Abstract
When inorganic phosphate is limiting, Arabidopsis has the facultative ability to metabolize exogenous nucleic acid substrates, which we utilized previously to identify insensitive phosphate starvation response mutants in a conditional genetic screen. In this study, we examined the effect of the phosphate analog, phosphite (Phi), on molecular and morphological responses to phosphate starvation. Phi significantly inhibited plant growth on phosphate-sufficient (2 mm) and nucleic acid-containing (2 mm phosphorus) media at concentrations higher than 2.5 mm. However, with respect to suppressing typical responses to phosphate limitation, Phi effects were very similar to those of phosphate. Phosphate starvation responses, which we examined and found to be almost identically affected by both anions, included changes in: (a) the root-to-shoot ratio; (b) root hair formation; (c) anthocyanin accumulation; (d) the activities of phosphate starvation-inducible nucleolytic enzymes, including ribonuclease, phosphodiesterase, and acid phosphatase; and (e) steady-state mRNA levels of phosphate starvation-inducible genes. It is important that induction of primary auxin response genes by indole-3-acetic acid in the presence of growth-inhibitory Phi concentrations suggests that Phi selectively inhibits phosphate starvation responses. Thus, the use of Phi may allow further dissection of phosphate signaling by genetic selection for constitutive phosphate starvation response mutants on media containing organophosphates as the only source of phosphorus.
Phosphorus is an essential structural constituent of many biomolecules and plays a pivotal role in energy conservation and metabolic regulation. Inorganic orthophosphate (Pi), the assimilated form of phosphorus, is often a limiting macronutrient in both terrestrial and aquatic ecosystems. As a consequence, assimilation, storage, and metabolism of Pi are highly regulated processes that directly affect plant growth (Theodorou and Plaxton, 1993; Raghothama, 1999). To cope with low Pi availability, plants have evolved sophisticated developmental and metabolic adaptations to enhance Pi acquisition from the rhizosphere. Such strategies include morphological changes in root architecture and associations with symbiotic mycorrhizal fungi to accelerate soil exploration as well as biochemical responses to chemically increase Pi availability from insoluble salt complexes and organophosphates present in recalcitrant soil matter (McCully, 1999; Raghothama, 1999). Despite numerous studies on adaptive responses to Pi limitation, little is known about the underlying molecular processes or regulatory genes that are involved in the Pi starvation response of plants.
On the other hand, genetic and molecular studies have provided much insight into the microbial response to Pi limitation. When faced with low Pi availability, both Escherichia coli and Saccharomyces cerevisiae activate a multigene emergency rescue system to scavenge traces of usable phosphorus from the surrounding medium. Both systems are known as a pho regulon and consist of at least 30 genes. Pi starvation leads to an increased expression of Pi acquisition enzymes and regulatory proteins such as nucleases, phosphatases, high-affinity Pi transporters, Pi-binding proteins, and Pi sensor protein kinases that monitor extracellular Pi availability (Torriani-Gorini, 1994). The pho regulon of S. cerevisiae provides a paradigm for Pi-responsive gene regulation in eukaryotes. Simple genetic screens have identified positive and negative regulators of the signaling pathway that controls induction and secretion of Pi starvation-inducible acid phosphatase (Lenburg and O'Shea, 1996).
In vascular plants, the existence of an analogous multigene Pi starvation-inducible rescue system has been proposed (Goldstein et al., 1988), and several putative components of a plant pho regulon have been described. These include Pi starvation-inducible acid phosphatases with broad substrate specificity (Duff et al., 1994; del Pozo et al., 1999; Baldwin et al., 2001), phosphoenolpyruvate phosphatase and pyrophosphate-dependent phosphofructokinase (Theodorou and Plaxton, 1993), ribonucleases (Green, 1994), phosphodiesterases (Abel et al., 2000), Pi transporters (Raghothama, 2000), and several genes of unknown function (Liu et al., 1997; Burleigh and Harrison, 1999). However, Pi sensory mechanisms and components of the signal transduction pathway(s) that integrate the Pi starvation response of plants have not been identified. To elucidate Pi starvation signaling pathway(s) in plants, we designed a conditional genetic screen in Arabidopsis that is based on the facultative ability of wild-type plants to utilize organophosphates such as nucleic acids in Pi-limiting conditions (Chen et al., 2000). We isolated several recessive mutant lines that grow on high Pi medium, but which show severe growth inhibition on medium containing nucleic acids as the only source of phosphorus. For a subset of these lines, the conditional growth defect is likely to be caused by reduced induction of multiple Pi-scavenging enzymes in response to low Pi, which suggests loss-of-function mutations in positive factors of plant pho gene regulation (Chen et al., 2000).
To explore the feasibility of a genetic selection system for constitutive Pi starvation response mutants, which may target repressors of Pi starvation-inducible genes, we examined in this study the effect of phosphite (Phi or phosphonate) on Pi starvation responses in Arabidopsis. The orthophosphite anion (H2PO3− or HPO32−) contains a nonacidic hydrogen atom and can be considered an analog of the Pi anion (H2PO4− or HPO42−). Previous work in Brassica napus and Brassica nigra showed that, although readily absorbed, Phi is relatively stable and not significantly oxidized or otherwise metabolized in plants (Carswell et al., 1996, 1997). In addition, Phi represses induction of the Pi starvation-inducible enzymes, phosphoenolpyruvate phosphatase and pyrophosphate-dependent phosphofructokinase, as well as high-affinity Pi uptake in conditions of Pi limitation, which led to the suggestion that Phi may be a useful inhibitor for dissecting Pi starvation response pathways (Carswell et al., 1996, 1997). Toward this goal, we extended these studies to Arabidopsis and examined the effect of Phi on several morphological and molecular Pi starvation responses. We studied the effect of Phi on plant growth in media supplemented with various phosphorus sources as well as the effect of Phi on root hair formation and Pi starvation-inducible nucleolytic enzyme activities and transcript levels. Our data suggest that Phi specifically attenuates developmental and molecular responses of Arabidopsis to low Pi availability.
