Abstract
Receptor tyrosine kinases (RTKs) regulate many cellular functions and are important targets in pharmaceutical development, particularly in cancer treatment. EGFR and EphA2 are two key RTKs that are associated with oncogenic phenotypes. Several studies have reported functional interplay between these receptors, but the mechanism of interaction is still unresolved. Here, we use a time-resolved fluorescence spectroscopy called PIE-FCCS to resolve EGFR and EphA2 interactions in live cells. We tested the role of ligands and found that EGF, but not ephrin A1 (EA1), stimulated heteromultimerization between the receptors. To determine the effect of anionic lipids, we targeted phospholipase C (PLC) activity to alter the abundance of phosphatidylinositol 4,5-bisphosphate (PIP2). We found that higher PIP2 levels increased homomultimerization of both EGFR and EphA2, as well as heteromultimerization. This study provides a direct characterization of EGFR and EphA2 interactions in live cells and shows that PIP2 can have a substantial effect on the spatial organization of RTKs.
EGFR and EphA2 interactions are affected by cellular PIP2 levels.
INTRODUCTION
Cell surface proteins are crucial for responding to stimuli and communicating with surrounding cells. Understanding the molecular level interactions of proteins in the plasma membrane is vital to targeting them in drug development (1, 2). The human genome contains 58 receptor tyrosine kinases (RTKs) that share similar downstream pathways (3, 4). RTKs play crucial roles in cell growth, migration, and differentiation and are common targets of cancer therapies (5, 6). RTKs also share common structural features, including a single transmembrane (TM) helix that connects an extracellular domain to an intracellular kinase domain. In the canonical model for RTK activation, kinase function is initiated by ligand-induced dimerization (4). However, many RTKs display alternate activation mechanisms including ligand-induced conformational changes to pre-existing dimers and the formation of higher-order oligomers (4, 7, 8). Increasingly, cross-interactions between RTKs are seen as an important aspect of their function (5). Many early efforts were made to characterize heterodimers within RTK subfamilies, and there has recently been additional focus on heterodimerization across subfamilies (5, 9).
The most studied cross-family interaction is between epidermal growth factor receptor (EGFR) and EphA2 (5). Individually, EGFR and EphA2 each have important cellular functions. EGFR is a member of the HER (or ErbB) family of RTKs and plays a role in cell proliferation, motility, growth, and differentiation (4, 7, 10). EphA2 is a member of the largest subfamily of RTKs, the Eph receptors, and is essential to development (11–13). EphA2 is implicated in many processes that affect cellular behaviors between cells, including adhesion and repulsion (11, 13, 14). In addition, both receptors are implicated in cancer. Although EphA2 expression is usually low in normal adult tissue, EphA2 overexpression in certain tumors is often correlated with poor prognosis (15–17). The Smith laboratory and collaborators recently showed how different multimerization interfaces contributed to distinct downstream signaling and oncogenic functions (18). EGFR contributes to the development and progression of cancer and is a validated target for therapeutic intervention in various solid tumors (10, 19–23). EphA2 has been implicated in anti-EGFR cancer therapy, where its role in drug resistance has been ascribed to direct interactions with EGFR and ErbB2 (24, 25).
Because of EGFR and EphA2’s relationship in cancer, recent efforts have focused on the characterization of their functional interaction (26–29). Binding between EphA2 and EGFR is proposed to activate Ras/mitogen-activated protein kinase and RhoA signaling pathways (27, 28). However, several studies have proposed that EGFR/EphA2 heterodimers can lead to the recruitment of different adaptor proteins and thus different biological effects (30, 31). For example, it was reported that Ephexin A1 bridges the interaction between EphA2 and EGFR (31), inducing oncogenic, noncanonical signaling through AKT pathways in the presence of EGF (27). In cases where EphA2 is overexpressed or phosphorylated at S897, there is an increased likelihood of interacting with EGFR (25, 26, 31). Despite these previous reports of EGFR-EphA2 cross-talk, there is insufficient experimental evidence to characterize the interactions and determine the factors that affect the interactions.
One factor that we expect to affect EGFR-EphA2 interactions is the membrane lipid environment, especially anionic lipids, which play an important role in RTK structure and function (32, 33). The juxtamembrane (JM) domain of all human RTKs contains several positively charged amino acid residues that interact with negatively charged lipids (34). Molecular dynamic simulations suggest that in the absence of ligands, negatively charged membrane lipids interact with and stabilize the inactive state of EGFR (35). Live-cell imaging of EGFR using dSTORM, showed that the lipid phosphatidylinositol 4,5-bisphosphate (PIP2) induces the formation of nanoclusters (36). Simulation studies suggested that EphA2 homomultimerization is affected by negatively charged membrane lipids, similar to the effect seen in EGFR (37, 38). Previous work from the Barrera laboratory revealed that dimerization of the isolated TM domain of EphA2 can be promoted by PIP2 (39). However, the effect of PIP2 on dimerization of full-length EphA2 has not yet been investigated. In addition, it is not known if lipids regulate the interaction between EphA2 and EGFR. In this study, we investigate the hypothesis that PIP2 contributes to the multimerization state of EphA2 and EGFR receptors.
Many biophysical techniques have been developed to characterize RTK multimers and their impact on cell signaling (5, 40, 41). Single-molecule (SM) fluorescence approaches are an important class of methods that have been developed to resolve membrane protein interactions in cells (41, 42). However, these SM methods are typically limited to very low expression levels (<1 molecule/μm2), far below the physiological expression level of many RTKs (42). Fluorescence fluctuation spectroscopy (FFS) has emerged as a valuable method for resolving membrane protein multimerization (43–46). FFS methods use time-dependent fluctuations to circumvent the optical diffraction limit. In this study, we use a two-color FFS technique called pulsed-interleaved excitation fluorescence cross-correlation spectroscopy (PIE-FCCS) (47) to examine the interaction between EGFR and EphA2 in cell membranes. PIE-FCCS can resolve membrane protein expression levels, diffusion coefficients, and the degree of cross-correlation (abbreviated as fc), which is a direct measure of how strongly the proteins diffuse together as dimers, trimers, and small oligomers (44). It is primarily sensitive to diffusing proteins and so does not detect immobile aggregates nor internalized proteins. As the degree of cross-correlation depends on correlated diffusion, we can interpret the codiffusing species as stable over the timescale of the transit time (~10−1 s), although it cannot resolve association lifetimes directly. The measurements described below were taken in live cells at physiological expression levels and thus provide direct evidence for membrane protein interactions in situ.
Our initial PIE-FCCS measurements did not detect interactions between EphA2 and EGFR in resting cells. This unexpected result was then investigated using various ligand-binding conditions and PIP2 modulation studies to determine the factors that affect EGFR-EphA2 interactions. By testing the effects of ligands and PIP2-modifying compounds, we found that both EGF and increased PIP2 levels led to significant EGFR-EphA2 heteromultimerization. We then tested the phosphorylation levels under these conditions and found that the PIP2-induced multimers were less active than ligand-induced multimers, suggesting that PIP2 was not stabilizing an active conformation of the receptors. Overall, our work demonstrates the important role of PIP2 in modulating the degree of interaction between EGFR and EphA2. The results suggest that other putative RTK interactions could also be affected by PIP2 levels in the plasma membrane.
RESULTS
EGFR does not associate with EphA2 in resting cells
To investigate the interaction between EGFR and EphA2, we performed PIE-FCCS measurements using two pulsed lasers and time-correlated single-photon counting (TCSPC) (Fig. 1A). We transfected cells with expression vectors encoding C-terminally tagged fluorescent proteins [eGFP and mCherry (mCH)], beginning with SRC and GCN4 as controls for monomer and dimer states, respectively (48). These control constructs have a short, lipidated peptide sequence for membrane anchoring and, in the case of GCN4, an α-helical leucine zipper motif for dimerization. We then collected data from individual COS-7 cells expressing the exogenous proteins in a physiological range of 100 to 1500 molecules/μm2 in the plasma membrane. From the correlation functions (Fig. 1, D to F), we obtained the fraction of cross-correlation (fc) and the effective diffusion coefficients of the eGFP and mCH-labeled proteins as described previously (44, 49, 50). The fc values are indicative of codiffusion of the two receptors. The median fc value of 0.01 for SRC (Fig. 1C, light gray) and 0.15 for GCN4 (Fig. 1C, medium gray) are indicative of their monomeric and dimeric state on the plasma membrane. The dimer fc value of 0.15 is smaller than that observed for a covalent dimer (e.g., duplex DNA; figs. S1 and S2). The fc values from live-cell measurements such as those used here have been explained previously using a stochastic model (44, 48). Briefly, the cross-correlation is affected by volume mismatch in the detection system, fluorescent protein dark states, and stochastic association of red and green fluorescent labels. We next coexpressed EGFR-mCH (Fig. 1B, top) and EphA2-GFP (Fig. 1B, bottom) under ligand-free conditions. The median fc value for EGFR and EphA2 was 0.04 (Fig. 1C, dark gray), indicating that they do not heteromultimerize in the absence of ligand. These unexpected results seem to contradict earlier reports of direct interactions between EGFR and EphA2 (26). In the next section, we investigate the hypothesis that PIP2 levels in the plasma membrane could affect the degree of interaction between these two RTKs.
Fig. 1. Sketch layout of PIE-FCCS and its role in membrane protien interaction.
(A) Diagram illustrates the optical path of dual-color PIE-FCCS. A super-continuum, pulsed laser-generated excitation beams (488 and 561 nm) that were directed into the sample by a dichroic mirror and microscope objective. The lasers were focused to a diameter of about 250 nm on the plasma membrane of live cells. Fluorescence emission was collected by the objective, filtered, and directed to single-photon counting modules connected to a TCSPC module for data recording. (B) Epifluorescence images are shown for COS-7 cells expressing EGFR-mCH (top) and EphA2-eGFP (bottom). (C) Summary data from independent experiments collected on five different days of single-cell measurements are shown for the controls and EGFR with EphA2 (***P < 0.0004). (D to F) Representative single-cell PIE-FCCS data are shown for a monomer control (SRC), dimer control (GCN4), and EGFR with EphA2 in COS-7 cells, respectively. In each graph, the green line is the autocorrelation function (ACF) for eGFP, the red line is the ACF for mCH, and the blue line is the cross-correlation function (CCF) between both protein constructs.
