaF/Gfi1 cells that constitutively expressed PUER in which PU.1 was fused to the estrogen receptor (ER) ligand binding domain, allowing PU.1 to be conditionally activated by 4-hydroxytamaxifen (4-HT). BaF/Gfi1/PUER cells were subsequently transfected with the reporter plasmid containing fragment I or fragment VII and cultured without or with Dox and 4-HT for 24 h, followed by ChIP with the anti-Gfi1 or anti-PU.1 antibody. PCR was then performed to examine the binding of PU.1 and Gfi1 to the endogenous Hemgn and to the promoter fragments I/VII with the primers utilized in the above ChIP experiments. As the PCR primers used to amplify the endogenous Hemgn core promoter did not amplify fragment VII, Gfi1 and PU.1 binding to endogenous Hemgn were examined in cells transfected with fragment VII. As shown in Fig. 5, like Gfi1 and Ikaros, PU.1 bound to the Hemgn core promoter and the 16-bp sequence was required for its binding.
Together, these results suggested a potential role of Ikaros and PU.1 in the regulation of Hemgn expression; however, it remained to be determined whether they are involved in Gfi1-mediated Hemgn upregulation.
Ikaros weakly activates Hemgn but is not essential for Hemgn upregulation by Gfi1
We first investigated the role of Ikaros in the regulation of Hemgn expression. BaF/Ik and BaF/Gfi1/Ik cells were transiently transfected with the reporter plasmid containing fragment I or fragment VII (Fig. 3A). Dox treatment resulted in a two-fold increase in the activity of fragment I, but not fragment VII in BaF/Ik cells (Fig. 6A). Interestingly, fragment I, but not fragment VII, was strongly activated (6 fold) upon Dox treatment of BaF/Gfi1/Ik cells. Consistent with promoter activation, Hemgn expression was upregulated by approximate two folds in BaF/Ik cells, more than 4.5 folds in BaF/Gfi1 cells, and about 10.8 folds in BaF/Gfi1/Ik cells following Dox treatment (Fig. 6B). To address whether Ikaros was required for Hemgn upregulation by Gfi1, we introduced the Dox-inducible Gfi1 into the mouse Ikaros null JE131 T cells (JE131/Gfi1). Notably, Hemgn expression was still upregulated by Gfi1 in the absence of Ikaros (Fig. 6C). Constitutive restoration of Ikaros expression in JE131/Gfi1 cells (JE131/Gfi1/Ik) modestly increased Hemgn expression, but significantly augmented Gfi1-mediated Hemgn upregulation. Together, these results indicated that Ikaros is not required for Gfi1-mediated Hemgn upregulation although it had a weak positive effect on, and collaborated with Gfi1 to enhance Hemgn expression.
Figure 6.
Ikaros collaborates with Gfi1 to activate Hemgn promoter.A, BaF/Ik and BaF/Gfi1/Ik cells were transfected with reporter plasmids containing fragment I or VII and then cultured without (Ctr) or with Dox (1 μg/ml) for 24 h prior to examination of luciferase activity. B, Hemgn mRNA levels were examined in BaF/Gfi1, BaF/Ik and BaF/Gfi1/Ik cells cultured in the absence (Ctr) or presence of Dox for 24 h. C, Hemgn mRNA levels were examined in JE131/Gfi1, JE131/Ik and JE131/Gfi1/Ik cells untreated or treated with Dox for 24 h (left panel). Data are shown as mean ± SD (n = 3). ∗∗p < 0.01; ∗∗∗p < 0.001. Expression of Gfi1 and Ikaros proteins was confirmed by Western blot analysis (right panel).
PU.1 represses Hemgn and is required for Hemgn upregulation by Gfi1
We next focused on the role of PU.1 in the regulation of Hemgn expression. In luciferase reporter assay, activation of PU.1 with 4-HT repressed the activity of Hemgn promoter fragment I in BaF/Gfi1/PUER cells irrespective of Dox treatment but had no effect on fragment VII (Fig. 7A), suggesting that the +47/63 bp region of the Hemgn promoter is critical for PU.1-mediated repression. Significantly, 4-HT treatment markedly downregulated Hemgn mRNA expression and completely abolished Gfi1-mediated Hemgn upregulation (Fig. 7B). As in BaF3 cells, PU.1 downregulated Hemgn expression in murine myeloid 32D cells and Lin- BM cells.
Figure 7.
PU.1 represses Hemgn and inhibits Gfi1-mediated activation of Hemgn.A, BaF/Gfi1/PUER cells were transfected with reporter plasmids containing fragment I or VII and cultured without or with Dox (1 μg/ml), 4HT (100 μM) or both for 24 h prior to evaluation of luciferase activity. B, BaF/Gfi1/PUER, 32D and Lin- BM cells transduced with PUER retroviral construct were cultured without or with 4HT, Dox or both as indicated for 24 h prior to examination of Hemgn mRNA levels by qRT-PCR. Data are shown as mean ± SD (n = 3). ∗∗p < 0.01; ∗∗∗p < 0.001.
Gfi1 has been shown to repress PU.1 and inhibit PU.1 function through direct protein-protein interaction (5, 34). Consistent with these studies, Gfi1 downregulated PU.1 expression in BaF/Gfi1 cells at both mRNA and protein levels (Fig. S1A). In contrast, PU.1 mRNA was significantly increased in Lin- BM cells from Gfi1−/− mice (Fig. S1B). To address whether Gfi1 upregulated Hemgn through repressing PU.1, we introduced the Dox-inducible Gfi1 into myeloid PUER cells, which were derived from PU.1−/− mice and transduced with PUER. In the absence of 4-HT, PUER is inactive. As shown in Fig. 8A, treatment of PUER cells with Dox downregulated rather than upregulated Hemgn. We further evaluated the effect of PU.1 knockdown on Hemgn upregulation by Gfi1. The expression of PU.1 was knocked down in BaF/Gfi1 cells using two different PU.1 shRNAs (Fig. 8B). PU.1 knockdown resulted in augmented expression of Hemgn mRNA but abolished Gfi1-mediated Hemgn upregulation.
Figure 8.
