Abstract
Filamentous fungal genomes contain two distantly related cyclic AMP-dependent protein kinase A catalytic subunits (PKAs), but only one PKA is found to play a principal role. In Aspergillus nidulans, PkaA is the primary PKA that positively functions in vegetative growth and spore germination but negatively controls asexual sporulation and production of the mycotoxin sterigmatocystin. In this report, we present the identification and characterization of pkaB, encoding the secondary PKA in A. nidulans. Although deletion of pkaB alone does not cause any apparent phenotypic changes, the absence of both pkaB and pkaA is lethal, indicating that PkaB and PkaA are essential for viability of A. nidulans. Overexpression of pkaB enhances hyphal proliferation and rescues the growth defects caused by ΔpkaA, indicating that PkaB plays a role in vegetative growth signaling. However, unlike ΔpkaA, deletion of pkaB does not suppress the fluffy-autolytic phenotype resulting from ΔflbA. While upregulation of pkaB rescues the defects of spore germination resulting from ΔpkaA in the presence of glucose, overexpression of pkaB delays spore germination. Furthermore, upregulation of pkaB completely abolishes spore germination on medium lacking a carbon source. In addition, upregulation of pkaB enhances the level of submerged sporulation caused by ΔpkaA and reduces hyphal tolerance to oxidative stress. In conclusion, PkaB is the secondary PKA that has a synthetic lethal interaction with PkaA, and it plays an overlapping role in vegetative growth and spore germination in the presence of glucose but an opposite role in regulating asexual sporulation, germination in the absence of a carbon source, and oxidative stress responses in A. nidulans.
In eukaryotic cells, protein kinases constitute one of the largest gene families and play a pivotal role in transducing external and/or internal signals to elicit appropriate cellular responses. Among these kinases, cyclic AMP (cAMP)-dependent protein kinase A (PKA) serves as a prototypical example. In the absence of cAMP, the PKA holoenzyme exists as an inactive equimolar hetero-tetramer composed of a homo-dimeric regulatory subunit and two associated catalytic subunits (reviewed in references 36 and 38). The regulatory subunit also functions in preventing the inactive PKA holoenzyme from entering the nucleus (6, 11). The cooperative binding of two cAMP molecules to each regulatory subunit of the enzyme results in dissociation of the active PKAs from the regulatory subunits. These active PKAs can phosphorylate downstream target proteins at serine or threonine residues (reviewed in reference 38). PKA signaling plays a central role in vegetative growth, development, mating, stress response, and pathogenicity in various fungi (reviewed in references 8, 21, and 22).
In the budding yeast Saccharomyces cerevisiae, three TPK genes encode the closely related PKA isoforms that share redundant and distinct functions in viability (39) and in pseudohyphal morphogenesis, respectively (26, 28). Similarly, two PKA isoforms play redundant and distinct roles in Candida albicans, i.e., PKAs are essential for (normal) growth and stress response while playing distinct roles in hyphal morphogenesis and invasive growth (3). In the fission yeast Schizosaccharomyces pombe, one PKA, Pka1, has been characterized and shown to function in spore germination, growth, and sexual development (15, 24).
Unlike yeasts, two distantly related PKAs exist in most (if not all) filamentous or dimorphic fungi, but only one PKA has been found to play a predominant role. For instance, in the plant pathogenic fungus Ustilago maydis, while Adr1 (class I PKA) was found to be crucial for development and pathogenicity, deletion of Uka1 (class II PKA) resulted in no detectable changes (9) (see Fig. 1D, below). Likewise, one PKA has been shown to play a primary role in normal growth, development, and pathogenicity in Magnaporthe grisea (1, 25, 43), Cryptococcus neoformans (7, 17), and Aspergillus fumigatus (23).
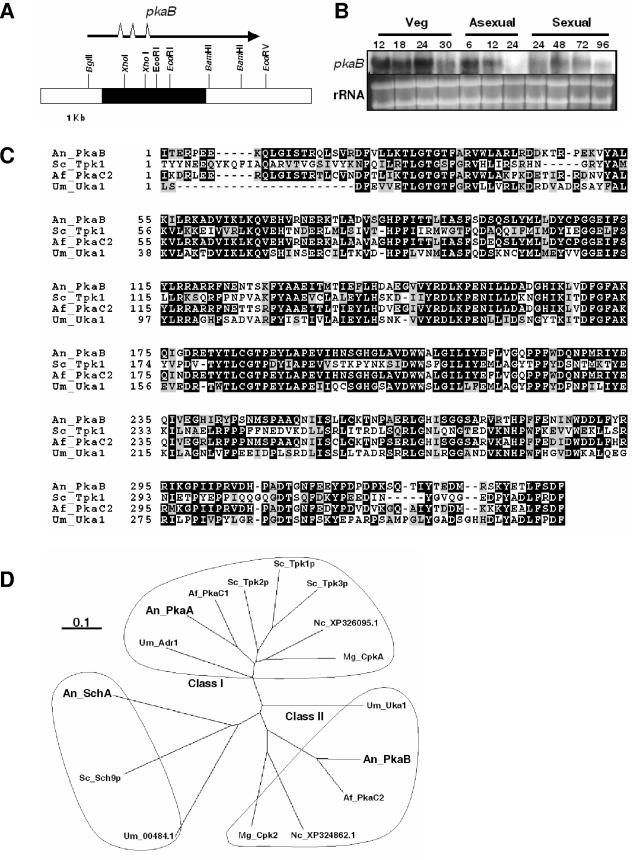
FIG.1.
Summary of pkaB, encoding the secondary PKA catalytic subunit. A. A partial restriction map of the pkaB gene region is shown. The pkaB ORF (filled box), the mRNA coding region (arrow), and the three introns (marked by discontinuities on the arrow) were determined by reverse transcription-PCR and random amplification of cDNA ends followed by sequence analyses. B. Steady-state mRNA levels of pkaB in various growth and developmental stages. Numbers indicate the times of incubation in liquid MMG (Veg) or solid MMG under the conditions inducing asexual development or sexual development. The pkaB mRNA levels were low at 24 and 96 h post-asexual and -sexual developmental induction. C. Amino acid sequence alignment of A. nidulans PkaB (An_PkaB), S. cerevisiae Tpk1P (Sc_Tpk1), A. fumigatus PkaC2 (Af_PkaC2), and U. maydis Uka1 (Um_Uka1), showing identities (black boxes) and similarities (gray boxes). The proteins were aligned with ClustalW (5) and displayed by BOXSHADE (http://www.ch.embnet.org/software/BOX_form.html). D. Phylogenetic tree of PkaA, PkaB, and SchA of A. nidulans, PkaC1 and PkaC2 of A. fumigatus, CpkA and Cpk2 of M. grisea, XP324862.1 and XP326095.1 of N. crassa, Tpk1p, Tpk2p, and Tpk3p of S. cerevisiae, Adr1, and Uka1 and UM00484.1 of U. maydis. The ClustalW alignment (5) of the fungal proteins was presented by TREEVIEW. The length of the bar represents an evolutionary distance of 0.1 amino acid substitutions per site.
In the model filamentous fungus Aspergillus nidulans, PkaA was shown to play a central role in various biological processes (10, 34). Deletion of pkaA resulted in delayed germination of conidiospores (asexual spores), highly restricted growth, and elevated asexual sporulation (conidiation) and production of the mycotoxin sterigmatocystin (ST). Additionally, genetic epistatic analysis showed that PkaA functions downstream of FadA, the Gα subunit for a heterotrimeric G protein that primarily mediates vegetative growth signaling (34, 44). This FadA-mediated signaling is negatively controlled by FlbA, an RGS protein, which is necessary for the commencement of development and ST production (16, 20, 44). Both the loss-of-function flbA and the constitutively active fadA [e.g., fadA(G42R) or fadA(Q204L)] mutations result in the absence of development and ST production as well as hyphal disintegration (autolysis) due to uncontrolled activation of vegetative growth signaling (16, 20, 44, 45). Deletion of pkaA partially restores conidiation in these proliferative mutants while retaining restricted growth (34), indicating that vegetative growth is primarily mediated via FadA and PkaA. However, deletion of pkaA does not completely eliminate PKA activity in A. nidulans (34), suggesting that multiple PKA catalytic subunits may exist in A. nidulans. In addition to PkaA, a homolog of the S. cerevisiae Sch9 protein named SchA has been studied in A. nidulans (10). Deletion of schA alone caused no apparent phenotypic changes, but the combination of null mutations in both pkaA and schA delayed conidiospore germination even further (10). A recent study showed that TPKs and Sch9 function in separate signaling cascades in S. cerevisiae (31), suggesting SchA and PkaA may function in distinct pathways in A. nidulans.