A PLC inhibitor increases EGFR homodimerization and EphA2 homomultimerization
PIP2 lipids play an important role in RTK signaling despite constituting a minor portion of the total phospholipid composition of cell membranes (51, 52). In cell signaling, PIP2 is broken down by phospholipase C (PLC) into the second messengers: Diacylglycerol (DAG) and inositol 1,4,5-triphosphate (IP3) (53). PLC-ƴ1 is expressed in most cell types and is activated downstream of RTKs (53). Several activators and inhibitors of PLC have been used to understand its role in cell signaling and development. U73122 is an inhibitor of PLC activity (54, 55). m-3M3FBS is a PLC activator (56, 57). In our current work, we use these two drug molecules to regulate PLC activity and thus increase (with U73122) or decrease (with 3M3FBS) PIP2 lipid density in the plasma membrane. As a control, we performed live-cell imaging experiments in an assay adapted from Balla et al. (58), which confirmed that U73122 increased while 3M3FBS decreased plasma membrane levels of PIP2 (fig. S3).
We began by investigating how EGFR and EphA2 ligands and PIP2-modifying drugs affect the homomultimerization of each receptor. For EGFR, we coexpressed EGFR-mCH and EGFR-eGFP in COS-7 cells. The PIE-FCCS measurements were recorded at expression densities between 100 and 1400 molecules/μm2, before and after treatment. In untreated cells, EGFR had a median fc value of 0.02 (Fig. 2A, white), consistent with it being primarily monomeric as reported by our laboratory previously (59, 60). We added an EGFR ligand, EGF, and observed a median fc value of 0.24 (Fig. 2A, gray), consistent with ligand-induced multimerization (60). The addition of EGF ligand to EGFR also led to receptor internalization, which can be seen in the epifluorescence images (Fig. 2C). PIE-FCCS data were then collected after increasing the level of PIP2 by treating the cells with a PLC inhibitor (U73122, 5 μM) for at least 15 min. The median fc value with the inhibitor was 0.10, indicating PIP2 induced dimerization of EGFR (Fig. 2A, blue). In contrast, decreasing PIP2 levels with a PLC activator (3M3FBS, 25 μM) resulted in a median fc value of 0.03 (Fig. 2A, green). Therefore, reduction of PIP2 did not significantly affect the multimerization nor diffusion of EGFR. The interpretation of the fc values is further supported by the diffusion coefficients of the receptors. The diffusion coefficients of EGFR decreased from 0.42 to 0.15 μm2/s with EGF treatment (Fig. 2B, gray) and to 0.36 μm2/s after U73122 treatment (Fig. 2B, blue). There was no statistically significant change in the diffusion coefficient with the PLC activator (0.40 μm2/s) compared to EGFR in the resting cells (0.42 μm2/s). Single-cell PIE-FCCS data (fig. S4) and fit parameters (table S1) can be found in the Supplementary Materials.
Fig. 2. PIE-FCCS investigation of the homomeric interactions of EGFR and EphA2.
(A) Self-assembly of EGFR was studied under various conditions. PIE-FCCS data were recorded, and the resulting fc values are reported before treatment (white), after the addition of EGF (gray), after treatment with the PLC inhibitor U73122 (blue), and after treatment with the PLC activator 3M3FBS (green). (B) Diffusion coefficients for EGFR are shown for each treatment condition. (C) Epifluorescence images of EGFR expressing COS-7 cells without (top) and with (bottom) EGF ligand treatment. (D) Homomultimerization state of EphA2 was recorded under the same treatments except the ligand treatment was with EA1. (E) Diffusion coefficients are shown for EphA2 under each treatment condition. (F) Epifluorescence images of EphA2 expressing COS-7 cells are shown without (top) and with (bottom) EA1 ligand treatment. In the box-and-whiskers plots, the whiskers indicate the maximum and minimum values; the box indicates the 25th to 75th percentile, and the line is the median value (median value shown as text). We performed one-way analysis of variance (ANOVA) tests with uncorrected Fisher’s least significant difference (LSD) post hoc tests to obtain adjusted and individual P values (*P < 0.0367, **P < 0.0034, and ****P < 0.0001). Diffusion coefficient data are represented as mean values ± SEM. NS, not significant.
To determine the effect of PIP2 levels on EphA2 homomultimerization, we expressed EphA2-GFP and EphA2-mCH in COS-7 cells. PIE-FCCS data were collected with and without ephrin A1 (EA1) or PLC regulators as described above. Before the ligand or drug treatment, the median fc value was 0.20, indicating substantial homomultimerization (Fig. 2D, white) as we have reported recently (61). The fc values for EphA2 receptors without ligand were larger than those for dimer controls (Fig. 1C) and more comparable to those for multimer controls (48). We next added EA1, an EphA2-specific ligand, and observed a median fc value of 0.37 (Fig. 2D, gray), indicative of ligand-induced multimerization. The EphA2 diffusion coefficient decreased from 0.29 (Fig. 2E, white) to 0.16 μm2/s (Fig. 2E, gray) with EA1 ligand treatment, which supports our median fc value interpretations. The addition of the PLC inhibitor (U73122, 5 μM) resulted in a larger median fc = 0.27 (Fig. 2D, blue) compared to resting cells. Similar to the EGFR results above, we interpret this as PIP2-induced multimerization of EphA2. We next treated cells with a PLC activator (3M3FBS, 25 μM), which resulted in a reduced median fc = 0.07. This result indicates that lower PIP2 reduced the ligand-independent multimerization of EphA2. The diffusion coefficients support these interpretations. The PLC inhibitor reduces protein mobility from 0.29 (Fig. 2E, white) to 0.24 μm2/s (Fig. 2E, blue), and the PLC activator enhanced it to 0.33 μm2/s (Fig. 2E, green). Together, these results demonstrate that PIP2 induces EGFR homodimerization and EphA2 homomultimerization. Single-cell PIE-FCCS data (fig. S5), and fit parameters (table S2) can be found in the Supplementary Materials.
PIP2 and EGF positively regulate EGFR-EPHA2 heteromultimerization
We next tested the effect of PIP2 on the interaction between the EGFR and EphA2 receptors. COS-7 cells were transfected with both receptors (EGFR-mCH and EphA2-eGFP), and PIE-FCCS measurements were performed as described above. During data collection, cells were selected with a nearly equal expression of both proteins. Under resting cell conditions, no statistically significant cross-correlation was observed between EGFR and EphA2, as indicated by the median fc value of 0.04 (Fig. 3A, white). We next incubated cells with epidermal growth factor (EGF) (500 ng/ml) or EA1 (500 ng/ml). PIE-FCCS data were collected between 15 and 90 min after ligand stimulation. After EGF treatment, the median fc value increased from 0.04 to 0.10, indicating the increased formation of heteromultimers. This increase in cross-correlation is consistent with previous reports that EGF can enhance the association of EGFR and EphA2 (31). Combined with our observations in the previous section, we conclude that the heteromultimers consisted of EGFR dimers and EphA2 multimers, which could have a size range of four to eight proteins. This interpretation is supported by the diffusion coefficient data in which we observe that the mobility of EGFR decreases significantly from 0.37 to 0.15 μm2/s, while the diffusion coefficient of EphA2 reduced from 0.25 to 0.20 μm2/s (Fig. 3, B and C). We next added the EphA2-specific ligand, EA1. In the presence of EA1, the median fc value was 0.02 (Fig. 3A, orange), statistically indistinguishable from the untreated cells, supporting the notion that EA1 does not lead to heteromultimerization between EGFR and EphA2. The reduction in the diffusion coefficient of the EphA2 receptor from 0.25 to 0.16 μm2/s is explained by EA1-induced homomultimerization of EphA2 (Fig. 3C).
Fig. 3. Heteromultimerization of EGFR and EphA2 in the presence of ligands and PLC drugs.
(A) PIE-FCCS data for EGFR-mCH and EphA2-eGFP coexpressed in COS-7 cells. Data were recorded before treatment (white), after the addition of EGF (gray) or EA1 (orange), and after treatment with the PLC inhibitor U73122 (blue) or with the PLC activator 3M3FBS (green). (B) Diffusion coefficients of EGFR-mCH after treatment with ligands and PLC-regulating drugs. (C) Diffusion coefficients of EphA2-eGFP after treatment with ligands and PLC-regulating drugs. (D) Representative epifluorescence images from each set of experiments. One-way ANOVA tests with uncorrected Fisher’s LSD post hoc tests show the individual P values (*P < 0.0259, **P < 0.003, and ****P < 0.0001). Diffusion coefficient data are represented as mean values ± SEM.
We next tested the effect of the PLC inhibitor and observed a significant change in the median fc value, from 0.04 to 0.15, indicative of heteromultimerization (Fig. 3A, green). The average diffusion coefficient of EGFR decreased from 0.37 to 0.28 μm2/s. The mobility of EphA2 receptors also decreased from 0.25 to 0.14 μm2/s. The median fc value remained unchanged after adding the PLC activator compared to resting cells, as shown in Fig. 3A (blue). The molecular diffusion of EGFR indicated no change in protein mobility (Fig. 3B, blue), consistent with the fc values. The mobility of the EphA2 receptor increased after treatment with the PLC activator, consistent with the disruption of homomultimerization explained in the previous section. Single-cell PIE-FCCS data (fig. S6) and fit parameters (table S3) can be found in the Supplementary Materials. Lifetime analysis of eGFP shows a slight reduction in lifetime with ligand (EGF) and PLC inhibitor, while mCH slightly increases. No statistically significant changes were observed with the EA1 ligand and PLC activator (fig. S8). A combined treatment of both ligand and PLC inhibitor (U73122) shows no signifcant increase in the median fc value of EGFR-EphA2 heteromultimer (fig. S9). To analyze the effect of the PLC protein drug effect on the JM and kinase domains of both proteins, we also performed a similar experiment with truncated constructs of EGFR and EphA2 with only the intracellular domains (ICDs). The ICD constructs showed no significant cross-correlation; however, the addition of PLC inhibitor (U73122) leads to a significant increase in cross-correlation similar to the full-length proteins (fig. S10F). These ICD data support the interpretation that PIP2 levels in the inner leaflet of the plasma membrane affect the conformation and dimerization propensity of EGFR and EphA2 through direct contacts with the ICD region of the proteins.