PU.1 is required for Gfi1-mediated upregulation of Hemgn.A, Hemgn mRNA (left panel) and protein (right panel) levels were examined in PUER/Gfi1 cells untreated or treated with Dox (1 μg/ml) for 24 h. B, BaF/Gfi1 cells were transduced with empty or two different PU.1 shRNA-expressing lentiviruses (498 and 501) and examined for Hemgn expression following Dox induction for 24 h (upper panel). PU.1 knockdown was confirmed by Western blot analysis (lower panel). C, BaF/Gfi1 cells were treated with Dox for the indicated times and examined for expression of Gfi1 and PU.1 proteins (upper panel) and Hemgn mRNA (middle panel). Lower panel: Graphical presentation of Gfi1, PU.1, and Hemgn expression. The levels of Gfi1 and PU.1 proteins were based on the densities of bands determined using Image J software. Data are shown as mean ± SD (n = 3). ∗∗p < 0.01; ∗∗∗p < 0.001.
If Gfi1 upregulated Hemgn through repressing PU.1, downregulation of PU1 should occur before Hemgn upregulation. We therefore examined the dynamics of Gfi1, PU.1 and Hemgn expression in BaF/Gfi1 cells following Dox treatment. Gfi1 protein expression was induced as early as 4 h following Dox induction whereas PU.1 downregulation occurred at approximately 6 to 9 h (Fig. 8C). Notably, upregulation of Hemgn mRNA was observed at 12 h post Dox treatment. Thus, PU.1 downregulation by Gfi1 correlated well with Hemgn upregulation in BaF/Gfi1 cells. Taken together, these results revealed an indirect regulatory mechanism of Hemgn upregulation by Gfi1, i.e., through repression of PU.1.
Hemgn has a role in Gfi1-mediated protection against DNA damage-induced apoptosis
Gfi1 has been shown to inhibit apoptosis induced by DNA damage in hematopoietic cells in part through post-translational modifications at the C-terminal domain of the p53 protein, leading to suppression of p53 activity (39). However, our previous study showed that Gfi1 also inhibited DNA damage-induced apoptosis in p53-deficient hematopoietic cells (18). Notably, it has been shown that Hemgn was induced in response to DNA damage and inhibited DNA damage-induced apoptosis (24, 36). We examined whether the protective activity of Gfi1 is mediated in part through Hemgn. The expression of Hemgn in BaF/Gfi1 cells was knocked down using two different shRNAs (Fig. 9D). A shRNA against human HEMGN was utilized to knock down its expression in the p53-deficient Ramos/Gfi1 cells. The cells were then cultured without or with Dox for 6 h followed by treatment with Doxo for 24 h. Apoptosis was evaluated by Annexin V staining for BaF/Gfi1 cells and by MTS assay for Ramos/Gfi1 cells. MTS assay instead of annexin V staining was used because Ramos/Gfi1 cells were treated with Doxo at 2 mg/ml, which interfered with flow cytometric analysis of annexin V-stained cells (18). Doxo treatment decreased the viabilities of BaF/Gfi1 and Ramos/Gfi1 cells (Fig. 9). Induction of Gfi1 expression with Dox led to an increase in cell viability by 24.5% in Doxo-treated BaF/Gfi1 cells and by 34.6% in Ramos/Gfi1 cells (Fig. 9, B and C). However, the increases in cell viability following Dox-induction of Gfi1 expression dropped to approximately 10% and 18% in BaF/Gfi1 and Ramos/Gfi1 cells, respectively, upon knockdown of Hemgn expression. To further demonstrate that suppression of DNA damage-induced apoptosis by Hemgn was independent of p53, we transduced Ramos cells with the Hemgn-expressing retroviral construct. As shown in Fig. S2, Hemgn overexpression inhibited apoptosis of Ramos cells treated with Doxo.
Figure 9.
Knockdown of Hemgn diminishes the protective effect of Gfi1 on DNA damage-induced apoptosis. BaF/Gfi1 and Ramos/Gfi1 cells were transduced with empty or Hemgn shRNA-expressing lentiviral constructs, followed by treatment with Dox (1 μg/ml) for 6 h prior to treatment with Doxo (200 ng/ml for BaF/Gfi1 cells and 2 mg/ml for Ramos/Gfi1 cells) for 24 h. A, the apoptosis of BaF/Gfi1 cells was examined by Annexin V assay. Shown is a representative flow cytometry experiment. The numbers denote percentages of live cells. B, Data from 3 independent flow cytometry experiments are presented. C, live Ramos/Gfi1 cells were quantitated using MTS assay. The numbers in (B) and (C) indicate increases in the viability of Doxo-treated cells cultured in the presence versus absence of Dox, and are shown as mean ± SD (n = 3). ∗∗p < 0.01; ∗∗∗p < 0.001. D, Hemgn expression was examined by Western blot analysis.
BM cells from Gfi1−/− mice showed increased sensitivity to DNA damage-induced apoptosis (4). As Hemgn expression in Gfi1−/− BM cells was decreased (Fig. 1C), we assessed whether restoration of Hemgn expression rescued the hypersensitivity of Gfi1−/− BM cells to DNA damage. Lin- BM cells from Gfi1−/− mice were transduced with an empty retroviral construct or Hemgn-expressing construct and sorted for GFP expression 48 h later. Hemgn expression was partially restored in the sorted BM cells (Fig. 10C). After treatment with Doxo, apoptosis was evaluated by Annexin V staining. The percentages of Annexin V-positive (apoptotic) cells were approximately 10% and 39% for WT and Gfi1−/− BM cells, respectively (Fig. 10B). Partial restoration of Hemgn expression in Gfi1−/− BM cells reduced the percentage of apoptotic cells to about 24%. These results reveal the role of Hemgn in Gfi1-mediated protection against DNA damage-induced apoptosis.
Figure 10.
Restoration of Hemgn expression in Gfi1−/−BM cells partially rescues the hypersensitivity to DNA damage. Lin− cells were isolated from WT and Gfi1−/− mice. Gfi1−/− cells were transduced with the empty or Hemgn-expressing retrovirus and sorted based on GFP. Cells were then treated with Doxo (50 ng/ml) for 24 h and examined for apoptosis by flow cytometry based on annexin V staining. A, shown is a representative experiment of flow cytometry. B, data from 3 independent flow cytometry experiments are presented. Data are shown as mean ± SD (n = 3). ∗∗p < 0.01; ∗∗∗p < 0.001. C, expression of Hemgn protein in the different cell populations was examined by Western blot analysis.
