Abstract
Clinical antibodies are an important class of drugs for the treatment of both chronic and acute diseases. Their manufacturability is subject to evaluation to ensure product quality and efficacy. One critical quality attribute is deamidation, a non-enzymatic process that is observed to occur during thermal stress, at low or high pH, or a combination thereof. Deamidation may induce antibody instability and lead to aggregation, which may pose immunogenicity concerns. The introduction of a negative charge via deamidation may impact the desired therapeutic function (i) within the complementarity-determining region, potentially causing loss of efficacy; or (ii) within the fragment crystallizable region, limiting the effector function involving antibody-dependent cellular cytotoxicity. Here we describe a transformative solution that allows for a comparative assessment of deamidation and its impact on stability and aggregation. The innovative streamlined method evaluates the intact protein in its formulation conditions. This breakthrough platform technology is comprised of a quantum cascade laser microscope, a slide cell array that allows for flexibility in the design of experiments, and dedicated software. The enhanced spectral resolution is achieved using two-dimensional correlation, co-distribution, and two-trace two-dimensional correlation spectroscopies that reveal the molecular impact of deamidation. Eight re-engineered immunoglobulin G4 scaffold clinical antibodies under control and forced degradation conditions were evaluated for deamidation and aggregation. We determined the site of deamidation, the overall extent of deamidation, and where applicable, whether the deamidation event led to self-association or aggregation of the clinical antibody and the molecular events that led to the instability. The results were confirmed using orthogonal techniques for four of the samples.
Keywords: Biotherapeutic, asparagine and glutamine deamidation, extent of deamidation, protein aggregation, protein stability, quantum cascade laser microspectroscopy, two-dimensional correlation spectroscopy, 2D-COS, co-distribution correlation spectroscopy, two-trace two-dimensional, 2T2D, correlation spectroscopy, high-performance liquid chromatography, HPLC, cation exchange
Introduction
Therapeutic proteins are a dominant product class of drugs for the treatment of many diseases. Therapeutic protein developers have invested a considerable amount of long-term effort into the early stage development pipeline, which currently includes ∼800 novel antibody therapeutics.1–4 These bioengineered proteins with different modalities require a complex drug development cycle and as such are subject to the evaluation of critical quality attributes (CQAs).5–12 Charge variance is one CQA that may have potential implications for other CQAs, such as target binding (hence efficacy), stability, and aggregation,13,14 and may cause an unwanted immune response in patients, raising safety concerns. Charge variants can be generated when the therapeutic protein is subjected to high temperatures or low or high pH conditions, by a non-enzymatic process known as deamidation in which asparagine (ASN) and glutamine (GLN) residues are converted to aspartate and glutamate, respectively.15–18
The impact that ASN and GLN deamidation may have on protein stability, aggregation, and immunogenicity risk is poorly understood, but the mechanism of deamidation for both residues is well established (Figure 1).18–20 Briefly, a nucleophilic attack from the adjacent backbone amide on the carbonyl of the side chain amide generates a cyclic intermediate (for ASN, succinimide; for GLN, glutarimide), followed by a hydrolysis step causing ring opening and the generation of the corresponding carboxylate side chain. Thus, a negatively charged residue is introduced at the ASN/GLN site. The hydrolytic step of the reaction may generate two different substitutions on the backbone, alpha or gamma, depending on which carbonyl of the ring is retained in the new side chain. Our novel approach allows for the real-time monitoring of these key functional groups within the mechanistic process of deamidation (Figure 1). The introduction of a negative charge can potentially introduce local instability in the protein and induce aggregation. Furthermore, when the location of the newly introduced negative charge is within the complementarity-determining region (CDR) of an antibody, the potential to impact efficacy is also a concern.
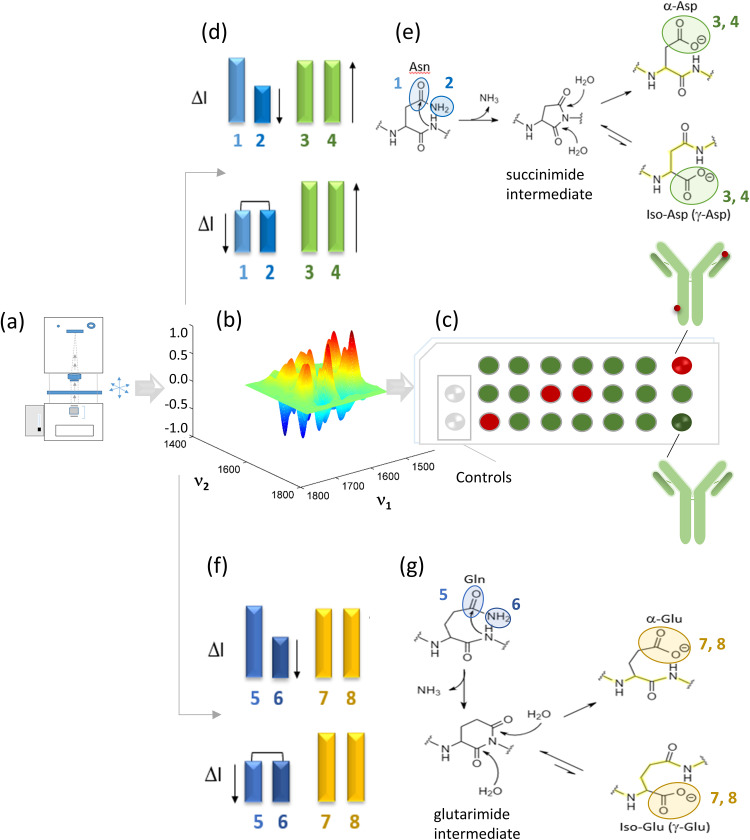
Figure 1.
Streamlined ASN and GLN deamidation comprehensive assessment: (a) QCLM equipped with a heated accessory, (b) 3D representation of a 2D-COS asynchronous plot used to monitor the deamidation process, (c) schematic representation of the slide cell array containing gray wells representing the negative controls or reference standard, green and red wells represent ideal and failed candidates, respectively. The failed candidates exhibit loss of stability, ASN or GLN deamidation, with regions prone to aggregation resulting in increased immunogenicity risk for which the deamidation sites may be localized within the CDRs or the Fc region shown with red spheres on the mAb model associated with the failed candidate. The method involves directly monitoring key signature peaks during the deamidation process: (d) bar graphs that summarize the intensity changes monitored in real time during the deamidation process: (1) ν(C=O) at 1678 cm–1, (2) δ(NH2) at 1612 cm–1 of the ASN side chain and (3 and 4) ν(COO–) at 1572 and 1567 cm–1 for the aspartate products, (e) ASN deamidation mechanism, and (f) bar graphs summarizing the real-time intensity changes monitored for the GLN deamidation process: (5) ν(C=O) at 1670 cm–1, (6) δ(NH2) at 1581 cm–1 of the GLN side chain and (7 and 8) ν(COO–) at 1554 and 1545 cm–1 for the glutamate products, and (g) the GLN deamidation mechanism. Highlighted in yellow is the backbone within each product.
To date, two different approaches have been implemented for predicting the probability of deamidation in a protein. The first is through amino acid sequence analysis and molecular simulations, a theoretically predictive approach based on work done by Robinson and Robinson. 16 In this approach, deamidation was determined to be limited by steric hindrance at the n + 1 position, and the probability of deamidation was G > S > T > N. These results were validated by molecular simulations evaluating the rotational freedom for these residues.16,21–24 The second approach uses empirical evaluation primarily based on liquid chromatography–mass spectroscopy analysis for the identification of ASN deamidation sites within the CDRs of antibodies.25,26 A confirmed deamidation site within the CDR has been associated with loss of efficacy due to decreased affinity for the antigen. Decreased affinity due to deamidation may also result in false negative results in target–antigen-binding assays, which can negatively impact exploratory research and the proteomic databases that provide the interaction networks on which drug discovery is based. Uncharacterized deamidation can also lead to false positive clusters in proteomic interaction networks. This is an area of current scientific interest due to its influence on the identification of novel drug targets.25–30 The fragment crystallizable (Fc) region, which may be designed to have active effector functions, may also exhibit limited antibody-dependent cell-mediated cytotoxicity (ADCC) due to deamidation.31–33 Such is the case for an immunoglobulin G1 (IgG1) antibody which lost more than half of its ADCC activity after Fc deamidation disrupted the binding between the IgG1 Fc and the Fcγ receptor IIIa. 31
Herein, we present a comparative assessment of ASN and GLN deamidation in eight clinical-stage monoclonal antibodies (mAbs) (Table I) using a novel platform incorporating quantum cascade laser microscopy (QCLM) within the mid-infrared (mid-IR) spectral region, innovative slide cell array and multiple correlation algorithms: two-dimensional correlation spectroscopy (2D-COS),34–36 two-dimensional co-distribution spectroscopy (2D-CDS),37,38 and two-trace two-dimensional (2T2D) correlation spectroscopy39,40 for the analysis of proteins. 41 Detailed information is provided in the Supplemental Material. This original approach involving the use of 2D-COS monitors changes in amino acid composition due to the deamidation process in real-time, providing a comprehensive body of evidence to understand the impact of deamidation. The asynchronous plots are used to determine correlations between out-of-phase peak intensity changes that detail the two-step deamidation process described above, where the loss of the ASN or GLN amide within the side chain is the first step, and the generation of the carboxylate side chain is the second. 41
Table I.
