Abstract
Diverse animal and plant viruses are able to translocate their virions between neighboring cells via intercellular connections. In this work, we analyze the virion assembly and cell-to-cell movement of a plant closterovirus and reveal a strong correlation between these two processes. The filamentous virions of a closterovirus possess a long body formed by the major capsid protein (CP) and a short tail formed by the minor capsid protein (CPm). Genetic and biochemical analyses show that the functions of these virion components are distinct. A virion body is required primarily for genome protection, whereas a tail represents a specialized device for cell-to-cell movement. Furthermore, tail assembly is mediated by the viral Hsp70 homolog (Hsp70h) that becomes an integral part of the virion. Inactivation of the ATPase domain of Hsp70h results in assembly of tailless virions that are incapable of translocation. A dual role for the viral molecular chaperone Hsp70h in virion assembly and transport, combined with the previous finding of this protein in intercellular channels, allowed us to propose a model of closteroviral movement from cell to cell.
Keywords: cell-to-cell movement/closterovirus/Hsp70/virion assembly
Introduction
Molecular chaperones that belong to the family of ∼70 kDa heat shock proteins (Hsp70) are found in all types of cellular organisms. The Hsp70s aid cell survival under heat shock (Lewis and Pelham, 1985) and function in protein folding, assembly of multiprotein complexes, translocation of proteins between cell compartments, etc. (Bukau and Horwich, 1998; Ellis and Hartl, 1999). Functionally distinct Hsp70s share the same type of structural organization (Boorstein et al., 1994) with the conserved N-terminal, ATPase domain (Flaherty et al., 1990) and more variable C-terminal domain (Zhu et al., 1996). Furthermore, the Hsp70s share the same basic mechanism of action. This mechanism involves repeated cycles of binding and release of the target proteins regulated by ATP hydrolysis (Pelham, 1986; Bukau and Horwich, 1998; Ellis and Hartl, 1999).
Originally, the molecular chaperones were discovered due to their role in bacteriophage infection (Georgopulos et al., 1990). Among those, a bacterial Hsp70 DnaK is required for assembly of the bacteriophage replication machinery. It was found thereafter that the diverse bacterial, plant and animal viruses rely on the multiple abilities of the Hsp70s (Sullivan and Pipas, 2001). Hsp70s were implicated in adenovirus replication (Glotzer et al., 2000), hepadnavirus assembly and reverse transcription (Hu et al., 1997), trafficking of the glycoproteins of influenza virus (Gething et al., 1986) and HIV (Knarr et al., 1999), polyomavirus assembly (Cripe et al., 1995), etc. Various animal and plant viruses induce expression of the Hsp70s (La Thangue et al., 1984; Phillips et al., 1991; Aranda et al., 1996; Liberman et al., 1999) or recruit Hsp70 via interaction with the virus-encoded J-domains (Kelley, 1999).
Certain animal viruses such as vaccinia (Frischknecht et al., 1999) and herpes simplex (Johnson et al., 2001), and virtually all plant viruses actively translocate between adjacent cells in the course of infection. Cell-to-cell movement of plant viruses proceeds through the plasmodesmata and involves functions of virus-encoded movement proteins (MPs) (Maule, 1994; Carrington et al., 1996; Lazarowitz and Beachy, 1999; Lucas, 1999; Chen et al., 2000; Citovsky and Zambryski, 2000; Voinnet et al., 2000). These MPs are able functionally to modify (Wolf et al., 1989; Oparka et al., 1999) or restructure the plasmodesmata (van Lent et al., 1991; Perbal et al., 1993). Some of the plant viruses are thought to bypass plasmodesmata in a virion form (Wellink and van Kammen, 1989; Dolja et al., 1994a; Santa Cruz et al., 1998), while others translocate their genomes in a complex with the MPs (Citovsky et al., 1990; Giesman-Cookmeyer and Lommel, 1993).
Beet yellows virus (BYV) is the prototype member of the Closteroviridae family of positive-strand RNA viruses (Dolja et al., 1994b; Karasev, 2000). The 15.5 kb genome of BYV codes for at least 10 proteins required for RNA replication, virion assembly and cell-to-cell movement (Figure 1A). Remarkably, closteroviruses encode the only known virus-specific Hsp70 homologs (Hsp70h) (Agranovsky et al., 1991). The Hsp70h of BYV was shown to localize to plasmodesmata (Medina et al., 1999) and to function as one of the viral MPs (Peremyslov et al., 1999). Other BYV MPs include a small hydrophobic protein (p6) and a 64 kDa protein (p64) (Alzhanova et al., 2000). In addition, the BYV leader proteinase was proposed to play an important, albeit indirect role in virus movement from cell to cell (Peng et al., 2001).
Fig. 1. (A) Genome map of the beet yellows virus (BYV). L-Pro, leader proteinase (the arrow below shows the site of L-Pro self-cleavage); MET, HEL and POL, the methyltransferase, RNA helicase and RNA polymerase domains of the BYV replicase, respectively; p6, a 6 kDa movement protein; p64, a 64 kDa movement protein; HSP70h, Hsp70 homolog; CPm and CP, the minor and major capsid proteins, respectively; p20 and p21, the 20 and 21 kDa proteins, respectively. (B) Electron micrograph of the BYV virion. The virion tail assembled by CPm was decorated with the CPm-specific antibodies and labeled with 10 nm gold particles (black dots). The virion body that is assembled by CP was not labeled. (C) Multiple alignment of the amino acid sequences of the CP and CPm of the BYV and grapevine leafroll-associated virus-2 (GLR) generated using the Macaw program (Schuler et al., 1991). The conserved amino acid motifs are boxed; the N-terminal motif unique to CPms (CPm box) is shown in gray. The four amino acid residues that are invariant among all capsid proteins of closteroviruses and conserved in most of the filamentous viruses are highlighted in bold and with asterisks. The numbers of amino acid residues in each protein are shown.
The rod-shaped and filamentous virions of plant viruses share helical symmetry, and are normally assembled from a single type of capsid protein. However, the filamentous virions of closteroviruses contain two types of the capsid proteins: major (CP) and minor (CPm). The CP forms a long virion body, whereas the CPm forms a short tail encompassing ∼5% of the virion length (Figure 1B) (Agranovsky et al., 1995; Tian et al., 1999). Both CP and CPm share conserved amino acid sequence motifs with other filamentous plant viruses (Dolja et al., 1991; Boyko et al., 1992). It was further revealed that both CP and CPm are required for BYV movement (Alzhanova et al., 2000), and that Hsp70h is physically associated with the virions (Tian et al., 1999; Napuli et al., 2000). Moreover, Hsp70h was recently implicated in the efficient assembly or stabilization of another closterovirus (Satyanarayana et al., 2000).
In this work, we examine the relationships between assembly of the BYV virions and translocation of the virus from cell to cell. We demonstrate that virion tails possess a specialized function in BYV cell-to-cell movement, and that Hsp70h is required for tail formation. A working model that implies a dual role for Hsp70h in assembly of the virion-associated movement device and in translocation of the virions towards and through the plasmodesmata is proposed.