PIP2 -induced EGFR and EphA2 heteromultimers are inactive
Once we determined how PLC drugs affected EGFR and EphA2 homo- and heteromultimerization, we investigated the activity of the multimers. Phosphorylation of key tyrosine residues is commonly used as an indicator that RTKs are in their active conformation. In addition, EphA2 S897 has been shown to be important for heterodimer formation and activity (31). We therefore used Western blotting to determine whether these phosphorylation sites were affected by the PLC drugs (Fig. 4). We began by treating A375 cells with EA1 and EGF to confirm the normal activation of endogenous proteins in our system. EGF caused a marked increase in EGFR pY1068 (fig. S10A), showing that EGFR was successfully activated. EGF treatment also caused an increase in EphA2 pS897 (Fig. 4C) correlating with EGFR-EphA2 heterodimerization. As expected, EA1 induced a strong increase in EphA2 pY588 (fig. S10B). We chose a 15-min treatment time to minimize the amount of EphA2 endocytosis and lysosomal degradation resultant from activation. However, there was a small reduction in total EphA2 levels (Fig. 4D) due to degradation (62). Together, these results demonstrate the successful activation of EGFR and EphA2 homo- and heteromultimers by their respective ligands.
Fig. 4. EGFR and EphA2 phosphorylation changes after treatment with ligand and PLC drugs.
(A) Representative Western blots after treatment with EA1 (15 min, 500 ng/ml), 3M3FBS (45 min, 25 μM), or 3M3FBS followed by EA1. (B) Representative Western blots after treatment with U73122 (15 min, 5 μM), EA1 (as above), EGF (15 min, 100 ng/ml), or U73122 followed by each ligand. (C) Quantification of EphA2 pS897 levels in all conditions normalized to the corresponding total EphA2 bands. N = 6 to 12. Statistical analysis was performed using a Kruskal-Wallis test [H(7) = 19.45, P = 0.007] with a Mann-Whitney U test for comparisons between groups. Significance values were adjusted by the Bonferroni correction for multiple tests. (D) Quantification of total EphA2 levels in all conditions normalized to actin. N = 6 to 12. Statistical analysis was performed using a Kruskal-Wallis test [H(7) = 38.44, P = 3.0 × 10−6] with a Mann-Whitney U test for comparisons between groups. Significance values were adjusted by the Bonferroni correction for multiple tests. *P < 0.05, **P < 0.01, and ***P < 0.001.
To determine the effects of PIP2 on EGFR and EphA2 phosphorylation, we treated A375 cells with 5 μM U73122 for 15 min or 25 μM 3M3FBS for 45 min. Neither up- nor down-regulation of PIP2 caused phosphorylation changes for EGFR pY1068 or EphA2 pY588, indicating that PIP2 changes do not trigger signaling associated to ligand-dependent activation (fig. S10). The PIE-FCCS data above show that PIP2 promotes self-assembly of each receptor (Fig. 2, A and D); however, the phosphorylation data indicate that these are inactive multimers. We were intrigued to find that a similar phenomenon occurred for EphA2 pS897: Regulation of PIP2 via PLC drugs had no effect on the level of EphA2 pS897 (Fig. 4C). This demonstrates that the PIP2-induced heteromultimers do not affect the availability of EphA2 as an Akt substrate (63). Together, the functional data support the conclusion that PIP2-induced EGFR-EphA2 heteromultimers trap the proteins in an inactive state.
We also investigated if ligand activation of EGFR and EphA2 was affected by PIP2 levels. We treated A375 cells first with the PLC modifiers, followed with EGF or EA1 treatment. Treatment with EA1 after down-regulation of PIP2 had no effect on the levels of EphA2 pY588 (fig. S10B). This suggests that EA1 can induce activation of EphA2 regardless of the initial oligomer size. Intriguingly, EA1 did not cause a decrease in total EphA2 in cells with decreased PIP2 (Fig. 4D). Although activation occurs normally, this suggests that there may be a difference in the ability of activated EphA2 to be degraded after ligand treatment. EGF treatment after down-regulation of PIP2 was not tested, as 3M3FBS had no effect on EGFR homodimerization nor EGFR-EphA2 heterodimerization (Figs. 2 and 3).
EA1 treatment of cells containing higher PIP2 caused EphA2 pY588 to increase to the same level as cells with basal amounts of PIP2 (fig. S10B). This result suggests that PIP2-mediated EphA2 homomultimers can still convert to active multimers successfully. Up-regulation of PIP2 before EA1 treatment decreased pS897 levels (Fig. 4C) similar to EGF treatment of cells containing up-regulated PIP2. There was no PIP2-induced change in EGFR pY1068 (fig. S10A), suggesting successful activation of EGFR homomultimers, similar to that of EphA2 activation. However, when PIP2 levels were increased before EGF treatment, the increase in EphA2 pS897 characteristic of EGF treatment alone did not occur (Fig. 4C). Because EphA2 pS897 was decreased by both EA1 and EGF in high PIP2 cells, we hypothesize that this change is due to conformational differences resulting from heterodimerization.
Computational predictions of the dimerization conformation of the EGFR and EphA2 intracellular domain dimer
On the basis of our data, we conclude that EGFR-EphA2 heteromultimers have two conformations: an inactive conformation that is not EphA2 S897 phosphorylated and an active conformation where S897 is phosphorylated. Our results additionally indicate that PIP2 stabilizes the inactive complex. There are no structures for EGFR-EphA2 heterodimers. However, recent advances in artificial intelligence allow us to obtain robust predictions of protein complexes. AlphaFold Multimer (AlFoM) is an extension of AlphaFold2 developed to predict protein-protein complexes (64). We applied AlFoM to predict the structure of the inactive and active EGFR-EphA2 heterodimer. We performed AlFoM simulations of the EGFR-EphA2 ICDs, which have been proposed to interact (65). The results show asymmetric kinase dimerization (Fig. 5A) similar to that observed for the active state of EGFR homodimers (66). To benchmark the predicted structures, they were overlayed with crystal structures for EGFR [Protein Data Bank (PDB): 2GS6] or EphA2 (PBD: 7KJA) (65, 67), and the root mean square deviation (RMSD) was 2.9 and 8.7 Å, respectively (fig. S11). The larger RMSD for EphA2 can be attributed to the change in the conformation of the sterile alpha motif (SAM) domain, but SAM domain movement is both expected and necessary for EphA2 activation. Therefore, to obtain a more accurate RMSD that reflects the structural similarities between the predicted and experimental structures while removing the confounding factor of the SAM domain orientational change, we split the EphA2 ICD into two pieces and repeated the RMSD calculation. EphA2 residues 559 to 875—which cover the kinase domain—overlay with our predicted structure with an RMSD of 1.8 Å, while SAM domain residues 904 to 976 overlay with an RMSD of only 0.8 Å (fig. S11), suggesting that the predicted structures are supported by experimentally determined structures. We additionally plotted the local distance difference test (lDDT) values per residue (fig. S12). The lDDT is a superposition-free score that evaluates local distance differences of all atoms in a model, where scores above 70 are generally considered to be modeled well (68). Our modeling scores show a relatively high level of confidence in these predictions over the structured portions of the proteins.
Fig. 5. AlFoM model of the heterodimer of EGFR residues 669 to 1210 (cyan) and EphA2 residues 559 to 976 (purple).
Predictions are shown for EphA2 WT (A) or for the S897E/S901E mutant (B) representative of phosphorylation at serine residues (red bubbles). Yellow residues indicate the active site of each kinase region. The purple halo encompasses the EphA2 SAM domain, while the cyan halo shows the EGFR JM domain. The red arrow indicates the EphA2 kinase active site, which is sterically blocked by the EGFR kinase domain in (A), but not in (B).
It was previously reported that EphA2 phosphorylation at S897 and S901 causes a configurational change of the SAM domain that allows a change in the function of the EphA2 homodimer (65). In addition, our Western blot data show that S897 is also an essential factor in signaling by the EGFR-EphA2 heterodimer. We therefore repeated the AlFoM prediction with a previously validated phosphomimetic mutant of EphA2 in which S897 and S901 have been mutated to glutamic acid (EphA2S897E/S901E; Fig. 5B) (65). In this prediction, the RMSD of EGFR with its crystal structure provided an RMSD of 2.9 Å. The two-part EphA2 comparison was repeated, and the kinase domain had an RMSD of 1.8 Å, while for the SAM domain was only 0.7 Å, showing excellent agreement between the crystal structure and the AlFoM model.
We were intrigued to find that pS897-mimetic mutations caused a large difference in the orientation of EGFR with respect to wild-type EphA2. Specifically, when EGFR interacts with EphA2WT, EGFR is positioned such that the active site of the EphA2 kinase domain (Fig. 5, yellow) is partially obstructed, while EGFR interacting with EphA2S897E/S901E results in a more open kinase active site. We further noticed that the EGFR JM helix is positioned differently in each structure. The positively charged residues in the EGFR JM strongly interact with negatively charged PIP2, and this interaction regulates EGFR homodimer stability and activity (69–72). We therefore hypothesized that this interaction might regulate the conformation of the EGFR-EphA2 heterodimer as well. To test this hypothesis, we performed additional PIE-FCCS measurements with EGFR and the EphA2S897E mutant that showed a higher median fc (0.16) value compared to EphA2WT (fig. S14A). On the basis of the AlFoM model results, we hypothesize that when PIP2 levels are increased, the JM is sequestered at the membrane, disallowing the flexibility required for EGFR to shift into its more open and active conformation.