Summary of background information for the clinical antibodies evaluated for extent of deamidation, loss of stability and aggregation.
| INN | Drug bank accession | Indication | Mechanism of action | Type | CDR | Antigen | Effector function | Scaffold originator | Scaffold current project |
|---|---|---|---|---|---|---|---|---|---|
| Gemtuzumab | DB00056 | Cancer: CD33 positive acute myeloid leukemia | Combination therapy with induction chemotherapy. Once, gemtuzumab binds CD33, the complex is internalized and the calicheamicin derivative is released binding the minor groove of double stranded DNA ultimately resulting in cell death. | ADC | ✓ | CD33 | — | IgG4 kappa | IgG4 |
| Pembrolizumab | DB09037 | Cancer: metastatic non-small cell lung carcinoma, head and neck squamous carcinoma, Hodgkin lymphoma, etc. | Direct binding to the cell surface receptor PD-1 and antagonizes its interaction with its known ligand PD-L1 and PD-L2. | mAb | ✓ | PD-1 receptor | — | IgG4 kappa | IgG4 |
| Evolocumab | DB09303 | Cardiovascular: LDL cholesterol reducing therapy | PCS9, inhibitor antibody | mAb | ✓ | PCS9 | — | IgG2 | IgG4 |
| Briakinumab | DB05459 | Autoimmune: T-cell driven autoimmune diseases | Binds and neutralizes p40 subunit found in both IL-12 and IL-23. As an inhibitor to their respective receptors and the subsequent downstream signaling. | mAb | ✓ | IL-12 | — | IgG1 kappa | IgG4 |
| Atezolimumab | DB11595 | Cancer: non-small lung cancer, small lung cancer, melanoma, etc. | PD-L1 inhibitor antibody to block PD-1 interaction | mAb | ✓ | PD-L1 | — | IgG1 | IgG4 |
| Nivolumab | DB09035 | Cancer: non-small lung cancer, melanoma, renal cell cancer, head and neck cancer, Hodgkin lymphoma, etc. | PD-1 blocking antibody | mAb | ✓ | PD-1 receptor | — | IgG4 | IgG4 |
| Trastuzumab | DB00072 | Cancer: Her2 positive breast, esophageal and gastric cancer, etc. | Binds to the Her2 receptor within the extracellular ligand binding domain blocking the cleavage of the extracellular domain to induce its receptor down-modulation and subsequently inhibit Her2-mediated intracellular cascades. Also, the Fc region mediates the activation of ADCC. | mAb | ✓ | Her2 receptor | ✓ |
IgG1 | IgG4 |
| Ustekinumab | DB05679 | Autoimmune: inhibitory conditions, Crohn's, Psoriasis, Psoriatic arthritis, etc. | The Fab binds the D1 domain of the p40 subunit of IL-12 and IL-23 in a 1:1 mol ratio. Ustekinumab only binds to IL-12 and IL-23 when unbound to the receptor. | mAb | ✓ | IL-12, IL-23 | — | IgG1 kappa | IgG4 |
INN: international nonproprietary name; CDR: complementarity-determining region; PD-1/PD-L1: programmed cell death protein; 1: ligand; Ig: immunoglobulin; IL: interleukin; mAb: monoclonal antibody; Fab: fragment antigen-binding; Her2: human epidermal growth factor 2; LDL: low-density lipoprotein; PCS9: proprotein convertase subtilisin Kexin type 9.
In general, a wide-field quantum cascade laser (QCL) transmission microscope is fit for studying intact biotherapeutics in their formulation condition, in that hyperspectral images are acquired in real-time, >250 times faster, collecting 16× more spectral data per pixel and as a result providing an enhanced signal-to-noise ratio when compared to a Fourier transform IR microscope. In addition, the intensity of the QCLs is greater than the synchrotron providing a plethora of photons reducing the need for a liquid-nitrogen-cooled mercury cadmium telluride detector, and instead a linear response room temperature focal plane array is used. In short, the QCLM hardware (ProteinMentor) extends these advantages by allowing for the real-time monitoring of an array of clinical antibodies (1 μL fixed volume per sample) under accurate thermal control. The technology has been quality by design for the biopharma industry to ensure a true multi-attribute platform that delivers on speed to market.
Our streamlined approach has other compelling advantages as well: (i) full-length protein is analyzed in its formulation condition; (ii) an innovative slide cell that allows for the comparative analysis of an array of proteins in solution; (iii) the ability to acquire hyperspectral images across a range of temperatures, allowing for the stepwise monitoring of the deamidation process; and (iv) the subsequent QCL IR microscopy difference spectra and two-dimensional (2D) correlation algorithms allow for a true multi-attribute assessment of an array of therapeutic proteins. Furthermore, the approach offers a greater understanding of the effect of ASN and GLN deamidation on the stability of therapeutic proteins, including at times the observation of protein aggregation.
Materials and Methods
Protein Engineering
All clinical antibodies (gemtuzumab, pembrolizumab, evolocumab, briakinumab, atezolizumab, nivolumab, trastuzumab, and ustekinumab; Table I) used in this study were engineered using human (Hs) IgG4 isotype mAbs, with the S228P heavy chain (HC) hinge-stabilizing mutation, regardless of the original drug isotype, with variable regions matching published sequences.42–44 The corresponding antibodies were produced by transient transfection of HC and light chain-containing plasmids into suspension-adapted HEK-293 cells (ATUM.bio). A cell culture supernatant was purified by protein A affinity chromatography (MabSelect SuRe pcc, GE Life Sciences) according to the manufacturer's protocol. Purified antibodies were formulated in phosphate-buffered saline at pH 7.4 and quantified by optical density at λmax = 280 nm using extinction coefficients calculated from the primary amino acid sequence.
Quantum Cascade Laser Microspectroscopy
The full-length clinical antibodies analyzed were in one of two conditions: (i) control, phosphate-buffered saline solution at pH 7.4 and stored at −80 °C until analysis and (ii) forced degraded, where the clinical antibody sample was formulated in 20 mM Tris (Millipore Sigma), 10 mM ethylenediaminetetraacetic acid (EDTA; Millipore Sigma) at pH 8.5, and stressed at high pH 8.5 at 40 °C for a one-week period, then stored at −80 °C until analysis. No further sample preparation was required. A 1 μL aliquot of each clinical antibody sample (control or forced degraded condition) was loaded onto a predefined well within the innovative CaF2 slide cell array and placed in the heated slide cell accessory with thermal control (±0.5 °C). Real-time hyperspectral images are acquired at different set temperatures within the range of 28–64 °C with 6 °C temperature intervals and 4 min equilibration periods. The resulting QCL IR spectra for the clinical antibodies were within the spectral region of 1785–1425 cm–1 at a spectral resolution of 4 cm–1. The QCL transmission microscope model ProteinMentor from Protein Dynamic Solutions, Inc. is comprised of broadly tunable external cavity QCLs, a four time low mag objective with a numerical aperture of 0.3, a 2 × 2 mm2 field of view providing a pixel size of 4.25 × 4.25 μm2, a linear response microbolometer focal plane array (480 × 480 pixels) camera, a heated slide cell accessory, and dedicated software for hyperspectral image and spectral data analysis (Supplemental Material). This platform provides significant advantages: (i) a series of real-time hyperspectral images of the sample in the well at the set temperature, evaluated under the exact same temperature conditions in H2O, (ii) enhanced signal-to-noise ratio within the entire spectral region of interest (1800–1400 cm–1), which includes both the amide I and II bands, and (iii) comparative assessment of the full-length clinical antibodies in their formulation conditions with amino acid-level resolution.45–52
Hyperspectral Image and QCL Microspectroscopy Spectral Data Analysis
The hyperspectral images were evaluated using the DataPDS, Correlation Dynamic and AmiCQ software suite from Protein Dynamic Solutions, Inc. QCLM spectral baseline correction, difference and weighted spectral calculations, overlays, 2D-CDS, 2D-COS, and 2T2D plots were generated using Correlation Dynamics software. Details of the 2D correlation algorithms34–40,53 and spectral analysis for the deamidation determination were initially described. 41
Cation Exchange Chromatography
Our platform technology identified the clinical antibodies having the highest deamidation and aggregation risk. We then proceeded to evaluate a selection of the clinical antibodies based on their availability (pembrolizumab, evolocumab, briakinumab, and trastuzumab) under control and forced degraded conditions using cation exchange (CEx) chromatography to verify the results obtained by the platform technology. These selected samples were buffer exchanged using 20 mM 2-morpholinoethanesulfonic acid (MES), pH 6.2, and ∼50 μg of the sample was loaded onto a Dinoex ProPac WCX-10 column with mobile phase A as 20 mM MES, pH 6.2, and mobile phase B as 20 mM MES, 300 mM NaCl, pH 6.2. For the selected samples (pembrolizumab, evolocumab, and trastuzumab), a gradient (0–68.2% B) was delivered from 2 to 60 min at a flow rate of 1.0 mL/min. However, for the briakinumab sample, a gradient (15–100% B) was delivered from 2 to 75 min at an identical flow rate. The ultraviolet absorbance was monitored at λmax = 280 nm.
Results and Discussion
This novel comparative assessment approach was achieved for the eight clinical antibodies that were generated using human IgG4 as the scaffold.8,42–44 Each antibody was subject to control conditions, defined as phosphate-buffered saline at pH 7.4 and storage at −80 °C until use, and forced degraded conditions, defined as 20 mM Tris, 10 mM EDTA at pH 8.5 with incubation at 40 °C for 1 week and then storage at −80 °C until use, following procedures established by Lu et al. 54 We also verified the concentration of these samples to ensure comparability (Table S1 and Figures S1 and S2, Supplemental Material).