RESULTS
Inhibition of Arabidopsis Growth by Phi
The capacity of plants to induce and secrete Pi-scavenging enzymes into the rhizosphere in response to limiting Pi concentrations is thought to sustain plant growth by solubilizing immobilized Pi and by degrading compounds that contain organically bound phosphorus. One example of this ability is Arabidopsis, which develops normally on synthetic media containing purified nucleic acid substrates as the only source of phosphorus. When grown on −Pi/+DNA or −Pi/+RNA media, Arabidopsis plants induce Pi starvation-inducible nucleolytic enzymes such as nucleases and acid phosphatase to metabolize exogenous nucleic acids (Chen et al., 2000). As a consequence, plants grown under such conditions do not show severe symptoms of Pi deficiency and are comparable in appearance and fresh weight with plants that are cultivated on +Pi medium (see Fig. 1).
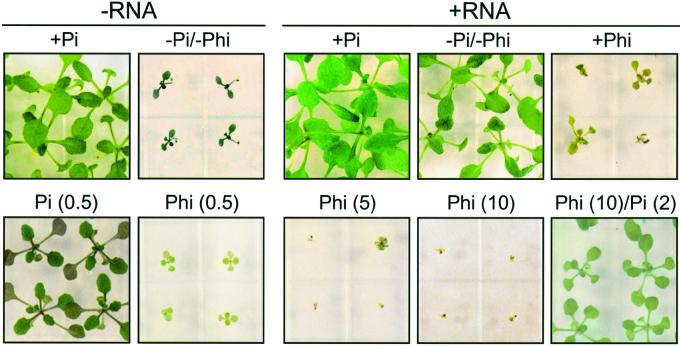
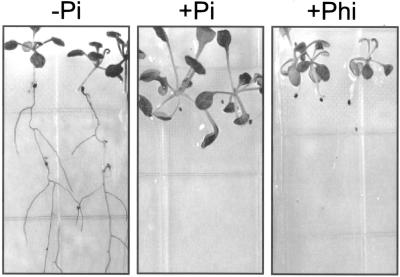
Figure 1.
Growth of Arabidopsis plants on media containing various sources of phosphorus. Seeds were germinated and grown for 18 d in media with the following phosphorus supplements or combinations thereof: −Pi; +Pi (2 mm); +Phi (2.5 mm, or as indicated, mm, in parentheses); or +RNA (2 mm phosphorus).
To study the effect of the Pi analog, Phi, on plant growth, we added increasing concentrations of Phi to +Pi, −Pi/+RNA, and −Pi media, and the fresh weight of pooled seedlings was measured 18 d after germination (Fig. 2). Addition of up to 2.5 mm Phi to +Pi medium (2 mm Pi) did not considerably reduce seedling fresh weight. However, higher Phi concentrations significantly inhibited plant growth when compared with growth on Phi-free +Pi medium (Student's t test, P < 0.05). Seedling fresh weight was reduced by 50% and 80% on +Pi medium containing 5 and 10 mm Phi, respectively. A more severe reduction of seedling fresh weight in response to increasing Phi concentrations was observed for growth on −Pi/+RNA (2 mm phosphorus) medium; fresh weight was reduced by 80% and 95% in the presence of 5 and 10 mm Phi, respectively (Student's t test, P < 0.05; Figs. 1 and 2). As expected, inhibition of seedling growth by Phi on −Pi/+RNA medium could be overcome by the addition of Pi (see Fig. 1). Similar data were obtained for growth on DNA-containing medium (data not shown). Seedlings grown in −Pi showed typical symptoms of Pi deficiency such as accumulation of anthocyanins, and their fresh weight was less than 10% compared with plants grown on +Pi or −Pi/+RNA media (Figs. 1 and 2). Addition of Phi to concentrations above 5 mm further decreased fresh weight of seedlings grown on −Pi medium (Student's t test, P < 0.05; Fig. 2). Enhanced root growth relative to shoot growth is a classical response of plants to Pi deficiency, which results in an increased root-to-shoot ratio. It is interesting to note that seedlings grown for 12 d on −Pi/+Phi medium failed to respond with a redistribution of growth to benefit the root system. Although total fresh weight amounts were similar for −Pi and −Pi/+Phi seedlings, the root-to-shoot ratio of −Pi/+Phi seedlings was about 8-fold lower than that of −Pi-grown seedlings (Student's t test, P < 0.05) and approached the root-to-shoot ratio of +Pi seedlings (data not shown).
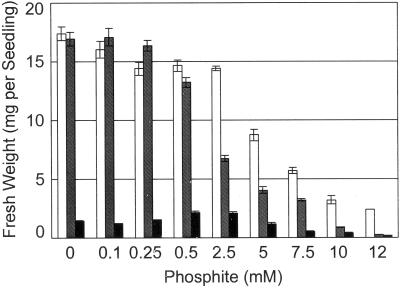
Figure 2.
Growth of Arabidopsis plants on media supplemented with increasing concentrations of Phi. Seeds were germinated and grown on vertically oriented agar plates for 18 d in +Pi (2 mm)/+Phi, white bars; −Pi/+RNA(2 mm phosphorus)/+Phi, hatched bars; and −Pi/+Phi, black bars. The fresh weight of 10 pooled seedlings was determined. The given fresh weights represent the mean of one seedling ± se (n = 6 pools of 10 seedlings).
Effect of Phi on Root Hair Formation
Low phosphorus availability stimulates lateral root formation as well as initiation and elongation of root hairs in many plant species, which may contribute to enhanced soil phosphorus acquisition. Increased root hair density and elongation is one of the morphological hallmarks of the Pi starvation response in Arabidopsis (Bates and Lynch, 2000). Therefore, we examined the effect of Phi on root hair formation of Arabidopsis plants grown in media with different sources of phosphorus. Roots of seedlings grown on −Pi medium developed root hairs and were more branched, whereas roots of +Pi seedlings were less branched and appeared largely devoid of root hairs (data not shown), which is consistent with previous reports (Bates and Lynch, 1996). Because plant growth is severely inhibited in our −Pi conditions (Chen et al., 2000), the effect of Pi limitation on root hair formation is best seen when roots of plants that were grown on −Pi/+RNA and +Pi/+RNA medium are compared. The external Pi concentration generated by the hydrolysis of exogenous RNA is sufficient to sustain continued plant growth but is seemingly low enough to promote pronounced development of root hairs. However, inclusion of Pi in the RNA-containing medium significantly reduces root hair density and elongation (Student's t test, P < 0.05; Fig. 3; Table I). Root hairs of plants grown on +Pi/+RNA medium are about 2-fold shorter and occur at a 3-fold lower density than those of −Pi/+RNA cultivated seedlings (Table I). It is important to note that Phi has a similar effect to Pi on root hair development. Both the length and the density of root hairs are significantly reduced by Phi (+Phi/−Pi/+RNA) when compared with seedlings grown on −Pi/+RNA medium (Student's t test, P < 0.05; Fig. 3; Table I).