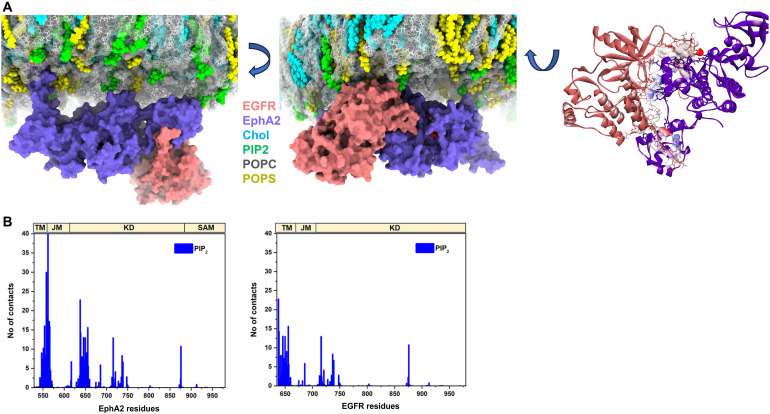
We next performed all-atom molecular simulations to analyze the behavior of the EGFR-EphA2 dimer in an explicit lipid bilayer environment. Over time, the heterodimer of EphA2 and EGFR underwent slow conformational transitions within the lipid bilayer, which appear reasonably converged by RMSD from the starting structures. The predominant conformation exhibited stable interactions between the ICDs of the two receptors. As depicted in Fig. 6, both EphA2 and EGFR engage with membrane lipids, specifically PIP2 and phosphatidylserine (POPS), which help stabilize the heterodimeric complex. A comparative analysis of lipid interactions revealed that EphA2 demonstrated a stronger preference for PIP2 binding than EGFR (10 residues for EGFR versus 14 of 15 residues for EphA2). Notably, the cationic residues within the JM region of EphA2 exhibited significantly higher interactions with PIP2 compared to the JM region of EGFR (fig. S15). This disparity can likely be attributed to the involvement of the EGFR JM region in the heterodimeric interface, forming a short helix and making it less available for interaction with the membrane. Conversely, the JM region of EphA2 remains more exposed, converting to an extended structure and allowing for stronger interactions with the lipid bilayer. The superimposition of the initial and final structures for both EphA2 and EGFR (fig. S16) indicates a more pronounced movement in the JM and SAM domains of EphA2 compared to EGFR. This suggests greater flexibility and conformational changes in these regions for EphA2 during the simulation. Moreover, this heterodimeric interface between EphA2 and EGFR is stabilized by several noncovalent interactions such as H-bonds, salt bridges, and hydrophobic interactions (table S4). Further experimental investigation into this model is required, but these structural predictions offer interesting possibilities for the effects of PIP2 on EGFR-EphA2 conformation and dimerization.
Fig. 6. Comparison of EGFR and EphA2 interactions with the lipid bilayer.
(A) Snapshots illustrating the predominant conformation of the EGFR-EphA2 heterodimer after molecular dynamics simulations. The intracellular view highlights the EGFR-EphA2 heterodimer’s association with the lipid bilayer, where EGFR is shown in salmon and EphA2 in purple-blue surfaces. Cholesterol, PIP2, and POPS are depicted as cyan, green, and yellow spheres, respectively, while POPC is displayed as gray lines for clarity. The image on the right shows the interaction interface surface of the heterodimer viewed from the intracellular side. (B) Comparison of the number of contacts between EGFR and EphA2 with PIP2 in the membrane over the course of the simulation. The number of contacts is averaged across four replica simulations, considering a 6-Å cutoff distance. KD, kinase domain; SAM, sterile alpha motif.
DISCUSSION
Decades of effort have characterized many RTK homodimer states, but there is still a poor understanding of heteromultimerization, including the conditions under which each multimer state is present. Our data indicate that EGFR-EphA2 heterocomplexes are not present under basal conditions or after EA1 treatment. Only in the presence of EGF do we detect heteromultimerization of EGFR and EphA2 (Fig. 3). Previous reports of EGFR-EphA2 interaction conditions are somewhat contradictory. One study reported that endogenous EphA2 and EGFR showed increased coimmunoprecipitation (co-IP) after EGF treatment, but only after 24 hours (26). Co-IP only occurred under basal conditions in cell lines where EGFR was constitutively activated/phosphorylated, which suggests that EGFR activation is necessary for heterodimer formation (26). A more recent study that used overexpressed proteins reported an increase in co-IP in as little as 5 to 30 min after EGF treatment (31). While it appears that the time at which heterodimerization occurs could be dependent on cell line, these studies agree with our findings that activation of EGFR by EGF induces formation of EGFR-EphA2 heteromultimers. However, another recent study reported the opposite: Fluorescence resonance energy transfer showed that heterodimers are strong under basal conditions and are decreased by EGF treatment (73). It is important to consider, although, that these experiments were predominantly performed with truncated versions of the receptors that do not contain the ICDs, suggesting that the ICDs participate directly in the conformational changes induced by EGF. The importance of the ICDs is supported by our findings that ICD-only constructs (fig. S10) also undergo PIP2 dependent homo- and heterodimerization, which is expected because PIP2 lipids predominantly localized to the inner leaflet of the plasma membrane. Future work will be essential to determine how protein mutations, expression levels, different cell lines, and variations in signaling networks affect ligand-mediated EGFR-EphA2 heterodimer formation.
Our data show that PIP2 is an important factor in membrane receptor assembly. Because EGFR is one of the most well-studied RTKs, a notable effort has been undertaken to understand how PIP2 affects EGFR homodimers. Both computational and experimental studies have shown that PIP2 binds to the JM segment via electrostatic interactions to stabilize the active conformation of EGFR homomultimers (70, 71). In addition, Western blot studies of EGFR phosphorylation suggest that PIP2 facilitates the activation of EGFR by EGF (72). However, while we observed homodimer stabilization under high PIP2 conditions (Fig. 2), our Western blot data did not support the hypothesis that PIP2 levels alone could drive ligand-independent EGFR phosphorylation (fig. S10). One possibility for this discrepancy is the difference in our methods used to disrupt PIP2. Michailidis et al. (72) worked in oocytes treated with wortmannin, a PI4K inhibitor that decreases PIP2 levels by blocking lipid synthesis, rather than affecting degradation like U73122 and 3M3FBS. In addition, PIP2 changes do not have an effect on EGFR phosphorylation in HeLa cells (72), indicating the effect might be cell line dependent. It is important to continue to study why PIP2 level changes have an effect in some cells but not in others, as this effort will result in a better understanding of the molecular mechanisms of RTK heterodimerization.
A large-scale molecular dynamics simulation study suggested that PIP2 binding to the JM region is conserved across RTKs (34). Hedger and coworkers (34) found that the conserved, positively charged JM residues of all 58 known RTKs bind to and induce clustering of anionic lipids, including PIP2. The Barrera laboratory recently showed that PIP2 specifically promotes dimerization of a construct that contains the TM domain and a short JM span of EphA2. This effect was specific, as it only occurred in the ligand-independent dimer conformation (39). However, before this study, it was not known if PIP2 played a role in the multimerization of full-length EphA2. Here, we report that PIP2 promotes the formation of EphA2 homomultimers (Fig. 2). Intriguingly, tyrosine autophosphorylation does not occur in these multimers (fig. S10), indicating that they are inactive. This result agrees with Stefanski et al.’s (39) hypothesis that PIP2 preferentially promotes ligand-independent multimerization. This is particularly interesting, as unliganded EphA2 oligomers promote tumorigenesis (18, 63, 74). On the basis of our data, we propose that tumors with elevated levels of PIP2 might suffer worse prognosis than those with lower levels. If this idea was correct, pharmacological regulation of PIP2 levels might be clinically beneficial to control EphA2-mediated cancers. Future work is necessary to test this hypothesis.
Because PIP2 promotes homodimerization of both EGFR and EphA2, it would be plausible to assume that it would reduce heteromultimerization simply by the laws of mass action. However, we were surprised to find that PIP2 also promotes heteromultimer formation (Fig. 3). Our study sets the stage for future work to determine how other anionic lipids such as phosphatidylserine or cholesterol, might regulate the heterodimerization of other RTKs, as well as the switch between homo- and heterodimerization. The phosphorylation data presented above suggest that PIP2 promotes an inactive form of the EGFR-EphA2 heteromultimer (Fig. 4C). Therefore, our data put forward a mechanism by which there may be more than one EGFR-EphA2 functional state. Because RTKs including EphA2 and EGFR have more than one homodimer conformation relating to active and inactive states (3, 63, 75, 76), it is not unreasonable to suggest that this extrapolates to heteromultimers.
In accordance with the finding presented here, we propose the following model outlined in Fig. 7. Under basal conditions, EGFR and EphA2 do not bind, and only form heterocomplexes in response to EGF activation. The EGF-induced heteromultimer is phosphorylated at EGFR Y1068, corresponding to EGFR kinase activity, and EphA2 S897, corresponding to kinase activation and the availability of EphA2 as a substrate for serine/threonine kinases (63). The AlFoM predictions and all-atom molecular simulations provide useful insights to the conformation of inactive and active heterodimers (Fig. 5). Low-cellular PIP2 allows the JM to move freely such that the kinase domains adopt an active conformation where the active pockets are free to perform phosphorylation. In contrast, increased levels of PIP2 cause the EGFR JM to bind tightly to the membrane. This conformation would promote an inactive heterodimer conformation, where the EphA2 active pocket is sterically blocked by the EGFR kinase domain, hindering EphA2 phosphorylation activity. It is also possible that PIP2 interactions work in concert with the rotomeric state of the helical JM domain of EGFR, which is important for ligand-induced conformational coupling across the membrane (77). These mechanistic details will require additional structural studies, but our model provides an important conceptual framework that will help guide future experiments. More work is needed to establish the mechanism for the conformational transition from an inactive to active heterodimer and how PIP2 affects EGFR-EphA2 heterodimers.
Fig. 7. Model for EGFR-EphA2 heterodimerization.
