Abstract
The cardioprotective effects of histone deacetylase (HDAC) inhibitors (HDIs) are at odds with the deleterious effects of HDAC depletion. Here, we use HDAC3 as a prototype HDAC to address this contradiction. We show that adult-onset cardiac-specific depletion of HDAC3 in mice causes cardiac hypertrophy and contractile dysfunction on a high-fat diet (HFD), excluding developmental disruption as a major reason for the contradiction. Genetically abolishing HDAC3 enzymatic activity without affecting its protein level does not cause cardiac dysfunction on HFD. HDAC3 depletion causes robust downregulation of lipid oxidation/bioenergetic genes and upregulation of antioxidant/anti-apoptotic genes. In contrast, HDAC3 enzyme activity abolishment causes much milder changes in far fewer genes. The abnormal gene expression is cardiomyocyte-autonomous and can be rescued by an enzyme-dead HDAC3 mutant but not by an HDAC3 mutant (Δ33–70) that lacks interaction with the nuclear-envelope protein lamina-associated polypeptide 2β (LAP2β). Tethering LAP2β to the HDAC3 Δ33–70 mutant restored its ability to rescue gene expression. Finally, HDAC3 depletion, not loss of HDAC3 enzymatic activity, exacerbates cardiac contractile functions upon aortic constriction. These results suggest that the cardiac function of HDAC3 in adults is not attributable to its enzyme activity, which has implications for understanding the cardioprotective effects of HDIs.
Key words: Histone deacetylase, Cardiac hypertrophy, Hypertrophic cardiomyopathy, HDACi, Catalytic-independent, Fatty acid oxidation, Pressure overload, Lamina
Graphical abstract
This study suggests that the cardiac function of HDAC3 in adults is not attributable to its enzyme activity, which has implications for understanding the cardioprotective effects of histone deacetylase inhibitors.

1. Introduction
Histone deacetylase (HDAC) inhibitors (HDIs) are cardioprotective in a growing list of preclinical animal models. HDIs that inhibit all classical zinc-dependent HDACs are known as pan-HDIs. Pan-HDI trichostatin A (TSA) attenuates cardiac hypertrophy and preserves contractile functions in transverse aortic constriction (TAC) or angiotensin II-induced hypertrophy animal models1, 2, 3. TSA is also protective against myocardial infarction-induced contractile dysfunction4,5. Suberoylanilide hydroxamic acid (SAHA), another pan-HDI, reduces myocardial fibrosis in the TAC model6 and attenuates myocardial injury in the isoproterenol model7. Givinostat (also known as ITF2357), another pan-HDI, improves diastolic function in the uninephrectomy/deoxycorticosterone acetate-induced diastolic dysfunction model and the heart failure with preserved ejection fraction model in Dahl salt-sensitive rats8,9. Mocetinostat, a benzamide-based inhibitor of Class I HDACs, improves contractile functions after myocardial infarction10 and attenuates cardiac hypertrophy in the TAC model11. However, the mechanism underlying HDIs-mediated cardiac benefits is not completely understood.
Opposite to the beneficial effects of HDIs in the heart, genetic mouse models with the whole-body or cardiac-specific knockout of HDACs often show cardiac defects12. The 11 classical zinc-dependent HDACs in mammals can be grouped into classes depending on their sequence homology: Class I HDACs (HDAC1, HDAC2, HDAC3, and HDAC8); Class IIa HDACs (HDAC4, HDAC5, HDAC7, and HDAC9); Class IIb HDAC (HDAC6 and HDAC10); and Class V HDAC (HDAC11). Knockout of both HDAC1 and HDAC2 in the heart results in neonatal lethality, accompanied by cardiac arrhythmias and dilated cardiomyopathy, while deletion of HDAC1 or HDAC2 individually does not lead to obvious phenotype13. Knockout of HDAC3 caused cardiac hypertrophy and affected lipid metabolism14,15. HDAC3 is also essential for epicardial development16. The mice with inactivated HDAC5 or HDAC9 were sensitive to stress signals, such as pressure overload and calcineurin stimulation17,18.
Thus, the beneficial effects of HDIs in preclinical heart disease models are at odds with the deleterious effects of HDAC depletion in genetic mouse models. We chose to focus on HDAC3 to address this question for several reasons. (1) HDAC3 is responsible for the enzyme activity of Class IIa HDACs. Class IIa HDACs have low intrinsic HDAC enzymatic activities due to a histidine substitution on the key catalytic tyrosine site19. As a result, most catalytic activity of Class IIa HDACs is attributed to HDAC3, which exists in the same multiprotein corepressor complexes20. (2) HDAC3 has strong enzyme activity and is considered a primary target of many HDIs. (3) Knockout of HDAC3 from different developmental stages generates different cardiac phenotypes. Knockout of HDAC3 in the heart at around embryonic day 9.5 using the α-myosin heavy chain (α-MHC)-Cre driver caused cardiac hypertrophy and lethal heart failure by the age of 3–4 months on a normal chow diet14. In contrast, knockout of HDAC3 in the heart postnatally using the muscle creatine kinase (MCK)-Cre driver does not have drastic cardiac phenotype on normal chow diet even at the age of over a year, and only caused cardiac hypertrophy and lethal heart failure when mice were fed with high-fat diet (HFD)15. These results suggest the developmental effects could account for the HDIs vs. HDAC deletion differences.
The enzyme activity of HDAC3 relies on forming a stable protein complex with nuclear receptor corepressor (NCoR or NCOR1) or silencing mediator for retinoid and thyroid receptor (SMRT or NCOR2). The deacetylation activation domain (DAD) in NCoR/SMRT binds to HDAC3 and causes a conformational change of HDAC3 protein, which makes the catalytic channel accessible to the substrate21,22. Purified HDAC3 protein does not show much enzyme activity by itself but gains robust enzyme activity after adding in purified DAD22,23. Conversely, whole-body knock-in of missense mutations in the DAD of NCoR/SMRT in mice (NS-DADm mice) abolishes the enzyme activity of HDAC3 without affecting its protein levels in multiple tissues24. Here, we address the function of HDAC3 in the adult heart and its functional reliance on enzyme activity.
2. Methods
2.1. Animals
Hdac3loxP/loxP/αMHC-MerCreMer mice were generated from crossing Hdac3 floxed mice (HDAC3loxP/loxP)15 with cardiac-specific tamoxifen-inducible MerCreMer driver mice (αMHC-MerCreMer, JAX #005657)25 (referred to as “iKO”), Hdac3loxP/loxP/MCK-Cre (KO) mice, and NS-DADm mice were previously described15,26,27. All mice were on the C57BL/6J genetic background. Male mice at the age of 3–4 months were used for echocardiography and histology experiments. Tamoxifen was dissolved in corn oil and injected intraperitoneally (20 mg/kg per day) for 5 consecutive days. The high-fat diet (HFD) containing 60 kcal % fat was purchased from Research Diets Inc (New Brunswick, NJ, USA). For euthanization, mice were placed in a chamber where carbon dioxide (CO2) was gradually introduced until the mice were unconscious and subsequently confirmed dead. To ensure animal welfare, we followed best practices for CO2 administration, including a gradual fill rate (≤30%–50% chamber volume per minute) to minimize distress, as recommended by the American Veterinary Medical Association (AVMA) Guidelines for the Euthanasia of Animals. During the euthanasia process, experimental animals did not exhibit pulmonary hemorrhage or other symptoms that affect animal welfare. The animal care and procedures were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) at the Baylor College of Medicine (AN-6583) and conformed to the NIH Guide for the Care and Use of Laboratory Animals.
2.2. TAC surgery
The chronic pressure overload model was induced by transverse aortic constriction (TAC) performed on 8-week-old male mice following the established procedure28. After anesthetization with intraperitoneal injection of a mixture of ketamine (100 mg/kg) and xylazine (20 mg/kg), mice were intubated and placed on a respirator. Midline sternotomy was performed, and the aorta constricted at the mid-aortic arch level with a 6/0 braided silk suture using a blunted 27.5-gauge needle as a calibrator. Sham control mice underwent the same surgical procedures without constriction of the aorta.