Discussion
Gfi1 is a transcriptional repressor critically involved in hematopoiesis. Transcriptional repression by Gfi1 is mediated mainly through interaction with LSD1. In this study, we examined whether Gfi1 may function to augment gene expression. We show here that Gfi1 upregulates Hemgn in both mouse and human hematopoietic cells. The +47/63 bp region of Hemgn promoter is required for Gfi1-mediated upregulation, which is dependent on its interaction with LSD1. This 16-bp region contains conserved binding sites for Gfi1 and Ikaros/PU.1. Using ChIP assay, we show that all three transcription factors bind to the 16-bp region. However, while Ikaros activates the Hemgn promoter and collaborates with Gfi1 to augment Hemgn expression, it is not required for Gfi1-mediated upregulation of Hemgn. In contrast, PU.1 represses Hemgn and blocks Gfi1-mediated Hemgn upregulation. Significantly, Gfi1 is unable to upregulate Hemgn in PU.1-knocked down and PU.1-deficient cells. In line with previous studies (5, 34), we show that Gfi1 represses PU.1 and that Gfi1-mediated PU.1 downregulation precedes Hemgn upregulation. These results are consistent with a mode of action in which Gfi1 represses PU.1, leading to Hemgn derepression.
Hemgn is a nuclear protein that is primarily expressed in HSCs and early progenitor cells (22, 23). Hemgn overexpression in human and mouse hematopoietic stem and progenitor cells (HSPCs) has been shown to enhance their proliferation, survival, and self-renewal (25, 27, 40, 41). Interestingly, like Gfi1−/− HSCs, Hemgn−/− HSCs are also impaired in their ability to reconstitute hematopoiesis in lethally irradiated recipient mice (24). An important function of Gfi1 is to inhibit stress-induced apoptosis, which is important for its role in hematopoiesis, in particular for maintaining the functional integrity of HSCs and the development of T lymphocytes (3, 4, 42). The anti-apoptotic activity of Gfi1 is also critical for its oncogenic potential in lymphoid leukemia and lymphoma (43). It has been shown that Gfi1 protects cells from DNA damage-induced apoptosis in part through inhibiting p53 function due to its involvement in post-translational modifications of p53 protein (39). However, our previous study indicated that Gfi1 may inhibit DNA damage-induced apoptosis via p53-independent mechanisms (18). In this study, we have shown that Hemgn knockdown partially abolished the protective effect of Gfi1 on DNA damage-induced apoptosis in both p53-expressing and p53-deficient cells. In contrast, partial restoration of Hemgn expression in Gfi1−/− BM cells diminished the hypersensitivity to DNA damage. It is of note that Hemgn has been shown to be upregulated and phosphorylated in response to DNA damage and inhibit stress-induced apoptosis (24, 33, 36). We have further shown here that Gfi1 and DNA damage act in collaboration to augment Hemgn expression in a p53-independent manner. Our data indicate that Hemgn upregulation contributes to the anti-apoptotic function of Gfi1 and suggests that Hemgn may have an important role in DNA damage response.
Our data also reveal Hemgn as a novel PU.1-repressed target gene. Interestingly, Hemgn has been shown to upregulate the expression of PU.1 in HSCs (34). PU.1 is a master regulator of hematopoiesis that is important for self-renewal of HSCs and critically implicated in regulating myeloid and lymphoid development in a graded manner (44, 45, 46, 47, 48). Notably, PU.1 has been shown to function as a potent tumor suppressor in myeloid and B cell lineages (49, 50, 51, 52). A reduction in PU.1 expression to 20% of normal levels induced the development of AML in mice (49). Conversely, increased expression of PU.1 led to growth arrest and apoptosis in leukemic, multiple myeloma, and lymphoma cells (51, 52, 53). The mechanisms by which PU.1 induces growth arrest and apoptosis are incompletely understood but may involve PU.1-mediated downregulation of c-Myc, Bcl-2, and IRF4 (52, 53). We show here that PU.1 represses Hemgn by binding to its +47/+63 region. As Hemgn enhances cell proliferation and survival, and its expression is increased in AML and ALL cells, it would be interesting to examine whether repression of Hemgn contributes to the role of PU.1 as a tumor suppressor.
We further show here that contrary to PU.1, Ikaros transcriptionally activates Hemgn and collaborates with Gfi1 to upregulate Hemgn expression. Interestingly, Ikaros has been shown to collaborate with Gfi1 to activate gene expression by co-occupying the regulatory regions of target genes, including NOTCH3 (54). However, Ikaros deficiency did not affect Gfi1-mediated Hemgn upregulation, indicating that Ikaros is not required for Hemgn upregulation by Gfi1. Like PU.1, Ikaros is a master regulator of hematopoiesis (55, 56, 57) and functions as an important tumor suppressor in the hematopoietic system, particularly in B cell lineage (57, 58, 59). Interestingly, in contradiction to its role as a tumor suppressor, immunomodulatory imide drugs (IMiDs), which induce Ikaros degradation resulting in reduced proliferation and survival of cancer cells, are frequently used in combination therapies for patients with multiple myeloma, lymphoma, and certain forms of myelodysplastic syndrome (57, 59). It appears that the activity of Ikaros may depend on cell types and developmental stages. For instance, Ikaros is involved in transcriptional repression of the CDK inhibitor Cdkn1a encoding p21WAF1/CIP1 in certain hematopoietic cells (57). It is tempting to evaluate whether Ikaros may positively regulate cell proliferation and survival by transcriptionally activating Hemgn in certain cells.