When PIP2 levels are normal, EGFR (blue) and EphA2 (orange) do not interact in the absence of ligand. When EGF is present, active heterodimer forms in which EGFR Y1068 and EphA2 S897 are phosphorylated. The active conformation of the ICDs allows a more open EphA2 kinase domain with the EGFR JM free to move away from the membrane. Under increased PIP2 conditions, PIP2 promotes inactive heterodimer formation. The inactive dimer is likely stabilized by the EGFR JM electrostatically interacting with the membrane, and the EGFR ICD is pulled toward the membrane to close the EphA2 kinase domain.
MATERIALS AND METHODS
Plasmids and cloning
EGFR was subcloned to eGFP-N1 and mCH-N1 vectors by Xho I and Age I digest as described previously (78). Human EphA2 cDNA (residues 1 to 971 based on NM_004431.4) with C-terminal GFP and mCH tag was purchased from Origene in gateway donor vectors as described previously (61). PH-PLCD1-GFP was a gift from T. Balla (Addgene, plasmid # 51407; http://n2t.net/addgene:51407; RRID: Addgene_51407) (79).
Cell culture and ligand stimulation
For the culture and expression of the receptor proteins, the COS-7 cells were transiently transfected using standard methods. This study used Dulbecco’s modified Eagle’s medium supplemented with 10% fetal bovine serum and 1% penicillin-streptomycin. Transfection of the plasmids was carried out with 70 to 80% confluent cells in 35-mm glass-bottom MatTek dishes. The cells were transiently transfected approximately 20 hours before the data collection with the protein of interest using Lipofectamine 2000 reagent (Thermo Fisher Scientific). A total of 2.5 μg of DNA (1:1) ratio of mCH and eGFP-tagged plasmids was used to express both fluorescent tags evenly and acquire a local density of 100 to 1500 receptors/μm2. Before measuring FCCS, the medium was changed to Opti-MEM I Reduced Serum Medium (Thermo Fisher Scientific) without phenol red. For each complex, measurement was taken for both the ligand-free and ligand-stimulated state using either recombinant human EGF (Sigma-Aldrich, St. Louis, MO) or EA1 as ligand. To stimulate receptor-expressing cells, a stock solution (20 μg/ml) was diluted to Opti-medium (500 ng/ml; imaging media) and added approximately 15 min before data collection. Data were collected for a maximum of 1 hour following stimulation.
We have previously published control constructs useful in interpreting fc values for monomers, dimers, and higher-order oligomers. Moreover, these control constructs indicate that the fluorescent proteins themselves do not promote or inhibit dimerization states of the proteins. According to a homotypic interaction, the fc values below 0.09 indicate monomers, 0.09 to 0.17 indicate dimers, and above 0.17 indicate higher-order oligomers.
Cells were incubated with different concentrations of PLC regulators to determine the optimum concentration. The PLC inhibitor (U73122) was incubated for 15 min at 1.0 to 5.0 μM concentration before data collection. Concentration ranges of 1.0 to 4.0 did not show any significant changes; however, concentration ranges of 5.0 did show a significant change. After 15 min of PLC activator (m-3M3FBS) stimulation, EFGR/EphA2 did not show a detectable fc value change with 5, 10, 15, and 20 μM concentration. However, the further experiments with 25 μM concentration and 45-min incubation, the median value of cross-correlation changed from 0.20 to 0.04, indicating monomeric forms of EphA2 receptors.
PIE-FCCS instrumentation, data collection, and analysis
The time-time autocorrelation function is used for single color FCS to analyze intensity fluctuations. The results were plotted on a semi-log axis to better view the time scale. In principle, ACF amplitude is inversely proportional to the average number of molecules in the observation area. In FCCS, two fluorescent probes are used to analyze the emission independently of one another using two separate spectrally distinct probes. As a result, both populations have a corresponding ACF, and molecular density can be determined independently. With PIE-FCCS, two laser pulses are phase-delayed, calculating the exact arrival time of each laser pulse’s emission photons.
The FCCS data were recorded using pulsed interleaved excitation and time-correlated single-photon detection with a custom inverted microscope setup. A supercontinuum pulsed laser (9.2-MHz repetition rate, SuperK NKT Photonics, Birker*d, Denmark) was split into two beams of 488 and 561 nm using a series of filter and mirror combinations. To achieve PIE, the beams are directed through separate optical fibers of varying lengths, causing a delay in arrival time between them. This eliminates spectral cross-talk between the detectors. Before entering the microscope, the beams were overlapped by a dichroic beam splitter (LM01-503-25, Semrock) and a customized filter block (zt488/561rpc, zet488/561 m, Chroma Technology). The overlapping beams of light were focused by the objective (×100 total internal reflection fluorescence) to a limited diffraction spot on the peripheral membrane of a COS-7 cell expressing the receptor constructs. In time-tagged time-resolved mode, photons were detected by individual avalanche photodiodes (Micro Photon Devices). To verify the alignment of the system, including the overlap of a confocal volume, a short fluorescently tagged DNA fragment was used. Before the experimental samples, positive and negative controls (Fig. 1, D and E, and figs. S1 and S2) were tested to compare the fit parameters.
The excitation beams were focused on the peripheral membranes to measure only membrane-bound receptors. Only the cell’s flat, peripheral membrane area was scanned to prevent fluorescence from cytosolic organelles or vesicles. The data collection was performed on one area of the membrane in each sample. Each area was assessed six times, with each acquisition lasting 10 s. MATLAB scripts were used to calculate each sample’s auto- and cross-correlation curves. As described in the previous work, we fit a single component, the two-dimensional (2D) diffusion model, to the averaged curves of six consecutive acquisitions per area
The intensity fluctuations were gated using PIE before cross-correlation and autocorrelation analyses. The intensity at time F(t) was compared to the intensity at a later time F(t + τ) in an autocorrelation analysis. As a function of time, self-similarity allowed for the interpretation of quantitative information, such as diffusion and particle number. In the equation above, the intensity fluctuations were divided into 10-s bins and normalized to the square of the average intensity. The autocorrelation functions were fit to the following model for 2D diffusion in the membrane that accounts for triplet relaxation and dark state dynamic
The diffusion coefficient of the sample has been calculated by using the standard formula given below
Last, the cross-correlation (fraction of correlation) was calculated by using the autocorrelation function of both fluorophores
For control samples for fraction of correlation (fc) calculation, we applied two samples for the calculation of fc value of membrane proteins. The first, a monomer, is the SRC-derived peptide fused directly to the fluorescent protein. The second, a dimer control, is a lipid-anchored peptide derived from SRC fused to the GCN4 (SRC-GCN4) α-helical dimerization motif and a C-terminal attached fluorescent protein, i.e., eGFP or mCH. The SRC and SRC-GCN4 constructs were used as negative and positive controls, respectively, for cross-correlation in the experiment. The absence of any cross-correlation in SRC indicates that PIE has successfully eliminated any artifacts that could have caused inaccurate cross-correlation readings and that the fluorescent proteins themselves did not play a role in promoting dimerization. The data from SRC-GCN4 demonstrate the maximum limit for a system that is strongly dimerized, which can be attributed to protein dark states and the existence of dimers having the same fluorescent protein tags. Laser powers were set to 0.350 and 0.80 μW for the 488- and 561-nm lasers, respectively.
To calibrate the highest fluorescence value of a sample, we used double-labeled DNA strands. These strands were also used as DNA standards to calibrate confocal volumes. A 40-nucleotide sequence with <50% G-C content (ACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATAT) was labeled with carboxy-tetramethylrhodamine (TAMRA) at the 5′ end and 6-carboxyfluorescein (FAM) at the 3′ end (Integrated DNA Technologies). An excess of the unlabeled strand was used to anneal the synthesized complementary strand pair as per the supplier’s protocols. The double-stranded DNA, labeled with both TAMRA and FAM (TAMRA-40-FAM), was diluted to a final concentration of 100 nM using 10 mM Tris-EDTA buffer. For the 3D sample data collection, laser powers were set to 7 and 7 μW for the 488- and 561-nm lasers, respectively.
Live-cell microscopy of PLCδ1PH-GFP
Live-cell imaging was performed to verify the effect of the PLC drugs in an assay adapted from Balla and Várnai (58). To prepare cells for imaging, A375 cells were plated at 60% confluency on a #1.5 glass-bottom eight-well chamber slide. The glass was coated in fibronectin (10 μg/ml) for 30 min at 37°C before plating, and cells were allowed to adhere for 24 hours. Cells were then transfected in OptiMEM medium with PLCδ1PH-GFP using a 3:1 ratio of polycation polyethylenimine to DNA with 0.2 μg of DNA per well and allowed to incubate overnight. Cells were washed using phosphate-buffered saline, starved of serum for 4 hours, and stained with Hoechst 33342 for 20 min to allow visualization of the nucleus. Cells were lastly placed in FluoroBrite medium for optimal imaging.
Time-lapse imaging of cells were performed using an inverted Zeiss LSM 900/Airyscan laser scanning confocal microscope (ZEISS) with a 63× oil immersion objective. For a given well, three to six spots containing multiple fluorescent cells were chosen. The well was then treated with 0.5% (v/v) dimethyl sulfoxide (DMSO), and the desired Z plane was found using DefiniteFocus. For cells planned for U73122 treatment, images were taken every 5 min for 20 min, and for cells planned for 3M3FBS treatment, images were taken every 10 min for 50 min. Following the experiment, DMSO was replaced with either U73122 or 3M3FBS, the DefiniteFocus was re-established, and time-lapse images were repeated as above.