2.3. Echocardiography
Non-invasive transthoracic echocardiography was performed using a VisualSonics Vevo 2100 system (Fujifilm VisualSonics, Inc., Toronto, Canada). Two-dimensional images were obtained at 2.5–3 frames/s using a 15 MHz probe from Fujifilm (RMV 707B, Visual Sonics) in the parasternal short-axis views to guide M-mode analysis at the midventricular level. Anesthesia was induced by 2.5% isoflurane and confirmed by a lack of response to firm pressure on one of the hind paws. During echocardiogram acquisition, isoflurane was adjusted to 1.5% to maintain a heart rate of 400–460 beats per minute. Parameters collected include: heart rate, left ventricular end-diastolic internal diameter (LVID; d), left ventricular end-systolic internal diameter (LVID; s), left ventricular end-diastolic anterior wall thickness (LVAW; d), left ventricular end-systolic anterior wall thickness (LVAW; s), left ventricular end-diastolic posterior wall thickness (LVPW; d), left ventricular end-systolic posterior wall thickness (LVPW; s), left ventricular ejection fraction (LVEF), left ventricular fractional shorting (LVFS). At the end of the procedures, all mice recovered from anesthesia without difficulties. We consistently performed left ventricular short-axis M-mode scans at the level of the bi-papillary muscles to assess LV systolic function, as well as to measure dimensions and wall thickness. To ensure the reliability of our measurements and address potential variability, we evaluated both inter-observer and intra-observer variability for all indices. This evaluation involved a blinded analysis of three randomly selected echocardiographic studies. The same observer analyzed the data on two separate occasions to assess intra-observer variability, while two independent observers analyzed the data to assess inter-observer variability. Indices of cardiac function were obtained from short-axis M-mode scans at the midventricular level, as indicated by the presence of papillary muscles in anesthetized mice. We calculated the ejection fraction (EF) using an M-mode echocardiographic image using Eq. (1):
| (1) |
where LV volume is calculated from the M mode with (7.0/(2.4 + LVID)) × LVID3 according to the operator manual of VisualSonics Vevo 2100 Imaging System.
2.4. Histology and image analysis
For Masson's trichrome staining, hearts were collected fixed in 4% paraformaldehyde overnight, dehydrated, paraffin-embedded, and prepared in 5-μm sections. Staining was performed according to standard procedures by the Neuropathology Core at Baylor College of Medicine. Wheat germ agglutinin (WGA) staining was performed on paraffin-embedded cross-sections of left ventricles, using tetramethylrhodamine isothiocyanate-conjugated wheat germ agglutinin (20 g/mL in PBS) (Sigma–Aldrich, Inc., St Louis, MO, USA), which was used to measure the cross-sectional area of cardiomyocytes. Cardiomyocyte diameter and fibrosis area were visualized with a Leica DMi8 automated fluorescence microscope (Leica Microsystems Inc., Deerfield, IL, USA) and quantified, in a blinded manner, using ImageJ software version 2.0, with five microscopic fields per heart.
2.5. Recombinant DNA and virus
Construction of Ad.U6.sgHDAC3.EF1.Cas9 was previously described29. The sequence of sgHDAC3 is GTAGAAATACGCCACGGTCT. pFUGW lentiviral plasmids expressing Flag-tagged HDAC3-WT, Y298H, Δ33–70, WT-LAP2β were generously provided by Dr. Rajan Jain (University of Pennsylvania). cDNA of wild type and mutant HDAC3 were PCR amplified from corresponding Lentivirus vectors and cloned into sites between T7 promoter and bGH poly (A) signal of pENTR-CMV (Addgene, Waltham, MA, USA) plasmids. pENTR-CMV-LAP2β-HA was generated by overlap extension PCR according to standard protocols. pAd plasmids and recombinant adenovirus vectors were then produced using the ViraPower adenoviral expression system (ThermoFisher Scientific, Waltham, MA, USA).
2.6. Cell culture, transfection, infection, and isotope tracing
AC16 cells were cultured in Dulbecco's modified Eagle's medium/Nutrient Mixture F-12 (DMEM/F12) (ThermoFisher Scientific) containing 10% fetal bovine serum (FBS), 100 U/mL penicillin, 100 ug/mL streptomycin, and plated on 100 mm cell culture dish at 37 °C under 5% CO2 in a humidified incubator. Human induced pluripotent stem cells (iPSCs) were maintained in mTeSR1 medium (STEMCELL Technologies Canada Inc., Vancouver, Canada) on Matrigel-coated cell culture plates (Corning, #354248; Corning, #3506). Cells were utilized between passages 20 and 80 and passaged every 3–5 days at a confluency of 85%–100% using Accutase (STEMCELL Technologies Canada Inc., Vancouver, Canada). The cardiac differentiation protocol was performed as previously described30. For adenovirus infection, AC16 cells were seeded into 6-well plates prior to infection. Adenoviruses were infected into cells when they reached 60%–70% confluence. Cell culture media was changed after 24 h. AC16 cells were infected for 48–72 h before all experiments. Control viruses are Ad-U6-GFP adenovirus with the same vector of Ad-U6-sgHDAC3. For drug treatment, cells were preincubated with suberoylanilide hydroxamic acid (SAHA) (Caymanchem, Ann Arbor, MI, USA) or Entinostat (MS275) (AdooQ Bioscience, Barranca Parkway, Canada) for 48 h at different dosages before associated experiments. HEK293 cells were cultured in DMEM with high glucose (Avantor, Inc., Radnor, PA, USA) containing 10% FBS, 100 U/mL penicillin, 100 μg/mL streptomycin, and plated on a 100 mm cell culture dish at 37 °C under 5% CO2 in humidified incubator. For transfections, HEK293 cells were seeded into 6-well plates prior to transfection. Plasmids were co-transfected into cells when they reached 60%–70% confluence with jetPRIME in vitro DNA transfection reagent (Avantor, Inc., Radnor, PA, USA) according to the manufacturer's instruction. LAP2β and HDAC3 plasmids were mixed at a 1:1 M ratio for transfection. The medium was changed after 4 h. After transfection for 48 h, samples were collected to perform co-immunoprecipitation. For fatty acids oxidation, AC16 cells were seeded into 6-well plates to 80%–90% confluence and were incubated in PBS with BSA-conjugated [9,10-3H(N)]-palmitate and carnitine for 2 h at 37 °C in the incubator. The resultant 3H2O in the incubation solution was separated from precursors using DOWEX 1X4-400 ion-exchange columns (Sigma–Aldrich Inc., St Louis, MO, USA) and was measured by a scintillation counter29. For non-radioactive FAO assay using a fluorescence-based fatty acid oxidation assay kit (Abcam Inc., Waltham, MA, USA), AC16 cells were seeded in a Matrigel-coated Costar 96 well assay plates (black wall with clear flat bottom; Corning Optical Communications Inc., Fort Worth, TX, USA) at a density of 5 × 104 cells/well in 200 μL culture medium and allowed to adhere to the plate for 24 h. Infections were similar to the above. The assay was performed according to the manufacturer's protocol for the baseline activity with oleate conjugated with BSA as the substrate. The oxygen consumption rate was measured with excitation 380 nm and emission 650 nm and was expressed as the initial rate of increase in the fluorescent signal, as recommended by the manufacturer.
2.7. Immunoprecipitation, HDAC assay, and Western blot
For immunoprecipitation, cells were lysed in the RIPA buffer containing 0.1% SDS, 1% NP40, phosphatase inhibitors, and protease inhibitors. Lysates were precleared twice for 1 h each with Protein G dynabeads (ThermoFisher Scientific) and immunoprecipitated overnight at 4° with Protein G dynabeads with anti-Flag M2 Affinity Gel (Sigma–Aldrich) or IgG (Santa Cruz Biotechnology Inc., Dallas, TX, USA). Beads were collected, washed 3 times in lysis buffer, and eluted into sample buffer (Bio-Rad Laboratories Inc., Hercules, CA, USA) containing 0.1 mol/L DTT and 10% 2-mercaptoethanol (Sigma–Aldrich). Samples were boiled for 10 min and analyzed by Western blot. For the HDAC assay, heart tissues were lysed in RIPA lysis buffer containing 0.1% SDS, 1% NP40, 0.5% sodium deoxycholate, and phosphatase/protease inhibitors. An equal amount of total protein from each sample was subjected to immunoprecipitation with HDAC3 antibodies (Abcam) followed by protein A agarose beads (ThermoFisher Scientific). After washing with lysis buffer, the beads were dried using an insulin syringe and mixed with the working solution containing a fluorescence-tagged acetylated peptide from the HDAC assay kit (Active Motif Inc., Carlsbad, Canada). The reaction was allowed to last for 40 min before quenching with the developing solution containing HDAC inhibitors, followed by fluorescence measurement in a plate reader. For Western blot, protein lysates from different resources were resolved by Tris-glycine SDS-PAGE, transferred to PVDF membrane, and blotted with antibodies against HDAC3 (Abcam), Histone H3 (Abcam), Histone H3 acetyl K27 (H3K27ac) (Abcam), NCOR127, TBLR1 (Novus Biologicals Inc., Centennial, Colorado, USA), HA (Abcam), GAPDH (Cell Signaling Technology Inc., Danvers, MA, USA). Images were acquired using LumiQuant AC600 (Acuronbio Technology Inc., Houston, TX, USA).