Experimental procedures
Cell lines and cell culture
The maintenance of murine pro-B Ba/F3, myeloid 32D, and human Burkitt lymphoma cell lines has been described before (18, 35, 60). Murine Ikaros-null T-cell leukemia cell line JE131 (61) was cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (P/S). PUER cells (62) were kindly provided by Dr Richard Dahl (Indiana University School of Medicine) and cultured in RPMI-1640 medium with 10% charcoal-stripped FBS and 1% P/S. Ba/F3, 32D and Ramos cells transduced with the Dox-inducible lentiviral expression construct pTMPrtTA-Gfi1-GFP encoding Gfi1 and GFP have been described before (18, 35, 63).
Expression constructs and stable gene delivery
The Dox-inducible Ikaros-expressing vector pTMPrtTA-Ik-GFP was generated by replacing the Gfi1 cDNA in pTMPrtTA-Gfi1-GFP with the Ikaros cDNA (61), kindly provided by Dr Andrew Wells (The Children’s Hospital of Philadelphia), and then introduced into Ba/F3 cells through lentiviral transduction as described (18, 63) to generate BaF/Ik cells. The GFP cDNA in pTMPrtTA-Ik-GFP was replaced with RFP cDNA to generate pTMPrtTA-Ik-RFP, which was then transduced into BaF/Gfi1 cells expressing the Dox-inducible Gfi1 (BaF/Gfi1) to generate BaF/Gfi1/Ik cells. The Dox-inducible pLentiGS-minCMV-TET-puromycin constructs containing Gfi1-LSD1 or Znf-LSD1 cDNA (64) were kindly provided by Dr Tim Somervaille (Cancer Research UK Manchester Institute, The University of Manchester) and transduced into Ba/F3 cells, followed by puromycin selection (2 μg/ml). For generation of JE131 cells reconstituted with Ikaros and expressing Dox-inducible Gfi1, cells were first transduced with pTMPrtTA-Gfi1-GFP and then with MIGR1-Ikaros (65). PUER cells expressing the Dox-inducible Gfi1 (PUER/Gfi1) were generated by lentiviral transduction with pTMPrtTA-Gfi1-GFP.
Construction of luciferase promoter constructs and luciferase reporter assay
A murine Hemgn promoter fragment spanning from −1972 bp to +63 bp (fragment I; see Fig. 3A) was amplified from mouse genomic DNA and cloned into the MluI and XhoI sites of the pGL3 basic vector (Promega, Madison, WI). The other Hemgn promoter fragments were derived from fragment I using the PCR-base technique. Mutations in Hemgn promoter fragment II were generated by site-directed mutagenesis using the Quick-change site-directed mutagenesis kit (Stratagene). The nucleotide sequences of all promoter fragments and mutants were confirmed by DNA sequencing. Ba/F3 cells were transfected with the reporter constructs containing the different Hemgn promoter fragments by electroporation. Approximately 2 hours post-transfection, cells were either left untreated or treated with Dox (1 μg/ml) and/or 4-HT (100 nM). Luciferase activities were measured 24 h post-transfection using a Molecular Devices Lmax luminometer. All luciferase values are shown as mean ± SD of three independent experiments.
Real-time reverse transcription polymerase chain reaction (qRT-PCR)
Total RNA was extracted with TRIzol reagent (ABP Biosciences) and reverse transcribed into cDNAs using the LunaScript RT SuperMix Kit (NEB) according to the manufacturer's instructions. The relative mRNA levels of various genes were quantified by qRT-PCR using iTaq Universal SYBR Green Supermix (Bio-Rad) and normalized to GAPDH mRNA expression as described previously (60).
Chromatin immunoprecipitation (ChIP assay) and plasmid ChIP assay
Cells were fixed with 1% formaldehyde for 10 min to stabilize protein-DNA complexes, followed by the addition of 0.125 M glycine to terminate the cross-linking process. Subsequent steps were performed using the Simple ChIP Enzymatic Chromatin IP Kit according to the manufacturer's protocol (Cell Signaling Technology). Immunoprecipitation (IP) was carried out using antibodies against Gfi1, Ikaros, PU.1, or normal rabbit IgG. The immunoprecipitated DNA was purified and quantified by qPCR as described above using primers listed in Table S1. For plasmid ChIP assay, cells were transfected with the reporter plasmids containing Hemgn fragment I (1972/+63 bp) or fragment VI (−1972/+47 bp) and treated with Dox and/or 4HT as described above. ChIP experiments were carried out as described above with qPCR performed using the primers shown in Table S1 to amplify the plasmid P1 and P2 regions (see Fig. 4C).
Mice and bone marrow cell isolation
Mice used in the experiments were bred and housed at the animal facility of The University of Toledo. All experiments involving mouse bone marrow (BM) cells were conducted in compliance with the guidelines approved by the Institutional Animal Care and Use Committee (IACUC) of The University of Toledo and were carried out according to the approved protocol. BM cells were isolated from six- to 8-week-old C57BL/6 Gfi1+/+ and Gfi1−/− mice (7) as previously described (66). Lin− cells were purified using the mouse Lineage Cell Depletion kit (Miltenyi Biotec) according to the manufacturer’s protocol and were cultured in IMDM media supplemented with 20% BIT9500 (Stemcell Technologies), 10 ng/ml IL-3, 20 ng/ml IL-6, and 25 ng/ml stem cell factor (SCF; Peprotech).
RNA interference
Ba/F3 or Ramos cells were infected with the pLKO.1 lentiviral construct containing short hairpin RNAs (shRNAs) against mouse and human Hemgn and selected in 2 μg/ml puromycin 48 h later as previously described (18, 63). The lentiviral constructs containing shRNAs targeting human Hemgn (67) (clone ID: TRCN0000430486) and mouse Hemgn (clone IDs: TRCN0000174895 and TRCN0000193835) were purchased from Sigma Aldrich. The lentiviral constructs containing shRNAs targeting mouse PU.1 (Clone IDs: TRCN0000009498 and TRCN0000009501) were obtained from Horizon Discovery.
Western blot analysis
Cells were lysed in SDS lysis buffer containing 1% SDS, 50 mM Tris–HCl (pH 8.0), and 10 mM EDTA (pH 8.0) as previously described (68). Proteins were separated by SDS-PAGE before being transferred onto polyvinylidene difluoride (PVDF) membranes. The membranes were then incubated with the appropriate antibodies, and signals were detected using enhanced chemiluminescence. The antibodies against Gfi1 (N-20), PU.1 (C-3), Hemogen (G-2), and Rabbit IgG were purchased from Santa Cruz Biotechnology. Anti-Ikaros antibody (D10E5) and monoclonal ANTI-FLAG M2 antibody were purchased from Cell Signaling Technology and Sigma-Aldrich, respectively. Anti-β-actin antibody (Cat#: 66,009–1) was obtained from Proteintech.