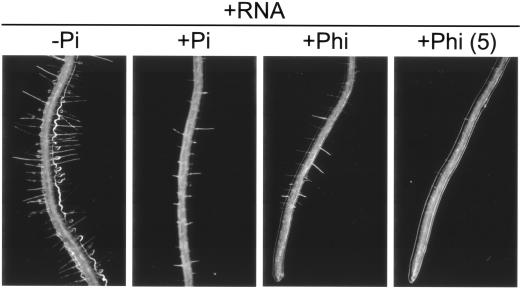
Figure 3.
Light microscope images of growing Arabidopsis roots. Plants were grown on vertically oriented agar plates for 21 d in +RNA (2 mm phosphorus) media containing the following supplements: −Pi, +Pi (2 mm), and +Phi (2.5 mm, or as indicated, mm, in parentheses). Addition of both Pi and Phi suppresses proliferation of root hairs (Student's t test, P < 0.05, see Table I).
Table I.
Inhibition of root hair formation by Pi and Phi
| Treatment | Root Hair Density | Root Hair Length |
|---|---|---|
| mm−1 | mm | |
| −Pi/+RNA | 11.50 ± 1.09 (100) | 0.267 ± 0.016 (100) |
| +Pi/+RNA | 3.18 ± 0.32 (27.7) | 0.136 ± 0.017 (50.9) |
| +Phi/+RNA | 4.50 ± 0.99 (39.1) | 0.176 ± 0.010 (65.9) |
Arabidopsis plants were grown on vertically oriented agar plates for 21 d in media with the following phosphorus supplements or combinations thereof: −Pi; +Pi (2 mm), +Phi (2.5 mm), +RNA (2 mm phosphorus). Values represent the mean root hair density ± se or mean root hair length ± se (n = 8–10 primary roots) and are expressed as percent of the respective −Pi/+RNA treatment (nos. in parenthesis). The differences between roots of −Pi/+RNA and +Pi/+RNA as well as between −Pi/+RNA and +Phi/−Pi/+RNA-grown seedlings are statistically significant (Student's t test, P < 0.05).
Effect of Phi on Leaf Pigment Accumulation
The accumulation of anthocyanins is characteristic of Pi-limited plants (Trull et al., 1997). As opposed to Arabidopsis plants grown on −Pi medium, which appear dark green-purple, the leaves of plants grown on −Pi/+Phi medium have a normal to light-green appearance (see Fig. 1). To determine the effect of Phi on anthocyanin accumulation, plants were grown for 3 d on +Pi medium and subsequently transferred to −Pi, +Pi, and −Pi/+Phi media for an additional 14 d of growth. As demonstrated by measurements of extractable anthocyanins, levels of anthocyanins in −Pi/+Phi-cultivated plants were approximately 10-fold lower than those grown in −Pi and were similar to those of +Pi-grown plants (Student's t test, P < 0.05). It is interesting that Phi concentrations as low as 0.25 mm were sufficient to prevent anthocyanin accumulation (Fig. 4A).
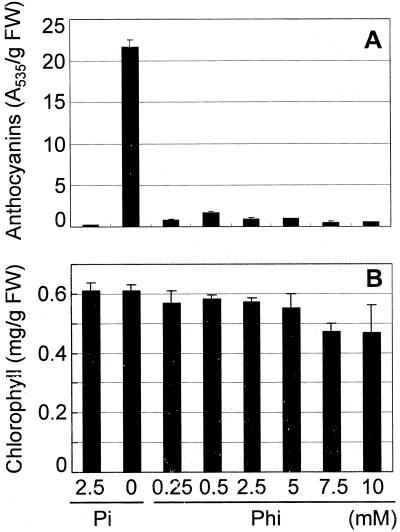
Figure 4.
Content of leaf pigments in Arabidopsis plants. Seedlings were grown for 3 d on +Pi medium and transferred for 14 d to medium containing the indicated concentrations of Pi and Phi. Anthocyanins (A) and chlorophylls (B) were extracted and quantified as described in “Materials and Methods.” Mean pigment content per gram fresh weight (±se) is given (n = 3 independent extractions of 100 shoots).
Plants grown at high Phi concentrations are light green in color, which may indicate a possible alteration in chlorophyll content (see Fig. 1). Therefore, we also measured chlorophyll a and b in leaves of plants grown in conditions of different phosphorus supply as described for the anthocyanin measurements. Although total chlorophyll content did not significantly differ between seedlings grown on −Pi, +Pi, and −Pi/+Phi (2.5 mm) media, chlorophyll levels were modestly decreased by Phi at concentrations above 5 mm Phi (Fig. 4B).
Phi Disrupts Induction of Nucleolytic Enzymes by Phosphate Limitation
Induction of Pi starvation-inducible enzymes such as acid phosphatase is a common molecular response of plants to limited Pi availability (Duff et al., 1994). This induction was demonstrated previously for Arabidopsis (Trull et al., 1997; Trull and Deikman, 1998; Chen et al., 2000). To examine the effect of Pi and Phi on Pi starvation-inducible acid phosphatase activity in roots, plants were germinated in high Pi medium and then transferred to +Pi, −Pi, and −Pi/+Phi media. Roots were histochemically stained for acid phosphatase activity using 5-bromo-4-chloro-3-indolyl-phosphate (BICP) as a chromogenic substrate. As shown in Figure 5, roots of Pi-starved plants showed intense staining, which indicates high acid phosphatase activity, whereas no detectable staining was observed for roots of plants that were grown on +Pi or −Pi/+Phi medium. This observation was further confirmed by quantitative enzyme assays (Table II). Spectrophotometric measurement of acid phosphatase specific activity in total plant extracts indicated a reduction by approximately 60% for plants grown on +Pi or −Pi/+Phi medium when compared with seedlings grown on −Pi medium. Similar results were obtained for total phosphodiesterase specific activity, whereas total ribonuclease specific activity was reduced by 90% under the same conditions (Student's t test, P < 0.05).