In this study, we have identified and characterized the pkaB gene encoding the secondary PKA in A. nidulans. PkaB is distantly related to PkaA and is most similar to other PKAs previously characterized as class II (see Fig. 1D, below). Although deletion of pkaB alone causes no apparent phenotypic changes in growth, development, spore germination, ST production, or stress responses, the absence of both PkaB and PkaA is lethal, indicating that PkaB and PkaA constitute the sole PKA catalytic subunits that are essential for the viability of A. nidulans. We further demonstrate that while PkaB and PkaA have overlapping functions in stimulating vegetative growth and spore germination in the presence of glucose, the two PKAs play opposite roles in regulating spore germination in the absence of an external carbon source, oxidative stress responses, and asexual sporulation in submerged culture.
MATERIALS AND METHODS
Aspergillus nidulans strains, growth conditions, and genetics.
Strains used in this study are listed in Table 1. Standard culture and genetics techniques were used (27). Fungal strains were grown on minimal solid or liquid medium with supplements (simplified as MM) as described previously (18). To examine the levels of colony growth, all strains were inoculated on solid MM with various carbon sources and incubated at 37°C for 3 days (MMG, with 1% glucose) or 6 days (MMT, with 100 mM threonine as a sole carbon source to induce overexpression of pkaB). To check the development of conidiophores in liquid submerged culture, liquid MMG was inoculated with the conidia of individual strains (108 spores/100 ml) and incubated at 37°C, 250 rpm. To examine the effect of overexpression or upregulation of pkaB by an ectopic copy of pkaB under the alcA promoter, relevant strains were grown in liquid MMG at 37°C, 250 rpm for 14 h. Subsequently, mycelia were collected, rinsed with liquid medium without a carbon source, divided into two equal parts, and transferred into liquid MMG or liquid MMT. Then, the cultures were shaken under the same conditions described above, and development of wild-type and mutant strains was examined under a microscope at 0, 6, and 12 h posttransfer.
TABLE 1.
A. nidulans strains used in this study
| Strain | Genotypea | Source or reference |
|---|---|---|
| FGSC4 | veA+ | FGSCb |
| FGSC26 | biA1 | FGSC |
| FGSC237 | pabaA1 yA2 trpC801 | FGSC |
| PW1 | biA1 argB2 methG1 | P. Weglenski |
| TKIS18.11 | pabaA1 yA2 ΔpkaA::argB+ ΔargB::trpC+trpC801 | 34 |
| TKIS20.1 | pabaA1 yA2 alcA(p)::pkaA::trpC+ | 34 |
| TSR1 | biA1 argB2 ΔpkaB::argB+methG1 | This study |
| TSR53 | biA1 argB2 ΔpkaB::argB+methG1 | This study |
| RSA53.126 | pabaA1 yA2 ΔpkaA::argB+ ΔargB::trpC+trpC801 | This study |
| RSA53.130 | biA1 ΔpkaA::argB+ ΔargB::trpC+trpC801 | This study |
| RSA53.111 | pabaA1 biA1 argB2 ΔpkaB::argB+ | This study |
| RSA53.143 | biA1 argB2 ΔpkaB::argB+ | This study |
| RSA37.1C.12 | biA1 ΔpkaA::argB+ ΔargB::trpC+ ΔpkaB::argB+alcA(p)::pkaA::trpC+ | This study |
| RNI5.1, -2, -3c | biA1 alcA(p)::pkaB::trpC+ | This study |
| RNI6 | pabaA1 yA2 ΔpkaA::argB+alcA(p)::pkaB::trpC+ | This study |
| RNI7 | pabaA1 yA2 ΔpkaA::argB+ ΔpkaB::argB+alcA(p)::pkaB::trpC+ | This study |
| RNI8 | biA1 alcA(p)::pkaA::trpC+ | This study |
All strains carry the veA1 mutation except FGSC4.
FGSC, Fungal Genetics Stock Center.
Multiple isogenic alcA(p)::pkaB strains. RNI5.1, RNI5.2, and RNI5.3 behaved identically.
To examine the pkaB mRNA levels in the wild type, samples from liquid submerged and postdevelopmental induction cultures were collected as described elsewhere (12). Briefly, 5 × 107 conidia of FGSC4 were inoculated in 100 ml of liquid MMG with 0.1% yeast extract in 250-ml flasks and incubated at 37°C, 250 rpm. For the vegetative growth phase, samples were collected from liquid submerged cultures at designated time points, squeeze dried, and stored at −80°C until subjected to total RNA isolation. For sexual and asexual developmental induction, 18-h vegetatively grown mycelia were transferred to solid MMG and the plates were either air exposed for asexual developmental induction or tightly sealed and shielded from light to induce sexual development.
Identification and structure of pkaB.
The pkaB gene was identified through tblastn search of the A. nidulans genome (The Broad Institute [http://www.genome.wi.mit.edu/annotation/fungi/aspergillus/index.html]) using Tpk1, Tpk2, and Tpk3 of S. cerevisiae. To examine the positions of introns, reverse transcription-PCR, followed by sequencing analyses of pkaB, was carried out. The Smart RACE kit (Clontech) was used to determine the 5′ and 3′ ends of the pkaB transcript.
Construction of pkaB deletion and pkaB overexpression strains.
The double-joint PCR method was used to generate the pkaB deletion mutant (46). A deletion construct containing argB with approximately 1 kb each of the 5′ and 3′ flanking regions of pkaB (amplified with primer pairs of OKH56 with -57 and OKH58 with -59 (Table 2) was introduced into the recipient strain, PW1 (biA1 argB2 methG1 veA1). The resulting transformants were randomly screened for deletion of pkaB as described previously (46). The pkaB overexpression construct was created by fusing the alcA promoter with the pkaB open reading frame (ORF). The alcA promoter was amplified by OJA106 and OJA107. The pkaB ORF with an alcA(p) tail was amplified by OSR16 and OSR18. The two DNA amplicons were fused as described elsewhere (46), and the fusion product was amplified by OJA108 and OSR17. The resulting alcA(p)::pkaB amplicon was cut with BamHI and ligated into the BamHI-cut pSH96 plasmid (42). The sequence-verified alcA(p)::pkaB plasmid was introduced into FGSC237, and the single-copy integration of plasmid into the trpC locus was confirmed by a series of PCR analyses. Three alcA(p)::pkaB transformants were isolated and sexually crossed with ΔpkaA (RSA53.130) or ΔpkaA ΔpkaB alcA(p)::pkaA (RSA37.1C.12) strains. From these crosses, alcA(p)::pkaB (RNI5.3), ΔpkaA alcA(p)::pkaB (RNI6), and ΔpkaA ΔpkaB alcA(p)::pkaB (RNI7) strains were isolated and used in subsequent overexpression studies.
TABLE 2.
Oligonucleotides used in this study
| Oligo | Sequence | Position or purpose |
|---|---|---|
| OKH60 | GAC TCT ATA CCA CCG TAC GCC GAT AT | Forward primer for argB+ |
| OKH61 | CAC CGG GTG CGA TTT GCC CCA TTT CC | Reverse primer for argB+ |
| OKH56 | AAC GAA CCC AGC GTC GAT CTA T | pkaB deletion 5′ forward |
| OKH57 | AGT CAA ATG AGG CCT CTA AAC TGG TCA ACT GTG CGC TTC TCC CTG TCA T | pkaB deletion 5′ reverse with complementary 5′ argB+ taila |
| OKH58 | AGC CAA GGT AGA TCC AGG CCT AAC ACA CA TTG CGG AAT GGT ATC ACA CGT C | pkaB deletion 3′ forward with complementary 3′ argB+ taila |
| OKH59 | TGG GCT TTG GTC CTC ATT GAG T | pkaB deletion 3′ reverse |
| OKH74 | CGT CCA CGC ACT TCC TCA | pkaB deletion construct 5′ nest |
| OKH75 | GGG ATA TCT GCC CGG CTA | pkaB deletion construct 3′ nest |
| OJA107 | ACA ATC AAT GTT CAA TGT AC | 5′ end of alcA(p) |
| OJA106 | TTT GAG GCG AGG TGA TAG GAT TGG A | 3′ end of alcA(p) |
| OJA108 | CG GGA TCC AGT GGT TCG GTA ATC | alcA(p) 5′ nested with BamHI taila |
| OSR16 | TCC AAT CCT ATC ACC TCG CCT CAA A CA TTC AAA TGG CTT CAG GAA | pkaB overexpression 5′-alcA(p) taila |
| OSR17 | TTC TTC GGC TAG GGA TCC | pkaB overexpression construct 3′ nested |
| OSR18 | TTG CTT TAT ACT CTG GCT TT | pkaB overexpression construct 3′ |
| OSR14 | ACT AGT ACT CCG TAC ATG C | pkaB forward primer for probe |
| OSR15 | TTG CTA GCC TTG GTA ATA | pkaB reverse primer for probe |
| OJA142 | CTG GCA GGT GAA CAA GTC | brlA forward primer for probe |
| OJA143 | AGA AGT TAA CAC CGT AGA | brlA reverse primer for probe |
Tail sequence is shown in italics.