Results
Computer analysis of the closteroviral capsid proteins
To explore further the relationships between the structures of the CP and CPm, we generated multiple alignment of these proteins encoded by two members of the closterovirus genus: BYV and grapevine leafroll-associated virus 2 (GLRaV-2) (Zhu et al., 1998). As illustrated in Figure 1C, these proteins share extensive similarity throughout their entire lengths, with the exception of the N-terminal regions. The signature amino acid motifs centered around invariant serine, arginine and aspartic acid residues that are found in all capsid proteins of the filamentous viruses (Dolja et al., 1991) were easily identifiable in CPs and CPms of BYV and GLRaV-2. In addition, the latter of these motifs harbors a glycine residue that is invariant among closteroviral capsid proteins (Zhu et al., 1998).
Several sequence motifs were found to be specific for each type of capsid protein. The most remarkable of these was a 28 residue motif present in the N-terminal regions of the BYV and GLRaV-2 CPms (gray box in Figure 1C). We will refer to this motif as a ‘CPm box’. Pair-wise comparisons revealed 41% sequence identity between the two CPms and 33% identity between the two CPs. In contrast, only 21 and 14% identity was found for the CP and CPm pairs within BYV and GLRaV-2, respectively. These results demonstrate that in addition to extensive conserved regions, the closteroviral CPs and CPms possess distinct structural features that may be involved in their functional specialization.
Virion body: roles in assembly and genome protection
To address the roles played by the major BYV CP in formation of the tailed virions and genome protection, we utilized a mutant in which an invariant, positively charged Arg114 (Figure 1C) was replaced with a negatively charged aspartic acid residue (mutant R114D). It was demonstrated previously that a structurally equivalent mutation in the potyviral CP resulted in complete inactivation of virion assembly (Dolja et al., 1994a). Moreover, limited characterization of the BYV R114D mutant suggested that it is assembly incompetent (Alzhanova et al., 2000). The mutant and wild-type full-length cDNA clones of BYV were transcribed in vitro, and corresponding RNAs were transfected into tobacco protoplasts. The resulting phenotypes of infection were analyzed at 4 days post-transfection.
Immunoblot analysis of the total protein extracts of the transfected protoplasts revealed that accumulation of the mutant CP was comparable with that of the wild-type CP (Figure 2, left panel). We next isolated partially purified virions from the protoplasts using differential ultracentrifugation. As expected, immunoblot analysis of the wild-type virions revealed a strong CP-specific signal. However, the mutant variant failed to produce any detectable signal (Figure 2, right panel), suggesting that the mutation abolished CP function in formation of the virion bodies. In full agreement with these results, the electron microscope (EM) analysis revealed BYV virions only in the protoplasts transfected with the wild-type RNA transcripts, whereas no virions were observed in mutant-transfected protoplasts (not shown). Taken together, these data indicate that replacement of the invariant amino acid residue resulted in accumulation of the non-functional CP subunits that failed to assemble the virion bodies.

Fig. 2. Immunoblot analysis of the total protein extracts and virions derived from the transfected protoplasts. M, mock-transfected protoplasts; WT, transfection with the wild-type RNA transcripts derived from pBYV-4; R114D, transcripts harboring the corresponding mutation in the CP gene. Anti-CP serum was used to detect protein bands.
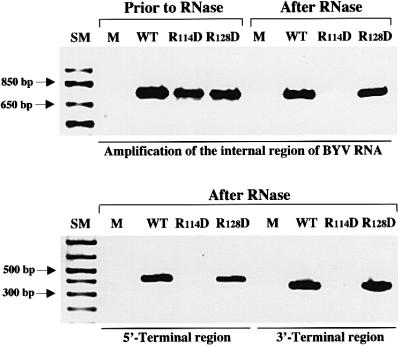
It was suggested recently that the tails cover the 5′-terminal region of the BYV RNA of ∼900 nucleotides in length (Zinovkin et al., 1999). Accordingly, it could be anticipated that in the ‘bodyless’ virions formed by the R114D mutant, this region of RNA will still be protected by the CPm, and will not be accessible for degradation by the RNases. To test this possibility, we employed RNase protection assays. The harvested protoplasts were homogenized and incubated at 37°C for 1.5 h in the presence of endogenous RNases and RNase T1 to allow degradation of the unassembled and unprotected viral RNA. The concentration of T1 was selected to ensure complete degradation of the unprotected BYV RNA under the experimental conditions (see Materials and methods). After incubation, the RNA was isolated and analyzed using RT–PCR and three sets of primers specific for the internal (∼750 nucleotides), 5′-terminal (∼430 nucleotides) and 3′-terminal (∼380 nucleotides) regions of the BYV RNA. The RNAs isolated prior to incubation were used as the controls. As shown in Figure 3, protoplast transfection with the wild-type transcripts resulted in protection of each of the tested genome regions. In contrast, the R114D mutant variant was unable to protect any of the BYV RNA regions. These results indicate that assembly of the virion body is critical for virus genome protection. Moreover, these results suggest at least two major possibilities, which are not mutually exclusive: (i) body formation is a prerequisite for tail assembly; and (ii) tails do not coat and protect an extended region of the BYV RNA.
Fig. 3. Analysis of RNA encapsidation using RNase protection assays. The protoplast homogenates were incubated in the presence of endogenous RNases and RNase T1, and the protection of encapsidated RNA was assessed using RT–PCR and primer sets specific for the internal, 5′-terminal and 3′-terminal regions of the BYV genome. The products of RT–PCR were separated in a 1% agarose gel and visualized under UV following staining with ethidium bromide. SM, size markers; R114D and R128D, transfection with the RNA transcripts harboring corresponding mutations in the CP and CPm genes, respectively. Other designations are as in Figure 2.
Virion tail is not required for genome protection
To determine the functions of the virion tail in virion assembly and genome protection, we focused initially on a detailed analysis of a CPm mutant R128D that is structurally analogous to the CP mutant R114D (Figure 1C). Immunoblot analysis of wild-type virions using anti-CPm serum detected a specific CPm band (Figure 4A, upper panel). In contrast, virions isolated from the protoplasts transfected with the R128D variant revealed no specific signal, indicating that this mutant is incapable of tail formation. This result confirmed that the invariant arginine residue is critical for assembly competence in both the CP and CPm. By analogy to the CP mutant R114D, we assumed that the phenotype of the CPm mutant R128D was due to protein misfolding rather than destabilization.
Fig. 4. (A) Protein composition of the virions isolated from transfected protoplasts. R128D, transfection with RNA transcripts harboring the corresponding mutation in the CPm gene. The type of antiserum used for analysis is shown above the panels. Other designations are as in Figure 2. (B) Length distribution profile of the wild-type (WT) and tailless (CPm mutant R128D) virions.
Interestingly, immunoblot analysis of the virion preparation using anti-CP antiserum was positive for this CPm mutant (Figure 4A, lower panel). Furthermore, the EM analysis revealed abundant BYV virions. Although these virions lacked the tails (see above), their size distribution was similar to that of the wild-type virions (Figure 4B). The possible 5% difference in length between the wild-type and tailless virions was probably beyond the resolution of our analysis. These data demonstrate that formation of the tails by CPm is not strictly required for assembly of the virion bodies by CP.
According to the proposal that CPm encapsidates the 5′-terminal, ∼900 nucleotide region of the BYV RNA (Zinovkin et al., 1999), it could be expected that in the tailless virions, this region will be unprotected and accessible for degradation by RNases. As in previous experiments, we used combined treatment with endogenous and T1 RNases and sets of primers specific for the internal, 5′-terminal and 3′-terminal regions of the BYV RNA. The RNase protection assays presented in Figure 3 clearly showed that each of these RNA regions was protected in the R128D mutant.