Collectively the Comparative Assessment Provided a Comprehensive Body of Evidence
All hyperspectral images were clear and featureless at a 4.3 μm spatial resolution (Figure S3, Supplemental Material) for (a) gemtuzumab, (b) pembrolizumab, (c) evolocumab, (d) briakinumab, (e) atezolizumab, (f) nivolumab, (g) trastuzumab, and (h) ustekinumab. Furthermore, these hyperspectral images were comprised of 230,400 QCL spectra enabling a comparison of the control and forced degraded. Baseline corrected QCL spectral data showed they were very similar, and almost no differences could be perceived within the temperature range between 28 and 64 °C (Figure 2a–2h). The quality of the spectral data obtained is clearly demonstrated, yet very little difference can be discerned. Difference spectra were generated by subtracting the first spectrum at the initial temperature from all subsequent spectra, which were then subject to both 2D-COS and co-distribution analysis. 41 The determination of ASN or GLN deamidation per secondary structure and overall extent of deamidation resulted from the direct analysis of the 2D-COS asynchronous plots (Figure 3a–3h). This evaluation also included the multi-attribute determination of the effect of deamidation using both 2D-COS and co-distribution analysis to ascertain decreased stability under both control and forced degraded conditions for each clinical antibody. The 2D correlation algorithms used to enhance the spectral resolution and enable the correlation between spectral peaks of interest are described in the Supplemental Material.34,41,53 Our results provided an unprecedented level of sensitivity and understanding, revealing both the site of deamidation and the effect that deamidation had on the molecular stability of the clinical antibody (Figure 3a–3h), including at times aggregation (Figure 4a–4h) and the potential for loss of efficacy due to the introduction of a negative charge within the CDR. This comparative multi-attribute approach has provided a comprehensive understanding of several CQAs which will be discussed below for each clinical antibody within the array. Furthermore, once the deamidation event was initiated, it continued during the temperature ramp, enabling the real-time monitoring of the amino acids associated with the ASN and GLN deamidation process for the determination of the extent and site of deamidation for the control and forced degraded samples (Tables II and III).
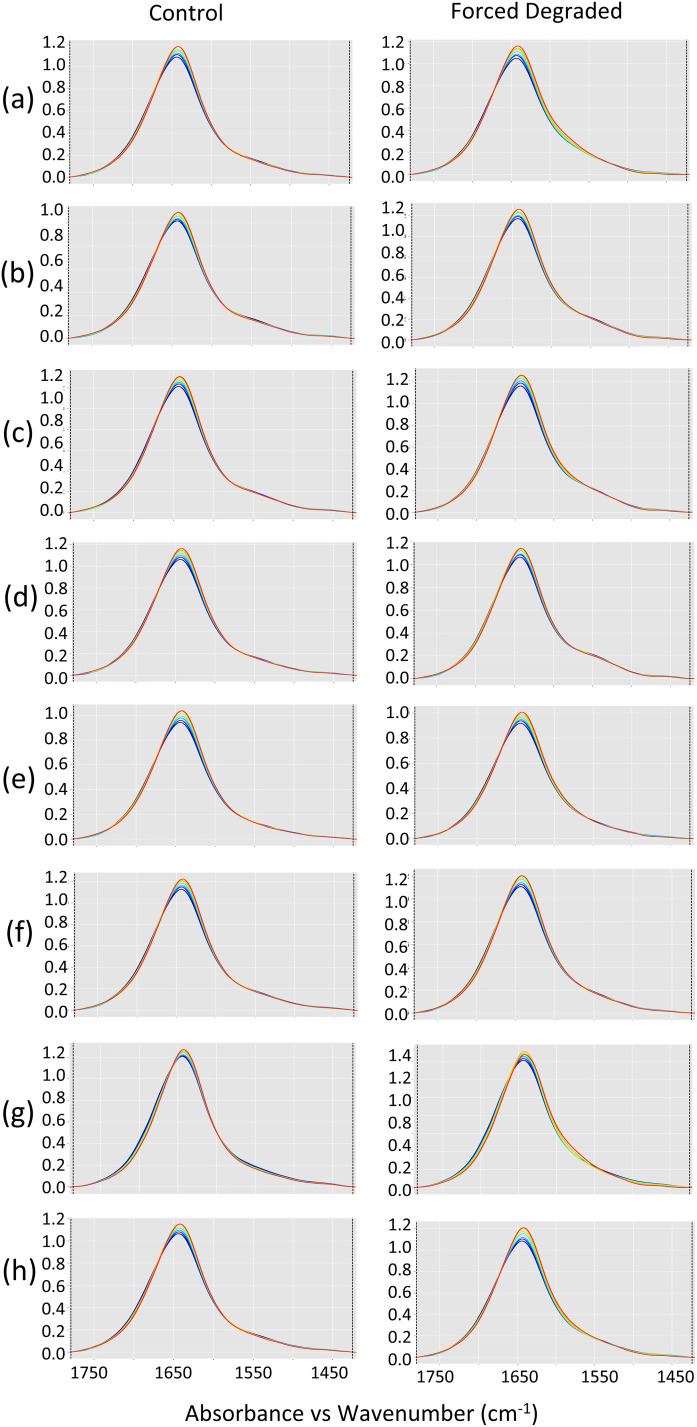
Figure 2.
Quantum cascade laser (QCL) microspectroscopy spectral overlay for all clinical antibody samples in the array within the spectral region of 1785–1425 cm–1 and during the temperature range of 28–64 °C with 6 °C temperature intervals. The columns are for the control and forced degraded samples and the rows (a) gemtuzumab, (b) pembrolizumab, (c) evolocumab, (d) briakinumab, (e) atezolizumab, (f) nivolumab, (g) trastuzumab, and (h) ustekinumab. Both the quality and consistency of the spectral data can be observed.
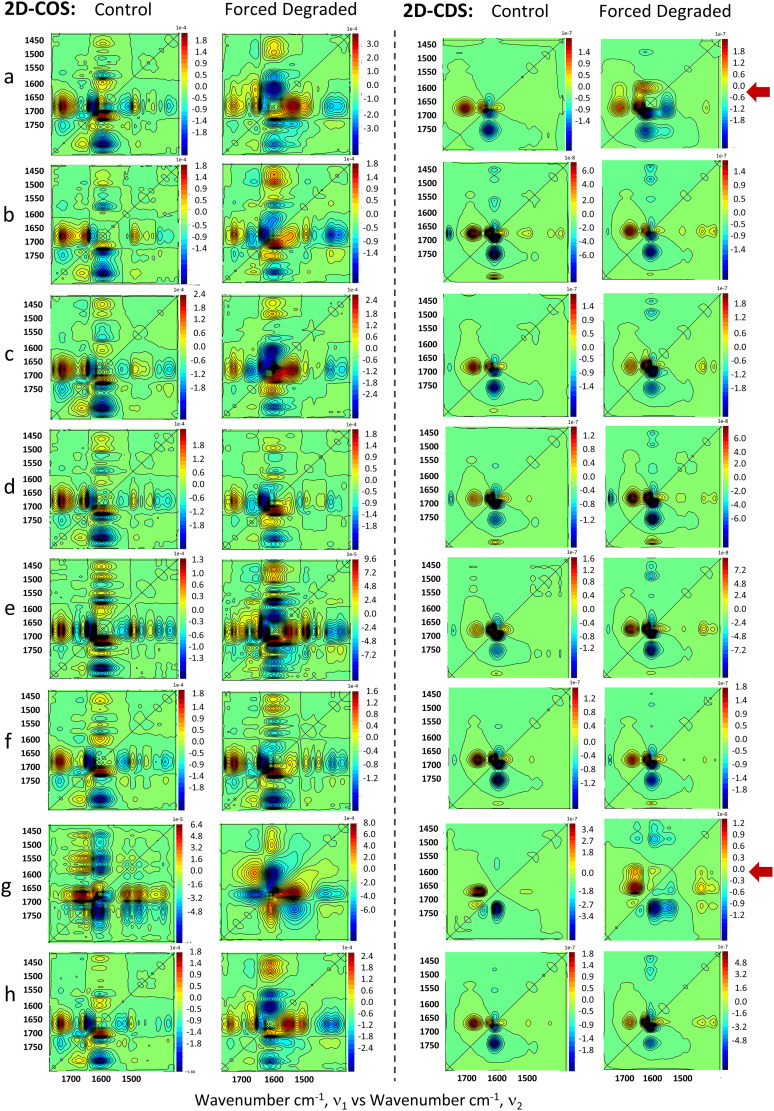
Figure 3.
Breakthrough IgG4 clinical antibody array comparative assessment during the real-time monitoring of ASN and GLN deamidation within the temperature range of 28–64 °C, that at times led to aggregation. The analysis involved the use of 2D correlation and co-distribution spectroscopies for: (a) gemtuzumab, (b) pembrolizumab, (c) evolocumab, (d) briakinumab, (e) atezolizumab, (f) nivolumab, (g) trastuzumab, and (h) ustekinumab. 2D-COS asynchronous plots are especially suited to the determination of the relationship of deamidation, aggregation and stability of the clinical antibodies. Adding further understanding is the co-distribution analysis, which allows the evaluation of the distribution of proteins in solution within the sample and defines if the event of deamidation or the deamidation that led to aggregation is a concern for the majority of the clinical antibodies within the sample analyzed. Evidence of aggregation–deamidation is highlighted with the red arrow and was observed in the forced degraded samples of (a) gemtuzumab and (g) trastuzumab compared to their respective controls.