Figure 5.
Staining for acid phosphatase activity. Seedlings were grown for 3 d on +Pi medium and transferred for 14 d to −Pi, +Pi (2.5 mm), or +Phi (2.5 mm) medium. Histochemical staining for acid phosphatase activity of roots with the BICP substrate was performed as described in “Materials and Methods.”
Table II.
Repression of Pi starvation-inducible enzymes by Pi and Phi
| Treatment | Acid Phosphatase | Phosphodiesterase | Ribonuclease |
|---|---|---|---|
| nmol min−1 mg−1 protein | ΔA260 min−1 mg−1 protein | ||
| −Pi | 219 ± 14 (100) | 24.7 ± 1.2 (100) | 1.32 ± 0.06 (100) |
| +Pi | 111 ± 18 (50.7) | 9.6 ± 1.1 (38.9) | 0.11 ± 0.05 (8.3) |
| +Phi | 67 ± 16 (30.6) | 9.2 ± 3.5 (37.2) | 0.16 ± 0.04 (12.1) |
Seedlings were grown for 3 d on +Pi medium and transferred for 14 d to −Pi, +Pi (2.5 mm), or +Phi (2.5 mm) medium. Spectrophotometric assays for acid phosphatase, phosphodiesterase, and ribonuclease were performed as described in “Materials and Methods”. Mean specific enzyme activities (±se) are given n = 3 independent extractions) and expressed as percent of the −Pi treatment (nos. in parentheses). Enzyme activities are significantly reduced in plants grown in +Pi and +Phi medium when compared with plants grown in −Pi medium (Student's t test, P < 0.05).
To compare the inhibitory effect of Pi and Phi on the induction of Pi starvation-inducible enzymes in more detail, we examined the effect of increasing concentrations of both anions on the expression of acid phosphatase and ribonuclease activities. Seeds were germinated for 3 d on +Pi medium and were transferred subsequently to media containing various concentrations of Pi or Phi. After 14 d of additional growth, plant extracts were analyzed by in-gel assays for both enzyme activities (Fig. 6). When compared with extracts of Pi-starved plants, increasing concentrations of Pi in the growth medium resulted in a steady reduction of extractable acid phosphatase and ribonuclease activities between 0.025 and 2.5 mm Pi. It is interesting that increasing Phi concentrations in the agar medium had a similar effect on both enzyme activities, which was most effective at concentrations above 0.05 mm Phi (Fig. 6). In summary, these data demonstrate repression by Phi of Pi starvation-inducible nucleolytic enzymes.
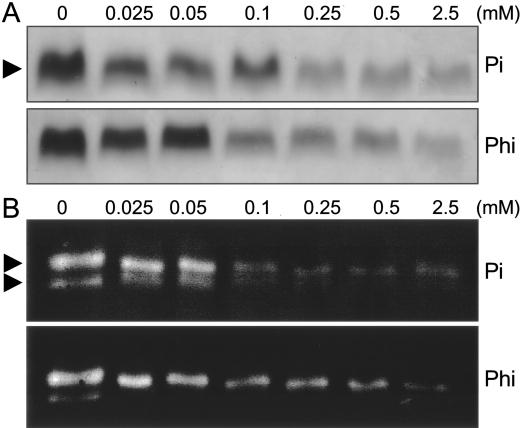
Figure 6.
Gel electrophoretic analysis of Pi starvation-inducible enzyme activities in Arabidopsis plants. Seedlings were grown for 3 d on +Pi medium and transferred for 14 d to medium containing the indicated concentrations of Pi and Phi. In-gel assays of acid phosphatase (A, 15 μg of protein per each lane) and ribonuclease (B, 80 of μg protein per each lane) with total plant protein extracts were performed as described in “Materials and Methods.” Arrowheads point to the 80-kD major acid phosphatase isoform, and to two major ribonucleases of 23 and 26 kD, corresponding to RNS1 and RNS2, respectively (Chen et al., 2000). One experiment representative of four independent trials is shown.
Phi Selectively Represses Phosphate Starvation-Inducible Gene Expression
Next, we examined whether Phi represses Pi starvation-inducible gene expression at the level of steady-state transcript levels. To monitor Pi-regulated gene expression, we used gene probes coding for an acid phosphatase (AtACP5), a Pi transporter (AtPT2), and a protein of unknown function (At4). These genes are induced by Pi limitation but are repressed in high Pi (Muchhal et al., 1996; del Pozo et al., 1999; Burleigh and Harrison, 1999). Seeds were germinated for 3 d on +Pi medium and then transferred to +Pi, −Pi, and −Pi/+Phi medium. After 14 d of growth, nucleic acids were extracted and steady-state mRNA levels examined by RNA gel-blot analysis. As shown in Figure 7, Pi starvation dramatically increased transcript levels of AtACP5, AtPT2, and At4 when compared with plants grown in Pi-sufficient condition. As expected, cultivation in −Pi/+Phi medium did not appreciably increase mRNA levels of the three genes tested, which were comparable with the transcript levels determined in +Pi-grown plants.
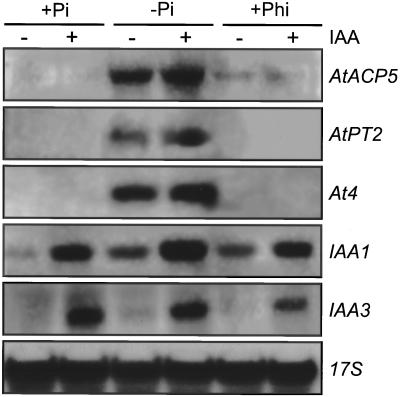
Figure 7.