2.8. RT-qPCR, RNA-seq, and data processing
For RT-qPCR, total RNA was extracted using TRIzol (Sigma–Aldrich) and RNeasy Mini Kit (Qiagen, Germantown, MD, USA). Reverse transcription and quantitative PCR were performed with the High Capacity RT kit, SYBR Green PCR Master Mix, and the Quant Studio 6 instrument (Life Science, Waltham, MA, USA) using the relative quantification method with standard curves. 18S RNA was used as the housekeeping denominator. mRNA expression levels are shown relative to WT mice or Sham-treated cells. RNA-seq was performed using total RNA (n = 3 in each group). Sequencing libraries were run on the BGI MGISEQ-2000 platform to an average depth of 60 million reads per sample. The sequencing data was filtered with SOAPnuke (v1.5.2) by removing reads containing the sequencing adapter. The resultant clean reads were obtained and stored in FASTQ format and mapped to the reference genome GRCm38.p6 using HISAT2 (v2.0.4). Bowtie2 (v2.2.5) was applied to align the clean reads to the reference coding gene set, and the expression level of genes was calculated with RSEM (v1.2.12). Differential expression analysis was performed using DESeq2 (v1.4.5). A gene was considered differentially expressed if the adjusted P-value was <0.05. We carried out functional annotation analysis using DAVID Bioinformatics Resources 6.7. Differentially expressed genes were used as input gene lists, and all genes expressed in the heart were used as the background. We looked for enrichment for genetic association with biological processes in Gene Ontology (GO) and KEGG pathways.
2.9. Statistical analysis
Results are presented as mean ± standard error of mean (SEM). Differences are analyzed by two-tailed unpaired Student's t-test for experiments with two groups and one-way analysis of variance (ANOVA) with Holm–Sidak post hoc analysis for multiple comparisons in experiments including ≥3 groups. All experiments were performed at least twice using distinct cohorts of mice or independent biological samples, except the RNA-seq, which was performed once. Statistical analyses were conducted using GraphPad Prism software 8.0. No statistical analysis was used to predetermine sample sizes. Instead, sample sizes were estimated based on previous publications and our previous experience required to obtain statistically significant results. The sample size for each group was indicated in the figures, figure legends, or the methods section above. Animals were excluded if they showed distress, infection, bleeding, or anorexia due to surgery or treatment. Experimental mice were randomly assigned to each experimental/control group. Investigators were blinded to the genotypes of the individual animals during experiments and results assessments. DEGs were identified by adjusting the P-values for multiple testing at an FDR (Benjamini Hochberg method) threshold of <0.05.
3. Results
3.1. Inducible HDAC3 depletion in adult hearts causes contractile dysfunctions with high-fat-feeding
One obvious explanation for the contradiction between the beneficial effects of HDIs and the deleterious effects of HDAC depletion is the differential effects on development. Conditional knockout of an HDAC can disrupt cardiac development, while the beneficial effects of HDIs were mainly observed in adult mice. Therefore, we first address whether the previously reported detrimental cardiac effects from HDAC3 knockout are due to disruption of the cardiac developmental processes. We previously reported postnatal depletion of HDAC3 in the heart using MCK-Cre15. However, the cardiac phenotype from this mouse model can still be confounded by the potential disruption of postnatal cardiac development. To eliminate the developmental effects, we crossbred Hdac3loxP/loxP with αMHC-MerCreMer to generate iKO mice.
Tamoxifen was administered at 7 weeks old in both iKO and the αMHC-MerCreMer mice as the wild-type control (WT). We confirmed the efficient depletion of HDAC3 by Western blot analyses (Fig. 1A). The iKO mice showed differential expression of metabolic genes on normal chow (Fig. 1B), similar to the Hdac3f/f/MCK-Cre mice, as we previously reported15. The iKO mice did not show cardiac defects on normal chow, which is expected from normal cardiac functions in the Hdac3loxP/loxP/MCK-Cre line (referred to as “KO”)15. Therefore, we fed iKO mice with a high-fat diet (HFD) starting at 7 weeks old. HFD for 2–3 months does not induce robust cardiac hypertrophy or systolic dysfunction in WT mice, as expected from previous studies31,32. The iKO mice showed normal body weight gain on HFD (Fig. 1C) but enlarged hearts at 16 weeks old (Fig. 1D). Myocardial gene expression of atrial natriuretic peptide (ANP) and B-type natriuretic peptide (BNP) was significantly elevated in iKO mice (Fig. 1E). iKO heart showed bigger cardiomyocytes (Fig. 1F and G) and widespread fibrosis in the heart (Fig. 1H and I). Echocardiography demonstrated that iKO mice developed severe cardiac hypertrophy at 4 months old after 2 months on HFD, with significant left ventricular wall thickening compared to WT mice (Fig. 1J and K). The iKO mice developed systolic dysfunction, as evidenced by impaired ejection fraction (EF) and fractional shortening (FS) (Fig. 1K). Thus, inducible depletion of cardiac HDAC3 in adult mice on top of 8–9 weeks of HFD feeding caused comparable cardiac dysfunction as postnatal HDAC3 depletion in combination with a similar duration of HFD feeding15. Therefore, the deleterious effects of HDAC3 depletion in the heart are not due to development disruption.
Figure 1.
Inducible HDAC3 depletion in adult hearts impairs contractile functions on a high-fat diet (HFD). (A) Western blot analysis of hearts from 4-month-old mice after tamoxifen injection. (B) RT-qPCR analysis of the heart from 4-month-old mice on chow diet. n = 5. (C) Body weight (BW) on HFD or chow diet. HFD started at 7 weeks old. n = 6. (D) Heart weight (HW) to body weight (BW) ratio of 4-month-old mice on HFD or chow diet. n = 6. (E) RT-qPCR analysis of myocardial ANP and BNP from 4-month-old mice on HFD. (F) Gross pictures and wheat germ agglutinin (WGA) staining of hearts from 4-month-old mice on HFD. Scale bar: 50 μm. (G) WGA quantification of cardiomyocyte cross-sectional area. n = 5. (H) Percentage of fibrosis area in trichrome staining transversal sections. n = 5. (I) Trichrome staining of hearts from 4-month-old mice on HFD. Scale bar: 50 μm. (J) Representative M-mode recordings of mouse hearts in echocardiography. (K) Echocardiography of geometry and systolic functions in 15-weeks-old mice on HFD, n = 6. All data are mean ± SEM. ∗P < 0.05 by t-test or one-way ANOVA with Holm-Sidak's post hoc.
3.2. Abolishing HDAC3 enzymatic activity does not cause cardiac defects on HFD
We next sought to address the role of HDAC3 enzyme activity in the heart without disrupting the HDAC3 protein level. The deacetylation activation domain (DAD) in NCoR/SMRT binds to HDAC3 and causes a conformational change of HDAC3 protein, making the catalytic channel accessible to the substrate21,22. Purified HDAC3 protein does not show much enzyme activity by itself but gains robust enzyme activity after adding in purified DAD22,23. Conversely, the whole-body knock-in NS-DADm mouse line harbors homozygous mutations in the DAD of both NCoR and SMRT (NCoR-Y478A and SMRT-Y470A) showed normal HDAC3 protein levels, but ablated HDAC3 enzymatic activity in multiple tissues24. Therefore, we used the NS-DADm mice to address the role of HDAC3 enzyme activity in the heart. Since iKO mice have a similar phenotype as the Hdac3loxP/loxP/MCK-Cre (KO) mice, we used KO mice to avoid potential complications from tamoxifen injection.