Nucleic acid isolation and manipulation.
Genomic DNA and total RNA isolation and Northern blot analyses were carried out as previously described (12, 32). The DNA probes used to examine the mRNA levels of pkaB and brlA were prepared by PCR amplification of the coding regions of individual genes using genomic DNA of FGSC4 as the template with appropriate oligonucleotide pairs (Table 2).
The pkaB mRNA levels in wild-type (FGSC26), ΔpkaB (RSA53.143), ΔpkaA (RSA53.130), alcA(p)::pkaB (RNI5.3), ΔpkaA alcA(p)::pkaB (RNI6), and ΔpkaA ΔpkaB alcA(p)::pkaB (RNI7) strains were examined as follows: (i) for conidia, total RNA was isolated from the freshly collected conidia of individual strains grown on solid MMG at 37°C for 5 days. (ii) In liquid medium without a carbon source, approximately 109 spores of each strain were inoculated in 100 ml of liquid medium without a carbon source and further incubated at 37°C, 250 rpm for 24 h. Each sample was then collected by using a filter sterilization unit (0.45 μm), transferred into a microcentrifuge tube, and subjected to total RNA isolation. (iii) In liquid MMG or MMT, approximately 5 × 107 or 108 spores of each strain were inoculated in 100 ml of liquid MMG or MMT, respectively, and further incubated at 37°C, 250 rpm for 24 h. Samples were then collected, transferred into microcentrifuge tubes, and subjected to total RNA isolation. Approximately 2 μg (from conidia) or 6 μg of total RNA was loaded per lane, separated by agarose electrophoresis, transferred to nylon membrane (Magnaprobe; Osmonics), and hybridized with the pkaB probe amplified by OSR14 and OSR15 (Table 2).
Germination of conidia.
To examine germination levels, the conidia (∼106 spores/plate) of wild-type (FGSC26), ΔpkaB (RSA53.143), ΔpkaA (RSA53.130), alcA(p)::pkaB (RNI5.3), ΔpkaA alcA(p)::pkaB (RNI6), and ΔpkaA ΔpkaB alcA(p)::pkaB (RNI7) strains were spread on solid MM containing glucose or threonine, or on the medium lacking a carbon source, and incubated at 37°C. The levels of germination (isotropic and germ tube formation) were examined every 2 h after 4 h of inoculation on solid MMG or MMT or after 8 h of inoculation on the medium lacking a carbon source.
Stress response.
To examine the sensitivity of hyphae to H2O2, the conidia of wild-type (FGSC26), ΔpkaB (RSA53.143), ΔpkaA (RSA53.130), alcA(p)::pkaB (RNI5.3), and alcA(p)::pkaA (RNI8) strains were incubated on solid MMG (∼250 spores/plate) in triplicate, and the plates were incubated at 37°C for 26 h. This 26-h incubation time was chosen because conidia of all the strains tested germinated and formed microscopically visible colonies without producing conidiophores. In the wild type, the first conidiophore from a single conidium is produced at 28 h of incubation at 37°C (44). The hyphae were overlaid with 10 ml of 0, 10, 50, 150, or 300 mM H2O2 solution and incubated at room temperature for 10 min. The H2O2 solution was then decanted, and the plates were washed four times with 15 ml of sterile double-distilled H2O. Plates were then incubated at 37°C for an additional 24 h. The number of surviving colonies was counted and calculated as a percentage of the control (untreated). To examine the sensitivity of conidia to H2O2, 1-ml aliquots of conidial suspensions containing ∼105 spores with various H2O2 concentrations (0, 0.25, 0.50, 0.75, and 1.0 M) were incubated at room temperature for 30 min. Spores were then collected by centrifugation at 10,000 × g for 1 min, washed twice with 1.5 ml of sterile double-distilled H2O, resuspended, diluted with sterile distilled water, and inoculated onto solid MMG. After incubation at 37°C for 24 h, the number of colonies was counted and calculated as a percentage of control. To examine the thermal tolerance of hyphae, the conidia of wild-type (FGSC26), ΔpkaB (RSA53.143), and alcA(p)::pkaB (RNI5.3) conidia (∼250/plate) were inoculated on solid MMG in triplicate. Plates were incubated at 37°C for 10 h and then shifted to 50°C. After 2, 3, or 4 h of incubation at 50°C, the plates were shifted back to 37°C for an additional 24 h, and the number of colonies was counted.
Microscopy.
The colony photographs were taken using a Sony DSC-F707 digital camera. Microscopic photographs were taken using an Olympus BH2 compound microscope installed with an Olympus DP-70 digital imaging system.
RESULTS
Identification of pkaB encoding the secondary PKA in A. nidulans.
Search of the A. nidulans genome employing Tpk1, Tpk2, and Tpk3 of S. cerevisiae resulted in the identification of the pkaB gene encoding an additional PKA (AN4717.2, chromosome III). The pkaB gene is composed of a 1,371-bp ORF with three (62-bp, 64-bp, and 54-bp) introns and is predicted to encode a 396-amino-acid polypeptide which shows 40 to ∼53% amino acid identity to yeast PKAs (Fig. 1A and C). The predicted PkaB protein contains a typical Ser/Thr protein kinase domain and an extension to the Ser/Thr-type protein kinase domain at the C terminus (14, 37). These domains are conserved in numerous PKA catalytic subunits. To ensure that pkaB was expressed, the steady-state mRNA levels were examined by Northern blot analysis. As shown in Fig. 1B, pkaB encodes a ∼2.4-kb transcript that accumulates at high levels during the vegetative growth phase and the early to middle phases of asexual/sexual development of A. nidulans.
PkaB shows high amino acid identity with other fungal PKAs: 82% with PkaC2 of A. fumigatus (GI 40641908 [23]), 53% with Tpk1p of S. cerevisiae (GI 6322297 [39]), and 49% with Uka1 of U. maydis (GI 2565330 [9]) (for alignment, see Fig. 1C). Phylogenetic analysis of Tpk1p, Tpk2p, Tpk3p, and Sch9p of S. cerevisiae (39, 40), PkaA and PkaB of A. nidulans (34), PkaC1 and PkaC2 of A. fumigatus (23), two putative PKAs, XP_324862.1 (GI 32408763) and XP_326295.1 (GI 32411629) of Neurospora crassa, two PKAs (CpkA and Cpk2) of Magnaporthe grisea (25) (GI 27802543 for Cpk2), and Uka1p, Adr1p, and UM00484.1 of U. maydis has been carried out (9) (Fig. 1D). As it is quite apparent, most filamentous fungi contain two distantly related PKAs (previously classified as I and II) and one Sch9-like kinase (10). Although PkaB was originally identified using yeast PKAs, it is distantly related to Tpk1p, Tpk2p, Tpk3p, and PkaA. Previous studies in U. maydis (9), M. grisea (25), C. neoformans serotype A (7), A. fumigatus (23), and A. nidulans (34) demonstrated that deletion of class I PKA resulted in profound effects, suggesting that class I PKA represents the predominant (primary) unit in these fungi (for an exception, see Discussion).
Deletion of both pkaB and pkaA is lethal.