To confirm further that the tailless, RNase-treated virions contain an intact, infectious RNA, these virions were used for the second round of protoplast transfection. Completely protected genomes in the wild-type virions served as a positive control, whereas unprotected RNA of the CP R114D mutant variant was used as a negative control. The results of these transfections were evaluated using immunoblot analysis and anti-CP serum. To ensure that the signal detected at 4 days post-transfection is due to the newly formed CP rather than to CP introduced with the inoculum, we conducted a time course experiment with wild-type virions. As seen in Figure 5A, the input CP is undetectable at time zero, whereas the newly made CP appears and increases on days 2–4 due to replication and expression of the virus genome.
Fig. 5. Infectivity of the virions tested in the second round of protoplast transfection. CP was detected in extracts of protoplasts using immunoblotting and anti-CP serum. (A) Time course of CP accumulation in protoplasts following transfection with wild-type virions isolated from protoplasts. The numbers correspond to days post-transfection. (B) Infectivity of the wild-type virions and tailless virions assembled by the wild-type CP in the presence of mutant CPm (R128D). Note that CP mutant R114D used as a negative control failed to produce any infectious virions.
As expected, the infectivity of the RNase-treated, wild-type virions was preserved in the second transfection, while no infection was observed for the R114D CP mutant, which lacks virion bodies (Figure 5B). Examination of the R128D CPm mutant revealed that the infectivity of the RNase-treated virions was comparable to that of the wild-type virions (Figure 5B). We concluded that the virion tail is dispensable for protection of genomic RNA. These results can be interpreted to mean either that tails do not normally encapsidate any RNA, or that CP is capable of encapsidation of the entire RNA, but only in the absence of functional CPm.
Correlation between assembly of the virion body and cell-to-cell movement
Our previous work suggested that each of the two BYV capsid proteins is involved in virus movement from cell to cell (Alzhanova et al., 2000). To address further the relationships between assembly of the virion body and BYV cell-to-cell movement, we used systematic alanine-scanning mutagenesis guided by the alignment shown in Figure 1C. Typically, negatively or positively charged amino acid residues distributed throughout the proteins’ sequence were replaced with alanine (Table I). The exceptions were the R114D mutation described above and two mutations targeting invariant Ser67 and Gly145 residues that were replaced with structurally dissimilar tryptophan residues. In total, we analyzed the phenotypes of 17 CP mutants.
Table I. Phenotypes of the CP mutants in the protoplast transfection and plant inoculation experiments.
| BYV variant | RNA protection | Virion morphology | Cell-to-cell movement (cells ± SD)a |
|---|---|---|---|
| Wild type | complete | normal | 4.6 ± 2.1 |
| E5A | complete | normal | 3.4 ± 2.2 |
| E14A | complete | normal | 4.0 ± 2.1 |
| D20A | partial | normal | 3.7 ± 1.7 |
| D28A | complete | normal | 3.4 ± 2.0 |
| D30A | complete | normal | 4.9 ± 2.2 |
| E37A | partial | normal | 3.9 ± 1.9 |
| E47A | partial | normal | 3.9 ± 1.9 |
| S67W | none | no virions | 1.0 ± 0 |
| R114D | none | no virions | 1.0 ± 0 |
| Q121A | partial | normal | 4.7 ± 2.0 |
| G145W | none | no virions | 1.0 ± 0 |
| D156A | none | no virions | 1.0 ± 0 |
| D166A | complete | normal | 5.7 ± 2.1 |
| Q169A | complete | normal | 5.1 ± 2.0 |
| E177A | complete | normal | 4.1 ± 2.3 |
| E183A | complete | normal | 3.6 ± 2.2 |
| R198A | partial | normal | 3.8 ± 2.0 |
aCell-to-cell movement was quantified as the mean diameter of the infection foci in cell numbers.
The assembly of the corresponding viruses was examined using protoplast transfection followed by RNase protection assay and EM analysis. In the RNase protection assay, eight mutants were indistinguishable from wild-type, five mutants showed a decreased level of protection and four mutants showed no protection (Table I). Strikingly, each of these protection-defective mutants was a replacement of one of the four invariant amino acid residues (cf. Table I and Figure 6). The results of the EM analysis agreed with those of the RNase protection assays (Table I). It should be noted, however, that virions of apparently normal morphology were observed even for the variants that exhibited reduced levels of genome protection.
Fig. 6. Diagrammatic summary of the phenotypes of CP and CPm mutants. Large unfilled boxes represent the conserved core parts of the CP and CPm, whereas lines represent variable N- and C-terminal regions. The CPm box is shown in gray; the letters S, R, G and D correspond to invariant serine, arginine, glycine and aspartic acid (Figure 1). Small vertical rectangles mark the positions of the mutants, with the upper rectangle for cell-to-cell movement phenotypes and the lower rectangles for the assembly phenotypes. The color code is shown at the bottom.
To examine the ability of these 17 BYV mutants to move from cell to cell, each mutation was introduced into a BYV–green fluorescent protein (GFP) variant tagged by insertion of the GFP reporter (Peremyslov et al., 1999), and corresponding RNA transcripts were inoculated mechanically to leaves of an indicator plant. Analysis of the virus movement revealed two dramatically different mutant phenotypes. All 13 mutants that were capable of virion formation were also competent in cell-to-cell movement (Table I). Statistical analysis indicated that the mean diameters of the infection foci formed by corresponding BYV–GFP variants were not significantly different from that of the parental BYV–GFP (P >0.1 for all variants). In contrast, each of the four mutants deficient in virion assembly was completely arrested to single, initially inoculated cells (Table I). To ensure that the movement-positive phenotypes of the tested variants were not due to in planta reversions of the mutations, we isolated RNAs from the virus progeny, amplified the CP region by RT–PCR and sequenced the corresponding products. In all cases, the original mutations were preserved in the progeny viruses (not shown). The results of this analysis imply a strong correlation between the abilities of CP to assemble virion bodies, to protect virus RNA and to support BYV movement from cell to cell.
Tail formation is a prerequisite for cell-to-cell movement
In total, we analyzed 21 CPm mutants, 18 of which were alanine-scanning mutants, whereas the remaining three were the R128D mutant described above, as well as S85W and G158W mutants harboring replacements of the invariant serine and glycine residues (Figure 1C). Most of the mutations were introduced into CPm in the positions structurally analogous to the positions of the CP mutations (Figure 6). The important exceptions are represented by the mutations located in the CPm box, which is unique for the minor capsid protein (Figure 1C).