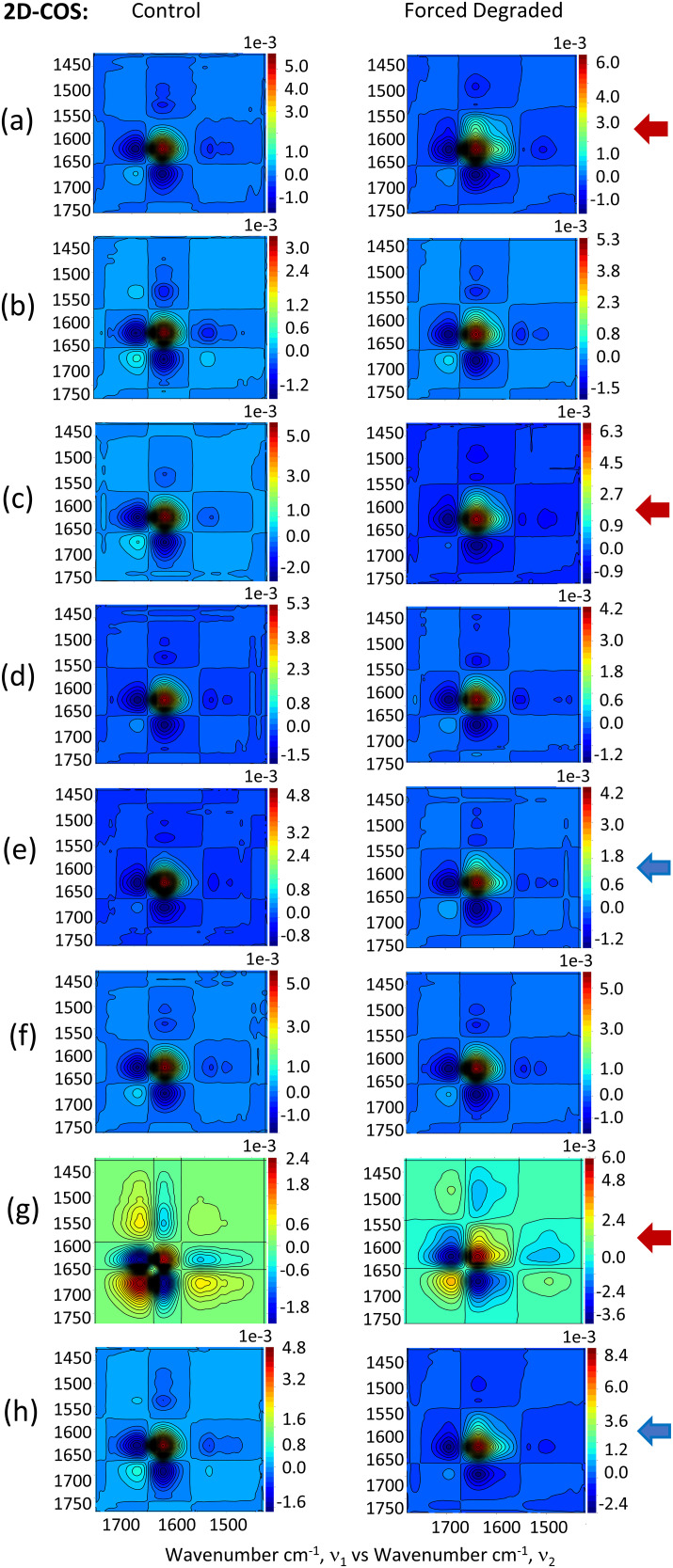
Figure 4.
Two-dimensional correlation spectroscopy (2D-COS) synchronous plots for all clinical antibodies in the array within the spectral region of 1785–1425 cm–1 and the temperature range of 28–64 °C. The columns are for the control and forced degraded samples and the rows: (a) gemtuzumab, (b) pembrolizumab, (c) evolocumab, (d) briakinumab, (e) atezolizumab, (f) nivolumab, (g) trastuzumab, and (h) ustekinumab. An auto peak characteristic of aggregation at 1614 cm–1 (red arrow) is observed for the forced degraded sample assigned to: (a) gemtuzumab, (c) evolocumab, and (g) trastuzumab. Furthermore, self-association events were more subtle auto peak features on the synchronous plots (blue arrow) also observed for the forced degraded samples: (e) atezolizumab and (h) ustekinumab.
Table II.
Summary of the extent of deamidation for all 8 clinical-stage antibodies under control conditions at pH 7.4.
| β-turn | β-turn | Hinge loop | α-helix | β-sheet | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1696 cm-1 | 1681 cm-1 | 1663 cm-1 | 1653 cm-1 | 1636 cm-1 | Overall | ||||||||
| aa a | Value b | Error | Value | Error | Value | Error | Value | Error | Value | Error | Extent c | Error d | Risk e |
| Gemtuzumab | |||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.049 | ±0.002 | 0.999 | ±0.033 | 0.682 | ±0.019 | 1.730 | ± 0.038 | Yes |
| Q | 0.091 | ±0.007 | 0.107 | ±0.006 | 0.737 | ±0.085 | 0.269 | ±0.019 | 0.223 | ±0.015 | 1.427 | ± 0.089 | Yes |
| Pembrolizumab | |||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.063 | ±0.002 | 0.532 | ±0.016 | 1.136 | ±0.047 | 1.731 | ± 0.050 | Yes |
| Q | 0.281 | ±0.012 | 0.249 | ±0.011 | 1.141 | ±0.085 | 0.244 | ±0.025 | 0.145 | ±0.022 | 2.060 | ± 0.093 | Yes |
| Evolocumab | |||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.075 | ±0.002 | 0.735 | ±0.030 | 1.114 | ±0.086 | 1.924 | ± 0.091 | Yes |
| Q | 0.078 | ±0.008 | 0.080 | ±0.007 | 0.638 | ±0.042 | 0.145 | ±0.012 | 0.222 | ±0.016 | 1.163 | ± 0.048 | Yes |
| Briakinumab | |||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.042 | ±0.001 | 0.719 | ±0.025 | 0.833 | ±0.026 | 1.594 | ± 0.036 | Yes |
| Q | 0.079 | ±0.006 | 0.077 | ±0.007 | 1.562 | ±0.156 | 0.158 | ±0.008 | 0.221 | ±0.009 | 2.097 | ± 0.157 | Yes |
| Atezolizumab | |||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.047 | ±0.003 | 0.716 | ±0.043 | 1.275 | ±0.123 | 2.038 | ± 0.130 | Yes |
| Q | 0.041 | ±0.009 | 0.063 | ±0.009 | 1.075 | ±0.207 | 0.304 | ±0.030 | 0.262 | ±0.024 | 1.745 | ± 0.269 | Yes |
| Nivolumab | |||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.054 | ±0.001 | 0.434 | ±0.010 | 2.196 | ±0.109 | 3.408 | ± 0.109 | Yes |
| Q | 0.028 | ±0.011 | 0.050 | ±0.010 | 1.806 | ±0.266 | 0.247 | ±0.011 | 0.182 | ±0.009 | 2.450 | ± 0.269 | Yes |
| Trastuzumab | |||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ± 0.000 | No |
| Q | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ± 0.000 | No |
| Ustekinumab | |||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.017 | ±0.002 | 0.483 | ±0.017 | 0.847 | ±0.060 | 1.347 | ± 0.062 | Yes |
| Q | 0.114 | ±0.014 | 0.127 | ±0.016 | 0.915 | ±0.048 | 0.185 | ±0.017 | 0.159 | ±0.014 | 1.500 | ± 0.057 | Yes |
a Amino acids: N, asparagine and Q, glutamine; b The value represents the number of asparagine or glutamine residues that deamidated during the thermal perturbation of the sample in the slide cell array; c Overall extent of deamidation is the sum of the values obtained for each secondary structure; d Sum error propagation for all the secondary structures; e Risk of deamidation.
Table III.
Summary of the extent of deamidation for all eight clinical-stage antibodies under forced degraded conditions pH 8.5.
| β-turn | β-turn | Hinge loop | α-helix | β-sheet | Overall extentc |
Errord | Riske | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1696 cm–1 | 1681 cm–1 | 1663 cm–1 | 1653 cm–1 | 1636 cm–1 | ||||||||||
| aaa | Valueb | Error | Value | Error | Value | Error | Value | Error | Value | Error | ||||
| Gemtuzumab | ||||||||||||||
| N | 1.627 | ±0.490 | 0.597 | ±0.091 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 2.224 | ±0.498 | Yes | |
| Q | 0.302 | ±0.022 | 0.266 | ±0.021 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.568 | ±0.030 | Yes | |
| Pembrolizumab | ||||||||||||||
| N | 0.016 | ±0.003 | 0.033 | ±0.007 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.049 | ±0.008 | No | |
| Q | 1.819 | ±0.204 | 1.357 | ±0.156 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 3.176 | ±0.259 | Yes | |
| Evolocumab | ||||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | No | |
| Q | 0.152 | ±0.028 | 0.030 | ±0.008 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.182 | ±0.029 | No | |
| Briakinumab | ||||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.032 | ±0.003 | 0.135 | ±0.007 | 1.583 | ±0.246 | 1.750 | ±0.246 | Yes | |
| Q | 0.263 | ±0.027 | 0.133 | ±0.016 | 0.664 | ±0.024 | 2.657 | ±0.474 | 1.204 | ±0.174 | 4.921 | ±0.506 | Yes | |
| Atezolizumab | ||||||||||||||
| N | 0.062 | ±0.008 | 0.306 | ±0.079 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.368 | ±0.079 | No | |
| Q | 1.089 | ±0.088 | 0.512 | ±0.043 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 1.601 | ±0.098 | Yes | |
| Nivolumab | ||||||||||||||
| N | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.057 | ±0.010 | 1.369 | ±0.788 | 1.426 | ±0.788 | Yes | |
| Q | 0.174 | ±0.031 | 0.175 | ±0.025 | 0.121 | ±0.012 | 0.527 | ±0.065 | 0.443 | ±0.090 | 1.440 | ±0.119 | Yes | |
| Trastuzumab | ||||||||||||||
| N | 0.894 | ±0.024 | 0.943 | ±0.030 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 1.837 | ±0.038 | Yes | |
| Q | 0.295 | ±0.010 | 0.299 | ±0.010 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.594 | ±0.014 | Yes | |
| Ustekinumab | ||||||||||||||
| N | 1.323 | ±0.211 | 0.500 | ±0.078 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 1.820 | ±0.225 | Yes | |
| Q | 0.412 | ±0.014 | 0.365 | ±0.021 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.000 | ±0.000 | 0.777 | ±0.025 | Yes | |
aAmino acids: N, asparagine; Q, glutamine.
bThe value represents the number of N or Q residues that deamidated during the thermal perturbation of the sample in the slide cell array.
cOverall extent of deamidation is the sum of the number of N or Q residues that deamidated (values) obtained for each secondary structure.
dSum error propagation for all the values determined for each secondary structure.
eRisk of deamidation.