The pkaB deletion mutant was isolated employing the double-joint PCR method by replacing the pkaB ORF with argB+ (46). Phenotypic characteristics of the ΔpkaB mutant were thoroughly examined for vegetative growth at various temperatures, asexual/sexual development, spore germination, ST production, growth on different external carbon sources, and tolerance to various stresses. However, no apparent phenotypic changes were caused by ΔpkaB. This is consistent with the previous reports that class II PKAs may have no clear function (9, 23). Based on these results, we hypothesized that PkaB may function as a secondary unit and may play a certain role in the absence of the primary PKA, PkaA. To test this hypothesis, we attempted to isolate the ΔpkaA ΔpkaB double mutant via a meiotic cross of ΔpkaA (RSA53.126) and ΔpkaB (TSR1 and TSR53) strains. Of the 80 randomly chosen progeny, 40 had the pkaA+ ΔpkaB or the ΔpkaA pkaB+ genotype, and no single ΔpkaA ΔpkaB mutant was isolated. Due to the presence of the endogenous argB mutations (ΔargB or argB2) in both deletion mutants (Table 1), no pkaA+ pkaB+ (wild-type) recombinants could grow on the medium lacking arginine. Thus, if all recombinant progeny were viable, one-third of progeny would be the pkaA+ ΔpkaB, ΔpkaA pkaB+, or ΔpkaA ΔpkaB mutant. If this were the case, the likelihood of not isolating the ΔpkaA ΔpkaB double mutant among 80 progeny by chance would be (1 − 0.33)80, or 1.22 × 10−14, i.e., almost 0% probability to miss the ΔpkaA ΔpkaB mutant. This strongly suggests that the ΔpkaA ΔpkaB mutant is nonviable and that PkaA and PkaB constitute the sole PKA catalytic subunits required for the viability of A. nidulans.
To further test the synthetic lethal interaction between PkaA and PkaB, a number of ΔpkaA and ΔpkaB strains carrying an ectopic copy of alcA(p)::pkaB or alcA(p)::pkaA were constructed and meiotically crossed with each other. Via random screening of progeny, we were able to isolate the ΔpkaA ΔpkaB mutants only if an ectopic copy of alcA(p)::pkaB or alcA(p)::pkaA was present (Table 1). These results further confirm that deletion of both pkaA and pkaB is lethal.
Overexpression of pkaB enhances vegetative growth and rescues the growth defects caused by ΔpkaA.
The synthetic lethal phenotype caused by the absence of both pkaB and pkaA implies that PkaB may function in hyphal growth and/or spore germination. To examine its potential role in activating vegetative growth, the effect of overexpression of pkaB on hyphal proliferation was examined. As shown in Fig. 2, overexpression of pkaB resulted in enhanced hyphal proliferation while repressing conidiation, resulting in a fluffy colony with delayed conidiation on inducing medium (MMT with or without yeast extract [YE]). Furthermore, overexpression of pkaB could partially rescue the growth defects caused by ΔpkaA (Fig. 2, lower middle panel). These results indicate that PkaB may stimulate vegetative growth, which indirectly represses conidiation in the air (see reduced conidiation in Fig. 2).
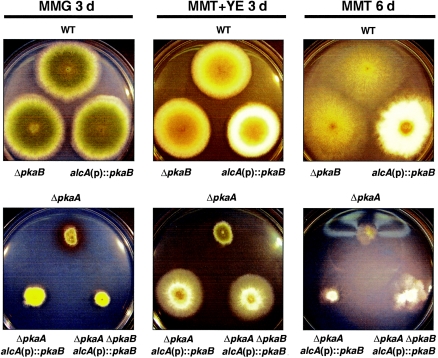
FIG. 2.
Overexpression of pkaB enhances vegetative growth. Photographs of the colonies of wild-type (WT; FGSC26), ΔpkaB (RSA53.143), alcA(p)::pkaB (RNI5.3), ΔpkaA (RSA53.130), ΔpkaA alcA(p)::pkaB (RNI6), and ΔpkaA ΔpkaB alcA(p)::pkaB (RNI7) strains grown on MMG (noninducing) or MMT plus YE (threonine plus 5 g/liter YE) for 3 days or on MMT for 6 days at 37°C are shown. Yeast extract was added to achieve both alcA induction and enhanced growth. While ΔpkaB caused no apparent phenotypic changes, overexpression of pkaB resulted in elevated hyphal proliferation and reduced asexual sporulation on both MMT plus YE and MMT (upper panel). Furthermore, overexpression of pkaB rescued the growth defects caused by ΔpkaA (lower panel).
To test for a potential role of PkaB in transducing FadA (Gα)-mediated vegetative signaling, we generated the ΔflbA ΔpkaB mutant and found that deletion of pkaB was not sufficient to suppress fluffy-autolytic phenotypes caused by the absence of FlbA function. This result suggests that either FadA-mediated vegetative signaling is primarily transmitted via PkaA, with PkaB functioning as a back-up unit, or that PkaB functions in a separate signaling branch (see Discussion).
Upregulation of pkaB rescues delayed germination caused by ΔpkaA in the presence of glucose.
Previously, it was shown that deletion of pkaA resulted in delayed spore germination (10). While deletion of pkaB had no effects on spore germination (data not shown), the synthetic lethality resulting from ΔpkaB ΔpkaA implied a potential role of PkaB in spore germination. Two copies of pkaB, an ectopic copy of alcA(p)::pkaB and the endogenous pkaB gene, in alcA(p)::pkaB strains resulted in ∼10-fold-elevated pkaB mRNA accumulation even when glucose (1%) was used as a carbon source (Fig. 3A). Such upregulation of pkaB allowed us to test the effects of elevated mRNA levels of pkaB on spore germination in the presence of glucose. Approximately 106 spores of wild-type and relevant mutant strains (Fig. 3B) were inoculated onto solid MMG, and conidial swelling (isotropic growth) and germ tube emergence were microscopically observed. More than 90% of the wild-type conidia began to form short germ tubes (data not shown) by 6 h, and all the conidia formed long germ tubes by 10 h (Fig. 3B). The conidia of the ΔpkaB and alcA(p)::pkaB mutants exhibited a germination pattern similar to that of the wild-type conidia (ΔpkaB not shown). As reported, deletion of pkaA severely delayed spore germination, i.e., only 40% of the ΔpkaA mutant conidia began to produce germ tubes at 10 h. As if PkaB could replace PkaA in spore germination, upregulation of pkaB could rescue germination defects caused by ΔpkaA, i.e., the conidia of the ΔpkaA alcA(p)::pkaB mutant exhibited a germination level similar to that of the wild-type conidia (Fig. 3B). In supporting dose-dependent effects of pkaB, the conidia of the ΔpkaA ΔpkaB alcA(p)::pkaB triple mutant did not germinate as effectively as those of the ΔpkaA alcA(p)::pkaB mutant (data not shown, but for pkaB accumulation see Fig. 3A). These results provide evidence that PkaB plays an overlapping role in spore germination in the presence of glucose.
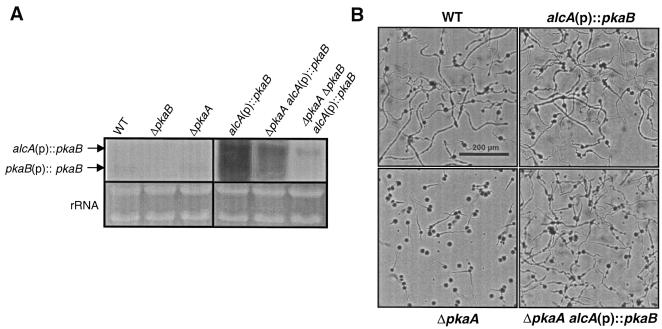
FIG. 3.
Upregulation of pkaB rescues germination defects caused by deletion of pkaA. A. The conidia of wild-type (WT; FGSC26), ΔpkaB (RSA53.143), ΔpkaA (RSA53.130), alcA(p)::pkaB (RNI5.3), ΔpkaA alcA(p)::pkaB (RNI6), and ΔpkaA ΔpkaB alcA(p)::pkaB (RNI7) strains were inoculated (108 spores/100 ml) into liquid MMG and incubated at 37°C, 250 rpm, for 24 h. Total RNA was isolated from each sample, 6 μg of total RNA was loaded per lane, and equal loading was confirmed by ethidium bromide staining of rRNA. Note that the presence of the alcA(p)::pkaB construct resulted in elevated pkaB mRNA levels (∼10-fold). While deletion of pkaA did not affect the pkaB mRNA levels in the absence of alcA(p)::pkaB, somehow ΔpkaA and ΔpkaA ΔpkaB caused reduced accumulation of the pkaB mRNA derived from the alcA promoter. Due to the differences in transcriptional termination, the pkaB mRNA derived from the alcA promoter is about 0.5 kb bigger than the endogenous pkaB transcript shown in wild-type (FGSC26) and ΔpkaA (RSA53.130) strains. Because wild-type and ΔpkaA strains showed extremely low pkaB mRNA levels compared to the strains containing alcA(p)::pkaB, different exposure conditions were employed to make the pkaB transcripts clearly visible in all the strains tested. It is important to note that the pkaB transcripts shown in alcA(p)::pkaB, ΔpkaA alcA(p)::pkaB, and ΔpkaA ΔpkaB alcA(p)::pkaB strains resulted from a 20-h exposure with one intensifying screen, whereas the pkaB transcripts shown in wild-type and ΔpkaA strains were visualized by a 24-h exposure with two intensifying screens. Therefore, the actual relative levels of the pkaB mRNA in wild-type and ΔpkaA strains must be reduced by twofold. B. The conidia of wild-type (FGSC26), alcA(p)::pkaB (RNI5.3), ΔpkaA (RSA53.130), and ΔpkaA alcA(p)::pkaB (RNI6) strains were inoculated on solid MMG and incubated at 37°C for 10 h. Note that the presence of an ectopic copy of alcA(p)::pkaB could rescue the germination defects caused by ΔpkaA.