Examination of the cell-to-cell movement of the CPm mutants allowed us to distinguish three statistically significant phenotypic classes (P <0.05 for any two mutants that belong to different classes). The first class contained nine mutants that were similar to the wild-type (Table II). Analyses of the corresponding progeny RNAs confirmed the retention of the original mutations (data not shown). The three second class mutants exhibited inefficient cell-to-cell movement, whereas the remaining nine mutants were restricted to single, initially inoculated cells (Table II). Comparison of the movement phenotypes of the CP and CPm mutants points to the following important implications. First, the mutations targeting each of the four invariant residues located in the central region of both the CP and CPm completely block cell-to-cell movement. Secondly, four mutations introduced into the N-terminal CPm box impaired or debilitated movement. Thirdly, four more CPm mutations located in the central and C-terminal protein regions (E66A, E135A, D180A and E182A) resulted in defective movement, in contrast to analogous CP mutations (E47A, Q121A, D167A and Q169A) which did not affect movement (Figure 6). Compared with CP, CPm appears to be more prone to mutations that affect cell-to-cell movement over its entire length. This conclusion poses the question: are these differences between CP and CPm due to differences in assembly requirements for virion body and tail, respectively?
Table II. Phenotypes of the CPm mutants in the protoplast transfection and plant inoculation experiments.
| BYV variant | Cell-to-cell movement (cells ± SD)a | Class | Tail formationb |
|---|---|---|---|
| Wild type | 5.2 ± 2.2? | I | yes |
| E5A | 5.0 ± 2.6 | I | |
| D9G | 5.1 ± 2.5 | I | |
| E15A | 5.3 ± 2.2 | I | |
| D19A | 5.3 ± 2.3 | I | |
| D29A | 1.0 ± 0 | III | no |
| E35A | 2.1 ± 1.3 | II | |
| K40A | 2.9 ± 1.6 | II | |
| E45A | 1.0 ± 0 | III | no |
| E48A | 3.9 ± 2.0 | I | |
| E55A | 4.4 ± 2.0 | I | |
| E66A | 3.1 ± 2.0 | II | |
| S85W | 1.0 ± 0 | III | |
| R128D | 1.0 ± 0 | III | no |
| E135A | 1.0 ± 0 | III | no |
| G158W | 1.0 ± 0 | III | |
| D169A | 1.0 ± 0 | III | |
| D180A | 1.0 ± 0 | III | no |
| E182A | 1.0 ± 0 | III | no |
| D195A | 5.1 ± 2.1 | I | |
| K205A | 3.4 ± 2.7 | I | |
| R214A | 3.5 ± 1.8 | I |
aCell-to-cell movement was quantified as the mean diameter of the infection foci in cell numbers.
bThe data are presented for analyzed variants.
To address this question, we tested the tail assembly of the five movement-deficient CPm mutants located in the N-terminal CPm box (D29A and E45A), central region (E135A) and C-terminal region (D180A and E182A) of CPm. To this end, virions were isolated from protoplasts transfected with corresponding BYV variants and analyzed using immunoblotting and anti-CPm serum. As shown in Figure 7 and Table II, none of these five mutants was competent in tail assembly. However, each mutant was capable of body formation (cf. panels in Figure 7). The R128D mutant, for which deficiency in tail assembly was demonstrated above (Figure 4), was also movement incompetent. These data imply a strong correlation between CPm functions in cell-to-cell movement and formation of virion tails, and suggest a higher sensitivity of CPm to destabilizing mutations relative to that of CP.
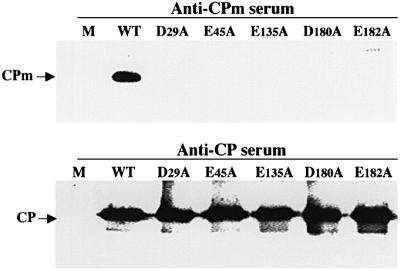
Fig. 7. Protein composition of the virions assembled in protoplasts by five movement-deficient CPm mutants. The names of mutants and type of antiserum used for analysis are shown above each panel.
Hsp70 homolog is required for tail assembly
Why are the tails less stable than the bodies? One possible explanation suggested by our results is that the tails do not contain RNA and hence are not stabilized by protein–RNA interactions. If so, tail formation may require the activity of some alternative factors. A natural candidate for the factor is BYV Hsp70h, which was found in tight association with the virions (Napuli et al., 2000). To test the possible role of Hsp70h in tail formation, we used three Hsp70h mutations. Among these, ΔXho lacks most of the Hsp70h open reading frame (ORF), whereas NoPh1 and NoCo1 are point mutations replacing Asp7 and Glu181, respectively, with lysine. The NoPh1 mutation targets a phosphate 1 motif, whereas NoCo1 targets a connect 1 motif (Bork et al., 1992). It was demonstrated previously that the analogous mutation in the phosphate 1 motif debilitates ATP hydrolytic activity of the bovine Hsp70 (Wilbanks et al., 1994). The mutation in the connect 1 motif of the bacterial Hsp70 results in uncoupling of the ATPase activity from substrate binding, and loss of function (Buchberger et al., 1994). Strikingly, the NoPh1 and NoCo1 mutations were shown to inactivate BYV Hsp70h function in cell-to-cell movement (Peremyslov et al., 1999). In addition to the Hsp70h mutants, we included in our analysis a mutant Nop6, in which expression of another BYV MP, p6, was inactivated due to replacement of the corresponding start codon (Alzhanova et al., 2000).
Immunoblot analysis using anti-CP serum showed that the amount of virions purified from the protoplasts transfected with each of the Hsp70h mutants and a p6 mutant was similar to that of the wild-type (Figure 8, lower panel). Strikingly, the anti-CPm serum revealed the presence of virion tails in a wild-type and in a p6 mutant, but in none of the Hsp70h mutants (Figure 8, upper panel).

Fig. 8. Protein composition of the virions assembled by four BYV variants possessing mutant movement protein Hsp70h or lacking movement protein p6. ΔXho, deletion of most of the Hsp70h ORF; NoPh1 and NoCo1, mutations in the phosphate 1 and connect 1 motifs of Hsp70h, respectively; Nop6, inactivation of the p6 start codon. The names of mutants and type of antiserum used for analysis are shown above the panels.
The morphology of the mutant virions was characterized further using EM. Figure 9A shows that full-size virions were detected in each of the three Hsp70h mutant variants. However, each of these mutants exhibited a reduced proportion of full-size virions (1200–1400 nm) compared with the wild-type. We have tested further whether the full-size virions formed by the Hsp70h mutants contained intact, infectious genomes. To this end, the virions isolated after original protoplast transfection with the RNA transcripts were treated with RNase T1, and used for a second transfection as described above. As seen from Figure 9B, the infectivity of mutant virions was comparable to the wild-type for each of the three Hsp70h mutants.
Fig. 9. (A) Length distribution profile of the BYV virions formed by three Hsp70h mutants compared with that of the wild-type virions. (B) Infectivity of the virions tested in the second round of protoplast transfection. The designations of the Hsp70h and p6 mutations are the same as in Figure 8.
These results demonstrate that Hsp70h is essential for the tail, but not the body assembly, although it may aid in efficient formation and/or stabilization of the full-size virion bodies. Comparison of the phenotypes of NoPh1 and NoCo1 mutants with those described for non-viral Hsp70s (Buchberger et al., 1994; Wilbanks et al., 1994) strongly suggests that ATP hydrolytic activity coupled to substrate binding is critical for the function of BYV Hsp70h in tail assembly. In addition, it can be concluded that another BYV movement protein, p6, is not required for virion assembly.