Images were processed and quantified using the ImageJ software. Four representative membrane areas and four representative cytosolic areas were chosen. The mean pixel intensity for each section was determined, and the normalized ratio of membrane to cytosol intensity for each single cell was calculated using the following equation
Western blot
A375 cells were plated to approximately 80% confluency in a 12-well culture dish and allowed to adhere for 24 hours. Cells were then starved of serum overnight. Treatments were made in serum-free DMEM, and cells were treated with 5 μM U73122 for 15 min or 25 μM 3M3FBS for 45 min and then treated with EA1 (500 ng/ml) or EGF (100 ng/ml) for 15 min. Control cells were treated with equal amounts of DMSO as PIP2-modifying drugs. Once treatments were complete, cells were scraped from the plate and lysed in TEN-T lysis buffer [50 mM tris (pH 7.5), 100 mM sodium chloride, 1 mM EDTA, and 1% Triton X-100] containing protease and phosphatase inhibitors for 30 min on ice. Lysates were centrifuged at 13,000g for 10 min to remove insoluble material. A total of 10 ng of protein was run on a 10% SDS–polyacrylamide gel electrophoresis gel and transferred to 0.45-μm nitrocellulose. Membranes were blocked with 5% milk for total protein blots or 3% bovine serum albumin for phospho-protein blots for 1 hr at room temperature and then incubated with primary antibodies (Table 1) overnight at 4 °C. Secondary antibodies (Table 1) were incubated 1 hr at room temperature and imaged for fluorescence on an Odyssey CLx Imaging System (LI-COR, Lincoln, Nebraska). Bands were quantified using Image Studio Lite. Phospho-protein bands were normalized to the corresponding total protein levels, while total protein bands were normalized to their corresponding actin loading control bands.
Table 1. List of antibodies used in Western blots.
Each antibody is listed by the name used in the text. The source of the antibody, solvent, dilution ratio, and host species are also provide.CST, Cell Signaling Technology. TBS, tris-buffered saline; BSA, bovine serum albumin.
| Antibody | Catalog # | Solvent | Dilution | Host species |
|---|---|---|---|---|
| Total EGFR | 4267S, CST | 5% milk in TBS | 1:1000 | Rabbit |
| pEGFR(Y1068) | 3777S, CST | 3% BSA in TBS | 1:1000 | Rabbit |
| Total EphA2 | 97535S, CST | 5% Milk | 1:1000 | Rabbit |
| pEphA2(Y588) | 12677, CST | 3% BSA | 1:1000 | Rabbit |
| pEphA2(S897) | 6347S, CST | 3% BSA | 1:1000 | Rabbit |
| Actin | 3700S, CST | 5% milk | 1:5000 | Mouse |
| Anti-mouse | 925-68020, LiCore | 5% milk | 1:5000 | Goat |
| Anti-rabbit | 926-32211, LiCore | 5% milk | 1:5000 | Goat |
AlphaFold Multimer
AlFoM (56) was used to predict ICD dimerization of:
EGFR(R669RRHIVRKRTLRRLLQERELVEPLTPSGEAPNQALLRILKETEFKKIKVLGSGAFGTVYKGLWIPEGEKVKIPVAIKELREATSPKANKEILDEAYVMASVDNPHVCRLLGICLTSTVQLIT-QLMPFGCLLDYVREHKDNIGSQYLLNWCVQIAKGMNYLEDRRLVHRDLAARNVLVKTPQHVKITDFGLAKLLGAEEKEYHAEGGKVPIKWMALESILHRIYTHQSDVWSYGVTVWELMTFGSKPYDGIPASEISSILEKGERLPQPPICTIDVYMIMVKCWMIDADSRPKFRELIIEFSKMARDPQRYLVIQGDERMHLPSPTDSNFYRALMDEEDMDDVVDADEYLIPQQGFFSSPSTSRTPLLSSLSATSNNSTVACIDRNGLQSCPIKEDSFLQRYSSDPTGALTEDSIDDTFLPVPEYINQSVPKRPAGSVQNPVYHNQPLNPAPSRDPHYQDPHSTAVGNPEYLNTVQPTCVNSTFDSPAHWAQKGSHQISLDNPDYQQDFFPKEAKPNGIFKGSTAENAEYLRVAPQSSEFIGA1210) and EphA2WT(H559RRRKNQRARQSPEDVY-FSKSEQLKPLKTYVDPHTYEDPNQAVLKFTTEIHPSCVTRQKVIGAGEFGEVYKGMLKTSSGKKEVPVAIKTLKAGYTEKQRVDFLGEAGIMGQFSHHNIIRLEGVISKYKPMMIITEYMENGALDKFLREKDGEFSVLQLVGMLRGIAAGMKYLANMNYVHRDLAARNILVNSNLVCKVSDFGLSRVLEDDPEATYTTSGGKIPIRWTAPEAISYRKFTSASDVWSFGIVMWEVMTYGERPYWELSNHEVMKAINDGFRLPTPMDCPSAIYQLMMQCWQQERARRPKFADIVSILDKLIRAPDSLKTLADFDPRVSIRLPSTSGSEGVPFRTVSEWLESIKMQQYTEHFMAAGYTAIEKVVQMTNDDIKRIGVRLPGHQKRIAYSLLGLKDQVNTVGIPI967).
For EphA2S897E/S901E, the two bolded S residues were replaced with E. The crystal structures of EGFR (PDB: 2GS6) (59) and EphA2 (PDB: 7KJA) (60) were obtained from www.rcsb.org/. For each simulation, five models were provided, and the Best Model was used for further analysis. RMSDs were calculated using the ChimeraX matchmaking tool.
Modeling of the EGFR-EphA2 heterodimer
We used Alphafold2 Multimer (64) to model the heterodimer of the intracellular region (ICR) of EphA2 and the kinase domain of EGFR. To gain deeper insights into the protein-lipid interface, we predicted the models of the TM-JM regions of both EphA2 (residues E530-H609) and EGFR (residues I646-I706) separately before joining them with the ICRs. In addition, we extended the TM-JM models by adding eight extra residues at the N terminus in an extended conformation (ϕ, ψ = ±120°), as these extramembrane proximal regions are known to enhance stability with the lipid bilayer. ATP and Mg2+ were also incorporated into the respective kinase domains to complete the model.
Set up for all-atom molecular dynamics simulation
To analyze the heterodimerization of EGFR-EphA2 system, we placed the starting model in the lipid bilayer perpendicular to the plane of the membrane, which has a lipid composition of POPC (55%), cholesterol (30%), POPS (10%), and PIP2 (5%). The system consists of 440 POPC, 240 cholesterol, 80 POPS, and 40 PIP2 lipids molecules, which were encased in a box with dimensions measuring 150 Å by 150 Å × 160 Å. The system was solvated using the TIP3P water model. To ensure neutrality, Na+ ions were added, and 150 mM NaCl was introduced using the CHARMM-GUI interface (80). Molecular dynamics simulations were performed using GROMACS 2021.5 (81) with the CHARMM36m force field (82). The long-range electrostatic interactions were managed using the particle mesh Ewald method, while hydrogen-containing covalent bonds were constrained using the SHAKE algorithm. The system underwent energy minimization through the steepest descent algorithm, consisting of 5000 steps. System equilibration followed a sequential process of six steps, wherein the harmonic restraints on proteins and lipids were gradually reduced. The initial two equilibration steps were conducted within the constant number of particles, volume, and temperature (NVT) ensemble, while the subsequent four steps occurred under the constant number of particles, pressure, and temperature (NPT) ensemble to maintain an isobaric-isothermal condition. A total of four replica simulations were conducted for 200 ns each.
Simulation data analysis
Trajectory analysis was conducted using the integrated modules within GROMACS. Subsequently, the data were visualized and plotted using Origin.
Acknowledgments
We are thankful to T. Simmons and A. Joshi for experimental advice and the use of the LSM 900/Airyscan microscope. We used the high-performance computing resources at CWRU for the molecular dynamics simulations.
Funding: This work was supported by the National Science Foundation grant CHE-1753060, the Human Frontiers of Science Program no. RGP0059/2019, and the NIH R35GM152126 (to A.W.S.), and NIH grants R35GM140846 (to F.N.B.) and R21AG084065 and R01EY029169 (to M.B.).
Author contributions: Conceptualization: A.W.S., P.K.S., and F.N.B. Performed experiments: P.K.S., J.A.R., and R.J.S. Data analysis: P.K.S., A.W.S., J.A.R., and F.N.B.; MD simulations: A.R.S. and M.B. Writing—original draft: P.K.S., A.W.S., J.A.R., and F.N.B. Writing—review and editing: A.W.S., F.N.B., P.K.S., and J.A.R. Supervision and project administration: A.W.S., F.N.B., and M.B. Funding acquisition: A.W.S., F.N.B., and M.B.
Competing interests: The authors declare that they have no competing interests.
Data and materials availability: All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Materials.