Apoptosis assay
Apoptosis was assessed using the Annexin V-PE apoptosis detection kit (BD Biosciences) according to previously established methods (69). In brief, 0.3 × 106 cells were collected and stained with Annexin V-PE and 7-amino-actinomycin (7-AAD). The cells were then analyzed by two-color flow cytometry using LSRFortessa (BD Biosciences) as previously described (18).
MTS assay
The experiments were conducted using the CellTiter 96 AQueous One Solution Cell Proliferation Assay kit following the manufacturer's instructions (Promega, Madison, WI), as described previously (18). Ramos cells (2 × 104) were plated in triplicate in 100 μl of RPMI 1640 medium in 96-well plates, with or without Doxorubicin (2 mg/ml) for 24 h. CellTiter 96 AQueous One Solution Reagent was added to each well, and incubated for 4 h. The absorbance was measured at a wavelength of 490 nm using a Lmax luminometer (Molecular Devices, Sunnyvale, CA).
Statistics
Statistical analysis was performed using a two-tailed student’s t test to compare the differences between any pair of data and to calculate the p-values. The data are depicted as mean ± SD in the figures. A p-value < 0.05 was considered significant and denoted by ∗, while p < 0.01 was represented by ∗∗, and p < 0.001 by ∗∗∗.
Data availability
The data generated in this study are available upon reasonable request from the corresponding authors.
Supporting information
This article contains supporting information.
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Acknowledgments
We thank Dr Richard Dahl (Department of Microbiology and Immunology, Indiana University School of Medicine, South Bend, IN) for providing the PUER cells and the PUER retroviral expression construct, Dr Andrew D. Wells (The Children's Hospital of Philadelphia, Philadelphia) for the MIGR1-FLAG-Ik1 retroviral construct, and Dr Tim C.P. Somervaille (Cancer Research UK Manchester Institute, The University of Manchester, Manchester, UK) for the pLentiGS-minCMV-TET-puromycin constructs containing Gfi1-LSD1 and Znf-LSD1 cDNA.
Author contributions
B. G., L. J. H., and F. D. investigation; B. G. and F. D. writing–original draft; B. G. and F. D. formal analysis; F. D. funding acquisition; F. D. conceptualization; B. G., F. D., and S. D. writing–review & editing; F. D. and S. D. resources.
Funding and additional information
This work was supported by grant R15 HL156109 (F. D.) from the National Institutes of Health. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Reviewed by members of the JBC Editorial Board. Edited by Brian D. Strahl
Supporting information
References
- 1.Grimes H.L., Chan T.O., Zweidler-McKay P.A., Tong B., Tsichlis P.N. The Gfi-1 proto-oncoprotein contains a novel transcriptional repressor domain, SNAG, and inhibits G1 arrest induced by interleukin-2 withdrawal. Mol. Cell Biol. 1996;16:6263–6272. doi: 10.1128/mcb.16.11.6263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Zeng H., Yücel R., Kosan C., Klein-Hitpass L., Möröy T. Transcription factor Gfi1 regulates self-renewal and engraftment of hematopoietic stem cells. EMBO J. 2004;23:4116–4125. doi: 10.1038/sj.emboj.7600419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hock H., Hamblen M.J., Rooke H.M., Schindler J.W., Saleque S., Fujiwara Y., et al. Gfi-1 restricts proliferation and preserves functional integrity of haematopoietic stem cells. Nature. 2004;431:1002–1007. doi: 10.1038/nature02994. [DOI] [PubMed] [Google Scholar]
- 4.Khandanpour C., Kosan C., Gaudreau M.C., Dührsen U., Hébert J., Zeng H., et al. Growth factor independence 1 protects hematopoietic stem cells against apoptosis but also prevents the development of a myeloproliferative-like disease. Stem Cells. 2011;29:376–385. doi: 10.1002/stem.575. [DOI] [PubMed] [Google Scholar]
- 5.Spooner C.J., Cheng J.X., Pujadas E., Laslo P., Singh H. A recurrent network involving the transcription factors PU.1 and Gfi1 orchestrates innate and adaptive immune cell fates. Immunity. 2009;31:576–586. doi: 10.1016/j.immuni.2009.07.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Möröy T., Khandanpour C. Growth factor independence 1 (Gfi1) as a regulator of lymphocyte development and activation. Semin. Immunol. 2011;23:368–378. doi: 10.1016/j.smim.2011.08.006. [DOI] [PubMed] [Google Scholar]
- 7.Hock H., Hamblen M.J., Rooke H.M., Traver D., Bronson R.T., Cameron S., et al. Intrinsic requirement for zinc finger transcription factor Gfi-1 in neutrophil differentiation. Immunity. 2003;18:109–120. doi: 10.1016/s1074-7613(02)00501-0. [DOI] [PubMed] [Google Scholar]
- 8.Zarebski A., Velu C.S., Baktula A.M., Bourdeau T., Horman S.R., Basu S., et al. Mutations in growth factor independent-1 associated with human neutropenia block murine granulopoiesis through colony stimulating factor-1. Immunity. 2008;28:370–380. doi: 10.1016/j.immuni.2007.12.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zhuang D., Qiu Y., Kogan S.C., Dong F. Increased CCAAT enhancer-binding protein epsilon (C/EBPepsilon) expression and premature apoptosis in myeloid cells expressing Gfi-1 N382S mutant associated with severe congenital neutropenia. J. Biol. Chem. 2006;281:10745–10751. doi: 10.1074/jbc.M510924200. [DOI] [PubMed] [Google Scholar]