Discussion
The universal function of virions is to protect viral genomes in the environment. In addition, virions of animal and bacterial viruses promote virus entry into host cells. Since plant viruses initiate infection through wounds in the rigid cell walls, it is assumed that there are no specific virion–cell interactions mediating virus entry (Lazarowitz, 2001). However, virions of certain plant viruses are required for symplastic spread of the infection from cell to cell. In this study, we analyzed assembly of the unusual, bipartite virions of a plant closterovirus, and addressed the relationship between virus assembly and cell-to-cell movement. We found that closteroviruses evolved into a remarkable biological system that recruited a molecular chaperone from the Hsp70 family to aid virion assembly and intercellular translocation.
A paradigm of helical virion assembly is provided by tobacco mosaic virus, whose virions are self-assembled from an RNA molecule and ∼2000 identical copies of the capsid protein (Butler and Klug, 1971; Namba and Stubbs, 1986). In contrast, the helical, tailed virions of closteroviruses are formed by two capsid proteins, CP and CPm. Moreover, recent work suggested that two additional proteins, Hsp70h and the ∼60 kDa protein, are present in the virions (Tian et al., 1999; Napuli et al., 2000), and assist in virion formation or stabilization (Satyanarayana et al., 2000). It was also proposed that the CPm is required for the plant-to-plant transmission of closteroviruses by insects (Tian et al., 1999).
In order to dissect structure–function relationships within closterovirus virions, we employed site-directed mutagenesis of the BYV CP, CPm and Hsp70h, followed by analyses of virion composition, morphology and protection of virion RNA from degradation. These analyses revealed that the assembly of virion bodies by CP does not require functional CPm or Hsp70h, and that the tailless virion body protects the entire virus genome. Conversely, inactivation of the CP results in a lack of protection, suggesting that a normal virion tail does not encapsidate an extended region of the RNA. It was also found that tail formation requires not only CPm, but also Hsp70h. Interestingly, although the inactivation of Hsp70h allows formation of the full-size virion bodies, it also results in the prevalence of incomplete BYV virions (Figure 9A). This effect was even more dramatic for the related citrus tristeza virus (Satyanarayana et al., 2000). These observations suggest that the Hsp70-assisted tail formation and attachment are required for virion stability and that tail disassembly may promote RNA uncoating.
We were next interested in the relationship between BYV assembly and cell-to-cell movement. Systematic analysis of 38 CP and CPm mutants clearly indicated that assembly of the tailed virion is a prerequisite for BYV movement. Each mutation introduced into CP or CPm that prevented assembly of the virion body or tail resulted in complete arrest of virus translocation. It appears, however, that the roles played by virion body and tail are distinct. Because CP is the only virion protein that is essential for genome protection, the major role of the body may be to prevent RNA degradation during its translocation inside and between cells. An additional role of the body is to provide a structural platform for tail attachment. In contrast, tails are not essential for body formation and genome protection, but are critical for virus movement. As a result, the tail may be considered a specialized movement device attached to an otherwise ordinary filamentous virion.
Taken together with the previous observations, current data allow us to propose the following working model of closteroviral assembly and movement (Figure 10). Similarly to other helical plant viruses, the virion body of a closterovirus is self-assembled from viral RNA and CP. The tails are formed by the CPm and Hsp70h, and attached to the body with the aid of Hsp70h. As a consequence, tails do not encapsidate and protect an extended region of the viral RNA. The fact that the Hsp70h mutants NoPh1 and NoCo1 with an inactivated or deregulated ATPase domain are deficient in tail assembly suggests the requirement for ATP hydrolysis that is coupled to substrate binding. Mature, stable virions are chaperoned toward plasmodesmata presumably via an association of Hsp70h with the cytoskeleton (Karasev et al., 1992). Anchoring of Hsp70h in the plasmodesmatal channel is mediated by an intrinsic localization signal (Medina et al., 1999). The subsequent virion translocation through the channel may involve mechanical force generated by Hsp70h, analogous to the ‘trapping and pulling’ model proposed for Hsp70-mediated translocation of proteins into mitochondria and endoplasmic reticulum (Pilon and Schekman, 1999). Relocalization of Hsp70h to plasmodesmata may trigger tail disassembly and destabilize the body. As a result, a short region of virion RNA may be exposed to interact with ribosomes and start a new cycle of infection.
Fig. 10. Hypothetical model of BYV assembly and cell-to-cell movement. CW, cell wall separating adjacent plant cells.
The key features of our model can be tested in the following experiments: (i) reproduction of the tail and body assembly in vitro from purified components; (ii) resolution of the virion’s fine structure; and (iii) genetic dissection of the Hsp70h roles in tail assembly, plasmodesmatal targeting and virion translocation. It is of particular importance to reveal the localization of each tail protein relative to its counterparts. Although the tail architecture presented in Figure 10 is hypothetical, preliminary data on Hsp70h interaction with the CPm are compatible with the model. To be complete, this model also needs to account for the roles played in virus assembly and/or movement by a virion-associated p64 and a membrane-associated p6.
Participation of the Hsp70s in virion assembly is not without precedent. Indeed, cellular Hsp70s were implicated in the assembly of such diverse animal viruses as hepadnaviruses (Hu et al., 1997), papovaviruses (Cripe et al., 1995), enteroviruses (Macejak and Sarnow, 1992) and poxviruses (Jindal and Young, 1992; reviewed in Sullivan and Pipas, 2001). It appears that closteroviruses went one step further and adopted Hsp70 to facilitate assembly of the tailed virions and virus translocation from cell to cell, setting the first example of a plant virus MP that is an integral part of the virion. It is, however, reminiscent of the gE/gI glycoproteins of herpes simplex virus. These virion envelope proteins are dispensable for entry of the extracellular virus, but are required for efficient virus spread from an infected cell to a neighboring cell (Johnson et al., 2001). Moreover, similarly to plasmodesmatal targeting of Hsp70h, gE and gI are targeted to gap junctions through which herpes simplex virus is translocated intercellularly. It seems possible that the cell-to-cell movement of other plant and animal viruses may also involve incorporation of the MPs into or in association with the virions.
Cellular Hsp70s were previously implicated in a variety of processes involving protein folding, assembly and disassembly of multiprotein complexes, and protein trafficking (Pelham, 1986; Bukau and Horvich, 1998). Our work provides a first example of an Hsp70 that functions in two basic processes of the virus life cycle: virion assembly and intercellular translocation. Why do closteroviruses need a specialized movement device that includes Hsp70h? One possible reason is that the closteroviruses are exceptionally long. Accordingly, their translocation may require additional energy that can be provided by the Hsp70h-mediated ATP hydrolysis. It appears that the tomato spotted wilt virus (TSWV), which belongs to the Bunyaviridae family of the enveloped, negative-strand RNA viruses, may adopt a similar movement strategy. It was found that TSWV MP interacts with the helical nucleocapsid and with plant Hsp40s (Soellick et al., 2000). Since a J-domain of the Hsp40s binds the Hsp70s (Kelley, 1999), this finding suggests the recruitment of plant Hsp70 into the TSWV movement machinery. Thus, the theme of molecular chaperones assisting in virus transport may be more general than previously thought.