RNA gel-blot analysis of auxin- and Pi starvation-inducible genes in Arabidopsis plants. Seedlings were grown for 3 d on +Pi (2.5 mm) medium, transferred for 14 d to +Pi (2.5 mm), −Pi, and −Pi/+Phi(10 mm) medium, and treated with (+) or without (−) 20 μm indole-3-acetic acid (IAA) for 2 h. Thirty micrograms of glyoxylated RNA was separated by electrophoresis, blotted, and probed with gene-specific probes for AtACP5, AtPT2, At4, IAA1, and IAA3. Hybridization with a rice (Oryza sativa) 17S rRNA probe (17S) was used as a control for equal loading.
To assess whether gene repression by Phi is a specific response rather than a nonspecific, toxic effect of the Pi analog, we examined auxin inducibility of two primary auxin response genes, IAA1 and IAA3 (Abel et al., 1995), in the same growth conditions. Plants were grown on +Pi, −Pi, and −Pi/+Phi media as described above and were additionally treated for 2 h with IAA or with incubation buffer only. RNA gel-blot analysis with the two IAA gene probes demonstrated that plants grown in −Pi/+Phi conditions retained the capacity to respond to the plant hormone by transcriptional activation of early auxin-inducible genes, similar to those plants that were grown in −Pi or +Pi conditions (Fig. 7). These data indicate selective repression of Pi starvation-inducible genes by Phi.
DISCUSSION
In this study, we examined the effect of the Pi analog, Phi, on plant growth and Pi starvation responses of Arabidopsis. We observed that Phi at concentrations above 2.5 mm significantly inhibited plant growth in phosphorus-sufficient (+Pi or −Pi/+RNA) and insufficient (−Pi) media, whereas lower Phi concentrations had only a marginal effect on seedlings grown in all conditions. Similar results were previously reported for B. nigra seedlings and B. napus cell suspension cultures. Phi-inhibited growth correlated with reduced internal Pi concentrations, indicating competitive action of Phi on Pi assimilation as a cause for growth inhibition (Carswell et al., 1996, 1997). 31P-NMR spectroscopic studies in B. nigra further revealed that Phi is metabolically stable and not converted to other compounds (Carswell et al., 1996). Thus, inhibition of Arabidopsis growth by Phi is likely to be a consequence of competitive inhibition of Pi assimilation and may further reflect an inability of Arabidopsis to readily utilize Phi via oxidation to Pi.
Although Phi apparently cannot substitute for Pi in terms of meeting the nutritional phosphorus requirements of plants, our data indicate that Phi can substitute for Pi in repressing typical molecular and developmental responses to Pi limitation. Biochemical adaptations to Pi starvation include increased synthesis of anthocyanins, presumably to adjust photosynthetic light reactions to the Pi-dependent Calvin cycle, and increased synthesis of enzymes for scavenging intra- and extracellular phosphorus (Trull et al., 1997; Bosse and Köck, 1998; Raghothama, 1999). We previously demonstrated that Pi starvation coordinately induces ribonuclease, phosphodiesterase, and acid phosphatase activities in tomato (Lycopersicon esculentum) cell suspension cultures, which sequentially participate in the complete hydrolysis of exogenous RNA to nucleosides and Pi (Nürnberger et al., 1990; Abel et al., 2000). Here, we found that Phi effectively prevents accumulation of anthocyanins and represses these nucleolytic enzymes in Pi limiting conditions at concentrations that are not growth inhibitory. Seedlings transferred to +Pi or +Phi medium expressed 50% to 90% lower activities of nucleolytic enzymes than seedlings transferred to −Pi medium. These results are in agreement with studies in Brassica sp., which showed that Phi reduced the induction of phosphoenolpyruvate phosphatase, pyrophosphate-dependent phosphofructokinase, and high-affinity Pi uptake in Pi-limiting conditions by 40% to 90% (Carswell et al., 1996, 1997). We extended our studies to compare the effect of Pi and Phi on steady-state levels of Pi-responsive mRNAs, which encode a Pi transporter, AtPT2 (Muchhal and Raghothama, 1999), an acid phosphatase, AtACP5 (del Pozo et al., 1999), and an unknown gene product, At4 (Burleigh and Harrison, 1999). Our data clearly indicate that Phi represses Pi starvation-inducible gene expression at the transcript level. It is important to note that in all conditions tested, Arabidopsis seedlings retained the capacity to respond to auxin with the induction of early auxin-inducible genes, IAA1 and IAA3. The ability to transcriptionally activate early auxin genes (Koshiba et al., 1995) suggests that repression of Pi-responsive gene expression by Phi is selective and not caused by a possible general cellular toxicity of the Pi analog. Selective attenuation of Pi starvation responses by Phi is further indicated by total chlorophyll levels, which did not appreciably change in plants grown on medium containing concentrations of up to 5 mm Phi. Likewise, Phi treatment of Brassica sp. seedlings and cell suspension cultures grown in +Pi and −Pi conditions did not significantly change total soluble protein concentrations or the activities of pyruvate kinase and ATP-dependent phosphofructokinase, enzymes that are not associated with the Pi starvation response of plants (Carswell et al., 1996, 1997).
Typical morphological effects of Pi limitation include dramatic changes in root architecture, presumably to accelerate soil exploration. Such developmental acclimations include enhanced root growth relative to the shoot, altered patterns of lateral root branching, development of proteoid roots, and increased root hair formation (Lynch, 1995; Schachtman et al., 1998). Microscopic analysis of root hair formation revealed that Phi suppressed, to a similar extent as Pi, the enhanced root hair elongation and root hair density of seedlings grown in Pi-deficient media. Although root hair formation in response to Phi has not been examined in other plants, similar studies in lupin (Lupinus albus) showed that Phi treatment of Pi-starved plants reduced the development of proteoid roots (Gilbert et al., 2000). We further observed that Phi altered the root-to-shoot ratio of seedlings developing in −Pi medium, which was reported previously for B. nigra plants (Carswell et al., 1996). Although the total fresh weights of −Pi- and +Phi-germinated, and growth-arrested, seedlings were comparable, which are likely determined by available seed phosphorus reserves such as phytic acid, the root-to-shoot ratio of +Phi seedlings was significantly reduced when compared with −Pi seedlings. Thus, the presence of Phi appears to cause a redistribution of seed phosphorus reserves to benefit shoot growth, rather than root growth as observed in −Pi/−Phi seedlings. A lower root-to-shoot ratio of +Phi seedlings is also consistent with the inhibitory effect of Phi on root hair formation, anthocyanin accumulation in the shoot, and induction of Pi starvation-inducible enzymes.