The streamlined deamidation approach takes advantage of the high sensitivity and selectivity of IR in combination with the 2D correlations involving the key signature peaks identified for the deamidation mechanism (Figure 1). Specifically, the analysis uses a 2D-COS asynchronous plot and the sequential order of molecular events to determine the site of deamidation.41,45–47 Briefly, the method relies on four factors: (i) the number of ASN and GLN residues that deamidated during the thermal perturbation (28–64 °C); (ii) the secondary structure location to which the deamidated ASN (N) or GLN (Q) residues were determined through the correlations established using the 2D-COS asynchronous plot; (iii) the perturbation of neighboring amino acids within the sequence of the antibody due to the introduction of the negatively charged residue (aspartate or glutamate) as observed from the sequential order of molecular events. Whereby, 10 of the 20 amino acids and the disruption of their weak interactions are monitored directly and can be correlated with the deamidated residues, thus allowing us to identify the amino acids within the site of deamidation, and (iv) the amino acid sequence to map the site of deamidation. Only the site that satisfies all four of the factors (criteria) mentioned above is determined to be the site of deamidation. If there is available high-resolution structure information, we model the antibody to confirm the aqueous solvent accessibility of the site of deamidation. More importantly, the clinical antibody is evaluated intact in its formulation condition, unlike other labor-intensive methods such as peptide mapping which require extensive sample preparation including the use of dithiothreitol or β-mercaptoethanol to reduce the disulfide bridges followed by enzymatic digestion which then requires separation of the resulting fragments by liquid chromatography. Also, the detection is done in the gas phase using mass spectrometry to identify the isoform of the iso-aspartate or iso-glutamate and the site is determined by the coverage of the sequence against the known sequence of the antibody. On many occasions, the peptide mapping assay is limited to the fragment antigen-binding (Fab) domain. More importantly, information regarding the impact deamidation may have on the secondary structure, aggregation potential or instability cannot be assessed using peptide mapping. Consequently, our streamline deamidation routine is a transformative comparative method and a true multi-attribute approach that provides an unprecedented understanding of the extent, site, and impact of deamidation on the clinical antibody.
Deamidation Results for the Clinical Antibodies Re-Engineered Using an IgG4 Scaffold
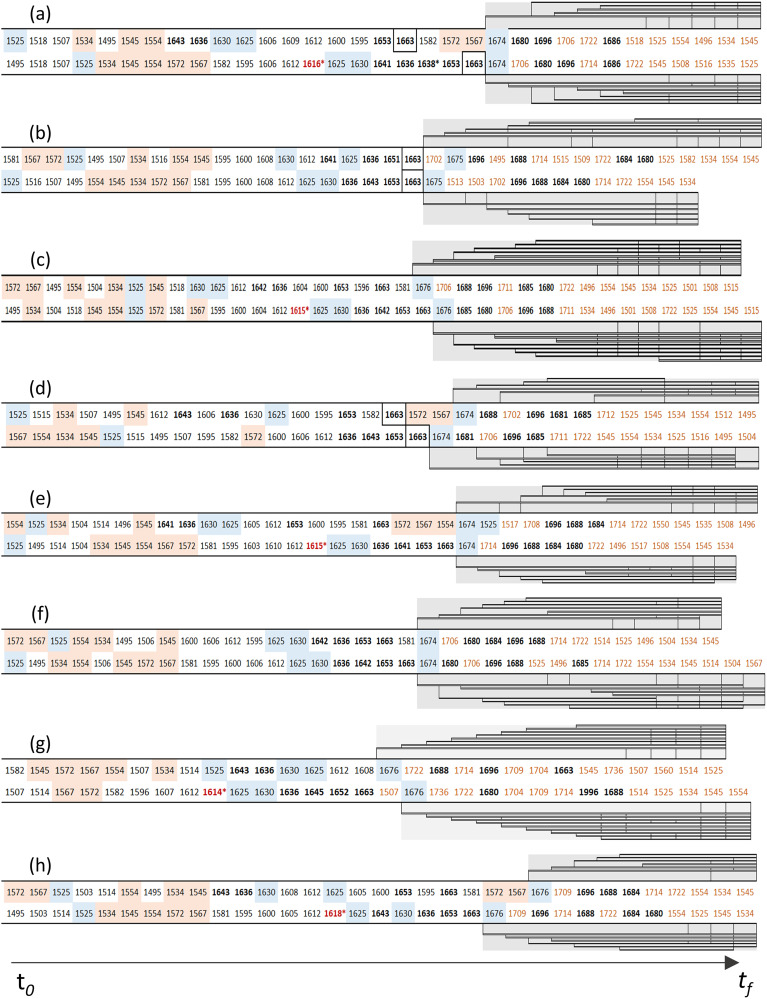
In general, the comprehensive analysis of 2D-COS asynchronous plots led to the determination of the extent and site of ASN and GLN deamidation for both the control and forced degraded samples (Tables II and III and Figure 3). The information contained in Tables II and III for the control and forced degraded samples, respectively, also contains the determination of the overall extent of deamidation for each clinical antibody sample. These reults allowed for comparison with other orthogonal techniques. The 2D-COS asynchronous plot is comprised of cross peaks that undergo out-of-phase or sequential intensity changes, allowing for the analysis of the signature cross peaks involved in the deamidation process. Specifically, the loss of the ASN side chain amide (1612 cm–1), followed by the generation of the deamidation product being the new carboxylate mode (1567 or 1572 cm–1) associated with the new aspartate. Similarly, for the GLN, the loss of the side chain amide (1581 cm–1) is followed by the generation of the deamidation product observed at (1545 or 1554 cm–1) associated with the new glutamate. In addition, we can ascertain the location of the specific ASN and GLN residues that underwent the deamidation process through its correlation with the specific peptide bond vibrational mode assigned to a secondary structure type. If one also considers that the event of deamidation causes perturbation of neighboring residues, then the sequential order of molecular events allows for the identification of the neighboring residues (Figure 5a–5h). This approach becomes a powerful combination associated with the deamidation process and allows for the determination of the extent and site of ASN or GLN deamidation.
Figure 5.
Sequential order of molecular events for all eight clinical antibodies during the temperature perturbation of 28–64 °C in the array: (a) gemtuzumab, (b) pembrolizumab, (c) evolocumab, (d) briakinumab, (e) atezolizumab, (f) nivolumab, (g) trastuzumab, and (h) ustekinumab. In each case, the sequential order of molecular events corresponds to: (top row) control and (bottom row) forced degraded samples.
For the Hs gemtuzumab, the deamidation results are summarized in Tables II and III and Figure 3a, and the sequential order of molecular events is in Figure 5a. Under the control conditions, the ASN site of deamidation was determined to be at HC N207 within the constant Fab region and HC N429 within the constant Fc region. The gemtuzumab site of GLN deamidation was at a light chain (LC) CDR3 Q93/94 for both sample conditions (control and forced degraded), suggesting a loss of efficacy. In addition, for the forced degraded sample, the ASN deamidation sites determined were HC CDR2 N55, which has the potential to induce loss of efficacy, and HC N207 within the constant Fab region. Furthermore, the combined effect of both GLN and ASN deamidation within the CDR regions induced the aggregation of the forced degraded clinical sample at pH 8.5 by destabilizing the β-sheets for both the HC and LC variable Fab regions, resulting in the potential for immunogenicity risk. Finally, our ASN deamidation results agree with Lu et al., 54 for Hs gemtuzumab at pH 8.5. Also, ASN deamidation for the forced degraded sample led to aggregation of the antibody, as observed from the 2D-COS synchronous and asynchronous plots and summarized in the sequential order of molecular events (Figures 2a, 4a, and 5a, respectively).
For the Hs pembrolizumab, the comparative analysis led to the determination of ASN and GLN deamidation for both the control and forced degraded samples. Deamidation impact was based on the available amino acid sequence and structural information for this clinical antibody (PDB ID 5DK3, shown in the Graphical Abstract), 44 and the extent of deamidation was defined by the 2D-correlation analysis shown in Figures 3b, 4b, and 5b. Under control conditions, deamidation was observed for both ASN and GLN summarized in Table II. The deamidation sites were HC N211 within the helical segment, HC N434 within the constant Fc region and the GLN sites were HC Q112 within the variable Fab region and LC CDR3 Q93, suggesting risk for loss of efficacy. Under forced degradation conditions, deamidation (Table III) was observed only for the GLN residues within the β-turns (1696 and 1681 cm–1), and the sites of deamidation were LC CDR3 Q93, HC Q112 within the variable Fab region in close proximity to the HC CDR3, and HC Q178 within the constant Fab region. A continuation of the GLN deamidation was observed for the forced degraded sample within the second dimer for the same GLN deamidation sites observed for the control sample due to its slower rate of deamidation, confirming the potential risk for loss of efficacy. One additional GLN site was also observed for the forced degraded sample when compared to the control. In addition, no aggregation or self-association was observed for pembrolizumab under either the control or the forced degraded conditions (Figure 3b). The extent of deamidation was also validated by analytical high-performance liquid chromatography (HPLC) CEx, data generated by Merrigen (Figure 4, Supplemental Material) for the same samples. We also performed 2T2D correlation analysis to determine the percent extent of deamidation of the forced degraded sample against the control and obtained comparable results that could directly be compared with the HPLC CEx summarized in Table IV.
Table IV.
Summary of QCLM and HPLC CEx chromatography results.
| mAb | % Extent of deamidation | |
|---|---|---|
| QCLMa | HPLC CExb | |
| Pembrolizumab IgG4 | 62.8 | 56.2 |
| Evolocumab IgG4 | 76.2 | 72.4 |
| Briakinumab IgG4 | 11.4 | 15.3* |
| Trastuzumab IgG4 | 92.4 | 89.5 |
CEx: cation exchange; HPLC: high-performance liquid chromatography; QCLM: quantum cascade laser microspectroscopy.
aReference and samples were evaluated in their original buffer conditions.
bReference and samples were five-fold diluted in 20 mM MES at pH 6.2 prior to injection.