Materials and methods
Generation of mutants and plant inoculation
Mutations in the CP and CPm genes were generated via site-directed mutagenesis and introduced into two variants of a full-length BYV cDNA clone, pBYV-4 (wild-type) and pBYV-GFP (tagged by insertion of the ORF encoding the GFP) as described (Peremyslov et al., 1999; Alzhanova et al., 2000). Among the 38 mutations targeting CP and CPm, 31 were alanine-scanning mutations, in which charged (aspartic acid, glutamic acid, arginine and lysine) or polar (glutamine) residues were replaced with alanine (Tables I and II). In the D9G mutation in the CPm gene, glycine was used instead of alanine to avoid changing the amino acid sequence in the C-terminal part of p64, the coding region of which overlaps that of CPm. Mutations R114D in the CP and R128D in the CPm are identical to CPD and CPmD mutations, respectively, which were described earlier (Alzhanova et al., 2000). In the remaining four mutations, invariant serine and glycine residues of the CP and CPm were replaced with tryptophan. Each of the mutations was verified by nucleotide sequencing. The nucleotide sequences of the primers used for mutagenesis, sequencing and RT–PCR (see below) are available upon request. The Nop6, a null mutation in the p6 gene (Peremyslov et al., 1998), as well as ΔXho, NoPh1 and NoCo1 mutations in the Hsp70h gene (Peremyslov et al., 1999), were generated previously.
Protoplast transfection, virion isolation and plant inoculation
Tobacco protoplasts were isolated from suspension culture and transfected via electroporation using ∼4 × 106 cells and products of 50 µl transcription reactions per transfection (Alzhanova et al., 2000). The in vitro transcriptions were conducted using SP6 RNA polymerase (Epicentere) and ∼1 µg of the DNA of pBYV-4 variants linearized at the unique SmaI site (Peremyslov et al., 1999). Protoplasts were held in the dark at room temperature for 4 days and harvested for further analysis. For virion isolation, 12 individual protoplast batches were pooled together and cells were pelleted at 1000 r.p.m. in a Marathon 6K (Fisher Scientific) centrifuge. The pellet was frozen in liquid nitrogen and thawed twice, and ground in 1 ml of extraction buffer (0.1 M Tris–HCl pH 7.4 with 0.5% Na2SO3, 0.5% 2-mercaptoethanol and 2% Triton X-100) with a Teflon pestle. After addition of 4 ml of buffer, extracts were incubated on ice for 2 h with periodic vortexing, and centrifuged at 10 000 r.p.m. in an Eppendorf centrifuge for 5 min. The volume of supernatant was adjusted to 10 ml with extraction buffer, layered over a 2 ml sucrose cushion (20% sucrose in TE buffer) and centrifuged at 100 000 g at 4°C for 1 h. The pellet was soaked overnight in 0.1 ml of TE buffer, resuspended in 0.9 ml of the TE buffer containing 2% Triton X-100, incubated at 4°C for 2 h with constant mixing and the resulting extract was clarified by 5 min centrifugation at 10 000 r.p.m. in an Eppendorf centrifuge. The supernatant was layered over 0.2 ml of the sucrose cushion and subjected to a second cycle of ultracentrifugation. The pelleted virions were soaked in 50 µl of TE buffer overnight and resuspended by pipeting.
To test the viral cell-to-cell movement, RNA transcripts of pBYV-GFP variants were inoculated manually to leaves of the indicator plant Claytonia perfoliata as described (Peremyslov et al., 1999). The green fluorescent infection foci were analyzed using an epifluorescent stereoscope (Leica MZFLIII) at 8 days post-inoculation. At least two experiments, each involving six inoculated leaves, were carried out for each variant.
RNase protection assays
The detailed protocol for the analysis of BYV RNA protection from degradation by endogenous protoplast RNases, including detection of the protected RNA by RT–PCR amplification, was published by Medina et al. (1999). In brief, three-quarters of harvested cells were subjected to one cycle of freezing–thawing in liquid nitrogen, resuspended in buffer containing DNase I, incubated for 1.5 h at 37°C to degrade the uncoated RNAs, and RNA was isolated using TRIZOL® reagent (Gibco-BRL). The remaining quarter of the cells were used to isolate untreated RNAs as a control. To ensure further the complete and uniform degradation of the unprotected RNAs, this procedure was modified to include exogenous RNase T1. In preliminary experiments, it was determined that incubation of ∼2 µg of pBYV-4 transcripts with 1 U of RNase T1 results in complete degradation of RNA in 15 min (not shown). Accordingly, the RNase protection assays were supplemented with 1 U of RNase T1 prior to 1.5 h incubation at 37°C. RT–PCR amplification of the protected RNAs was conducted using pairs of primers that were designed to amplify the internal (BYV nucleotides 3459–4208), 5′-terminal (BYV nucleotides 1–428) or 3′-terminal (BYV nucleotides 15 090–15 468) regions of the BYV RNA. An alternative protocol included isolation of the virions from transfected protoplasts (see above) followed by a treatment with RNase T1 (1 U per 50 µl of virion suspension; 15 min at 37°C). The resulting reaction mixtures were used directly for electroporation into a fresh batch of the protoplasts.
Analyses of virion composition and morphology
The protein composition of the virions isolated from protoplasts was analyzed using rabbit anti-CP (Napuli et al., 2000) or anti-CPm (He et al., 1998) serum diluted 1:2000 and 1:1000, respectively, and an ECL western blotting detection kit as recommended by the manufacturer (Amersham/Pharmacia Biotech). Transmission electron microscopy was conducted as described earlier (Napuli et al., 2000). For analysis of virion length distribution, the virions from protoplast extracts were trapped on anti-CP serum-treated grids, and the lengths of 100 or more randomly selected particles were measured for each variant using NIH Imaging software. The immunodecoration and immunogold labeling of the virion tails were conducted using anti-CPm serum followed by treatment with anti-rabbit immunoglobulin conjugated to 10 nm gold particles (Ted Pella, Inc.) as described by Tian et al. (1999).
Acknowledgments
Acknowledgements
We thank Drs Hugh Pelham, Bill Dawson and Jim Carrington for stimulating discussions and critical reading of the manuscript, and Dr Tatineni Satyanarayana for helping to develop serial transfection of protoplasts. This work was supported by grants from the National Institutes of Health (R1GM53190B) and United States Department of Agriculture (CSREES 2001-35319-10875) to V.V.D.