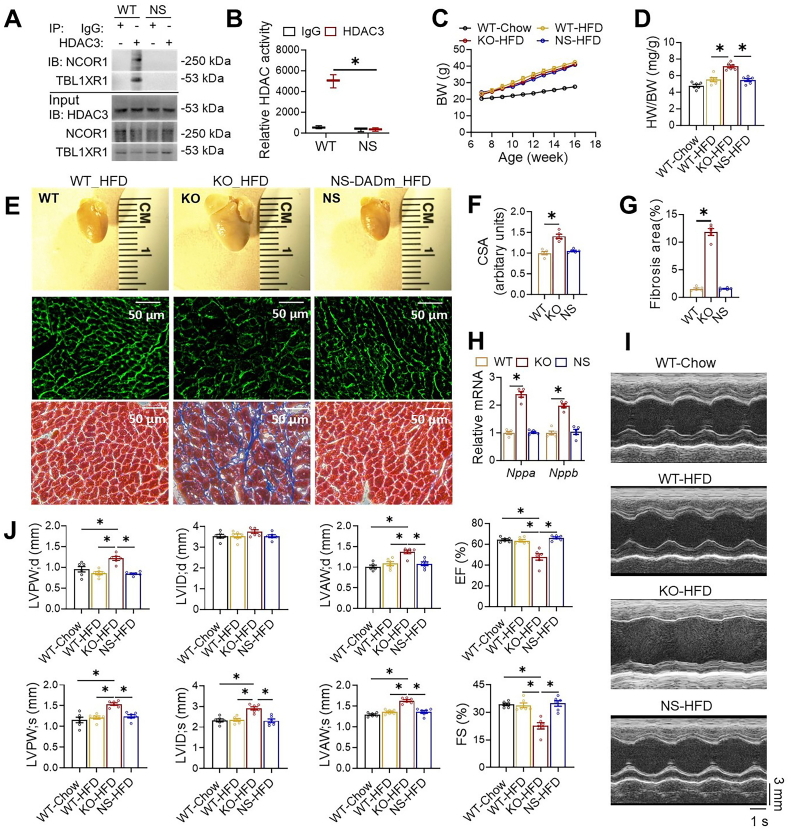
To measure the deacetylase enzyme activity of HDAC3 on any target, not just histone targets, we used HDAC3-specific antibodies to immunoprecipitate HDAC3 from the total protein lysates of the heart and subjected the immunoprecipitates to Western blot analysis and a peptide-based enzyme activity assay (Fig. 2A and B). While HDAC3 protein levels remain normal in the adult NS-DADm heart, the interaction between HDAC3 and NCOR1 or TBL1XR1, another stable component of the NCOR complex, was disrupted (Fig. 2A). In addition, the HDAC3 deacetylase activity was abolished in the HDAC3 immunoprecipitates from the NS-DADm heart (Fig. 2B). NS-DADm mice gained similar body weight on HFD as WT and KO mice (Fig. 2C). As we have reported before15, the KO heart was markedly enlarged compared to WT after feeding HFD for 9 weeks (Fig. 2D and E), with enlarged cardiomyocyte size (Fig. 2E and F), widespread interstitial fibrosis (Fig. 2E and G), and elevated myocardial expression of ANP and BNP (Fig. 2H). Echocardiography showed that KO mice developed cardiac hypertrophy and systolic dysfunction (Fig. 2I and J). In contrast, NS-DADm mice showed no defects after HFD (Fig. 2D–J). These data demonstrate that abolishing HDAC3 enzymatic activity without affecting its protein levels does not affect cardiac functions on HFD. Thus, the detrimental effects of HDAC3 depletion on contractile functions in the presence of HFD are not due to the abolishment of HDAC3 enzyme activity.
Figure 2.
Abolishing HDAC3 enzymatic activity does not cause cardiac defects on HFD. (A) Immunoblot (IB) analysis of immunoprecipitates (IP) and input lysates from hearts of 4-month-old WT and NS-DADm mice with the indicated antibodies. (B) Fluorescence-based HDAC enzyme assay using lysates of mice hearts after immunoprecipitation with HDAC3 antibody or normal IgG. n = 3. (C) Body weight (BW) on HFD or chow diet. HFD started at 7 weeks old. n = 6. (D) Heart weight (HW) to body weight (BW) ratio of 4-month-old mice on HFD or chow diet. n = 6. (E) Gross pictures, wheat germ agglutinin (WGA) staining, and trichrome staining of hearts from 4-month-old mice fed on HFD. Scale bar: 50 μm. (F) WGA quantification of cardiomyocyte cross-sectional area on samples from HFD-fed mice. n = 5. (G) Percentage of fibrosis area in trichrome staining transversal sections. n = 5. (H) RT-qPCR analysis of the myocardial ANP and BNP from 4-month-old mice fed on HFD. n = 5. (I) Representative M-mode recordings of mouse hearts in echocardiography of 4-month-old mice. (J) Echocardiography of geometry and systolic functions of 4-month-old mice, n = 6. All data are mean ± SEM. ∗P < 0.05 by t-test or one-way ANOVA with Holm-Sidak's post hoc.
3.3. Distinct transcriptomic changes between HDAC3 depletion and loss of HDAC3 enzyme activity
To address whether differential cardiac effects between HDAC3 depletion and loss of HDAC3 enzyme activity is due to differential transcriptomic effects, we performed RNA-seq analyses in the hearts of KO mice and NS-DADm mice. We harvested hearts at 6 weeks old on normal chow to exclude potential confounding effects of contractile dysfunction on cardiac gene expression. There were about 17 times more differentially expressed genes (DEGs) in the KO vs. WT hearts than in the NS-DADm vs. WT hearts (Fig. 3A). The downregulated DEGs (KO vs. WT) were enriched in mitochondrial fatty acid metabolism (Fig. 3B), while the upregulated DEGs (KO vs. WT) were enriched in cell survival and antioxidant response (Fig. 3C). Most downregulated metabolic genes were not significantly altered in the NS-DADm vs. WT control (Fig. 3D and E), which were further confirmed by RT-qPCR analyses (Fig. 3F). Both KO and NS-DADm hearts showed upregulation of genes involved in cell survival and antioxidant response compared to their respective WT controls, but the fold-changes in the NS-DADm vs. WT comparison were much less than those in the KO vs. WT comparison (Fig. 3G–I). The lack of differential expression in metabolic genes in NS-DADm is in keeping with the lack of contractile dysfunctions in NS-DADm. These results suggest that HDAC3 uses an enzyme-independent mechanism to regulate lipid metabolism, which could contribute to the detrimental effects of HDAC3 depletion. We do not suggest that these lipid metabolic genes are direct target genes of HDAC3, as this question is not relevant to whether HDAC3 function requires its deacetylase enzyme activity.
Figure 3.
Enzyme-independent regulation of fatty acid oxidation genes by HDAC3 in cardiomyocytes. (A) Overlap of differentially expressed genes (DEGs) in the NS-DADm (NS) and KO hearts compared to their respective WT controls on normal chow of 6-week-old mice, identified by RNA-seq. DEGs cutoff: q < 0.05 and |log2Fold-Change| >1. (B) Gene ontology analysis of the pooled downregulated genes (KO vs. WT and NS-DADm vs. WT, q < 0.05). (C) Gene ontology analysis of the pooled upregulated genes (KO vs. WT and NS-DADm vs. WT, q < 0.05). (D) A heatmap of fold-change top-downregulated genes in the KO/WT and NS-DADm/WT comparisons. (E) A heatmap of adjusted P values (Padj) in the KO/WT and NS-DADm/WT comparisons. (F, G) RT-qPCR analysis of mRNA from the heart of 6 weeks mice. n = 5. ∗P < 0.05 by one-way ANOVA with Holm-Sidak's post hoc. (H) A heatmap of fold-change top-upregulated genes in the KO/WT and NS-DADm/WT comparisons. (I) A heatmap of Padj values in the KO/WT and NS-DADm/WT comparisons.
3.4. Discrete cell-autonomous effects between HDAC3 depletion and HDAC enzyme inhibition
Considering that many systemic or paracrine factors could contribute to the gene expression changes in vivo, we sought to address whether the differential effects of HDAC3 depletion and HDAC3 enzyme inactivation on gene expression and metabolism are cell-autonomous in an in vitro cell culture model. We first examined the effects of HDAC3 deletion in AC16 cardiomyocytes. Western blot analysis confirmed the efficient depletion of HDAC3 by adenovirus-mediated delivery of single-guiding RNAs (sgRNA) and Cas9 targeting HDAC3 (Fig. 4A). Global histone acetylation levels were not robustly altered by knocking out a single HDAC, which is consistent with previous studies33,34. RT-qPCR analysis demonstrated that HDAC3 depletion in AC16 cells downregulated genes in fatty acid oxidation and upregulated genes in anti-apoptosis and antioxidant responses (Fig. 4B and C). The metabolic flux analysis with the 3H-palmitate isotope tracer showed that HDAC3 depletion reduced the fatty acid oxidation rate (Fig. 4D). These results suggest that the effects of HDAC3 depletion on gene expression and lipid metabolism can be recapitulated in cultured cardiomyocytes.
Figure 4.