*Higher salt gradient was used to try to elute and evaluate the acidic peaks.
For Hs evolocumab samples, the extent of ASN and GLN deamidation (Tables II and III and Figure 3c) and the sequential order of molecular events (Figure 5c) were used in the determination of the site of deamidation. Evolocumab ASN deamidation was observed at HC CDR2 N55 or N57 within the variable Fab region under control conditions consistent with Lu et al. 54 In addition, GLN deamidation was observed for HC Q65. Furthermore, evolocumab in its forced degraded form did not exhibit a continued risk of deamidation during the thermal perturbation, suggesting all the available ASN and GLN residues for deamidation had indeed deamidated during the period of incubation, which led to the aggregation of the forced degraded sample. Another possibility for the Hs evolocumab forced degraded sample was that the observed aggregation resulted in less solvent accessibility and thus limited continued deamidation. The results obtained were consistent with HPLC CEx data for the same control and forced degraded samples shown in Figure S5 (Supplemental Material; data generated by Merrigen). The forced degraded sample chromatogram was characteristic of acidic charge variants, and no residual main peak was observed. In addition, we performed 2T2D correlation analysis to determine the percent extent of deamidation and obtained results that were comparable tothe HPLC CEx summarized in Table IV.
For Hs briakinumab samples, the ASN and GLN deamidation quantitative results are summarized in Tables II and III and Figure 3d resulting in the determination of the sequential order of molecular events as shown in Figure 5d. GLN deamidation occurred within the CDR, potentially impacting efficacy under both control and forced degraded conditions. Briakinumab was observed to undergo both GLN and ASN deamidation in real time, with the forced degraded sample at pH 8.5 exhibiting a greater extent of deamidation. Both clinical antibody samples exhibited HC CDR3 Q107 and LC CDR3 Q90 deamidation within the variable Fab region, potentially increasing the risk of loss of target specificity/affinity and therefore loss of efficacy. The forced degraded sample also exhibited GLN deamidation at HC Q173 within the constant Fab region and LC Q38/39 within the variable Fab region. Briakinumab ASN deamidation sites under controlled conditions were determined at HC N84 within the Fab variable region and HC N310 within the constant Fc region, while under the forced degraded conditions the sites for ASN deamidation were HC N429 near the C-terminal end and LC CDR2 N32. The second site of ASN deamidation also impacts target affinity. We have also confirmed events of deamidation by HPLC CEx chromatography (Figure S6, Supplemental Material), and we performed 2T2D correlation analysis to determine the extent of deamidation, which produced results comparable to HPLC CEx results summarized in Table IV.
For Hs atezolizumab, ASN and GLN deamidation results (Tables II and III and Figure 3e) and the sequential order of molecular events (Figure 5e) for both the control and forced degraded samples showed the CDRs are not at risk for deamidation. For the control sample, ASN deamidation was determined at HC N209 within the constant Fab region. GLN deamidation for the control sample was at LC Q199 within the constant Fab region. Meanwhile, for the forced degraded sample, no ASN deamidation was observed. GLN deamidation was determined at LC Q155 within the constant Fab region. Also, the forced degraded sample at pH 8.5 underwent self-association, which perturbed hydrogen bonding interactions involving the positively charged residues (lysine and arginine) as observed from the sequential order of molecular events.
For Hs nivolumab, the control sample was observed to deamidate (Table II) and (Figures 3f and 5f) within multiple sites: HC N204 within the constant Fab region and HC N308 and N382 within the constant Fc region of the antibody. GLN deamidation was also observed for the control sample at HC CDR3 Q105 and LC Q100 within the variable Fab region near LC CDR3, suggesting a potential loss of efficacy. The forced degraded sample (Table III) had a site for ASN deamidation at HC N427 and a GLN deamidation site at LC Q199 within the constant Fab region. Also, no aggregation was observed for both clinical antibody samples.
For Hs trastuzumab, the CDR did not exhibit risk for ASN or GLN deamidation (Tables II and III and Figures 3g and 5g). The Hs trastuzumab IgG4 isotype real-time assessment did not deamidate under the control conditions. For the forced degraded sample, an ASN deamidation site was determined at HC N434 within the constant Fc region. GLN deamidation risk was observed for LC Q199 within the constant Fab region. We re-examined the sample months later and ran multiple evaluations; the results were consistent with a propensity for ASN and GLN deamidation for the forced degraded sample (Figure 6). Furthermore, the forced degraded trastuzumab ASN deamidation led to the aggregation of the clinical antibody at pH 8.5. We have also confirmed events of deamidation by HPLC CEx chromatography (Figure S7, Supplemental Material), and we performed 2T2D correlation analysis to determine the percent extent of deamidation, summarized in Table IV.
Figure 6.
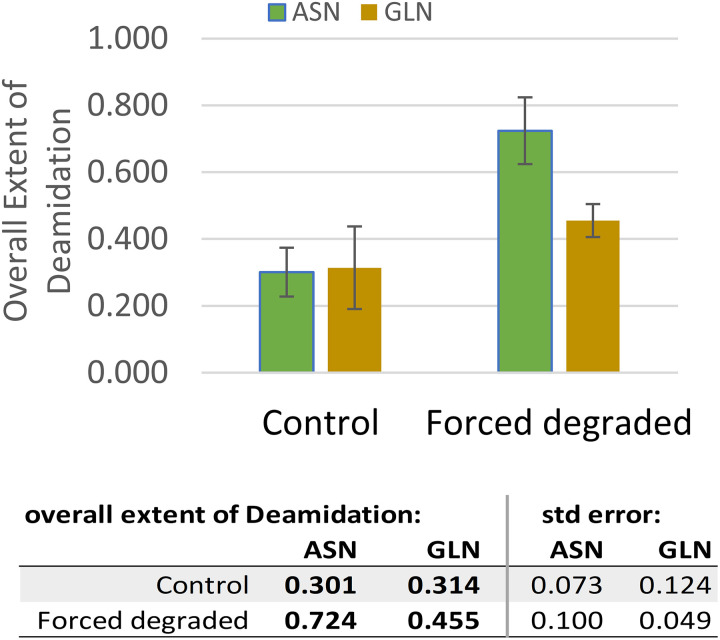
Bar graph summarizing the overall ASN and GLN extent of deamidation for the control and forced degraded trastuzumab samples after six months of storage at 2–8 °C. The results presented were for a series of five separate evaluations after the 2–8 °C storage period. Specifically, an average of five separate runs for triplicate samples resulting in a total of n = 30 determinations for which a statistically significant result was obtained. These results confirm our initial evaluation in Tables II and III, whereby the risk of deamidation for trastuzumab occurs after the sample is subject to a period of forced degradation.
Hs ustekinumab underwent both ASN and GLN deamidation (Tables II and III) and (Figures 3h and 5h). An ASN deamidation site was found to be common to both control and forced degraded samples: HC N210 within the constant Fab region. For the forced degraded sample, a second site was determined at HC N433 located within the constant Fc region. GLN deamidation for the control sample was determined to be at HC CDR3 Q104 within the variable Fab region, while for the forced degraded sample the GLN deamidation site at the LC Q199 was within the constant Fab region. The event of deamidation within the forced degraded sample at pH 8.5 led to self-association, which perturbed hydrophobic interactions involving tryptophan as observed in the co-distribution and 2D-COS analysis.
Confirmation of the Findings by CEx
Quantum cascade laser microspectroscopy was determined to be orthogonal to HPLC CEx (Table IV). The QCLM percent extent of deamidation values were within 4–6% when compared to HPLC CEx, with one advantage: all the samples were evaluated in a single array within 10 min. Furthermore, follow-on evaluations of trastuzumab using real-time monitoring of the deamidation process provided predictive results that can be used to evaluate the stability of the clinical-stage antibody.
Evidence of Deamidation Leading to Aggregation
An increased overall extent of deamidation was correlated to aggregation. For this, we also performed an asynchronous co-distribution (2D-CDS) analysis, 41 ,45–47 which allowed the assessment of aggregation within the population of clinical antibodies in each sample and verified via 2D-COS synchronous plot analysis. For gemtuzumab in solution under forced degraded conditions (Figures 3a and 4a), we observed aggregation, which was correlated to the ASN deamidation event, but no such aggregation was seen under control conditions. Similarly, for evolocumab (Figures 3c and 4c) and trastuzumab (Figures 3g and 4g), deamidation led to aggregation for the forced degraded sample. Furthermore, self-association was observed for forced degraded atezolizumab (Figure 4e) and ustekinumab (Figure 4h). Meanwhile, for pembrolizumab (Figures 3b and 4b), briakinumab (Figures 3d and 4d), and nivolumab (Figures 3f and 4f), peak patterns were similar for the control and forced degraded samples, and thus no aggregation was evident due to deamidation.
A novel multi-attribute method that provides a comprehensive real-time evaluation of an array of full-length clinical antibodies in solution was employed to determine the site and extent of ASN and GLN deamidation. These results were validated and proved to be orthogonal to HPLC CEx (herein) and independently by Lu et al. 54 using LC–MS. More importantly, our approach allowed for the investigation of the degradation brought about by thermal or high pH conditions for the assessment of the stability of the re-engineered IgG4 S228P clinical antibodies via the deamidation pattern and its effects on aggregation.