Overexpression of pkaB delays spore germination.
The effect of overexpression on conidial germination was examined by direct inoculation of the conidia of wild-type, ΔpkaB, alcA(p)::pkaB, ΔpkaA, ΔpkaA alcA(p)::pkaB, and ΔpkaA ΔpkaB alcA(p)::pkaB strains on MMT. Because it was not possible to determine the pkaB mRNA levels under these experimental conditions, the pkaB mRNA levels were assessed by incubating the conidia of individual strains in liquid MMT at 37°C, 250 rpm, for 24 h (see Materials and Methods). The pkaB transcript levels in strains with alcA(p)::pkaB were about 20-fold higher than those in wild-type and ΔpkaA strains (not shown). Somewhat surprisingly, we found that overexpression of pkaB on solid MMT severely delayed conidial germination (Fig. 4A). Because threonine is not one of the most favorable carbon sources for A. nidulans, germination of the conidia of both wild-type and ΔpkaB strains on solid MMT was delayed about 4 h, i.e., ∼90% of the wild-type and ΔpkaB conidia formed germ tubes within 10 h of incubation (Fig. 4A; ΔpkaB not shown). However, the majority of the conidia of the alcA(p)::pkaB, ΔpkaA, and ΔpkaA alcA(p)::pkaB mutants did not germinate by 10 h (Fig. 4A), and 80% of the conidia of the three mutants began to produce short germ tubes at 13 h (not shown). These results suggest that overexpression of PkaB negatively affects spore germination.
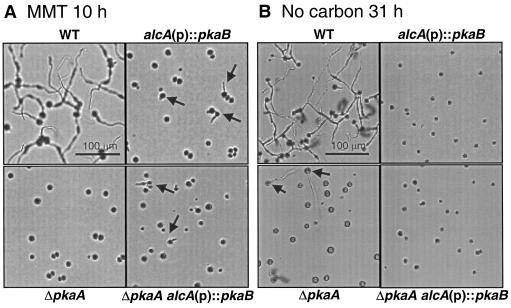
FIG. 4.
Inhibitory roles of pkaB in germination. A. The conidia of wild-type (WT; FGSC26), alcA(p)::pkaB (RNI5.2), ΔpkaA (RSA53.130), and ΔpkaA alcA(p)::pkaB (RNI6) strains were inoculated on solid MMT (inducing medium) and incubated at 37°C for 10 h. Note that while most of the wild-type conidia germinated, only a few mutant conidia germinated. The arrows show the alcA(p)::pkaB mutant conidia forming a germ tube. Although culture conditions were not identical, the pkaB mRNA levels in alcA(p)::pkaB (RNI5.2) and ΔpkaA alcA(p)::pkaB (RNI6) strains grown in liquid MMT for 24 h (see Materials and Methods) were about 20-fold higher than those of the wild type (not shown). B. The conidia of wild-type (WT; FGSC26), alcA(p)::pkaB (RNI5.2), ΔpkaA (RSA53.130), and ΔpkaA alcA(p)::pkaB (RNI6) strains were inoculated on the solid medium lacking a carbon source and incubated at 37°C for 31 h. An electrophoresis-grade agarose was used as a solidifying agent to avoid the introduction of unwanted energy sources. At 31 h, most of the wild-type conidia were able to germinate and about 20% of the ΔpkaA mutant conidia could form germ tubes (arrows). However, upregulation of pkaB caused by alcA(p)::pkaB completely blocked germination of spores regardless of the presence or absence of pkaA. Although culture conditions were different, the pkaB mRNA levels in alcA(p)::pkaB (RNI5.2) and ΔpkaA alcA(p)::pkaB (RNI6) strains incubated in the liquid medium lacking a carbon source at 37°C for 24 h (see Materials and Methods) were about fivefold higher than those of the wild type (data not shown).
Upregulation of pkaB blocks spore germination in the absence of a carbon source.
We then examined germination of the conidia of wild-type, alcA(p)::pkaB, ΔpkaA, and ΔpkaA alcA(p)::pkaB strains on the solid medium without a carbon source. Under these conditions, a few wild-type conidia began to germinate at 10 h of incubation, and most formed elongated thin germ tubes at 31 h (Fig. 4B). However, while conidia of the ΔpkaA mutant exhibited isotropic growth (swelling) and germinated at low levels (∼20%), the alcA(p)::pkaB mutant conidia showed no sign of germination (e.g., swelling of spores) at 31 h (Fig. 4B). Moreover, no conidia of the alcA(p)::pkaB mutant were able to germinate even after 5 days of incubation (not shown). Similar results were obtained when the conidia of the ΔpkaA alcA(p)::pkaB (Fig. 4B) and ΔpkaA ΔpkaB alcA(p)::pkaB (not shown) mutants were tested. Collectively, while both PkaB and PkaA positively function in spore germination in the presence of glucose, PkaB negatively controls conidial germination in the absence of external carbon sources.
Upregulation of pkaB enhances submerged sporulation caused by ΔpkaA.
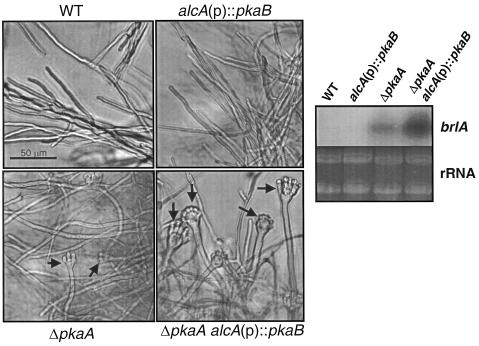
The ΔpkaA mutant was shown to produce conidiophore-like structures in some liquid submerged culture conditions, where wild-type strains do not sporulate (34). To test the effect of ΔpkaB and upregulation of pkaB on sporulation in liquid submerged cultures, wild-type, ΔpkaB, alcA(p)::pkaB, ΔpkaA, ΔpkaA alcA(p)::pkaB, and ΔpkaA ΔpkaB alcA(p)::pkaB strains were cultured in liquid MMG at 37°C, 250 rpm, and the status of development was microscopically observed. While the wild type and the ΔpkaB mutant did not sporulate, the ΔpkaA mutant produced a few conidiophore-like structures at 20 h (Fig. 5). Interestingly, the ΔpkaA alcA(p)::pkaB and ΔpkaA ΔpkaB alcA(p)::pkaB (not shown), but not alcA(p)::pkaB, mutants elaborated near-complete conidiophore-like structures abundantly at 20 h. We then examined the mRNA levels of brlA, encoding a key transcriptional activator for conidiophore development (2). As shown in Fig. 5, while derepression of the brlA expression under submerged culture conditions was caused by both ΔpkaA and ΔpkaA alcA(p)::pkaB, the level of brlA mRNA was much higher in the ΔpkaA alcA(p)::pkaB mutant, indicating that upregulation of pkaB results in elevated derepression of brlA in the absence of pkaA. These results suggest that PkaB may play an opposite role in controlling conidiophore development in liquid submerged culture.
FIG. 5.
Upregulation of pkaB enhances submerged sporulation caused by ΔpkaA. The conidia (108/100 ml) of wild-type (WT; FGSC26), alcA(p)::pkaB (RNI5.2), ΔpkaA (RSA53.130), and ΔpkaA alcA(p)::pkaB (RNI6) strains were inoculated in liquid MMG and incubated at 37°C, 250 rpm, for 20 h. The arrows indicate conidiophore-like structures formed in ΔpkaA and ΔpkaA alcA(p)::pkaB strains. The steady-state mRNA levels of brlA were examined at 26 h of incubation. Transcripts of brlA were detectable in the ΔpkaA and ΔpkaA alcA(p)::pkaB mutants. Note that brlA is highly accumulated by upregulation of pkaB in the absence of pkaA. For the pkaB mRNA levels in individual strains, see Fig. 3A.
Upregulation of pkaB or ΔpkaA reduces the hyphal tolerance to oxidative stress.