Supplementary Materials
This PDF file includes:
Figs. S1 to S16
Tables S1 to S4
REFERENCES AND NOTES
- 1.Santos R., Ursu O., Gaulton A., Bento A. P., Donadi R. S., Bologa C. G., Karlsson A., Al-Lazikani B., Hersey A., Oprea T. I., Overington J. P., A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Groves J. T., Kuriyan J., Molecular mechanisms in signal transduction at the membrane. Nat. Struct. Mol. Biol. 17, 659–665 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Paul M. D., Hristova K., The transition model of RTK activation: A quantitative framework for understanding RTK signaling and RTK modulator activity. Cytokine Growth Factor Rev. 49, 23–31 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lemmon M. A., Schlessinger J., Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Paul M. D., Hristova K., The RTK Interactome: Overview and perspective on RTK heterointeractions. Chem. Rev. 119, 5881–5921 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Du Z., Lovly C. M., Mechanisms of receptor tyrosine kinase activation in cancer. Mol. Cancer 17, 58 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lemmon M. A., Schlessinger J., Ferguson K. M., The EGFR family: Not so prototypical receptor tyrosine kinases. Cold Spring Harb. Perspec. Biol. 6, a020768 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kovacs T., Zakany F., Nagy P., It takes more than two to tango: Complex, hierarchal, and membrane-modulated interactions in the regulation of receptor tyrosine kinases. Cancers 14, 994 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Graus-Porta D., Beerli R. R., Daly J. M., Hynes N. E., Miescher F., ErbB-2, the preferred heterodimerization partner of all ErbB receptors, is a mediator of lateral signaling for interaction with polypeptide ligands. A direct con-of receptor dimers and stimulation of the intrinsic kinase. EMBO J. 16, 1647–1655 (1997). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Normanno N., De Luca A., Bianco C., Strizzi L., Mancino M., Maiello M. R., Carotenuto A., De Feo G., Caponigro F., Salomon D. S., Epidermal growth factor receptor (EGFR) signaling in cancer. Gene 366, 2–16 (2006). [DOI] [PubMed] [Google Scholar]
- 11.Pasquale E. B., Eph-ephrin bidirectional signaling in physiology and disease. Cell 133, 38–52 (2008). [DOI] [PubMed] [Google Scholar]
- 12.Pasquale E. B., Eph receptors and ephrins in cancer: Bidirectional signalling and beyond. Nat. Rev. Cancer 10, 165–180 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Liang L. Y., Patel O., Janes P. W., Murphy J. M., Lucet I. S., Eph receptor signalling: From catalytic to non-catalytic functions. Oncogene 38, 6567–6584 (2019). [DOI] [PubMed] [Google Scholar]
- 14.Song W., Kim L. C., Han W., Hou Y., Edwards D. N., Wang S., Blackwell T. S., Cheng F., Brantley-Sieders D. M., Chen J., Phosphorylation of PLCg1 by EphA2 receptor tyrosine kinase promotes tumor growth in lung cancer. Mol. Cancer Res. 18, 1735–1743 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhao P., Jiang D., Huang Y., Chen C., EphA2: A promising therapeutic target in breast cancer. J. Genet. Genomics 48, 261–267 (2021). [DOI] [PubMed] [Google Scholar]
- 16.Tröster A., Jores N., Mineev K. S., Sreeramulu S., DiPrima M., Tosato G., Schwalbe H., Targeting EPHA2 with kinase inhibitors in colorectal cancer. ChemMedChem 18, e202300420 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wilson K., Shiuan E., Brantley-Sieders D. M., Oncogenic functions and therapeutic targeting of EphA2 in cancer. Oncogene 40, 2483–2495 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shi X., Lingerak R., Herting C. J., Ge Y., Kim S., Toth P., Wang W., Brown B. P., Meiler J., Sossey-Alaoui K., Buck M., Himanen J., Hambardzumyan D., Nikolov D. B., Smith A. W., Wang B., Time-resolved live-cell spectroscopy reveals EphA2 multimeric assembly. Science 382, 1042–1050 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Stern D. F., Tyrosine kinase signalling in breast cancer: ErbB family receptor tyrosine kinases. Breast Cancer Res. 2, 176–183 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Codony-Servat J., Codony-Servat C., Cardona A. F., Giménez-Capitán A., Drozdowskyj A., Berenguer J., Bracht J. W. P., Ito M., Karachaliou N., Rosell R., Cancer stem cell biomarkers in EGFR-mutation–positive non–small-cell lung cancer. Clin. Lung Cancer 20, 167–177 (2019). [DOI] [PubMed] [Google Scholar]
- 21.Paez J. G., Jänne P. A., Lee J. C., Tracy S., Greulich H., Gabriel S., Herman P., Kaye F. J., Lindeman N., Boggon T. J., Naoki K., Sasaki H., Fujii Y., Eck M. J., Sellers W. R., Johnson B. E., Meyerson M., EGFR mutations in lung cancer: Correlation with clinical response to gefitinib therapy. Science 304, 1497–1500 (2004). [DOI] [PubMed] [Google Scholar]
- 22.Sigismund S., Avanzato D., Lanzetti L., Emerging functions of the EGFR in cancer. Mol. Oncol. 12, 3–20 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Tebbutt N., Pedersen M. W., Johns T. G., Targeting the ERBB family in cancer: Couples therapy. Nat. Rev. Cancer 13, 663–673 (2013). [DOI] [PubMed] [Google Scholar]
- 24.Cioce M., Fazio V. M., Epha2 and egfr: Friends in life, partners in crime. can epha2 be a predictive biomarker of response to anti-egfr agents? Cancers 13, 700 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Brantley-Sieders D. M., Zhuang G., Hicks D., Wei B. F., Hwang Y., Cates J. M. M., Coffman K., Jackson D., Bruckheimer E., Muraoka-Cook R. S., Chen J., The receptor tyrosine kinase EphA2 promotes mammary adenocarcinoma tumorigenesis and metastatic progression in mice by amplifying ErbB2 signaling. J. Clin. Invest. 118, 64–78 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Larsen A. B., Pedersen M. W., Stockhausen M. T., Grandal M. V., van Deurs B., Poulsen H. S., Activation of the EGFR gene target EphA2 inhibits epidermal growth factor-induced cancer cell motility. Mol. Cancer Res. 5, 283–293 (2007). [DOI] [PubMed] [Google Scholar]
- 27.Oh D., Chen Z., Biswas K. H., Bai F., Ong H. T., Sheetz M. P., Groves J. T., Competition for shared downstream signaling molecules establishes indirect negative feedback between EGFR and EphA2. Biophys. J. 121, 1897–1908 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hu R., Li Y., Guo Y., Li X., Du S., Liao M., Hou H., Sun H., Zhao S., Su J., Chen X., Yin M., BRD4 inhibitor suppresses melanoma metastasis via the SPINK6/EGFR-EphA2 pathway. Pharmacol. Res. 187, 106609 (2023). [DOI] [PubMed] [Google Scholar]
- 29.Swidergall M., Solis N. V., Millet N., Huang M. Y., Lin J., Phan Q. T., Lazarus M. D., Wang Z., Yeaman M. R., Mitchell A. P., Filler S. G., Activation of EphA2-EGFR signaling in oral epithelial cells by Candida albicans virulence factors. PLOS Pathogens 17, e1009221 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.De Robertis M., Loiacono L., Fusilli C., Poeta M. L., Mazza T., Sanchez M., Marchionni L., Signori E., Lamorte G., Vescovi A. L., Garcia-Foncillas J., Fazio V. M., Dysregulation of EGFR pathway in EphA2 cell subpopulation significantly associates with poor prognosis in colorectal cancer. Clin. Cancer Res. 23, 159–170 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kim J., Chang I. Y., You H. J., Interactions between EGFR and EphA2 promote tumorigenesis through the action of Ephexin1. Cell Death Dis. 13, 528 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.McLaughlin S., Smith S. O., Hayman M. J., Murray D., An electrostatic engine model for autoinhibition and activation of the epidermal growth factor receptor (EGFR/ErbB) family. J. Gen. Physiol. 126, 41–53 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.McLaughlin S., Wang J., Gambhir A., Murray D., PIP2 and proteins: Interactions, organization, and information flow. Annu. Rev. Biophys. Biomol. Struct. 31, 151–175 (2002). [DOI] [PubMed] [Google Scholar]
- 34.Hedger G., Sansom M. S. P., Koldsø H., The juxtamembrane regions of human receptor tyrosine kinases exhibit conserved interaction sites with anionic lipids. Sci. Rep. 5, 9198 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Arkhipov A., Shan Y., Das R., Endres N. F., Eastwood M. P., Wemmer D. E., Kuriyan J., Shaw D. E., Architecture and membrane interactions of the EGF receptor. Cell 152, 557–569 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wang Y., Gao J., Guo X., Tong T., Shi X., Li L., Qi M., Wang Y., Cai M., Jiang J., Xu C., Ji H., Wang H., Regulation of EGFR nanocluster formation by ionic protein-lipid interaction. Cell Res. 24, 959–976 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chavent M., Seiradake E., Jones E. Y., Sansom M. S. P., Structures of the EphA2 receptor at the membrane: Role of lipid interactions. Structure 24, 337–347 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Chavent M., Karia D., Kalli A. C., Domański J., Duncan A. L., Hedger G., Stansfeld P. J., Seiradake E., Jones E. Y., Sansom M. S. P., Interactions of the EphA2 Kinase Domain with PIPs in Membranes: Implications for receptor function. Structure 26, 1025–1034.e2 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Stefanski K. M., Russell C. M., Westerfield J. M., Lamichhane R., Barrera F. N., PIP2 promotes conformation-specific dimerization of the EphA2 membrane region. J. Biol. Chem. 296, 100149 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Petazzi R. A., Aji A. K., Chiantia S., Fluorescence microscopy methods for the study of protein oligomerization. Prog. Mol. Biol. Transl. Sci. 169, 1–41 (2020). [DOI] [PubMed] [Google Scholar]
- 41.Zhang X., Yin J., Pan W., Li Y., Li N., Tang B., Imaging strategies for receptor tyrosine kinase dimers in living cells. Anal. Bioanal. Chem. 415, 67–82 (2023). [DOI] [PubMed] [Google Scholar]
- 42.Asher W. B., Geggier P., Holsey M. D., Gilmore G. T., Pati A. K., Meszaros J., Terry D. S., Mathiasen S., Kaliszewski M. J., McCauley M. D., Govindaraju A., Zhou Z., Harikumar K. G., Jaqaman K., Miller L. J., Smith A. W., Blanchard S. C., Javitch J. A., Single-molecule FRET imaging of GPCR dimers in living cells. Nat. Methods 18, 397–405 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Digman M. A., Gratton E., Lessons in fluctuation correlation spectroscopy. Annu. Rev. Phys. Chem. 62, 645–668 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Christie S., Shi X., Smith A. W., Resolving membrane protein–protein interactions in live cells with pulsed interleaved excitation fluorescence cross-correlation spectroscopy. Acc. Chem. Res. 53, 792–799 (2020). [DOI] [PubMed] [Google Scholar]