In summary, our study in Arabidopsis supports and extends previous work on the biological effects of Phi in Brassica sp. (Carswell et al., 1996, 1997) and in tomato (Varadarajan and Raghothama, 2000). Our data provide compelling new evidence for the hypothesis that Phi specifically interferes with the manifestation of a wide range of biochemical and developmental Pi starvation responses. Although Phi is not a substrate in enzymatic phosphoryl transfer reactions (Guest and Grant, 1991), other Pi-binding proteins, such as plasmalemma Pi transporters or signaling components involved in monitoring Pi status, apparently do not discriminate between Pi and Phi (Carswell et al., 1997). These properties of Phi and its dual, concentration-dependent effect in plants suggest the value of Phi as a molecular tool to manipulate and investigate Pi sensing in plants. Although Phi at concentrations below 2.5 mm attenuates Pi starvation responses in a similar manner to Pi and does not significantly affect growth of Arabidopsis, higher Phi concentrations clearly inhibit plant development, likely by competing with Pi assimilation as shown for Brassica sp. (Carswell et al., 1996, 1997). The dual effect of Phi on Pi starvation responses and plant growth may be exploited in a genetic approach to dissect Pi starvation response pathways. We have previously isolated a number of conditional, insensitive Pi starvation response mutants in Arabidopsis by screening for plants that fail to develop normally on nucleic acid-containing media but that can be rescued on +Pi medium (Chen et al., 2000). In a converse manner, addition of Phi to nucleic acid-containing media at concentrations that inhibit Pi starvation responses and plant growth may allow for the selection of mutants that constitutively express Pi starvation-inducible nucleolytic enzymes. The Pi derived from exogenous nucleic acid substrates may out-compete the growth-inhibitory effect of Phi and allow for growth of constitutive Pi starvation response mutants (see Fig. 1). Such constitutive mutants will greatly contribute to our understanding of the signaling pathways that mediate molecular and developmental responses of plants to Pi limitation.
MATERIALS AND METHODS
Plant Material and Growth Conditions
Wild-type Arabidopsis ecotype Columbia was obtained from the Arabidopsis Biological Resource Center (Ohio State University, Columbus). Seeds were surface sterilized and germinated on 0.8% (w/v) phytagar medium containing 5 mm KNO3, 2.5 mm KH2PO4 (adjusted to pH 5.5 with KOH), 2 mm MgSO4, 2 mm Ca(NO3)2, 50 μm Fe-EDTA, 70 μm H3BO3, 14 μm MnCl2, 0.5 μm CuSO4, 1 μm ZnSO4, 0.2 μm NaMoO4, 10 μm NaCl, and 0.01 μm CoCl2. In addition, 2.5 mm MES [2-(N-morpholino)-ethanesulfonic acid]-KOH (pH 5.5) and 0.5% (w/v) Suc were included. This medium is referred to as high or +Pi medium. For low or −Pi medium, KH2PO4 was omitted from the above nutrient solution. For +Phi medium, a stock solution of filter-sterilized 100 mm potassium Phi (pH 5.5) freshly prepared from phosphorous acid (Aldrich Chemicals, Milwaukee, WI) and KOH, was added to autoclaved −Pi medium to the specified Phi concentrations. For RNA-containing (+RNA) medium, filter-sterilized purified RNA was substituted for KH2PO4 and added to the autoclaved medium to a final concentration of 0.6 mg mL−1, equaling about 2 mm total phosphorus (Chen et al., 2000). Phytagar (commercial grade) was purchased from Gibco-BRL (Gaithersburg, MD) and contributed about 25 μm total phosphorus to the final medium (catalog no. 10695-039, lot no. 1021483). After stratification at 4°C for 2 to 3 d, seeds were germinated in a controlled environmental chamber at 22°C under illumination with fluorescent and incandescent light at a photon fluence rate of approximately 60 μmol m−2 s−1 for 16 h daily. To facilitate fresh weight determination and analysis of roots, plants were grown on vertically oriented agar plates as indicated.
Pigment Analysis
Anthocyanins were determined according to Lange et al. (1971). Seedlings were homogenized in propanol:HCl:water (18:1:81) and further extracted in a boiling water bath for 3 min. After centrifugation, the absorbance of the supernatant was measured at 535 and 650 nm. The absorbance due to anthocyanins was calculated as A = A535 − A650. Chlorophyll content was analyzed according to Chapman (1988). Seedlings were ground in 80% (w/v) acetone, and the absorbance of the supernatant was measured at 663 and 646 nm. Chlorophyll concentration was calculated as follows: chlorophyll a (mg L−1) = 12.21 A663 − 2.81 A646, chlorophyll b (mg L−1) = 20.13 A646 − 5.03 A663, and total chlorophyll (mg L−1) = 17.3 A646 + 7.18 A663 (Chapman, 1988).
Histochemical Staining
For histochemical staining of acid phosphatase activity in roots, plants were grown on agar plates of the indicated media composition. The agar surface subsequently was overlayed with 0.6% (w/v) phytagar containing 0.008% (w/v) BICP (Sigma, St. Louis). Plates were incubated overnight and photographs were taken.
Root Analysis
Images of the roots were captured with a Cool Snap high-performance CCD camera and imported into the public domain National Institutes of Health Image program for quantitative analysis of root hair length and density.
Electrophoretic Analysis of Ribonucleases and Acid Phosphatases
Total proteins of Arabidopsis seedlings were isolated as described by Yen and Green (1991) and equal amounts per treatment were separated by SDS-PAGE using the discontinuous buffer system of Laemmli (1970). Proteins were allowed to renature during repeated washes of gels with 25% (w/v) isopropanol. Ribonuclease isoenzymes separated in 12% (w/v) polyacrylamide gels and acid phosphatase isoenzymes separated in 7.5% (w/v) polyacrylamide gels were detected according to Yen and Green (1991) and Trull et al. (1997), respectively.