- 10.Lin Y., Wu Y., Li J., Dong C., Ye X., Chi Y.I., et al. The SNAG domain of Snail1 functions as a molecular hook for recruiting lysine-specific demethylase 1. EMBO J. 2010;29:1803–1816. doi: 10.1038/emboj.2010.63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Möröy T., Khandanpour C. Role of GFI1 in epigenetic regulation of MDS and AML pathogenesis: mechanisms and therapeutic implications. Front. Oncol. 2019;9:824. doi: 10.3389/fonc.2019.00824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gilks C.B., Bear S.E., Grimes H.L., Tsichlis P.N. Progression of interleukin-2 (IL-2)-dependent rat T cell lymphoma lines to IL-2-independent growth following activation of a gene (Gfi-1) encoding a novel zinc finger protein. Mol. Cell Biol. 1993;13:1759–1768. doi: 10.1128/mcb.13.3.1759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Grimes H.L., Gilks C.B., Chan T.O., Porter S., Tsichlis P.N. The Gfi-1 protooncoprotein represses Bax expression and inhibits T-cell death. Proc. Natl. Acad. Sci. U. S. A. 1996;93:14569–14573. doi: 10.1073/pnas.93.25.14569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zhu J., Guo L., Min B., Watson C.J., Hu-Li J., Young H.A., et al. Growth factor independent-1 induced by IL-4 regulates Th2 cell proliferation. Immunity. 2002;16:733–744. doi: 10.1016/s1074-7613(02)00317-5. [DOI] [PubMed] [Google Scholar]
- 15.Yücel R., Karsunky H., Klein-Hitpass L., Möröy T. The transcriptional repressor Gfi1 affects development of early, uncommitted c-Kit+ T cell progenitors and CD4/CD8 lineage decision in the thymus. J. Exp. Med. 2003;197:831–844. doi: 10.1084/jem.20021417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pargmann D., Yücel R., Kosan C., Saba I., Klein-Hitpass L., Schimmer S., et al. Differential impact of the transcriptional repressor Gfi1 on mature CD4+ and CD8+ T lymphocyte function. Eur. J. Immunol. 2007;37:3551–3563. doi: 10.1002/eji.200737130. [DOI] [PubMed] [Google Scholar]
- 17.Vadnais C., Chen R., Fraszczak J., Hamard P.J., Manfredi J.J., Möröy T. A novel regulatory circuit between p53 and GFI1 controls induction of apoptosis in T cells. Sci. Rep. 2019;9:6304. doi: 10.1038/s41598-019-41684-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Du P., Tang F., Qiu Y., Dong F. GFI1 is repressed by p53 and inhibits DNA damage-induced apoptosis. PLoS One. 2013;8 doi: 10.1371/journal.pone.0073542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Petrusca D.N., Toscani D., Wang F.M., Park C., Crean C.D., Anderson J.L., et al. Growth factor independence 1 expression in myeloma cells enhances their growth, survival, and osteoclastogenesis. J. Hematol. Oncol. 2018;11:123. doi: 10.1186/s13045-018-0666-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Petrusca D.N., Mulcrone P.L., Macar D.A., Bishop R.T., Berdyshev E., Suvannasankha A., et al. GFI1-Dependent repression of SGPP1 increases multiple myeloma cell survival. Cancers (Basel) 2022;14:772. doi: 10.3390/cancers14030772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Vadnais C., Chen R., Fraszczak J., Yu Z., Boulais J., Pinder J., et al. GFI1 facilitates efficient DNA repair by regulating PRMT1 dependent methylation of MRE11 and 53BP1. Nat. Commun. 2018;9:1418. doi: 10.1038/s41467-018-03817-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yang L.V., Nicholson R.H., Kaplan J., Galy A., Li L. Hemogen is a novel nuclear factor specifically expressed in mouse hematopoietic development and its human homologue EDAG maps to chromosome 9q22, a region containing breakpoints of hematological neoplasms. Mech. Dev. 2001;104:105–111. doi: 10.1016/s0925-4773(01)00376-8. [DOI] [PubMed] [Google Scholar]
- 23.Yang L.V., Heng H.H., Wan J., Southwood C.M., Gow A., Li L. Alternative promoters and polyadenylation regulate tissue-specific expression of Hemogen isoforms during hematopoiesis and spermatogenesis. Dev. Dyn. 2003;228:606–616. doi: 10.1002/dvdy.10399. [DOI] [PubMed] [Google Scholar]
- 24.Zhao K., Liu J.F., Zhu Y.X., Dong X.M., Yin R.H., Liu X., et al. Hemgn protects hematopoietic stem and progenitor cells against transplantation stress through negatively regulating IFN-γ signaling. Adv. Sci. (Weinh) 2022;9 doi: 10.1002/advs.202103838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Li C.Y., Zhan Y.Q., Xu C.W., Xu W.X., Wang S.Y., Lv J., et al. EDAG regulates the proliferation and differentiation of hematopoietic cells and resists cell apoptosis through the activation of nuclear factor-kappa B. Cell Death Differ. 2004;11:1299–1308. doi: 10.1038/sj.cdd.4401490. [DOI] [PubMed] [Google Scholar]
- 26.Lü J., Xu W.X., Wang S.Y., Jiang Y., Li C.Y., Cai W.M., et al. [Overexpression of EDAG-1 in NIH3T3 cells leads to malignant transformation] Sheng Wu Hua Xue Yu Sheng Wu Wu Li Xue Bao (Shanghai) 2002;34:95–98. [PubMed] [Google Scholar]
- 27.Chen D.L., Hu Z.Q., Zheng X.F., Wang X.Y., Xu Y.Z., Li W.Q., et al. EDAG-1 promotes proliferation and invasion of human thyroid cancer cells by activating MAPK/Erk and AKT signal pathways. Cancer Biol. Ther. 2016;17:414–421. doi: 10.1080/15384047.2016.1156259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li Z., Huang X., Liu A., Xu J., Lai J., Guan H., et al. Circ_PSD3 promotes the progression of papillary thyroid carcinoma via the miR-637/HEMGN axis. Life Sci. 2021;264 doi: 10.1016/j.lfs.2020.118622. [DOI] [PubMed] [Google Scholar]
- 29.Zhang M.J., Ding Y.L., Xu C.W., Yang Y., Lian W.X., Zhan Y.Q., et al. Erythroid differentiation-associated gene interacts with NPM1 (nucleophosmin/B23) and increases its protein stability, resisting cell apoptosis. FEBS J. 2012;279:2848–2862. doi: 10.1111/j.1742-4658.2012.08663.x. [DOI] [PubMed] [Google Scholar]