Discrete cell-autonomous effects between HDAC3 depletion and HDAC enzyme inhibition. (A) Western blot analysis of lysates from AC16 cells with HDAC3 knock-down. (B, C) RT-qPCR analysis of the AC16 cells with HDAC3 knock-down. n = 4. (D) Fatty acid oxidation (FAO) of the AC16 cells with HDAC3 knock-down. n = 4. (E, F) Western blot analysis of lysates from AC16 cells administrated with 2 μmol/L MS275 or SAHA at the indicated dosages, along with vehicle control (mock). (G, H) RT-qPCR analysis of the AC16 cells treated with 2 μmol/L SAHA or MS275. (I, J) Fatty acid oxidation (FAO) of the AC16 cells treated with 2 μmol/L SAHA or MS275. (K, L) RT-qPCR analysis of the AC16 cells treated with 2 μmol/L SAHA or MS275. All data are mean ± SEM. ∗P < 0.05 by t-test.
We next examined the effects of HDIs, such as suberoylanilide hydroxamic acid (SAHA) and entinostat (MS-275), in AC16 cells. Despite increased global histone acetylation (Fig. 4E and F), SAHA and MS-275 did not affect the expression of lipid oxidation genes (Fig. 4G and H) and did not alter the fatty acid oxidation rate (Fig. 4I and J). These results are consistent with the lack of significant changes in fatty acid oxidation genes in vivo in NS-DADm mice. HDIs also upregulated some of the genes involved in cell survival and antioxidant responses in AC16 cells (Fig. 4K and L), which is consistent with the upregulation of these genes in the NS-DADm vs. WT hearts. We performed similar Western blot and RT-qPCR analyses in induced pluripotent stem cell (iPSC)-derived cardiomyocytes (iPSC-CM) and observed similar results (Supporting Information Fig. S1). These results demonstrated that the differential effects of HDAC3 depletion vs. HDAC3 enzyme inhibition are cardiomyocyte-autonomous.
3.5. HDAC3-LAP2β interaction in the enzyme-independent function of HDAC3 in cardiomyocytes
The enzyme-dependent repression of cell survival and antioxidant genes by HDAC3 is generally in line with the canonical view of how HDACs repress gene transcription through histone deacetylation and chromatin remodeling35,36, although some of these genes could be indirectly altered due to the altered expression of the direct HDAC3 target genes. Rather than dissecting the direct vs. indirect targets of HDAC3 enzyme-dependent target genes, we want to focus on the enzyme-independent target genes because the mechanism is more intriguing. Recent studies suggest that the interaction of HDAC3 with an inner nuclear membrane protein LAP2β is required for the enzyme-independent function of HDAC3 in restricting the precocious cardiac progenitor differentiation37, and LAP2 dysfunction protein is associated with abnormal lipid metabolism in the hepatocyte38. Therefore, we wonder whether a similar mechanism is responsible for the enzyme-independent function of HDAC3 in lipid metabolism in mature cardiomyocytes.
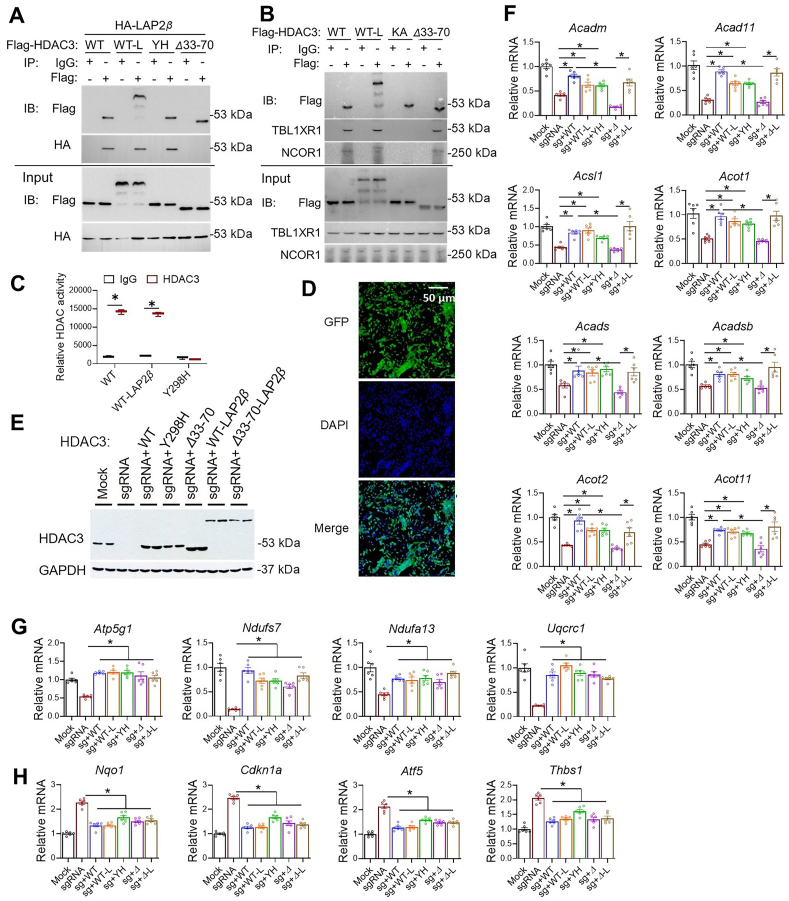
Previous studies indicated the interaction between HDAC3 and LAP2β was mediated by the 38-amino acid domain of HDAC3 (33–70)37. We confirmed that Flag-tagged HDAC3 wild-type (WT) interacts with HA-tagged LAP2β, and a deletion mutant of HDAC3 (Δ33–70) abolished such interaction (Fig. 5A) without affecting interaction with endogenous NCOR1 or TBL1XR1 (Fig. 5B). In comparison, the missense mutation of HDAC3 on the catalytic site (Y298H or YH) retained interaction with LAP2β (Fig. 5A) but abolished the enzyme activity (Fig. 5C). Another missense mutation of HDAC3 (K25A or KA) from our previous report34 disrupted interaction with NCOR1/TBL1XR1 and served as a negative control for the co-immunoprecipitation assay (Fig. 5B). LAP2β still binds to the HDAC3 K25A mutant (Supporting Information Fig. S2A), suggesting that the HDAC3–LAP2β interaction is independent of the HDAC3–NCOR interaction and thus likely remains intact in NS-DADm mice. HDAC inhibitors. SAHA or MS-275 did not affect the binding of HDAC3 with LAP2β (Fig. S2B). We also fused HDAC3 WT or Δ33–70 to LAP2β (WT-L and Δ33–70-L) (Fig. 5A–C) and used these mutants in a rescue experiment on top of HDAC3 depletion in AC16 cells. All HDAC3 constructs were engineered to evade the sgRNA targeting regions. We used adenovirus vectors to deliver HDAC3 constructs into AC16 cells, which can efficiently infect nearly all cells in the culture (Fig. 5D). The virus dosage was adjusted so that all exogenous HDAC3 was expressed at a similar level as the endogenous HDAC3 (Fig. 5E).
Figure 5.
HDAC3–LAP2β interaction in the enzyme-independent function of HDAC3 in cardiomyocytes. (A, B) Immunoblot (IB) analysis of immunoprecipitates (IP) and input lysates from HEK293 cells transfected with indicated plasmids with the indicated antibodies. (C) Fluorescence-based HDAC enzyme assay using lysates of HEK293 cells transfected with indicated constructs after immunoprecipitation with HDAC3 antibody or normal IgG, n = 3. (D) Fluorescence microscopy of AC16 cells infected with adenovirus expressing GFP. (E) Western blot analysis of AC16 cells infected with the indicated adenovirus vectors. (F–H) RT-qPCR analysis of the AC16 cells infected with the indicated adenovirus vectors. n = 4. All data are mean ± SEM. ∗P < 0.05 by t-test or one-way ANOVA with Holm-Sidak's post hoc.
Re-expression of HDAC3 WT in AC16 cells rescued KO-induced changes in lipid oxidation gene expression (Fig. 5F). The enzyme-dead Y298H mutant behaved similarly to WT in almost all genes tested (Fig. 5F–H). These results are consistent with the in vivo results in NS-DADm mice and the in vitro data with HDIs. These results demonstrated that HDAC3-mediated regulation of gene expression in cardiomyocytes is independent of HDAC3 enzyme activity. The HDAC3 Δ33–70 mutant did not rescue lipid oxidation gene expression (Fig. 5F), but artificially tethering LAP2β to the HDAC3 Δ33–70 mutant regained the ability to rescue lipid oxidation gene expression. Consistent with the gene expression results, the fatty acid oxidation assay showed a similar pattern of rescue (Fig. S2C). These results suggest that the interaction with LAP2β is crucial for the HDAC3-mediated regulation of lipid oxidation. Notably, HDAC3 Δ33–70 mutant rescued some oxidative phosphorylation genes (Fig. 5G) and genes involved in cell survival and antioxidant responses (Fig. 5H) as well as HDAC3 WT, suggesting that the regulation of these genes by HDAC3 does not require LAP2β. In summary, HDAC3 modulates gene expression in cardiomyocytes through an enzyme-independent mechanism. Its interaction with LAP2β contributes significantly to the regulation of downstream target genes, although not all of them are influenced by this interaction. It is unclear what caused such gene-specific regulation and whether the interaction with LAP2β is the major contributor to the in vivo function of HDAC3.