The method involves a streamlined evaluation of an array of clinical antibody samples by directly monitoring the event of deamidation within the full-length protein in its predetermined formulation conditions, requiring only 1 μL of sample per well. These clinical samples were assessed for deamidation, decreased stability, and aggregation in both control and forced degraded samples. Three key differentiators provided by this technology are (i) the comparative assessment of an array of full-length proteins in solution under controlled conditions; (ii) no additional sample preparation, thus effectively streamlining the analysis; and (iii) the highly sensitive and selective QCL IR spectral data analysis that provides an in-depth understanding of the molecular factors that contribute to safety and efficacy concerns further down the development pipeline.
There are significant advantages to the streamlined workflow that are not limited to sample size or formulation conditions.41,45–47 Directly monitoring the deamidation process in real time allows for the determination of loss of stability that may lead to aggregation. This empirical approach provides an unprecedented level of understanding of the impact of GLN and ASN deamidation on the stability of a therapeutic protein/antibody. By determining the extent and site of deamidation, one can assess the impact on (i) target/antigen binding, which has implications for drug discovery/proteomics as well as efficacy; (ii) local and overall decreased stability resulting in decreased shelf-life of the antibody; (iii) the potential for self-association/aggregation, which can lead to decreased product yields; and (iv) potential unwanted immune response, leading to immunogenicity. The anti-drug antibody response is currently being evaluated by our team through in vitro studies. The combined risks to patients result in both efficacy and safety concerns. 14
Our approach involves the analysis of 2D-COS data, which enhances the spectral resolution. The correlation between key signature peaks enables the accurate determination of the extent of deamidation. The unprecedented ability to directly monitor in real-time the key signature peaks directly associated with the vibrational modes41,48–52 involved in the deamidation process during thermal stress provides amino acid-level resolution. The 2D-COS asynchronous analysis delivers enhanced spectral resolution, a quantitative determination of the extent of deamidation, a molecular description of the weak interactions and conformational changes that occur within the full-length protein during the thermal stress, and spatial and temporal resolution of the deamidation process, resulting in the determination of the site of deamidation. The spectra also show the conformational changes that are associated with the destabilization process during thermal stress. Furthermore, the approach allows the monitoring of 10 of the 20 types of amino acid residues in relation to neighboring residues that are perturbed during the deamidation process, facilitating the empirical determination of the site of deamidation within the full-length clinical antibody. The 2D-CDS analysis assesses the extent of deamidation in the population of clinical antibodies within the sample that have been thermally stressed. It allows for the evaluation of the deamidation process when inducing significant aggregation. Eight examples were presented, the combination of which illustrates the predictive nature of the results provided for GLN and ASN deamidation, its impact on stability, and its role in the aggregation of the product. We are extending this study to evaluate the effect that ASN/GLN deamidation may have on immunogenicity risk. Finally, 2T2D is a new algorithm developed by Dr. Isao Noda39,40 that allowed for the comparative assessment of the forced degraded sample against the control at room temperature. 47 This deamidation screen comparability assessment is orthogonal to HPLC CEx chromatography.
Conclusion
To our knowledge, we are the first to provide in a single experiment the multi-attribute comparability assessment of an array of intact clinical antibodies providing insight into several CQAs that impact safety and efficacy. Specifically, eight clinical antibodies were analyzed under both control and forced degraded conditions for the in-depth understanding of the risks of ASN and GLN deamidation: by determining the extent and site of deamidation, its impact on aggregation and stability. The results were validated independently by LC–MS (peptide mapping). Furthermore, our deamidation screening method was verified for four of the eight samples studied under control and forced degraded conditions by HPLC CEx. The significance of this work lies in the potential for providing developability and comparability assessment of biotherapeutics. Especially, when one considers the commercial/manufacturing setting whereby batch-to-batch consistency of these CQAs is monitored or investigated for the implementation of process control strategies.
Supplemental Material
Supplemental material, sj-docx-1-asp-10.1177_00037028241231824 for Streamlined Multi-Attribute Assessment of an Array of Clinical-Stage Antibodies: Relationship Between Degradation and Stability by Belinda Pastrana, Elizabeth Culyba, Sherly Nieves, Stephen L. Sazinsky, Eduardo I. Canto and Isao Noda in Applied Spectroscopy
Acknowledgments
The authors would like to acknowledge Dr. Jin Xu from Merrigen (Lowell, Massachusetts) for performing the analytical CEx analysis and Dr. Melissa Stauffer (Scientific Editing Solutions, Walworth, WI) for editing the manuscript.
Footnotes
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding: The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work presented herein was made possible by support from the National Science Foundation (SBIR PII Award 1632420 (BP)) and Verseau Therapeutics (EC and SS).
ORCID iDs: Belinda Pastrana https://orcid.org/0000-0002-2114-1182
Isao Noda https://orcid.org/0000-0002-0081-4739
Supplemental Material: All Supplemental material cited in this research article is available in the online version.
References
- 1.U.S. Food and Drug Administration. “Purple Book Database of Licensed Biological Products”. https://purplebooksearch.fda.gov/ [accessed Jan 21 2024].
- 2.Kaplon H., Chenoweth A., Crescioli S., Reichert J.M.. “Antibodies to Watch in 2022”. mAbs. 2022. 14(1): 2014296. 10.1080/19420862.2021.2014296 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kaplon H., Reichert J.M.. “Antibodies to Watch in 2021”. mAbs. 2021. 13(1): 1860476. 10.1080/19420862.2020.1860476 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Strohl W.R.. “Current Progress in Innovative Engineered Antibodies”. Protein Cell. 2018. 9(1): 86–120. 10.1007/S13238-017-0457-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yang X., Xu W., Dukleska S., Benchaar S., et al. “Developability Studies Before Initiation of Process Development: Improving Manufacturability of Monoclonal Antibodies”. mAbs. 2013. 5(5): 787–94. 10.4161/MAbs.25269 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Yang C., Gao X., Gong R.. “Engineering of Fc Fragments with Optimized Physicochemical Properties Implying Improvement of Clinical Potentials for Fc-Based Therapeutics”. Front. Immunol. 2018. 8: 1860. 10.3389/fimmu.2017.01860 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bailly M., Mieczkowski C., Juan V., Metwally E., et al. “Predicting Antibody Developability Profiles Through Early-Stage Discovery Screening”. mAbs. 2020. 12(1): 1743053. 10.1080/19420862.2020.1743053 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Xu Y., Wang D., Mason B., Rossomando T., et al. “Structure, Heterogeneity, and Developability Assessment of Therapeutic Antibodies”. mAbs. 2019. 11(2): 239–264. 10.1080/19420862.2018.1553476 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Carter P.J., Lazar G.A.. “Next Generation Antibody Drugs: Pursuit of the ‘High-Hanging Fruit”. Nat. Rev. Drug Discov. 2018. 17(3): 197–223. 10.1038/nrd.2017.227 [DOI] [PubMed] [Google Scholar]
- 10.Dos Santos M.L., Quintilio W., Manieri T.M., Tsuruta L.R., et al. “Advances and Challenges in Therapeutic Monoclonal Antibodies Drug Development”. Braz. J. Pharm. Sci. 2018. 54(Special Issue): 1–15. 10.1590/S2175-97902018000001007 [DOI] [Google Scholar]
- 11.Xu J., Ou J., McHugh K.P., Borys M.C., et al. “Upstream Cell Culture Process Characterization and In-Process Control Strategy Development at Pandemic Speed”. mAbs. 2022. 14(1): 2060724. 10.1080/19420862.2022.2060724 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Leung D., Wurst J., Liu T., Martinez R., et al. “Antibody Conjugates-Recent Advances and Future Innovations”. Antibodies. 2020. 9(1): 2. 10.3390/Antib9010002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Obrezanova O., Arnell A., De La Cuesta R.G., Berthelot M.E., et al. “Aggregation Risk Prediction for Antibodies and Its Application to Biotherapeutic Development”. mAbs. 2015. 7(2): 352–363. 10.1080/19420862.2015.1007828 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Rosenberg A.S.. “Effects of Protein Aggregates: An Immunologic Perspective”. Am. Assoc. Pharm. Sci. J. 2006. 8(3): 501–507. 10.1208/aapsj080359 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Robinson A.B.. “Evolution and the Distribution of Glutaminyl and Asparaginyl Residues in Proteins”. Proc. Natl. Acad. Sci. U.S.A. 1974. 71(3): 885–888. 10.1073/pnas.71.3.885 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Robinson A.B., Robinson L.R.. “Distribution of Glutamine and Asparagine Residues and Their Near Neighbors in Peptides and Proteins”. Proc. Natl. Acad. Sci. U.S.A. 1991. 88(20): 8880–8884. 10.1073/pnas.88.20.8880 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Robinson N.E., Robinson A.B.. “Deamidation of Human Proteins”. Proc. Natl. Acad. Sci. U.S.A. 2001. 98(22): 12409–12413. 10.1073/pnas.221463198 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kato K., Nakayoshi T., Kurimoto E., Oda A.. “Mechanisms of Deamidation of Asparagine Residues and Effects of Main-Chain Conformation on Activation Energy”. Int. J. Mol. Sci. 2020. 21(19): 1–14. 10.3390/ijms21197035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kato A., Kuratani M., Yanagisawa T., Ohtake K., et al. “Extensive Survey of Antibody Invariant Positions for Efficient Chemical Conjugation Using Expanded Genetic Codes”. Bioconjug. Chem. 2017. 28(8): 2099–2108. 10.1021/acs.bioconjchem.7b00265 [DOI] [PubMed] [Google Scholar]