Enzyme Assays
Plant extracts were prepared according to Yen and Green (1991), using 100 mm Tris-acetate, 100 mm potassium acetate, 250 mm sodium ascorbate, 2 mm EDTA, 5 mm DTT, 10% (w/v) glycerol (pH 8.0) as extraction buffer. Acid phosphatase activity was measured in a total volume of 0.06 mL containing 100 mm acetic acid-NaOH (pH 5.5) 5 mm p-nitrophenyl phosphate as the substrate and appropriate amounts of plant extract (Chen et al., 2000). Phosphodiesterase activity was measured in 50 mm acetic acid-NaOH (pH 6.0) and 5 mm MgCl2 using bis(p-nitrophenyl) phosphate as the substrate (Abel et al., 2000). Reactions were incubated in microtiter wells at 37°C, terminated by the addition of 200 μL of 1 m Na2CO3, and assayed spectrophotometrically. Acid phosphatase and phosphodiesterase activities of plant extracts were calculated using a molar extinction coefficient of 18,300 cm2 nmol−1 for p-nitrophenol at 405 nm. Ribonuclease activity was determined spectrophotometrically by the release of ethanol-soluble hydrolysis products according to Abel and Glund (1987). One ribonuclease unit is defined as the amount of enzyme causing an increase in ΔA260 of 1.0 min−1 cm−1 mL−1 (Wilson, 1982). For calculation of specific enzyme activities, assays were performed under conditions of demonstrated linearity with respect to time and protein amount. Five to 10 μg of total protein extract typically were used per reaction.
Other Assays
Protein was assayed according to the method of Bradford (1976) using bovine serum albumin as the standard. The method of Ames (1966) was used to determine the total phosphorus content of phytagar and the Pi concentration of growth media.
Isolation of Nucleic Acids and RNA Hybridization Analysis
Total nucleic acids were isolated using the phenol/chloroform extraction procedure described by Theologis et al. (1985). RNA hybridization analysis with various 32P-labeled probes, AtACP5 (del Pozo et al., 1999), At4 (Burleigh and Harrison, 1999), AtPT2 (Muchhal et al., 1996), and IAA1 and IAA3 (Abel et al., 1995), was carried out as described by Abel et al. (1995). Equal RNA loading was assessed by hybridization with a rice (Oryza sativa) 17S ribosomal DNA probe.
Statistical Analyses
Means ± se were calculated and Student's t tests were carried out. All differences were significant to at least a value of P < 0.05, which indicates that the probability that the differences in the two sample means were due to chance alone was less than 5%.
ACKNOWLEDGMENTS
We thank Drs. Maria Harrison, Kaschandra Raghothama, and Javier Paz-Ares for plasmid DNAs of At4, AtPT2, and AtACP5, respectively.
Footnotes
This work was supported by the United States Department of Energy and by the Conselho Nacional de Desenvolvimento Científico e Tecnológico, Brazil (predoctoral fellowship to C.A.D.).
Article, publication date, and citation information can be found at www.plantphysiol.org/cgi/doi/10.1104/pp.010396.
LITERATURE CITED
- Abel S, Glund K. Ribonuclease in plant vacuoles: purification and molecular properties of the enzyme from cultured tomato cells. Planta. 1987;172:71–78. doi: 10.1007/BF00403030. [DOI] [PubMed] [Google Scholar]
- Abel S, Nguyen MD, Theologis A. The PS-IAA4/5-like family of early auxin-inducible mRNAs in Arabidopsis. J Mol Biol. 1995;251:533–549. doi: 10.1006/jmbi.1995.0454. [DOI] [PubMed] [Google Scholar]
- Abel S, Nürnberger T, Ahnert V, Krauss G-J, Glund K. Induction of an extracellular cyclic nucleotide phosphodiesterase as an accessory ribonucleolytic activity during phosphate starvation of cultured tomato cells. Plant Physiol. 2000;122:543–552. doi: 10.1104/pp.122.2.543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ames BN. Assay of inorganic phosphate, total phosphate and phosphatases. Methods Enzymol. 1966;8:115–118. [Google Scholar]
- Baldwin JC, Karthikeyan AS, Raghothama KG. LEPS2, a phosphorus starvation-induced novel acid phosphatase from tomato. Plant Physiol. 2001;125:728–737. doi: 10.1104/pp.125.2.728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bates TR, Lynch JP. Stimulation of root hair elongation in Arabidopsis by low phosphorus availability. Plant Cell Environ. 1996;19:529–538. [Google Scholar]
- Bates TR, Lynch JP. The efficiency of Arabidopsis (Brassicaceae) root hairs in phosphorus acquisition. Am J Bot. 2000;18:964–970. [PubMed] [Google Scholar]
- Bosse D, Köck M. Influence of phosphate starvation on phosphohydrolases during development of tomato seedlings. Plant Cell Environ. 1998;21:325–332. [Google Scholar]
- Bradford MM. A rapid and sensitive method for the quantification of microgram quantities of proteins utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Burleigh SH, Harrison MJ. The down-regulation of Mt4-like genes by phosphate fertilization occurs systemically and involves phosphate translocation to the shoots. Plant Physiol. 1999;119:241–248. doi: 10.1104/pp.119.1.241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carswell MC, Grant BR, Plaxton WC. Disruption of the phosphate-starvation response of oilseed rape suspension cells by the fungicide phosphonate. Planta. 1997;203:67–74. doi: 10.1007/s00050166. [DOI] [PubMed] [Google Scholar]
- Carswell MC, Grant BR, Theodorou ME, Harris J, Niere JO, Plaxton WC. The fungicide phosphonate disrupts the phosphate-starvation response in Brassica nigra seedlings. Plant Physiol. 1996;110:105–110. doi: 10.1104/pp.110.1.105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chapman DJ. Qualitative analysis of pigments. In: Lobban CS, Chapman DJ, Kremer BP, editors. Experimental Phycology: A Laboratory Manual. New York: Cambridge University Press; 1988. pp. 93–103. [Google Scholar]