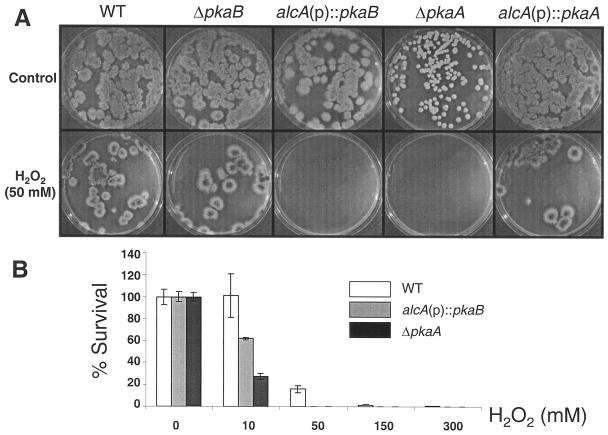
Finally, we examined a potential role of PkaB in the stress response. When the conidia or the 10-h-cultured hyphae of wild-type, ΔpkaB, and alcA(p)::pkaB strains (see Materials and Methods) were tested for thermal tolerance, mutant and wild-type strains exhibited no detectable differences. We then examined the sensitivities of the conidia and the hyphae of wild-type, ΔpkaB, alcA(p)::pkaB, ΔpkaA, and alcA(p)::pkaA strains to oxidative stress. When treated with various concentrations of H2O2 for 30 min, no differences in survival rates between the wild-type and mutant spores were observed. However, survival rates of the hyphae of ΔpkaA and alcA(p)::pkaB [but not alcA(p)::pkaA or ΔpkaB] strains (Fig. 6A) on solid MMG decreased drastically when treated with H2O2. While almost 100% of the wild-type hyphae survived, only 60% of the alcA(p)::pkaB and 30% of the ΔpkaA hyphae formed visible colonies after treatment with 10 mM H2O2 (Fig. 6B). Furthermore, no alcA(p)::pkaB and ΔpkaA colonies survived after treatment with 50 mM or higher concentrations of H2O2 (Fig. 6A and B). This suggests that upregulation of pkaB or the absence of pkaA reduces the tolerance of hyphae to oxidative stress and that PkaB and PkaA play opposite roles in the hyphal tolerance to oxidative stress.
FIG. 6.
Upregulation of pkaB or ΔpkaA reduces hyphal tolerance to oxidative stress. The hyphae of wild-type (WT; FGSC26), ΔpkaB (RSA53.111), alcA(p)::pkaB (RNI5.2), ΔpkaA (RSA53.126), and alcA(p)::pkaA (RNI8) strains grown on solid MMG at 37°C for 26 h were treated with 0, 10, 50, 150, and 300 mM H2O2 for 10 min in triplicate. The plates were then incubated at 37°C for an additional 24 h. A. The colonies that survived through 0 mM (control) or 50 mM H2O2 treatment are shown. Note that no alcA(p)::pkaB or ΔpkaA mutants survived after treatment, whereas the alcA(p)::pkaA and ΔpkaB mutants did not exhibit differences in survival rates compared to the wild type. B. The bar graph shows the relative survival rates of the hyphae of wild-type, alcA(p)::pkaB, and ΔpkaA strains.
DISCUSSION
All cells have the capacity to sense and respond to the environment via signal transduction cascades. Membrane-bound heterotrimeric G proteins (G proteins) play a key role in transduction of the extracellular cues into cells to elicit the appropriate physiological and biochemical responses. In various fungi, the G protein-cAMP-dependent protein kinase (PKA) signaling pathway is a primary mechanism for vegetative growth, morphological switch, nutrient sensing, mating, stress response, secondary metabolism and pathogenicity (reviewed in references 8, 21, and 22). In this study, we have identified and characterized the second PKA catalytic subunit in the model ascomycete, A. nidulans. One of the critical findings in our study is that the absence of both pkaA and pkaB is lethal, which indicates that PkaA and PkaB constitute the sole PKA catalytic subunits essential for growth of A. nidulans. Moreover, our intensive genetic studies unveiled (partially) the complicated functions of this secondary PKA in controlling diverse physiological processes.
Previous studies showed that the cellular PKA activity is largely dependent on class I PKA in various filamentous fungi, such as A. nidulans (34), C. neoformans serotype A (7), M. grisea (1), and A. fumigatus (23). As it plays a predominant role in these fungi, mutational inactivation of class I PKA results in profound effects. However, we found that, compared to those of ascomycete fungi, the two PKAs in basidiomycetes appear to be more closely related (see the distance between Adr1 and Uka1 of U. maydis in Fig. 1D). Our additional phylogenetic analysis employing Pka1 and Pka2 of the A and D serotypes of C. neoformans (17) further confirmed that the two basidiomycete PKAs are somewhat closely related (data not shown, but similar to Adr1 and Uka1 of U. maydis in Fig. 1D) and suggested that the predominant PKA unit may not be clearly distinguished in some basidiomycete fungi. In fact, a recent study showed that the predominant PKA is different depending on the serotype of C. neoformans (17). While loss of Pka1 (similar to Um_Adr1, class I [Fig. 1D]) in serotype A resulted in failure in mating, melanin, and capsule production as well as virulence establishment (7), mutational inactivation of Pka2 (similar to Um_Uka1, class II) in serotype D affected mating, haploid fruiting, and virulence factor (melanin) production (17).
Double deletion analyses have been carried out in two basidiomycete fungi, C. neoformans and U. maydis. Deletion of both pka1 and pka2 in serotype D did not block fungal growth, indicating that PKA is not essential for the viability of C. neoformans (17). Similarly, in U. maydis, while the Δuka1 Δadr1 double mutant formed noticeably less mycelial mass and lower levels of tumor than the adr1 single mutant, the viability remained unaffected, indicating that PKAs are not absolutely required for growth of this fungus (9). On the contrary, our studies evidently demonstrate that mutational inactivation of both PkaB and PkaA is lethal and that PKAs are essential for the viability of A. nidulans. A similar result was obtained in M. grisea, where the double mutant of cpkA and cpk2 is lethal (J. R. Xu, personal communication). This is somewhat discordant to the previous findings that the A. nidulans adenylate cyclase, CyaA, is not essential for fungal growth (10). As a possible explanation, while PKA catalytic subunits are generally activated by cAMP binding to the PKA regulatory subunit (PkaR; C. d'Enfert, unpublished data), PKA can be activated by other mechanisms in addition to CyaA-cAMP signaling, and/or PKAs may maintain a basal level activity even in the absence of cAMP.
The fact that deletion of both pkaB and pkaA is lethal implies that PkaB plays a role in fungal growth and/or spore germination. Similar to overexpression of pkaA (34), overexpression of pkaB enhanced hyphal proliferation and reduced conidiation in air-exposed solid MMT, indicating that PkaB and PkaA have an overlapping role in vegetative growth and developmental regulation. Moreover, overexpression or upregulation of pkaB could partially rescue the growth and germination defects caused by ΔpkaA, suggesting that PkaB can at least partially replace PkaA functions in certain physiological processes. Vegetative growth signaling mediated by FadA (Gα)-FlbA (RGS) is in part transduced by PkaA (34). Deletion of pkaB could not suppress fluffy-autolytic phenotypes caused by the absence of FlbA. This finding suggests that either PkaB functions in the FadA-mediated signaling pathway but its role in vegetative growth signaling is minute, or that PkaB primarily functions in a separate signaling branch (Fig. 7; see below).
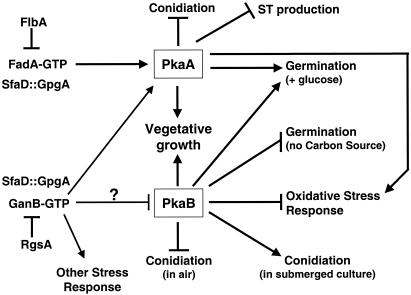
FIG. 7.
Proposed model for PKA-mediated signaling in A. nidulans. We propose that the two PKAs are differently regulated by G protein signaling and that overlapping and opposite roles of PkaA (primary PKA) and PkaB (secondary PKA) control various biological processes. FlbA-FadA (Gα)-controlled vegetative growth signaling is in part transduced by PkaA, which in turn represses conidiation and ST production (34). The demonstration that GanB and SfaD::GpgA, as well as PkaA, are required for proper germination of spores (10, 19, 30, 33) indicates that PkaA may be activated by both GanB and FadA. RgsA-controlled GanB signaling is also associated with activation of the stress response (13). The fact that ΔpkaA delayed conidial germination and reduced hyphal tolerance to oxidative stress further supports the idea that PkaA may be activated by GanB. While PkaB can stimulate vegetative growth and activate conidial germination in the presence of glucose, upregulation of pkaB blocks germination of spores in the absence of a carbon source and reduces hyphal tolerance to oxidative stress. This suggests that the resulting cellular responses for PkaB and GanB-PkaA signaling may be opposite. Thus, if PkaB functions downstream of GanB, the activity of PkaB may be negatively regulated by GanB. GanB-mediated signaling may activate other stress responses via a different (possibly a mitogen-activated protein kinase) downstream signaling cascade.