- 20.Catak S., Monard G., Aviyente V., Ruiz-López M.F.. “Reaction Mechanism of Deamidation of Asparaginyl Residues in Peptides: Effect of Solvent Molecules”. J. Phys. Chem. A. 2006. 110(27): 8354–8365. 10.1021/jp056991q [DOI] [PubMed] [Google Scholar]
- 21.Asomugha C.O., Gupta R., Srivastava O.P.. “Structural and Functional Roles of Deamidation of N146 and/or Truncation of NH2- or COOH-Termini in Human Αβ-Crystallin”. Mol. Vis. 2011. 17: 2407–2420. [PMC free article] [PubMed] [Google Scholar]
- 22.Sydow J.F., Lipsmeier F., Larraillet V., Hilger M., et al. “Structure-Based Prediction of Asparagine and Aspartate Degradation Sites in Antibody Variable Regions”. PLoS One. 2014. 9(6): e100736. 10.1371/journal.pone.0100736 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Yan Q., Huang M., Lewis M.J., Hu P.. “Structure Based Prediction of Asparagine Deamidation Propensity in Monoclonal Antibodies”. mAbs. 2018. 10(6): 901–912. 10.1080/19420862.2018.1478646 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kempkes L.J.M., Martens J., Grzetic J., Berden G., et al. “Deamidation Reactions of Asparagine- and Glutamine-Containing Dipeptides Investigated by Ion Spectroscopy”. J. Am. Soc. Mass. Spectrom. 2016. 27(11): 1855–1869. 10.1007/S13361-016-1462-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Guerrero C., Milenković T., Pržulj N., Kaiser P., et al. “Characterization of the Proteasome Interaction Network Using a QTAX-Based Tag-Team Strategy and Protein Interaction Network Analysis”. Proc. Natl. Acad. Sci. U.S.A. 2008. 105(36): 13333–13338. 10.1073/Pnas.0801870105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ramberger E., Suarez-Artiles L., Pérez-Hernández D., Haji M., et al. “A Universal Peptide Matrix Interactomics Approach to Disclose Motif-Dependent Protein Binding”. Mol. Cell. Proteomics. 2021. 20: 100135. 10.1016/j.mcpro.2021.100135 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Foglierini M., Pappas L., Lanzavecchia A., Corti D., et al. “Ancestree: An Interactive Immunoglobulin Lineage Tree Visualizer”. PLoS Comput. Biol. 2020. 16(7): 1–10. 10.1371/journal.pcbi.1007731 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Gil N., Fajardo E.J., Fiser A.. “Discovery of Receptor-Ligand Interfaces in the Immunoglobulin Superfamily”. Proteins: Struct., Funct., Bioinf. 2020. 88(1): 135–142. 10.1002/Prot.25778 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Scott-Taylor T.H., Axinia S.C., Amin S., Pettengell R.. “Immunoglobulin G; Structure and Functional Implications of Different Subclass Modifications in Initiation and Resolution of Allergy”. Immun. Inflamm. Dis. 2018. 6(1): 13–33. 10.1002/iid3.192 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Chiu M.L., Goulet D.R., Teplyakov A., Gilliland G.L.. “Antibody Structure and Function: The Basis for Engineering Therapeutics”. Antibodies. 2019. 8(4): 55. 10.3390/antib8040055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.De Taeye S.W., Bentlage A.E.H., Mebius M.M., Meesters J.I., et al. “Fcγr Binding and ADCC Activity of Human IgG Allotypes”. Front. Immunol. 2020. 11(May): 1–16. 10.3389/fimmu.2020.00740 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hintersteiner B., Lingg N., Janzek E., Mutschlechner O., et al. “No Microheterogeneity of Therapeutic Monoclonal Antibodies Is Governed by Changes in the Surface of the Protein”. Biotechnol. J. 2016. 11(12): 1617–1627. 10.1002/biot.201600504 [DOI] [PubMed] [Google Scholar]
- 33.Schmid I., Bonnington L., Gerl M., Bomans K., et al. “Assessment of Susceptible Chemical Modification Sites of Trastuzumab and Endogenous Human Immunoglobulins at Physiological Conditions”. Commun. Biol. 2018. 1: 28. 10.1038/S42003-018-0032-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Noda I.. “Recent Advancement in the Field of Two-Dimensional Correlation Spectroscopy”. J. Mol. Struct. 2008. 883–884(1–3): 2–26. 10.1016/j.molstruc.2007.11.038 [DOI] [Google Scholar]
- 35.Noda I.. “Techniques of Two-Dimensional (2D) Correlation Spectroscopy Useful in Life Science Research”. Biomed. Spectrosc. Imaging. 2015. 4(2): 109–127. 10.3233/Bsi-150105 [DOI] [Google Scholar]
- 36.Noda I.. “Vibrational Two-Dimensional Correlation Spectroscopy (2DCOS) Study of Proteins”. Spectrochim. Acta, Part A. 2017. 187: 119–129. 10.1016/J.Saa.2017.06.034 [DOI] [PubMed] [Google Scholar]
- 37.Noda I.. “Techniques Useful in Two-Dimensional Correlation and Co-Distribution Spectroscopy (2DCOS and 2DCDS) Analyses”. J. Mol. Struct. 2016. 1124: 29–41. 10.1016/J.Molstruc.2016.01.089 [DOI] [Google Scholar]
- 38.Noda I.. “Two-Dimensional Correlation and Co-Distribution Spectroscopy (2D-COS and 2D-CDS) Analyses of Planar Spectral Image Data”. J. Mol. Struct. 2020. 1211: 128068. 10.1016/J.Molstruc.2020.128068 [DOI] [Google Scholar]
- 39.Noda I.. “Two-Trace Two-Dimensional (2T2D) Correlation Spectroscopy: A Method for Extracting Useful Information from a Pair of Spectra”. J. Mol. Struct. 2018. 1160: 471–478. 10.1016/J.MOLSTRUC.2018.01.091 [DOI] [Google Scholar]
- 40.Noda I.. “Closer Examination of Two-Trace Two-Dimensional (2T2D) Correlation Spectroscopy”. J. Mol. Struct. 2020. 1213: 128194. 10.1016/J.MOLSTRUC.2020.128194 [DOI] [Google Scholar]
- 41.Pastrana B., Meuse C.W.. “Two-Dimensional Correlation and Two-Dimensional Co-Distribution Spectroscopies of Proteins”. In: Meyers R., editor. Encyclopedia of Analytical Chemistry. West Sussex: John Wiley and Sons, Ltd, 2022, Pp. 1–31. 10.1002/9780470027318.A9792. [DOI] [Google Scholar]
- 42.Jain T., Sun T., Durand S., Hall A., et al. “Biophysical Properties of the Clinical-Stage Antibody Landscape”. Proc. Natl. Acad. Sci. U.S.A. 2017. 114(5): 944–949. 10.1073/Pnas.1616408114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Davies A.M., Sutton B.J.. “Human IgG4: A Structural Perspective”. Immunol. Rev. 2015. 268(1): 139–159. 10.1111/Imr.12349 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Scapin G., Yang X., Prosise W.W., Mccoy M., et al. “Structure of Full-Length Human Anti-PD1 Therapeutic IgG4 Antibody Pembrolizumab”. Nat. Struct. Mol. Biol. Nature Publishing Group, 2015. 22(12): 953–958. 10.1038/Nsmb.3129 [DOI] [PubMed] [Google Scholar]
- 45.Pastrana-Rios B., Noda I.. System and Method for Determining Deamidation and Immunogenicity of Polypeptides. US Patent 17/288,476. Filed 2021. Patent Pending.
- 46.Pastrana-Ríos B., Rodriguez-Toro J.. Method and System for Spectral Data Analysis. US Patent 10714209B2. Filed 2019. Issued 2020.
- 47.Pastrana-Rios B.. Rapid Deamidation Screening. US Patent 63/509,340. Filed 2023. Patent Pending.
- 48.Hernández B., Pfluger F., Adenier A., Kruglik S.G., Ghomi M.. “Vibrational Analysis of Amino Acids and Short Peptides in Hydrated Media. IV. Amino Acids with Hydrophobic Side Chains”. J. Phys. Chem. B. 2010. 114(46): 15319–15330. 10.1021/jp106786j [DOI] [PubMed] [Google Scholar]
- 49.Venyaminov S.Y., Kalnin N.N.. “Quantitative IR Spectrophotometry of Peptide Compounds in Water (H2O) Solutions. I. Spectral Parameters of Amino Acid Residue Absorption Bands”. Biopolymers. 1990. 30(13–14): 1243–1257. 10.1002/bip.360301309 [DOI] [PubMed] [Google Scholar]
- 50.Barth A.. “Infrared Spectroscopy of Proteins”. Biochim. Biophys. Acta Bioenerg. 2007. 1767(9): 1073–1101. 10.1016/j.bbabio.2007.06.004 [DOI] [PubMed] [Google Scholar]
- 51.Nieves S., Pastrana B.. “Quantum Cascade Laser Infrared Microscopy and 2D IR Correlation Spectroscopy Improves Crystallization Screening of a Protein Complex”. Spectroscopy. 2019. 34(11): 34–39. [Google Scholar]
- 52.Pastrana B., Nieves S., Li W., Liu X., Dimitrov D.S.. “Developability Assessment of an Isolated CH2 Immunoglobulin Domain”. Anal. Chem. 2021. 93(3): 1342–1351. 10.1021/acs.analchem.0c02663 [DOI] [PubMed] [Google Scholar]
- 53.Noda I., Ozaki Y.. Two-Dimensional Correlation Spectroscopy: Applications in Vibrational and Optical Spectroscopy. Chichester, UK: John Wiley and Sons, Ltd., 2004. 10.1002/0470012404 [DOI] [Google Scholar]
- 54.Lu X., Nobrega R.P., Lynaugh H., Jain T., Barlow K., Boland T., et al. “Deamidation and Isomerization Liability Analysis of 131 Clinical-Stage Antibodies”. mAbs. 2019. 11(1): 45–57. 10.1080/19420862.2018.1548233 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental material, sj-docx-1-asp-10.1177_00037028241231824 for Streamlined Multi-Attribute Assessment of an Array of Clinical-Stage Antibodies: Relationship Between Degradation and Stability by Belinda Pastrana, Elizabeth Culyba, Sherly Nieves, Stephen L. Sazinsky, Eduardo I. Canto and Isao Noda in Applied Spectroscopy