- Chen DL, Delatorre CA, Bakker A, Abel S. Conditional identification of phosphate-starvation-response mutants in Arabidopsis. Planta. 2000;211:13–22. doi: 10.1007/s004250000271. [DOI] [PubMed] [Google Scholar]
- del Pozo JC, Allona I, Rubio V, Leyva A, de la Pena A, Aragoncillo C, Paz-Ares J. A type 5 acid phosphatase gene from Arabidopsis is induced by phosphate starvation and by some other types of phosphate mobilizing/oxidative stress conditions. Plant J. 1999;19:579–589. doi: 10.1046/j.1365-313x.1999.00562.x. [DOI] [PubMed] [Google Scholar]
- Duff SMG, Sarath G, Plaxton WC. The role of acid phosphatases in plant phosphorus metabolism. Physiol Plant. 1994;90:791–800. [Google Scholar]
- Gilbert GA, Knight A, Vance CP, Allan DL. Proteoid root development of phosphorus deficient lupin is mimicked by auxin and phosphonate. Ann Bot. 2000;85:921–928. [Google Scholar]
- Goldstein AH, Danon A, Beartlein DA, McDaniel RG. Phosphate starvation inducible metabolism in Lycopersicon esculentum: II. Characterization of the phosphate starvation-inducible-excreted acid phosphatase. Plant Physiol. 1988;87:716–720. doi: 10.1104/pp.87.3.716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Green PJ. The ribonucleases of higher plants. Annu Rev Plant Physiol Plant Mol Biol. 1994;45:421–445. [Google Scholar]
- Guest D, Grant BR. The complex actions of phosphonates as antifungal agents. Biol Rev. 1991;66:159–187. [Google Scholar]
- Koshiba T, Ballas N, Wong L-M, Theologis A. Transcriptional regulation of PS-IAA4/5 and PS-IAA6 early gene expression by indoleacetic acid and protein synthesis inhibitors in pea (Pisum sativum) J Mol Biol. 1995;253:396–413. doi: 10.1006/jmbi.1995.0562. [DOI] [PubMed] [Google Scholar]
- Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lange H, Shropshire W, Mohr H. An analysis of phytochrome-mediated anthocyanin synthesis. Plant Physiol. 1971;47:649–655. doi: 10.1104/pp.47.5.649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lenburg ME, O'Shea EK. Signaling phosphate starvation. Trends Biochem Sci. 1996;21:383–387. [PubMed] [Google Scholar]
- Liu C, Muchhal US, Raghothama KG. Differential expression of TPS11, a phosphate starvation-induced gene in tomato. Plant Mol Biol. 1997;33:867–874. doi: 10.1023/a:1005729309569. [DOI] [PubMed] [Google Scholar]
- Lynch J. Root architecture and plant productivity. Plant Physiol. 1995;109:7–13. doi: 10.1104/pp.109.1.7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCully ME. Roots in soil: unearthing the complexities of roots and their rhizosphere. Annu Rev Plant Physiol Mol Biol. 1999;50:219–243. doi: 10.1146/annurev.arplant.50.1.695. [DOI] [PubMed] [Google Scholar]
- Muchhal US, Pardo JM, Raghothama KG. Phosphate transporters from the higher plant Arabidopsis. Proc Natl Acad Sci USA. 1996;93:10519–10523. doi: 10.1073/pnas.93.19.10519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muchhal US, Raghothama KG. Transcriptional regulation of plant phosphate transporters. Proc Natl Acad Sci USA. 1999;96:5868–5872. doi: 10.1073/pnas.96.10.5868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nürnberger T, Abel S, Jost W, Glund K. Induction of an extracellular ribonuclease in cultured tomato cells upon phosphate starvation. Plant Physiol. 1990;92:970–976. doi: 10.1104/pp.92.4.970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raghothama KG. Phosphate acquisition. Annu Rev Plant Physiol Plant Mol Biol. 1999;50:665–693. doi: 10.1146/annurev.arplant.50.1.665. [DOI] [PubMed] [Google Scholar]
- Raghothama KG. Phosphate transport and signaling. Curr Opn Plant Biol. 2000;3:182–187. [PubMed] [Google Scholar]
- Schachtman DP, Reid RJ, Ayling SM. Phosphorus uptake by plants: from soil to cell. Plant Physiol. 1998;116:447–453. doi: 10.1104/pp.116.2.447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theodorou ME, Plaxton WC. Metabolic adaptations of plant respiration to nutritional phosphate deprivation. Plant Physiol. 1993;101:339–344. doi: 10.1104/pp.101.2.339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theologis A, Huynh TV, Davis RW. Rapid induction of specific mRNAs by auxin in pea epicotyl tissue. J Mol Biol. 1985;183:53–68. doi: 10.1016/0022-2836(85)90280-3. [DOI] [PubMed] [Google Scholar]
- Torriani-Gorini A. The Pho regulon of Escherichia coli. In: Torriani-Gorini A, Yagil E, Silver S, editors. Phosphate in Microorganisms: Cellular and Molecular Biology. Washington, DC: American Society for Microbiology; 1994. pp. 1–4. [Google Scholar]
- Trull MC, Deikman J. An Arabidopsis mutant missing one acid phosphatase isoform. Planta. 1998;206:544–550. doi: 10.1007/s004250050431. [DOI] [PubMed] [Google Scholar]
- Trull MC, Guiltinan MJ, Lynch JP, Deikman J. The responses of wild-type and ABA mutant Arabidopsis plants to phosphorus starvation. Plant Cell Environ. 1997;20:85–92. [Google Scholar]
- Varadarajan D, Raghothama KG (2000) Phosphite, a structural analog of phosphate, suppresses Pi starvation in-duced molecular responses in tomato (abstract no. 470). Annual American Society of Plant Physiologists Meeting, July 15–19, 2000, San Diego
- Wilson CM. Plant nucleases: biochemistry and development of multiple molecular forms: isoenzymes. Curr Top Biol Med Res. 1982;6:33–54. [PubMed] [Google Scholar]
- Yen Y, Green PJ. Identification and properties of the major ribonucleases of Arabidopsis. Plant Physiol. 1991;97:1487–1493. doi: 10.1104/pp.97.4.1487. [DOI] [PMC free article] [PubMed] [Google Scholar]