Spore germination is the first crucial step for fungal growth and colony formation. At least two signaling pathways (G protein-cAMP and Ras) are associated with controlling spore germination in A. nidulans (10). Fillinger et al. first reported that cAMP signaling controls the early events of conidial germination in response to external carbon sources (10). Disruption of cyaA and pkaA caused delayed spore germination, and the double deletion mutant exhibited more pronounced delay than each single mutant. Because no cAMP was detectable in the ΔcyaA mutant, their results support the idea that PkaA (and probably PkaB) in A. nidulans may retain some activity in the absence of cAMP and that additional signaling pathways, including RasA, are involved in the spore germination process (10). Our findings indicate that two PKAs play overlapping roles in spore germination in the presence of glucose. However, in contrast, overexpression of pkaB or ΔpkaA delayed germination on solid MMT [inducing conditions for alcA(p)::pkaB]. Furthermore, upregulation of pkaB completely blocked spore germination on the medium without an external carbon source (Fig. 4B). These results suggest that, whereas PkaB can activate germination when glucose (a favorable carbon source) is present, one of its important regulatory roles might be to negatively control spore germination under unfavorable conditions, such as a lack of external energy sources. Balanced activities of two PKAs may play an important role in regulating proper spore germination.
Besides its functions in controlling fungal growth and nutrient sensing, cAMP-PKA has been found to be involved in stress responses in fungi. In S. cerevisiae, the G-protein-coupled receptor Gpr1 and the Gα subunit Gpa2 stimulate cAMP production by adenylyl cyclase in response to glucose and other fermentable carbon sources, thereby activating TPKs (29). This Gpr1-Gpa2-cAMP-PKA signaling is negatively regulated by Rgs2 (an RGS protein [41]). Activities of TPKs antagonize the induction of the general stress response in S. cerevisiae, which is mediated by Msn2p and Msn4p, positive transcriptional factors for the stress response pathway (35). Deletion of RGS2 causes prolonged activation of Gpa2-PKA and thereby increases the sensitivity of cells to thermal stress, while overexpression of RGS2 causes significant elevation in heat resistance (41). Recent studies showed that GanB (a Gα similar to Gpa2 [4]) functions in sensing external carbon sources and controlling spore germination via a cAMP/PKA pathway in A. nidulans, where SfaD-GpgA (Gβγ) is predicted to function in activating GanB (19, 30, 33). Furthermore, we demonstrated that GanB signaling activates thermal and oxidative stress responses, which are negatively controlled by RgsA (an RGS protein similar to Rgs2 in yeast [13]). Contrary to the yeast stress response, deletion of rgsA results in the elevated thermal and oxidative stress tolerances of both hyphae and spores (13). In this study we demonstrated that, while no pkaB mutations (deletion and/or overexpression) affected thermal tolerance of hyphae and spores, or tolerance of spores against oxidative stress, ΔpkaA or alcA(p)::pkaB resulted in significantly reduced tolerance of hyphae to oxidative stress. Our findings indicate that, while RgsA-GanB activities may regulate the general stress response in A. nidulans, the PKA-mediated stress response might be limited to oxidative stress and that PkaA and PkaB play opposite roles in conferring hyphal tolerance to oxidative stress.
Deletion or dominant-negative mutation of ganB as well as deletion of pkaA result in formation of conidiophore-like structures in liquid submerged culture (4, 34). We found that, while overexpression of pkaB alone did not cause sporulation in liquid submerged culture by itself, it enhanced the submerged sporulation phenotype caused by ΔpkaA. Collectively, based on the overlapping and opposite roles of the two PKAs in various physiological processes and the role of GanB-cAMP-PKA signaling in controlling spore germination, stress response, and asexual development, it can be speculated that PkaA and PkaB function downstream of GanB. If this were the case, GanB signaling might stimulate PkaA activity while inhibiting PkaB function. Currently available information is integrated and summarized in the model presented in Fig. 7. In this model, it is proposed that both FadA and GanB activate PkaA. PkaA functions as the predominant unit for vegetative growth, germination, and regulation of asexual development and ST production. PkaB is the secondary PKA and is likely inhibited by GanB signaling, whereas overexpression and/or upregulation of pkaB may overcome the GanB-mediated repressive effects. Understanding the exact functions and the genetic position of PkaB will require further studies.
Acknowledgments
We are grateful to the Broad Institute for the genome database, Kap-Hoon Han for helpful discussion, Nancy Keller for providing pkaA mutant strains, and Ann Larson and Ellin Doyle for critically reviewing the manuscript.
This work was supported by a Hatch grant and National Science Foundation grant MCB-0421863 to J.H.Y.
REFERENCES
- 1.Adachi, K., and J. E. Hamer. 1998. Divergent cAMP signaling pathways regulate growth and pathogenesis in the rice blast fungus Magnaporthe grisea. Plant Cell 10:1361-1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Adams, T. H., M. T. Boylan, and W. E. Timberlake. 1988. brlA is necessary and sufficient to direct conidiophore development in Aspergillus nidulans. Cell 54:353-362. [DOI] [PubMed] [Google Scholar]
- 3.Bockmühl, D. P., S. Krishnamurthy, M. Geards, A. Sonneborn, and J. F. Ernst. 2001. Distinct and redundant roles of the two protein kinase A isoforms Tpk1P and Tpk2P in morphogenesis and growth of Candida albicans. Mol. Microbiol. 42:1243-1257. [DOI] [PubMed] [Google Scholar]
- 4.Chang, M.-H., K.-S. Chae, D.-M. Han, and K.-Y. Jahng. 2004. The GanB Gα-protein negatively regulates asexual sporulation and plays a positive role in conidial germination in Aspergillus nidulans. Genetics 167:1305-1315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chenna, R., H. Sugawara, T. Koike, R. Lopez, T. J. Gibson, D. G. Higgins, and J. D. Thompson. 2003. Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 31:3497-3500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Colledge, M., and J. D. Scott. 1999. AKAPs: from structure to function. Trends Cell Biol. 9:216-221. [DOI] [PubMed] [Google Scholar]
- 7.D'Souza, C. A., J. A. Alspaugh, C. Yue, T. Harashima, G. M. Cox, J. R. Perfect, and J. Heitman. 2001. cAMP-dependent protein kinase controls mating and virulence of the fungal pathogen Cryptococcus neoformans. Mol. Cell. Biol. 21:3179-3191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.D'Souza, C. A., and J. Heitman. 2001. Conserved cAMP signaling cascades regulate fungal development and virulence. FEMS Microbiol. Rev. 25:349-364. [DOI] [PubMed] [Google Scholar]
- 9.Dürrenberger, F., K. Wong, and W. J. Kronstad. 1998. Identification of a cAMP-dependent protein kinase catalytic subunit required for virulence and morphogenesis in Ustilago maydis. Proc. Natl. Acad. Sci. USA 95:5684-5689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Fillinger, S., M. K. Chaveroche, K. Shimizu, N. P. Keller, and C. d'Enfert. 2002. cAMP and ras signalling independently control spore germination in the filamentous fungus Aspergillus nidulans. Mol. Microbiol. 44:1001-1016. [DOI] [PubMed] [Google Scholar]
- 11.Griffioen, G., and J. M. Thevelein. 2002. Molecular mechanisms controlling localization of protein kinase A. Curr. Genet. 41:199-207. [DOI] [PubMed] [Google Scholar]
- 12.Han, K.-H., J.-A. Seo, and J.-H. Yu. 2004. A putative G protein-coupled receptor negatively controls sexual development in Aspergillus nidulans. Mol. Microbiol. 51:1333-1345. [DOI] [PubMed] [Google Scholar]