- 45.Sankaran J., Wohland T., Current capabilities and future perspectives of FCS: Super-resolution microscopy, machine learning, and in vivo applications. Commun. Biol. 6, 699 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Bacia K., Kim S. A., Schwille P., Fluorescence cross-correlation spectroscopy in living cells. Nat. Methods 3, 83–89 (2006). [DOI] [PubMed] [Google Scholar]
- 47.Müller B. K., Zaychikov E., Bräuchle C., Lamb D. C., Pulsed interleaved excitation. Biophys. J. 89, 3508–3522 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Kaliszewski M. J., Shi X., Hou Y., Lingerak R., Kim S., Mallory P., Smith A. W., Quantifying membrane protein oligomerization with fluorescence cross-correlation spectroscopy. Methods 140-141, 40–51 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Comar W. D., Schubert S. M., Jastrzebska B., Palczewski K., Smith A. W., Time-resolved fluorescence spectroscopy measures clustering and mobility of a G protein-coupled receptor opsin in live cell membranes. J. Am. Chem. Soc. 136, 8342–8349 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Smith A. W., Detection of rhodopsin dimerization in situ by PIE-FCCS, a time-resolved fluorescence spectroscopy. Methods Mol. Biol. 1271, 205–219 (2015). [DOI] [PubMed] [Google Scholar]
- 51.Balla T., Phosphoinositides: Tiny lipids with giant impact on cell regulation. Physiol. Rev. 93, 1019–1137 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.van den Bogaart G., Meyenberg K., Risselada H. J., Amin H., Willig K. I., Hubrich B. E., Dier M., Hell S. W., Grubmuller H., Diederichsen U., Jahn R., Membrane protein sequestering by ionic protein-lipid interactions. Nature 479, 552–555 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Kadamur G., Ross E. M., Mammalian phospholipase C. Annu. Rev. Physiol. 75, 127–154 (2013). [DOI] [PubMed] [Google Scholar]
- 54.Smith R. J., Sam L. M., Justen J. M., Bundy G. L., Bala G. A., Bleasdale J. E., Receptor-coupled signal transduction in human polymorphonuclear neutrophils: Effects of a novel inhibitor of phospholipase C-dependent processes on cell responsiveness. J. Pharmacol. Exp. Ther. 253, 688–697 (1990). [PubMed] [Google Scholar]
- 55.Bleasdale J. E., Thakur N. R., Gremban R. S., Bundy G. L., Fitzpatrick F. A., Smith R. J., Bunting S., Selective inhibition of receptor-coupled phospholipase C-dependent processes in human platelets and polymorphonuclear neutrophils. J. Pharmacol. Exp. Ther. 255, 756–768 (1990). [PubMed] [Google Scholar]
- 56.Horowitz L. F., Hirdes W., Suh B.-C., Hilgemann D. W., Mackie K., Hille B., Phospholipase C in living cells : Activation, inhibition, Ca2+ requirement, and regulation of M current. J. Gen. Physiol. 126, 243–262 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Bae Y.-S., Lee T. G., Park J. C., Hur J. H., Kim Y., Heo K., Kwak J.-Y., Suh P.-G., Ryu S. H., Identification of a compound that directly stimulates phospholipase C activity. Mol. Pharmacol. 63, 1043–1050 (2003). [DOI] [PubMed] [Google Scholar]
- 58.Balla T., Várnai P., Visualizing cellular phosphoinositide pools with GFP-fused protein-modules. Sci. STKE 2002, pl3 (2002). [DOI] [PubMed] [Google Scholar]
- 59.Endres N. F., Das R., Smith A. W., Arkhipov A., Kovacs E., Huang Y., Pelton J. G., Shan Y., Shaw D. E., Wemmer D. E., Groves J. T., Kuriyan J., Conformational coupling across the plasma membrane in activation of the EGF receptor. Cell 152, 543–556 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Huang Y., Bharill S., Karandur D., Peterson S. M., Marita M., Shi X., Kaliszewski M. J., Smith A. W., Isacoff E. Y., Kuriyan J., Molecular basis for multimerization in the activation of the epidermal growth factor receptor. eLife 5, e14107 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.X. Shi, R. Lingerak, C. J. Herting, Y. Ge, S. Kim, P. Toth, C. Cuizon, J. Zheng, L. Chao, K. Sossey-Alaoui, M. Buck, S. Singh, V. Varadan, J. Himanen, D. Hambardzumyan, D. Nikolov, A. W. Smith, B. Wang, Cell Surface multimeric assemblies regulate canonical and noncanonical EphA2 receptor tyrosine kinase signaling. bioRxiv 439330 [Preprint] (2021). 10.1101/2021.04.11.439330. [DOI]
- 62.Gomez-Soler M., Gehring M. P., Lechtenberg B. C., Zapata-Mercado E., Ruelos A., Matsumoto M. W., Hristova K., Pasquale E. B., Ligands with different dimeric configurations potently activate the EphA2 receptor and reveal its potential for biased signaling. iScience 25, 103870 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Miao H., Li D. Q., Mukherjee A., Guo H., Petty A., Cutter J., Basilion J. P., Sedor J., Wu J., Danielpour D., Sloan A. E., Cohen M. L., Wang B., EphA2 mediates ligand-dependent inhibition and ligand-independent promotion of cell migration and invasion via a reciprocal regulatory loop with Akt. Cancer Cell 16, 9–20 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.R. Evans, M. O’Neill, A. Pritzel, N. Antropova, A. Senior, T. Green, A. Žídek, R. Bates, S. Blackwell, J. Yim, O. Ronneberger, S. Bodenstein, M. Zielinski, A. Bridgland, A. Potapenko, A. Cowie, K. Tunyasuvunakool, R. Jain, E. Clancy, P. Kohli, J. Jumper, D. Hassabis, Protein complex prediction with AlphaFold-Multimer. bioRxiv 463034 [Preprint] (2021). 10.1101/2021.10.04.463034. [DOI]
- 65.Lechtenberg B. C., Gehring M. P., Light T. P., Horne C. R., Matsumoto M. W., Hristova K., Pasquale E. B., Regulation of the EphA2 receptor intracellular region by phosphomimetic negative charges in the kinase-SAM linker. Nat. Commun. 12, 7047 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Jura N., Endres N. F., Engel K., Deindl S., Das R., Lamers M. H., Wemmer D. E., Zhang X., Kuriyan J., Mechanism for activation of the EGF receptor catalytic domain by the juxtamembrane segment. Cell 137, 1293–1307 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Zhang X., Gureasko J., Shen K., Cole P. A., Kuriyan J., An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor. Cell 125, 1137–1149 (2006). [DOI] [PubMed] [Google Scholar]
- 68.Mariani V., Biasini M., Barbato A., Schwede T., lDDT: A local superposition-free score for comparing protein structures and models using distance difference tests. Bioinformatics 29, 2722–2728 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Kim D. H., Triet H. M., Ryu S. H., Regulation of EGFR activation and signaling by lipids on the plasma membrane. Prog. Lipid Res. 83, 101115 (2021). [DOI] [PubMed] [Google Scholar]
- 70.Maeda R., Sato T., Okamoto K., Yanagawa M., Sako Y., Lipid-protein interplay in dimerization of juxtamembrane domains of epidermal growth factor receptor. Biophys. J. 114, 893–903 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Halim K. B. A., Koldsø H., Sansom M. S. P., Interactions of the EGFR juxtamembrane domain with PIP2-containing lipid bilayers: Insights from multiscale molecular dynamics simulations. Biochim. Biophys. Acta 1850, 1017–1025 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Michailidis I. E., Rusinova R., Georgakopoulos A., Chen Y., Iyengar R., Robakis N. K., Logothetis D. E., Baki L., Phosphatidylinositol-4,5-bisphosphate regulates epidermal growth factor receptor activation. Pflugers Arch. 461, 387–397 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Paul M. D., Grubb H. N., Hristova K., Quantifying the strength of heterointeractions among receptor tyrosine kinases from different subfamilies: Implications for cell signaling. J. Biol. Chem. 295, 9917–9933 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Paraiso K. H., Das Thakur M., Fang B., Koomen J. M., Fedorenko I. V., John J. K., Tsao H., Flaherty K. T., Sondak V. K., Messina J. L., Pasquale E. B., Villagra A., Rao U. N., Kirkwood J. M., Meier F., Sloot S., Gibney G. T., Stuart D., Tawbi H., Smalley K. S., Ligand-independent EPHA2 signaling drives the adoption of a targeted therapy-mediated metastatic melanoma phenotype. Cancer Discov. 5, 264–273 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Maruyama I. N., Mechanisms of activation of receptor tyrosine kinases: Monomers or dimers. Cells 3, 304–330 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Songtawee N., Bevan D. R., Choowongkomon K., Molecular dynamics of the asymmetric dimers of EGFR: Simulations on the active and inactive conformations of the kinase domain. J. Mol. Graph. Model. 58, 16–29 (2015). [DOI] [PubMed] [Google Scholar]
- 77.Deepto Mozumdar A. D., Zhang J. Y., Rafizadeh D. N., Schepartz A., Discrete coiled coil rotamers form within the EGFRvIII juxtamembrane Domain. Biochemistry 59, 3965–3972 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Du Z., Brown B. P., Kim S., Ferguson D., Pavlick D. C., Jayakumaran G., Benayed R., Gallant J. N., Zhang Y. K., Yan Y., Red-Brewer M., Ali S. M., Schrock A. B., Zehir A., Ladanyi M., Smith A. W., Meiler J., Lovly C. M., Structure–function analysis of oncogenic EGFR Kinase Domain Duplication reveals insights into activation and a potential approach for therapeutic targeting. Nat. Commun. 12, 1382 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Várnai P., Balla T., Visualization of phosphoinositides that bind pleckstrin homology domains: Calcium- and agonist-induced dynamic changes and relationship to myo-[3H]inositol-labeled phosphoinositide pools. J. Cell Biol. 143, 501–510 (1998). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Lee J., Cheng X., Swails J. M., Yeom M. S., Eastman P. K., Lemkul J. A., Wei S., Buckner J., Jeong J. C., Qi Y., Jo S., Pande V. S., Case D. A., Brooks C. L. III, MacKerell A. D. Jr., Klauda J. B., Im W., CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Abraham M. J., Murtola T., Schulz R., Páll S., Smith J. C., Hess B., Lindahl E., GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015). [Google Scholar]
- 82.Huang J., Rauscher S., Nawrocki G., Ran T., Feig M., de Groot B. L., Grubmüller H., MacKerell A. D., CHARMM36m: An improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figs. S1 to S16
Tables S1 to S4