References
- Agranovsky A.A., Boyko,V.P., Karasev,A.V., Koonin,E.V. and Dolja,V.V. (1991) Putative 65 kDa protein of beet yellows closterovirus is a homologue of HSP70 heat shock proteins. J. Mol. Biol., 217, 603–610. [DOI] [PubMed] [Google Scholar]
- Agranovsky A.A., Lesemann,D.E., Maiss,E., Hull,R. and Atabekov,J.G. (1995) ‘Rattlesnake’ structure of a filamentous plant RNA virus built of two capsid proteins. Proc. Natl Acad. Sci. USA, 92, 2470–2473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alzhanova D.V., Hagiwara,Y., Peremyslov,V.V. and Dolja,V.V. (2000) Genetic analysis of the cell-to-cell movement of beet yellows closterovirus. Virology, 267, 192–200. [DOI] [PubMed] [Google Scholar]
- Aranda M.A., Escaler,M., Wang,D. and Maule,A.J. (1996) Induction of HSP70 and polyubiquitin expression associated with plant virus replication. Proc. Natl Acad. Sci. USA, 93, 15289–15293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boorstein W.R., Ziegelhoffer,T. and Craig,E.A. (1994) Molecular evolution of the HSP70 multigene family. J. Mol. Evol., 38, 1–17. [DOI] [PubMed] [Google Scholar]
- Bork P., Sander,C. and Valencia,A. (1992) An ATPase domain common to procaryotic cell cycle proteins, sugar kinases, actin and Hsp70 heat shock proteins. Proc. Natl Acad. Sci. USA, 89, 7290–7294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boyko V.P., Karasev,A.V. Agranovsky,A.A., Koonin,E.V. and Dolja,V.V. (1992) Capsid protein gene duplication in a filamentous RNA virus of plants. Proc. Natl Acad. Sci. USA, 89, 9156–9160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buchberger A., Valencia,A., McMacken,R., Sander,C. and Bukau,B. (1994) The chaperone function of DnaK requires the coupling of ATPase activity with substrate binding through residue E171. EMBO J., 13, 1687–1695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bukau B. and Horwich,A.L. (1998) The HSP70 and HSP60 chaperone machines. Cell, 92, 351–366. [DOI] [PubMed] [Google Scholar]
- Butler P.J.G. and Klug,A. (1971) Assembly of the particle of tobacco mosaic virus from RNA and discs of protein. Nature, 229, 47–50. [DOI] [PubMed] [Google Scholar]
- Carrington J.C., Kasschau,K.D., Mahajan,S.K. and Schaad,M.C. (1996) Cell-to-cell and long-distance transport of viruses in plants. Plant Cell, 8, 1669–1681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen M.-H., Sheng,J., Hind,G., Handa,A.K. and Citovsky,V. (2000) Interaction between the tobacco mosaic virus movement protein and host cell pectin methylesterases is required for viral cell-to-cell movement. EMBO J., 19, 913–920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Citovsky V. and Zambryski,P. (2000) Systemic transport of RNA in plants. Trends Plant Sci., 5, 52–54. [DOI] [PubMed] [Google Scholar]
- Citovsky V., Knorr,D., Schuster,G. and Zambryski,P. (1990) The P30 movement protein of tobacco mosaic virus is a single-stranded nucleic acid binding protein. Cell, 60, 637–647. [DOI] [PubMed] [Google Scholar]
- Cripe T.P., Delos,S.E., Estes,P.A. and Garcea,R.L. (1995) In vivo and in vitro association of hsc70 with polyomavirus capsid proteins. J. Virol., 69, 7807–7813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dolja V.V., Boyko,V.P., Agranovsky,A.A. and Koonin,E.V. (1991) Phylogeny of capsid proteins of rod-shaped and filamentous plant viruses: two families with distinct patterns of sequence and probably structure conservation. Virology, 184, 79–86. [DOI] [PubMed] [Google Scholar]
- Dolja V.V., Haldeman,R., Robertson,N., Dougherty,W.G. and Carrington,J.C. (1994a) Distinct functions of the capsid protein in assembly and cell-to-cell movement of tobacco etch potyvirus in plants. EMBO J., 13, 1482–1491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dolja V.V., Karasev,A.V. and Koonin,E.V. (1994b) Molecular biology and evolution of closteroviruses: sophisticated build-up of large RNA genomes. Annu. Rev. Phytopathol., 32, 261–285. [Google Scholar]
- Ellis R.J. and Hartl,F.-U. (1999) Principles of protein folding in the cellular environment. Curr. Opin. Struct. Biol., 9, 102–110. [DOI] [PubMed] [Google Scholar]
- Flaherty K.M., DeLuca-Flaherty,C. and McKay,D.B. (1990) Three-dimensional structure of the ATPase fragment of a 70 K heat-shock cognate protein. Nature, 346, 623–628. [DOI] [PubMed] [Google Scholar]
- Frischknecht F., Moreau,V., Rottger,S., Gonfloni,S., Reckmann,I., Superti-Furga,G. and Way,M. (1999) Actin-based motility of vaccinia virus mimics receptor tyrosine kinase signalling. Nature, 401, 926–929. [DOI] [PubMed] [Google Scholar]
- Georgopulos C., Ang,D., Liberek,K. and Zylicz,M. (1990) Properties of the Escherichia coli heat shock proteins and their role in bacteriophage λ growth. In Morimito,R.I., Tissieres,A. and Georgopulos,C. (eds), Stress Proteins in Biology and Medicine. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, pp. 191–221.
- Gething M.-J., McCammon,K. and Sambrook,J. (1986) Expression of wild-type and mutant forms of influenza hemagglutinin: the role of folding in intracellular transport. Cell, 46, 939–950. [DOI] [PubMed] [Google Scholar]
- Giesman-Cookmeyer D. and Lommel,S.A. (1993) Alanine scanning mutagenesis of a plant virus movement protein identifies three functional domains. Plant Cell, 5, 973–982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Glotzer J., Saltic,M., Chlocca,S., Michou,A., Moseley,P. and Cotten,M. (2000) Activation of heat-shock response by adenovirus is essential for virus replication. Nature, 407, 207–211. [DOI] [PubMed] [Google Scholar]
- He X., Harper,K., Grantham,G., Yang,C.H. and Creamer,R. (1998) Serological characterization of the 3′-proximal encoded proteins of beet yellows closterovirus. Arch. Virol., 143, 1349–1363. [DOI] [PubMed] [Google Scholar]
- Hu J., Toft,D.O. and Seeger C. (1997) Hepadnavirus assembly and reverse transcription require a multi-component chaperone complex which is incorporated into nucleocapsids. EMBO J., 16, 59–68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jindal S. and Young,R.A. (1992) Vaccinia virus infection induces a stress response that leads to association of Hsp70 with viral proteins. J. Virol., 66, 5357–5362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson D.C., Webb,M., Wisner,T.W. and Brunetti,C. (2001) Herpes simplex virus gE/gI sorts nascent virions to epithelial cell junctions, promoting virus spread. J. Virol., 75, 821–833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karasev A.V. (2000) Genetic diversity and evolution of closteroviruses. Annu. Rev. Phytopathol., 38, 293–324. [DOI] [PubMed] [Google Scholar]
- Karasev A.V., Kashina,A.S., Gelfand,V.I. and Dolja,V.V. (1992) HSP70-related 65K protein of beet yellows virus is a microtubule-binding protein. FEBS Lett., 304, 12–14. [DOI] [PubMed] [Google Scholar]
- Kelley W.L. (1999) Molecular chaperones: how J-domains turn on Hsp70s. Curr. Biol., 9, R305–R308. [DOI] [PubMed] [Google Scholar]
- Knarr G., Modrow,S., Todd,A., Gething,M.J. and Buchner,J. (1999) BiP-binding sequences in HIV gp160. Implications for the binding specificity of BiP. J. Biol. Chem., 274, 29850–29857. [DOI] [PubMed] [Google Scholar]
- LaThangue N.B., Shriver,K., Dawson,C. and Chan,W.L. (1984) Herpes simplex virus infection causes the accumulation of a heat shock protein. EMBO J., 3, 267–277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lazarowitz S.G. (2001) Plant viruses. In Knipe,D.M. and Howley,P.M. (eds), Fields Virology. Vol. 1. Lippincott Williams & Wilkins, Philadelphia, PA, pp. 533–598.