- 30.Li C.Y., Zhan Y.Q., Li W., Xu C.W., Xu W.X., Yu D.H., et al. Overexpression of a hematopoietic transcriptional regulator EDAG induces myelopoiesis and suppresses lymphopoiesis in transgenic mice. Leukemia. 2007;21:2277–2286. doi: 10.1038/sj.leu.2404901. [DOI] [PubMed] [Google Scholar]
- 31.Yang L.V., Wan J., Ge Y., Fu Z., Kim S.Y., Fujiwara Y., et al. The GATA site-dependent hemogen promoter is transcriptionally regulated by GATA1 in hematopoietic and leukemia cells. Leukemia. 2006;20:417–425. doi: 10.1038/sj.leu.2404105. [DOI] [PubMed] [Google Scholar]
- 32.Jiang J., Yu H., Shou Y., Neale G., Zhou S., Lu T., et al. Hemgn is a direct transcriptional target of HOXB4 and induces expansion of murine myeloid progenitor cells. Blood. 2010;116:711–719. doi: 10.1182/blood-2009-07-235341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Griaud F., Williamson A.J., Taylor S., Potier D.N., Spooncer E., Pierce A., et al. BCR/ABL modulates protein phosphorylation associated with the etoposide-induced DNA damage response. J. Proteomics. 2012;77:14–26. doi: 10.1016/j.jprot.2012.06.003. [DOI] [PubMed] [Google Scholar]
- 34.Dahl R., Iyer S.R., Owens K.S., Cuylear D.D., Simon M.C. The transcriptional repressor GFI-1 antagonizes PU.1 activity through protein-protein interaction. J. Biol. Chem. 2007;282:6473–6483. doi: 10.1074/jbc.M607613200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zhang Y., Hu N., Dong F. Gfi1-Mediated repression of c-Fos, Egr-1 and Egr-2, and inhibition of ERK1/2 signaling contribute to the role of Gfi1 in granulopoiesis. Sci. Rep. 2019;9:737. doi: 10.1038/s41598-018-37402-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Karim S., Mirza Z., Chaudhary A.G., Abuzenadah A.M., Gari M., Al-Qahtani M.H. Assessment of radiation induced therapeutic effect and cytotoxicity in cancer patients based on transcriptomic profiling. Int. J. Mol. Sci. 2016;17:250. doi: 10.3390/ijms17020250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Casey M.J., Call A.M., Thorpe A.V., Jette C.A., Engel M.E., Stewart R.A. The scaffolding function of LSD1/KDM1A reinforces a negative feedback loop to repress stem cell gene expression during primitive hematopoiesis. iScience. 2023;26 doi: 10.1016/j.isci.2022.105737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Tymms M.J., Kola I. Regulation of gene expression by transcription factors Ets-1 and Ets-2. Mol. Reprod. Dev. 1994;39:208–214. doi: 10.1002/mrd.1080390214. [DOI] [PubMed] [Google Scholar]
- 39.Fraszczak J., Vadnais C., Rashkovan M., Ross J., Beauchemin H., Chen R., et al. Reduced expression but not deficiency of GFI1 causes a fatal myeloproliferative disease in mice. Leukemia. 2019;33:110–121. doi: 10.1038/s41375-018-0166-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.An L.L., Li G., Wu K.F., Ma X.T., Zheng G.G., Qiu L.G., et al. High expression of EDAG and its significance in AML. Leukemia. 2005;19:1499–1502. doi: 10.1038/sj.leu.2403808. [DOI] [PubMed] [Google Scholar]
- 41.Zhao K., Zheng W.W., Dong X.M., Yin R.H., Gao R., Li X., et al. EDAG promotes the expansion and survival of human CD34+ cells. PLoS One. 2018;13 doi: 10.1371/journal.pone.0190794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Karsunky H., Mende I., Schmidt T., Möröy T. High levels of the onco-protein Gfi-1 accelerate T-cell proliferation and inhibit activation induced T-cell death in Jurkat T-cells. Oncogene. 2002;21:1571–1579. doi: 10.1038/sj.onc.1205216. [DOI] [PubMed] [Google Scholar]
- 43.Khandanpour C., Phelan J.D., Vassen L., Schütte J., Chen R., Horman S.R., et al. Growth factor independence 1 antagonizes a p53-induced DNA damage response pathway in lymphoblastic leukemia. Cancer Cell. 2013;23:200–214. doi: 10.1016/j.ccr.2013.01.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Iwasaki H., Somoza C., Shigematsu H., Duprez E.A., Iwasaki-Arai J., Mizuno S., et al. Distinctive and indispensable roles of PU.1 in maintenance of hematopoietic stem cells and their differentiation. Blood. 2005;106:1590–1600. doi: 10.1182/blood-2005-03-0860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Mak K.S., Funnell A.P., Pearson R.C., Crossley M. PU.1 and haematopoietic cell fate: dosage matters. Int. J. Cell Biol. 2011;2011 doi: 10.1155/2011/808524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.de la Luz Sierra M., Sakakibara S., Gasperini P., Salvucci O., Jiang K., McCormick P.J., et al. The transcription factor Gfi1 regulates G-CSF signaling and neutrophil development through the Ras activator RasGRP1. Blood. 2010;115:3970–3979. doi: 10.1182/blood-2009-10-246967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Friedman A.D. Transcriptional control of granulocyte and monocyte development. Oncogene. 2007;26:6816–6828. doi: 10.1038/sj.onc.1210764. [DOI] [PubMed] [Google Scholar]
- 48.Turkistany S.A., DeKoter R.P. The transcription factor PU.1 is a critical regulator of cellular communication in the immune system. Arch. Immunol. Ther. Exp. (Warsz). 2011;59:431–440. doi: 10.1007/s00005-011-0147-9. [DOI] [PubMed] [Google Scholar]
- 49.Rosenbauer F., Wagner K., Kutok J.L., Iwasaki H., Le Beau M.M., Okuno Y., et al. Acute myeloid leukemia induced by graded reduction of a lineage-specific transcription factor, PU.1. Nat. Genet. 2004;36:624–630. doi: 10.1038/ng1361. [DOI] [PubMed] [Google Scholar]