3.6. Distinction between HDAC3 depletion and activity suppression upon pressure overload
We have used HFD-induced obesity as a pathological stressor to study HDAC3 enzyme-dependent and -independent functions. To address whether the insights obtained from the HFD model are generalizable, we sought to use transverse aortic constriction (TAC), a common model for pressure overload-induced cardiac hypertrophy and heart failure. We subjected WT, KO, and NS-DADm mice to TAC at 7–8 weeks old. All groups of mice had similar body weights (Fig. 6A). After TAC, KO mice, but not NS-DADm mice, showed more prominent cardiac hypertrophy than WT mice at 12 weeks old (Fig. 6B–G). Specifically, KO mice after TAC showed enlarged hearts (Fig. 6B and G), enlarged cardiomyocytes (Fig. 6B and C), prominent interstitial fibrosis (Fig. 6D and E), and elevated ANP and BNP gene expression in the heart compared to WT (Fig. 6F). Echocardiography analysis showed that KO mice, but not NS-DADm mice, displayed cardiac dilation and severe systolic dysfunction, as evidenced by severe ventricular wall and reduced ejection fraction (Fig. 6H and I). These findings suggest that the adverse effects of HDAC3 depletion on cardiac contractile functions are independent of its enzyme activity. Abolishing HDAC3 enzyme activity per se, without altering its protein levels, does not increase susceptibility to overload-induced hypertrophy or contractile dysfunctions.
Figure 6.
Distinct cardiac outcomes between HDAC3 depletion and activity suppression upon pressure overload. (A) Body weight (BW) after transverse aortic constriction (TAC) or sham surgery at 2-month-old. n = 9. (B) Gross pictures and wheat germ agglutinin (WGA) staining of hearts from 3-month-old mice after TAC. Scale bar: 50 μm. (C) WGA quantification of cardiomyocyte cross-sectional area. n = 5. (D) Percentage of fibrosis area in trichrome staining transversal sections. n = 5. (E) Trichrome staining of cross-sections of hearts from 3-month-old mice after TAC. Scale bar: 50 μm. (F) RT-qPCR analysis of the myocardial ANP and BNP from 3-month-old mice after TAC. n = 5. (G) Heart weight (HW) to body weight (BW) ratio of 3-month-old mice after TAC or sham surgery. n = 9. (H) Echocardiography of geometry and contractile functions of 3-month-old mice, n = 9. (I) Representative M-mode recordings of mouse hearts in echocardiography of 3-month-old mice. All data are mean ± SEM. ∗P < 0.05 by t-test or one-way ANOVA with Holm-Sidak's post hoc.
4. Discussion
The dissection of enzyme-dependent and -independent functions of HDAC3 contributes to our understanding of the apparent paradox between the beneficial effects of HDIs and the deleterious effects of HDAC depletion on cardiac functions. The results suggest that HDAC3 uses an enzyme-independent, LAP2β-dependent mechanism to maintain active oxidative metabolism of the myocardium. Depletion of HDAC3 proteins represses fatty acid oxidation and compromises myocardial bioenergetics, which can contribute to contractile dysfunctions. This enzyme-independent function of HDAC3 in adult hearts is in keeping with previous studies showing the enzyme-independent function of HDAC3 in cardiac development in mice37,39 and cardiac contractility in drosophila40. The involvement of LAP2β in lipid metabolism is in line with several recent studies suggesting the active roles of the nuclear envelope in regulating lipid metabolic genes38,41,42. In particular, overexpression of a mutant LAP2 protein with disrupted interaction with nucleoplasmic lamin A caused more lipid accumulation than WT LAP2 in the hepatocytes38. We also tested the alternative explanation that the adverse effect in genetic mouse models could be due to developmental disruption, which is absent in HDI treatment in adult animals. We refuted this explanation because the adult-onset HDAC3 knockout mice showed similar progressive cardiac dysfunctions as the previously reported postnatal HDAC3 knockout mice15.
The absence of defects in HDAC inhibitor-treated cells (compared to vehicle) or NS-DADm mice (compared to WT) suggests that the adverse cardiac outcomes of HDAC3 depletion are not attributable to the loss of its enzymatic activity. This notion is separate from whether the beneficial effects of HDAC inhibitors (compared to vehicle-treated WT) arise from HDAC3 inhibition. Since we did not see functional improvement in NS-DADm mice compared to WT, our data suggest that the beneficial effects of HDAC inhibitors are likely independent of HDAC3. Pan-HDAC inhibitors’ cardioprotective effects may be from blocking enzyme activity of other HDACs or off-target effects. For example, the HDI ITF2357 (givinostat) was shown to attenuate cardiac myofibril relaxation8. SAHA was shown to regulate mitochondrial metabolism through post-translational modification of mitochondrial enzymes43. The expression of mitochondrial fatty acid oxidation genes was not changed after HDI treatment43, which is in line with our transcriptomic results. HDIs-mediated cardioprotection is reported to be associated with anti-apoptosis mechanisms35,36,44 and mildly enhanced oxidative stress45, which is consistent with the upregulated anti-apoptotic and antioxidant genes in NS-DADm hearts or after HDIs treatment in our study. However, we did not observe an improvement in cardiac functions in NS-DADm mice, probably due to complications of HDAC3 enzyme activity abolishment in other tissues.
The current study adds to a growing list of enzyme-independent functions of HDAC3. We and others have previously shown that some in vivo functions of HDAC3 are not dependent on its enzyme activity. For example, liver-specific knockout of HDAC3 led to hepatosteatosis and upregulated lipogenic gene expression. Mutation of the catalytic tyrosine abolished the enzyme activity of HDAC3 but can still rescue hepatosteatosis and similarly repress lipogenic gene expression as wild-type (WT) HDAC334. The NS-DADm mice showed only mild hepatosteatosis and limited upregulation of lipogenic genes in the liver compared to HDAC3 depletion24. These results suggest that the function of HDAC3 in liver lipid metabolism is largely independent of its enzyme activity. Similarly, the function of HDAC3 in cardiac development and spermatogenesis is not dependent on its enzyme activity, as we and others have shown37,46. However, the enzyme dependency is highly tissue-specific, as the function of HDAC3 in skeletal muscle fuel metabolism is indeed through its enzymatic activity26,29. The function of the brain HDAC3 in neurocognition is also dependent on its enzyme activity27,47. Along the same line, the function of HDAC3 in B-cell development and survival requires its enzymatic activity48. These results suggest that the degree to which the function of HDAC3 requires its enzyme activity is highly dependent on the tissue or cell type. Therefore, whether the HDACs function in each context requires their enzyme activity should be tested independently.
The current study has several limitations. We only characterized male mice because we did not see obvious sex differences when characterizing the original KO mice on HFD. It is intriguing that some downstream target genes are sensitive to disruptions in HDAC3–LAP2β binding while others are not. We do not know the mechanism underlying the gene-specific regulation. We speculate that the chromatin context, especially what transcription factors and coregulators occupy the neighboring genomic loci, as well as the epigenomic marks and the accessibility of the loci, could all play a role in this gene-specific effect. Another possible limitation is that we did not run chromatin immunoprecipitation (ChIP) when assessing protein hyper-acetylation due to HDAC3 depletion or inhibition. ChIP is more sensitive than Western blot since it can detect loci-specific histone modifications. However, compared to the enzyme activity assay we did, the more laborious ChIP does not address non-histone targets that could be equally important for the phenotypic changes. The key scientific question of our study is not how HDAC3 regulates gene expression through histone modifications but whether HDAC3 function in the heart depends on its enzyme activity on any histone or non-histone substrate. We did not compare HDIs with genetic manipulation of HDAC3 in vivo because HDIs can have effects independent of HDAC3 or any other HDACs. In addition, HDIs administered in vivo are cleared from the body after a few hours, which is different from the continuous abolishment of HDAC3 enzyme activity in iKO or NS-DADm mice. Thus, direct comparisons between HDIs and HDAC3 iKO or NS-DADm mice for physiological outcomes could be confounded by multiple factors. Therefore, we only used HDIs in vitro to assess short-term transcriptional effects. We also did not know whether interaction with LAP2β is sufficient for the function of HDAC3 in vivo since it is technically challenging to use AAV to achieve transgene expression with a widespread and homogeneous pattern that matches the endogenous protein level. Considering that the endogenous HDAC3 protein level in the heart is relatively low, interpreting the results of such experiments could be confounded by overexpression artifacts. Finally, the current study focuses on the function of HDAC3 in cardiomyocytes in HFD or TAC pathological models. Whether HDAC3 plays a role in other cell types in these conditions is beyond the scope of the current study.