- Lazarowitz S.G. and Beachy,R.N. (1999) Viral movement proteins as probes for intracellular and intercellular trafficking in plants. Plant Cell, 11, 535–548. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis M.J. and Pelham,H.R. (1985) Involvement of ATP in the nuclear and nucleolar functions of the 70 kd heat shock protein. EMBO J., 4, 3137–3143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liberman E., Fong,Y.L., Selby,M.J., Choo,Q.L., Cousens,L., Houghton,M. and Yen,T.S. (1999) Activation of the grp78 and grp94 promoters by hepatitis C virus E2 envelope protein. J. Virol., 73, 3718–3722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lucas W.J. (1999) Plasmodesmata and the cell-to-cell transport of proteins and nucleoprotein complexes. J. Exp. Bot., 50, 979–987. [Google Scholar]
- Macejak D.G. and Sarnow,P. (1992) Association of heat shock protein 70 with enterovirus capsid precusor P1 in infected human cells. J. Virol., 66, 1520–1527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maule A.J. (1994) Plant–virus movement: de novo processing or redeployed machinery? Trends Microbiol., 2, 305–306. [DOI] [PubMed] [Google Scholar]
- Medina V., Peremyslov,V.V., Hagiwara,Y. and Dolja,V.V. (1999) Subcellular localization of the HSP70-homolog encoded by beet yellows closterovirus. Virology, 260, 173–181. [DOI] [PubMed] [Google Scholar]
- Namba K. and Stubbs,G. (1986) Structure of tobacco mosaic virus at 3.6 Å resolution: implications for assembly. Science, 231, 1401–1406. [DOI] [PubMed] [Google Scholar]
- Napuli A.J., Falk,B.W. and Dolja,V.V. (2000) Interaction between Hsp70 homolog and filamentous virions of the beet yellows virus. Virology, 274, 232–239. [DOI] [PubMed] [Google Scholar]
- Oparka K.J. et al. (1999) Simple, but not branched, plasmodesmata allow the nonspecific trafficking of proteins in developing tobacco leaves. Cell, 97, 743–754. [DOI] [PubMed] [Google Scholar]
- Pelham H.R.B. (1986) Speculations on the functions of the major heat shock and glucose-regulated proteins. Cell, 46, 959–961. [DOI] [PubMed] [Google Scholar]
- Peng C.-W., Peremyslov,V.V., Mushegian,A.R., Dawson,W.O. and Dolja,V.V. (2001) Functional specialization and evolution of leader proteinases in the family Closteroviridae. J. Virol., 75, 12153–12160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perbal M.-C., Thomas,C.L. and Maule,A.J. (1993) Cauliflower mosaic virus gene I product (P1) forms tubular structures which extend from the surface of infected protoplasts. Virology, 195, 281–285. [DOI] [PubMed] [Google Scholar]
- Peremyslov V.V., Hagiwara,Y. and Dolja,V.V. (1998) Genes required for replication of the 15.5-kilobase RNA genome of a plant closterovirus. J. Virol., 72, 5870–5876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peremyslov V.V., Hagiwara,Y. and Dolja,V.V. (1999) HSP70 homolog functions in cell-to-cell movement of a plant virus. Proc. Natl Acad. Sci. USA, 96, 14771–14776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phillips B., Abravaya,K. and Morimoto,R.I. (1991) Analysis of the specificity and mechanism of transcriptional activation of the human Hsp70 gene during infection by DNA viruses. J. Virol., 65, 5680–5692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pilon M. and Schekman,R. (1999) Protein translocation: how Hsp70 pulls it off. Cell, 97, 679–682. [DOI] [PubMed] [Google Scholar]
- SantaCruz S., Roberts,A.G., Prior,D.A.M., Chapman,S. and Oparka,K.J. (1998) Cell-to-cell and phloem-mediated transport of potato virus X: the role of virions. Plant Cell, 10, 495–510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Satyanarayana T., Gowda,S., Mawassi,M., Albiach-Marti,M.R., Ayllon,M.A., Robertson,C., Garnsey,S.M. and Dawson,W.O. (2000) Closterovirus encoded HSP70 homolog and p61 in addition to both coat proteins function in efficient virion assembly. Virology, 278, 253–265. [DOI] [PubMed] [Google Scholar]
- Schuler G.D., Altschul,S.F. and Lipman,D.J. (1991) A workbench for multiple alignment construction and analysis. Proteins, 9, 180–190. [DOI] [PubMed] [Google Scholar]
- Soellick T.R., Uhrig,J.F., Bucher,G.L. and Schreier,P.H. (2000) The movement protein NSm of tomato spotted wilt tospovirus (TSWV): RNA binding, interaction with the TSWV N protein and identification of interacting plant proteins. Proc. Natl Acad. Sci. USA, 97, 2373–2378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sullivan C.S. and Pipas,J.M. (2001) The virus–chaperone connection. Virology, 287, 1–8. [DOI] [PubMed] [Google Scholar]
- Tian T., Rubio,L., Yeh,H.-H., Crawford,B. and Falk,B.W. (1999) Lettuce infectious yellows virus: in vitro acquisition analysis using partially purified virions and the whitefly, Bemisia tabaci.J. Gen. Virol., 80, 1111–1117. [DOI] [PubMed] [Google Scholar]
- vanLent J., Storms,M., van der Meer,F., Wellink,J. and Goldbach,R. (1991) Tubular structures involved in movement of cowpea mosaic virus are also formed in infected cowpea protoplasts. J. Gen. Virol., 72, 2615–2623. [DOI] [PubMed] [Google Scholar]
- Voinnet O., Lederer,C. and Baulcombe,D.C. (2000) A viral movement protein prevents spread of the gene silencing signal in Nicotiana benthamiana. Cell, 103, 157–167. [DOI] [PubMed] [Google Scholar]
- Wellink J. and van Kammen,A. (1989) Cell-to-cell transport of cowpea mosaic virus requires both the 58/48K proteins and the capsid protein. J. Gen. Virol., 70, 2279–2286. [Google Scholar]
- Wilbanks S.M., DeLuca-Flaherty,C. and McKay,D. (1994) Structural basis of the 70-kilodalton heat shock cognate protein ATP hydrolytic activity. J. Biol. Chem., 269, 12893–12898. [PubMed] [Google Scholar]
- Wolf S., Deom,C.M., Beachy,R.N. and Lucas,W.J. (1989) Movement protein of tobacco mosaic virus modifies plasmodesmatal size exclusion limit. Science, 246, 377–379. [DOI] [PubMed] [Google Scholar]
- Zhu H.-Y., Ling,K.-S., Goszczynski,D.E., McFerson,J.R. and Gonsalves,D. (1998) Nucleotide sequence and genome organization of grapevine leafroll-associated virus-2 are similar to beet yellows virus, the closterovirus type member. J. Gen. Virol., 79, 1289–1298. [DOI] [PubMed] [Google Scholar]
- Zhu X., Zhao,X., Burkholder,W.F., Gragerov,A., Ogata,C.M., Gottesman,M.E. and Hendrikson,W.A. (1996) Structural analysis of substrate binding by the molecular chaperone DnaK. Science, 272, 1606–1614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zinovkin R., Jelkman,W. and Agranovsky,A.A. (1999) The minor coat protein of beet yellows closterovirus encapsidates the 5′ terminus of RNA in virions. J. Gen. Virol., 80, 269–272. [DOI] [PubMed] [Google Scholar]