- 50.Steidl U., Rosenbauer F., Verhaak R.G., Gu X., Ebralidze A., Otu H.H., et al. Essential role of Jun family transcription factors in PU.1 knockdown-induced leukemic stem cells. Nat. Genet. 2006;38:1269–1277. doi: 10.1038/ng1898. [DOI] [PubMed] [Google Scholar]
- 51.Yuki H., Ueno S., Tatetsu H., Niiro H., Iino T., Endo S., et al. PU.1 is a potent tumor suppressor in classical Hodgkin lymphoma cells. Blood. 2013;121:962–970. doi: 10.1182/blood-2012-05-431429. [DOI] [PubMed] [Google Scholar]
- 52.Ueno N., Nishimura N., Ueno S., Endo S., Tatetsu H., Hirata S., et al. PU.1 acts as tumor suppressor for myeloma cells through direct transcriptional repression of IRF4. Oncogene. 2017;36:4481–4497. doi: 10.1038/onc.2017.79. [DOI] [PubMed] [Google Scholar]
- 53.Oikawa T., Yamada T., Kihara-Negishi F., Yamamoto H., Kondoh N., Hitomi Y., et al. The role of Ets family transcription factor PU.1 in hematopoietic cell differentiation, proliferation and apoptosis. Cell Death Differ. 1999;6:599–608. doi: 10.1038/sj.cdd.4400534. [DOI] [PubMed] [Google Scholar]
- 54.Sun W., Guo J., McClellan D., Poeschla A., Bareyan D., Casey M.J., et al. GFI1 cooperates with IKZF1/IKAROS to activate gene expression in T-cell acute lymphoblastic leukemia. Mol. Cancer Res. 2022;20:501–514. doi: 10.1158/1541-7786.MCR-21-0352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Davis K.L. Ikaros: master of hematopoiesis, agent of leukemia. Ther. Adv. Hematol. 2011;2:359–368. doi: 10.1177/2040620711412419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Heizmann B., Kastner P., Chan S. The Ikaros family in lymphocyte development. Curr. Opin. Immunol. 2018;51:14–23. doi: 10.1016/j.coi.2017.11.005. [DOI] [PubMed] [Google Scholar]
- 57.Affar M., Bottardi S., Quansah N., Lemarié M., Ramón A.C., Affar E.B., et al. IKAROS: from chromatin organization to transcriptional elongation control. Cell Death Differ. 2023 doi: 10.1038/s41418-023-01212-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Payne M.A. Zinc finger structure-function in Ikaros marvin A payne. World J. Biol. Chem. 2011;2:161–166. doi: 10.4331/wjbc.v2.i6.161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Cippitelli M., Stabile H., Kosta A., Petillo S., Gismondi A., Santoni A., et al. Role of aiolos and Ikaros in the antitumor and immunomodulatory activity of IMiDs in multiple myeloma: better to lose than to find them. Int. J. Mol. Sci. 2021;22:1103. doi: 10.3390/ijms22031103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zhang Y., Dong F. Gfi1 upregulates c-Myc expression and promotes c-Myc-driven cell proliferation. Sci. Rep. 2020;10 doi: 10.1038/s41598-020-74278-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Kathrein K.L., Lorenz R., Innes A.M., Griffiths E., Winandy S. Ikaros induces quiescence and T-cell differentiation in a leukemia cell line. Mol. Cell Biol. 2005;25:1645–1654. doi: 10.1128/MCB.25.5.1645-1654.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Walsh J.C., DeKoter R.P., Lee H.J., Smith E.D., Lancki D.W., Gurish M.F., et al. Cooperative and antagonistic interplay between PU.1 and GATA-2 in the specification of myeloid cell fates. Immunity. 2002;17:665–676. doi: 10.1016/s1074-7613(02)00452-1. [DOI] [PubMed] [Google Scholar]
- 63.Liu Q., Basu S., Qiu Y., Tang F., Dong F. A role of Miz-1 in Gfi-1-mediated transcriptional repression of CDKN1A. Oncogene. 2010;29:2843–2852. doi: 10.1038/onc.2010.48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Maiques-Diaz A., Spencer G.J., Lynch J.T., Ciceri F., Williams E.L., Amaral F.M.R., et al. Enhancer activation by pharmacologic displacement of LSD1 from GFI1 induces differentiation in acute myeloid leukemia. Cell Rep. 2018;22:3641–3659. doi: 10.1016/j.celrep.2018.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Song C., Ge Z., Ding Y., Tan B.H., Desai D., Gowda K., et al. IKAROS and CK2 regulate expression of BCL-XL and chemosensitivity in high-risk B-cell acute lymphoblastic leukemia. Blood. 2020;136:1520–1534. doi: 10.1182/blood.2019002655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Hu N., Qiu Y., Dong F. Role of Erk1/2 signaling in the regulation of neutrophil versus monocyte development in response to G-CSF and M-CSF. J. Biol. Chem. 2015;290:24561–24573. doi: 10.1074/jbc.M115.668871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Guo X., Zhao Y., Kim J., Dean A. Hemogen/BRG1 cooperativity modulates promoter and enhancer activation during erythropoiesis. Blood. 2022;139:3532–3545. doi: 10.1182/blood.2021014308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Basu S., Liu Q., Qiu Y., Dong F. Gfi-1 represses CDKN2B encoding p15INK4B through interaction with Miz-1. Proc. Natl. Acad. Sci. U. S. A. 2009;106:1433–1438. doi: 10.1073/pnas.0804863106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Qiu Y., Zhang Y., Hu N., Dong F. A truncated granulocyte colony-stimulating factor receptor (G-CSFR) inhibits apoptosis induced by neutrophil elastase G185R mutant: implication for understanding CSF3R gene mutations in severe congenital neutropenia. J. Biol. Chem. 2017;292:3496–3505. doi: 10.1074/jbc.M116.755157. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data generated in this study are available upon reasonable request from the corresponding authors.