Data access
RNA-seq data are available in Gene Expression Omnibus (GSE249901, access token: mxszegsuxbwjbiz).
Author contributions
Sichong Qian collected most of the data. Chen Zhang performed transverse aortic constriction surgery and provided training on echocardiography. Wenbo Li constructed plasmids and maintained mouse lines. Shiyang Song, Guanqiao Lin and Yueli Wang helped with some of the echocardiography analysis. Zixiu Cheng assisted in analyzing and uploading transcriptomics data. Wenjun Zhou performed WGA staining. Huiqi Yin maintained the mouse lines. Sichong Qian, Haiyang Li, Ying H Shen, and Zheng Sun obtained funding. Zheng Sun conceived the study. Sichong Qian, Chen Zhang, Wenbo Li, Shiyang Song, Guanqiao Lin, Ying H Shen, and Zheng Sun analyzed and interpreted the data. Sichong Qian and Zheng Sun wrote the manuscript with input from the other authors.
Conflicts of interest
The authors disclose no competing financial conflict of interest.
Acknowledgments
We thank Dr. Mitchell Lazar (University of Pennsylvania) for the NS-DADm and Hdac3loxP/loxP mouse lines, Dr. Rajan Jain (University of Pennsylvania) for LAP2β-related lentiviral plasmids, Dr. Chris Ward at Baylor College of Medicine (BCM) Mouse Metabolism and Phenotyping Core (R01DK114356 and UM1HG006348, USA) for echocardiography instrument, and BCM Neuropathology Core (P50103555, USA) for histology analyses. The investigators were supported in part by the National Natural Science Foundation of China (82200411) and Beijing Nova Program (20240484662, China). The laboratories of the authors were supported by grants from the American Heart Association (AHA 30970064, USA) and the National Institute of Health (NIH HL153320, USA).
Footnotes
Peer review under the responsibility of Chinese Pharmaceutical Association and Institute of Materia Medica, Chinese Academy of Medical Sciences.
Supporting information to this article can be found online at https://doi.org/10.1016/j.apsb.2025.05.002.
Contributor Information
Ying H. Shen, Email: hyshen@bcm.edu.
Zheng Sun, Email: zheng.sun@bcm.edu.
Appendix A. Supporting information
The following is the Supporting Information to this article.
References
- 1.Kong Y., Tannous P., Lu G., Berenji K., Rothermel B.A., Olson E.N., et al. Suppression of class I and II histone deacetylases blunts pressure-overload cardiac hypertrophy. Circulation. 2006;113:2579–2588. doi: 10.1161/CIRCULATIONAHA.106.625467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ooi J.Y., Tuano N.K., Rafehi H., Gao X.M., Ziemann M., Du X.J., et al. HDAC inhibition attenuates cardiac hypertrophy by acetylation and deacetylation of target genes. Epigenetics. 2015;10:418–430. doi: 10.1080/15592294.2015.1024406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Demos-Davies K.M., Ferguson B.S., Cavasin M.A., Mahaffey J.H., Williams S.M., Spiltoir J.I., et al. HDAC6 contributes to pathological responses of heart and skeletal muscle to chronic angiotensin-II signaling. Am J Physiol Heart Circ Physiol. 2014;307:H252–H258. doi: 10.1152/ajpheart.00149.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Zhang L., Qin X., Zhao Y., Fast L., Zhuang S., Liu P., et al. Inhibition of histone deacetylases preserves myocardial performance and prevents cardiac remodeling through stimulation of endogenous angiomyogenesis. J Pharmacol Exp Therapeut. 2012;341:285–293. doi: 10.1124/jpet.111.189910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zhang L., Chen B., Zhao Y., Dubielecka P.M., Wei L., Qin G.J., et al. Inhibition of histone deacetylase-induced myocardial repair is mediated by c-kit in infarcted hearts. J Biol Chem. 2012;287:39338–39348. doi: 10.1074/jbc.M112.379115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Renaud L., Harris L.G., Mani S.K., Kasiganesan H., Chou J.C., Baicu C.F., et al. HDACs regulate miR-133a expression in pressure overload-induced cardiac fibrosis. Circ Heart Fail. 2015;8:1094–1104. doi: 10.1161/CIRCHEARTFAILURE.114.001781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Khurana I., Maxwell S., Royce S., Mathiyalagan P., Karagiannis T., Mazarakis N., et al. SAHA attenuates Takotsubo-like myocardial injury by targeting an epigenetic Ac/Dc axis. Signal Transduct Targeted Ther. 2021;6:159. doi: 10.1038/s41392-021-00546-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jeong M.Y., Lin Y.H., Wennersten S.A., Demos-Davies K.M., Cavasin M.A., Mahaffey J.H., et al. Histone deacetylase activity governs diastolic dysfunction through a nongenomic mechanism. Sci Transl Med. 2018;10 doi: 10.1126/scitranslmed.aao0144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Travers J.G., Wennersten S.A., Peña B., Bagchi R.A., Smith H.E., Hirsch R.A., et al. HDAC inhibition reverses preexisting diastolic dysfunction and blocks covert extracellular matrix remodeling. Circulation. 2021;143:1874–1890. doi: 10.1161/CIRCULATIONAHA.120.046462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nural-Guvener H., Zakharova L., Feehery L., Sljukic S., Gaballa M. Anti-fibrotic effects of class I hdac inhibitor, mocetinostat is associated with IL-6/Stat3 signaling in ischemic heart failure. Int J Mol Sci. 2015;16:11482–11499. doi: 10.3390/ijms160511482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kim G.J., Jung H., Lee E., Chung S.W. Histone deacetylase inhibitor, mocetinostat, regulates cardiac remodelling and renin–angiotensin system activity in rats with transverse aortic constriction-induced pressure overload cardiac hypertrophy. Rev Cardiovasc Med. 2021;22:1037–1045. doi: 10.31083/j.rcm2203113. [DOI] [PubMed] [Google Scholar]
- 12.Lu J., Qian S., Sun Z. Targeting histone deacetylase in cardiac diseases. Front Physiol. 2024;15 doi: 10.3389/fphys.2024.1405569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Montgomery R.L., Davis C.A., Potthoff M.J., Haberland M., Fielitz J., Qi X., et al. Histone deacetylases 1 and 2 redundantly regulate cardiac morphogenesis, growth, and contractility. Genes Dev. 2007;21:1790–1802. doi: 10.1101/gad.1563807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Montgomery R.L., Potthoff M.J., Haberland M., Qi X., Matsuzaki S., Humphries K.M., et al. Maintenance of cardiac energy metabolism by histone deacetylase 3 in mice. J Clin Investig. 2008;118:3588–3597. doi: 10.1172/JCI35847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sun Z., Singh N., Mullican S.E., Everett L.J., Li L., Yuan L., et al. Diet-induced lethality due to deletion of the Hdac3 gene in heart and skeletal muscle. J Biol Chem. 2011;286:33301–33309. doi: 10.1074/jbc.M111.277707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jang J., Song G., Pettit S.M., Li Q., Song X., Cai C.L., et al. Epicardial hdac3 promotes myocardial growth through a novel microRNA pathway. Circ Res. 2022;131:151–164. doi: 10.1161/CIRCRESAHA.122.320785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Chang S., McKinsey T.A., Zhang C.L., Richardson J.A., Hill J.A., Olson E.N. Histone deacetylases 5 and 9 govern responsiveness of the heart to a subset of stress signals and play redundant roles in heart development. Mol Cell Biol. 2004;24:8467–8476. doi: 10.1128/MCB.24.19.8467-8476.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zhang C.L., McKinsey T.A., Chang S., Antos C.L., Hill J.A., Olson E.N. Class II histone deacetylases act as signal-responsive repressors of cardiac hypertrophy. Cell. 2002;110:479–488. doi: 10.1016/s0092-8674(02)00861-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lahm A., Paolini C., Pallaoro M., Nardi M.C., Jones P., Neddermann P., et al. Unraveling the hidden catalytic activity of vertebrate class IIa histone deacetylases. Proc Natl Acad Sci U S A. 2007;104:17335–17340. doi: 10.1073/pnas.0706487104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Fischle W., Dequiedt F., Hendzel M.J., Guenther M.G., Lazar M.A., Voelter W., et al. Enzymatic activity associated with class II HDACs is dependent on a multiprotein complex containing HDAC3 and SMRT/N-CoR. Mol Cell. 2002;9:45–57. doi: 10.1016/s1097-2765(01)00429-4. [DOI] [PubMed] [Google Scholar]