- 13.Han, K.-H., J.-A. Seo, and J.-H. Yu. 2004. Regulators of G-protein signaling in Aspergillus nidulans: RgsA downregulates stress response and stimulate asexual sporulation through attenuation of GanB (Gα) signaling. Mol. Microbiol. 53:529-540. [DOI] [PubMed] [Google Scholar]
- 14.Hanks, S. K., and T. Hunter. 1995. Protein kinases 6. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification. FASEB J. 9:576-596. [PubMed] [Google Scholar]
- 15.Hatanaka, M., and C. Shimoda. 2001. The cyclic AMP/PKA signal pathway is required for initiation of spore germination in Schizosaccharomyces pombe. Yeast 18:207-217. [DOI] [PubMed] [Google Scholar]
- 16.Hicks, J. K., J.-H. Yu, N. P. Keller, and T. H. Adams. 1997. Aspergillus sporulation and mycotoxin production both require inactivation of the FadA Gα protein-dependent signaling pathway. EMBO J. 16:4916-4923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hicks, J. K., C. A. D'Souza, G. M. Cox, and J. Heitman. 2004. Cyclic AMP-dependent protein kinase catalytic subunits have divergent roles in virulence factor production in two varieties of the fungal pathogen Cryptococcus neoformans. Eukaryot. Cell 3:14-26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Käfer, E. 1977. Meiotic and mitotic recombination in Aspergillus and its chromosomal aberrations. Adv. Genet. 19:33-131. [DOI] [PubMed] [Google Scholar]
- 19.Lafon, A., J.-A. Seo, K.-H. Han, J.-H. Yu, and C. d'Enfert. The heterotrimeric G-protein GanB(α)-SfaD(β)-GpgA(γ) is a carbon source sensor involved in early cAMP-dependent germination in Aspergillus nidulans. Genetics, in press. [DOI] [PMC free article] [PubMed]
- 20.Lee, B. N., and T. H. Adams. 1994. Overexpression of flbA, an early regulator of Aspergillus asexual sporulation, leads to activation of brlA and premature initiation of development. Mol. Microbiol. 14:323-334. [DOI] [PubMed] [Google Scholar]
- 21.Lee, N., C. A. D'Souza, and J. W. Kronstad. 2003. Of smuts, blasts, mildews and blights: cAMP signaling in phytopathogenic fungi. Annu. Rev. Phytopathol. 41:399-427. [DOI] [PubMed] [Google Scholar]
- 22.Lengeler, K. B., R. C. Davidson, C. A. D'Souza, T. Harashima, W.-C. Shen, P. Wang, X.-W. Pan, M. Waugh, and J. Heitman. 2000. Signal transduction cascades regulating fungal development and virulence. Microbiol. Mol. Biol. Rev. 64:746-785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Liebmann, B., M. Muller, A. Braun, and A. A. Brakhage. 2004. The cyclic AMP-dependent protein kinase a network regulates development and virulence in Aspergillus fumigatus. Infect. Immun. 72:5193-5203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Maeda, T., Y. Watanabe, H. Kunitomo, and M. Yamamoto. 1994. Cloning of the pka1 gene encoding the catalytic subunit of the cAMP-dependent protein kinase in Schizosaccharomyces pombe. J. Biol. Chem. 269:9632-9637. [PubMed] [Google Scholar]
- 25.Mitchell, T. K., and R. A. Dean. 1995. The cAMP-dependent protein kinase catalytic subunit is required for appressorium formation and pathogenesis by the rice blast pathogen Magnaporthe grisea. Plant Cell 7:1869-1878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Pan, X., and J. Heitman. 1999. Cyclic AMP-dependent protein kinase regulates pseudohyphal differentiation in Saccharomyces cerevisiae. Mol. Cell. Biol. 19:5352-5356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pontecorvo, G., J. A. Roper, L. M. Hemmons, K. D. Macdonald, and A. W. J. Bufton. 1953. The genetics of Aspergillus nidulans. Adv. Genet. 5:141-238. [DOI] [PubMed] [Google Scholar]
- 28.Robertson, L. S., and G. R. Fink. 1998. The three yeast A kinases have specific signaling functions in pseudohyphal growth. Proc. Natl. Acad. Sci. USA 95:13783-13787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Rolland, F., J. H. De Winde, K. Lemaire, E. Boles, J. M. Thevelein, and J. Winderickx. 2000. Glucose-induced cAMP signalling in yeast requires both a G-protein coupled receptor system for extracellular glucose detection and a separable hexose kinase-dependent sensing process. Mol. Microbiol. 38:348-358. [DOI] [PubMed] [Google Scholar]
- 30.Rosén, S., J.-H. Yu, and T. H. Adams. 1999. The Aspergillus nidulans sfaD gene encodes a G protein β subunit that is required for normal growth and repression of sporulation. EMBO J. 18:5592-5600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Roosen, J., K. Engelen, K. Marchal, J. Mathys, G. Griffioen, E. Cameroni, J. M. Thevelein, C. De Virgilio, B. De Moor, and J. Winderickx. 2005. PKA and Sch9 control a molecular switch important for the proper adaptation to nutrient availability. Mol. Microbiol. 55:862-880. [DOI] [PubMed] [Google Scholar]
- 32.Seo, J.-A., Y. Guan, and J.-H. Yu. 2003. Suppressor mutations bypass the requirement of fluG for asexual sporulation and sterigmatocystin production in Aspergillus nidulans. Genetics 165:1083-1093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Seo, J.-A., K.-H. Han, and J.-H. Yu. Multiple roles of a heterotrimeric G protein γ subunit in governing growth and development of Aspergillus nidulans. Genetics, in press. [DOI] [PMC free article] [PubMed]
- 34.Shimizu, K., and N. P. Keller. 2001. Genetic involvement of a cAMP-dependent protein kinase in a G protein signaling pathway regulating morphological and chemical transitions in Aspergillus nidulans. Genetics 157:591-600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Smith, A., P. W. Mary., and S. Garrett. 1998. Yeast PKA represses Msn2p/Msn4p-dependent gene expression to regulate growth, stress response and glycogen accumulation. EMBO J. 17:3556-3564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Taylor, S. S., J. A. Buechler, and W. Yonemoto. 1990. cAMP-dependent protein kinase: framework for a diverse family of regulatory enzymes. Annu. Rev. Biochem. 59:971-1005. [DOI] [PubMed] [Google Scholar]
- 37.Taylor, S. S., and E. Radzio-Andzelm. 1994. Three protein kinase structures define a common motif. Structure 2:345-355. [DOI] [PubMed] [Google Scholar]
- 38.Taylor, S. S., J. Yang, J. Wu, N. M. Haste, E. Radzio-Andzelm, and G. Anand. 2004. PKA: a portrait of protein kinase dynamics. Biochimica. Biophysica. Acta 1697:259-269. [DOI] [PubMed] [Google Scholar]
- 39.Toda, T., S. Cameron, P. S. Sass, M. P. Zoller, and M. Wigler. 1987. Three different genes in S. cerevisiae encode the catalytic subunits of the cAMP-dependent protein kinase. Cell 50:277-287. [DOI] [PubMed] [Google Scholar]
- 40.Toda, T., S. Cameron, P. Sass, and M. Wigler. 1988. SCH9, a gene of Saccharomyces cerevisiae that encodes a protein distinct from, but functionally and structurally related to cAMP-dependent protein kinase catalytic subunits. Genes Dev. 2:517-527. [DOI] [PubMed] [Google Scholar]
- 41.Versele, M., J. H. de Winde, and J. M. Thevelein. 1999. A novel regulator of G protein signalling in yeast, Rgs2, downregulates glucose-activation of the cAMP pathway through direct inhibition of Gpa2. EMBO J. 18:5577-5591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wieser, J., and T. H. Adams. 1995. flbD encodes a Myb-like DNA-binding protein that coordinates initiation of Aspergillus conidiophore development. Genes Dev. 9:491-502. [DOI] [PubMed] [Google Scholar]
- 43.Xu, J. R., M. Urban, J. A. Sweigard, and J. E. Hamer. 1997. The CPKA gene of Magnaporthe grisea is essential for appressorial penetration. Mol. Plant Microbe. Interact. 10:187-194. [Google Scholar]
- 44.Yu, J.-H., J. Wieser, and T. H. Adams. 1996. The Aspergillus FlbA RGS domain protein antagonizes G-protein signaling to block proliferation and allow development. EMBO J. 15:5184-5190. [PMC free article] [PubMed] [Google Scholar]
- 45.Yu, J.-H., S. Rosèn, and T. H. Adams. 1999. Extragenic suppressors of loss-of-function mutations in the Aspergillus FlbA regulator of G-protein signaling domain protein. Genetics 151:97-105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yu, J.-H., Z. Hamari, K.-H. Han, J.-A. Seo, Y. Reyes-Domínguez, and C. Scazzochio. 2004. Double-joint PCR: a PCR-based molecular tool for gene manipulation in filamentous fungi. Fungal Gen. Biol. 41:973-981. [DOI] [PubMed] [Google Scholar]