- 21.Arrar M., Turnham R., Pierce L., de Oliveira C.A.F., McCammon J.A. Structural insight into the separate roles of inositol tetraphosphate and deacetylase-activating domain in activation of histone deacetylase 3. Protein Sci. 2013;22:83–92. doi: 10.1002/pro.2190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Watson P.J., Fairall L., Santos G.M., Schwabe J.W.R. Structure of HDAC3 bound to co-repressor and inositol tetraphosphate. Nature. 2012;481:335–340. doi: 10.1038/nature10728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Guenther M.G., Barak O., Lazar M.A. The SMRT and N-CoR corepressors are activating cofactors for histone deacetylase 3. Mol Cell Biol. 2001;21:6091–6101. doi: 10.1128/MCB.21.18.6091-6101.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.You S.H., Lim H.W., Sun Z., Broache M., Won K.J., Lazar M.A. Nuclear receptor corepressors are required for the histone deacetylase activity of hdac3 in vivo. Nat Struct Mol Biol. 2013;20:182–187. doi: 10.1038/nsmb.2476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sohal D.S., Nghiem M., Crackower M.A., Witt S.A., Kimball T.R., Tymitz K.M., et al. Temporally regulated and tissue-specific gene manipulations in the adult and embryonic heart using a tamoxifen-inducible Cre protein. Circ Res. 2001;89:20–25. doi: 10.1161/hh1301.092687. [DOI] [PubMed] [Google Scholar]
- 26.Song S., Wen Y., Tong H., Loro E., Gong Y., Liu J., et al. The HDAC3 enzymatic activity regulates skeletal muscle fuel metabolism. J Mol Cell Biol. 2018;11:133–143. doi: 10.1093/jmcb/mjy066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhou W., He Y., Rehman A.U., Kong Y., Hong S., Ding G., et al. Loss of function of NCOR1 and NCOR2 impairs memory through a novel GABAergic hypothalamus-CA3 projection. Nat Neurosci. 2019;22:205–217. doi: 10.1038/s41593-018-0311-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.deAlmeida A.C., van Oort R.J., Wehrens X.H.T. Transverse aortic constriction in mice. J Vis Exp. 2010;38:1729. doi: 10.3791/1729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Hong S., Zhou W., Fang B., Lu W., Loro E., Damle M., et al. Dissociation of muscle insulin sensitivity from exercise endurance in mice by HDAC3 depletion. Nat Med. 2017;23:223–234. doi: 10.1038/nm.4245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lin Y., Zhang F., Chen S., Zhu X., Jiao J., Zhang Y., et al. Binary colloidal crystals promote cardiac differentiation of human pluripotent stem cells via nuclear accumulation of SETDB1. ACS Nano. 2023;17:3181–3193. doi: 10.1021/acsnano.3c00009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Brainard R.E., Watson L.J., Demartino A.M., Brittian K.R., Readnower R.D., Boakye A.A., et al. High fat feeding in mice is insufficient to induce cardiac dysfunction and does not exacerbate heart failure. PLoS One. 2013;8 doi: 10.1371/journal.pone.0083174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Tadinada S.M., Weatherford E.T., Collins G.V., Bhardwaj G., Cochran J., Kutschke W., et al. Functional resilience of C57BL/6J mouse heart to dietary fat overload. Am J Physiol Heart Circ Physiol. 2021;321:H850–H864. doi: 10.1152/ajpheart.00419.2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Stengel K.R., Bhaskara S., Wang J., Liu Q., Ellis J.D., Sampathi S., et al. Histone deacetylase 3 controls a transcriptional network required for B cell maturation. Nucleic Acids Res. 2019;47:10612–10627. doi: 10.1093/nar/gkz816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sun Z., Feng D., Fang B., Mullican S.E., You S.H., Lim H.W., et al. Deacetylase-independent function of HDAC3 in transcription and metabolism requires nuclear receptor corepressor. Mol Cell. 2013;52:769–782. doi: 10.1016/j.molcel.2013.10.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zhang L.X., Zhao Y., Cheng G., Guo T.L., Chin Y.E., Liu P.Y., et al. Targeted deletion of NF-κB p50 diminishes the cardioprotection of histone deacetylase inhibition. Am J Physiol Heart Circ Physiol. 2010;298:H2154–H2163. doi: 10.1152/ajpheart.01015.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhao T.C., Cheng G., Zhang L.X., Tseng Y.T., Padbury J.F. Inhibition of histone deacetylases triggers pharmacologic preconditioning effects against myocardial ischemic injury. Cardiovasc Res. 2007;76:473–481. doi: 10.1016/j.cardiores.2007.08.010. [DOI] [PubMed] [Google Scholar]
- 37.Poleshko A., Shah P.P., Gupta M., Babu A., Morley M.P., Manderfield L.J., et al. Genome-nuclear lamina interactions regulate cardiac stem cell lineage restriction. Cell. 2017;171 doi: 10.1016/j.cell.2017.09.018. 573-87.e14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Brady G.F., Kwan R., Ulintz P.J., Nguyen P., Bassirian S., Basrur V., et al. Nuclear lamina genetic variants, including a truncated LAP2, in twins and siblings with nonalcoholic fatty liver disease. Hepatology. 2018;67:1710–1725. doi: 10.1002/hep.29522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Lewandowski S.L., Janardhan H.P., Trivedi C.M. Histone deacetylase 3 coordinates deacetylase-independent epigenetic silencing of transforming growth factor-β1 (TGF-β1) to orchestrate second heart field development. J Biol Chem. 2015;290:27067–27089. doi: 10.1074/jbc.M115.684753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ren J., Zeng Q., Wu H., Liu X., Guida M.C., Huang W., et al. Deacetylase-dependent and -independent role of HDAC3 in cardiomyopathy. J Cell Physiol. 2023;238:647–658. doi: 10.1002/jcp.30957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Bahmanyar S., Schlieker C. Lipid and protein dynamics that shape nuclear envelope identity. Mol Biol Cell. 2020;31:1315–1323. doi: 10.1091/mbc.E18-10-0636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Sołtysik K., Ohsaki Y., Tatematsu T., Cheng J., Fujimoto T. Nuclear lipid droplets derive from a lipoprotein precursor and regulate phosphatidylcholine synthesis. Nat Commun. 2019;10:473. doi: 10.1038/s41467-019-08411-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wallner M., Eaton D.M., Berretta R.M., Liesinger L., Schittmayer M., Gindlhuber J., et al. HDAC inhibition improves cardiopulmonary function in a feline model of diastolic dysfunction. Sci Transl Med. 2020;12:1729. doi: 10.1126/scitranslmed.aay7205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Granger A., Abdullah I., Huebner F., Stout A., Wang T., Huebner T., et al. Histone deacetylase inhibition reduces myocardial ischemia-reperfusion injury in mice. FASEB J. 2008;22:3549–3560. doi: 10.1096/fj.08-108548. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Zhao T.C., Zhang L.X., Cheng G., Liu J.T. gp-91 mediates histone deacetylase inhibition-induced cardioprotection. Biochim Biophys Acta. 2010;1803:872–880. doi: 10.1016/j.bbamcr.2010.04.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yin H., Kang Z., Zhang Y., Gong Y., Liu M., Xue Y., et al. HDAC3 controls male fertility through enzyme-independent transcriptional regulation at the meiotic exit of spermatogenesis. Nucleic Acids Res. 2021;49:5106–5123. doi: 10.1093/nar/gkab313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Norwood J., Franklin J.M., Sharma D., D'Mello S.R. Histone deacetylase 3 is necessary for proper brain development. J Biol Chem. 2014;289:34569–34582. doi: 10.1074/jbc.M114.576397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Stengel K.R., Barnett K.R., Wang J., Liu Q., Hodges E., Hiebert S.W., et al. Deacetylase activity of histone deacetylase 3 is required for productive VDJ recombination and B-cell development. Proc Natl Acad Sci U S A. 2017;114:8608–8613. doi: 10.1073/pnas.1701610114. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.