Abstract
The human survival motor neuron (SMN) gene is the spinal muscular atrophy-determining gene, and a knockout of the murine Smn gene results in preembryonic lethality. Here we show that SMN can directly interact in vitro and in vivo with the large nonstructural protein NS1 of the autonomous parvovirus minute virus of mice (MVM), a protein essential for viral replication and a potent transcriptional activator. Typically, SMN localizes within nuclear Cajal bodies and diffusely in the cytoplasm. Following transient NS1expression, SMN and NS1 colocalize within Cajal bodies. At early time points following parvovirus infection, NS1 fails to colocalize with SMN within Cajal bodies; however, during the course of MVM infection, dramatic nuclear alterations occur. Formerly distinct nuclear bodies such as Cajal bodies, promyelocytic leukemia gene product (PML) oncogenic domains (PODs), speckles, and autonomous parvovirus-associated replication (APAR) bodies are seen aggregating at later points in infection. These newly formed large nuclear bodies (termed SMN-associated APAR bodies) are active sites of viral replication and viral capsid assembly. These results highlight the transient nature of nuclear bodies and their contents and identify a novel nuclear body formed during infection. Furthermore, simple transient expression of the viral nonstructural proteins is insufficient to induce this nuclear reorganization, suggesting that this event is induced specifically by a step in the viral infection process.
Minute virus of mice (MVM) is an autonomous parvovirus consisting of a single-stranded genome approximately 5 kb in length (12, 13). The genome contains two overlapping transcription units controlled by two viral promoters: P4 and P38 (2, 16, 32, 33). Expression of the viral nonstructural proteins, NS1 and NS2, is driven by the P4 promoter, while P38-derived transcripts encode the viral structural proteins, VP1 and VP2 (2, 16, 32, 33). NS1 performs critical functions in viral gene expression and genome replication. NS1 is also required for MVM-associated cytotoxicity, although the mechanism(s) is unclear (3). Upon infection, parvoviruses cannot induce cells into S phase, so the virus must rely on actively dividing cells to naturally progress through the cell cycle; when the cells enter S phase, a productive, lytic infection can progress (12, 13). Cells infected by autonomous parvoviruses are prevented from further progress in the cell cycle (39-42) yet remain viable and sustain virus replication for many hours before they die (49).
Recently, a novel subnuclear compartment was identified following infection by either of two highly related parvoviruses, MVM and H-1 (5, 14). This structure, termed autonomous parvovirus-associated replication (APAR) bodies, was identified at 15 h postinfection and was found to be distinct from most of the classically described nuclear bodies such as Cajal bodies, promyelocytic leukemia gene product (PML) oncogenic domains (PODs), and speckles (14). APAR bodies were shown to be active sites of viral replication and to contain cellular factors such as cyclin A, DNA polymerases δ and α, proliferating cell nuclear antigen (PCNA), and replication protein A (4).
The eukaryotic nucleus is highly organized and contains numerous subnuclear compartments (34). Although nuclear bodies have been identified and classified based on their constituents, mounting evidence suggests that nuclear bodies and their contents are highly dynamic and can fluctuate in response to a variety of internal and external stimuli (28, 35, 46, 47). The functions of the various nuclear structures have been partially ascribed to their molecular constituents. For example, Cajal bodies are enriched in mature snRNPs and contain factors such as the survival motor neuron gene product (SMN), which is involved in snRNP biogenesis (9, 22, 29). PODs are distinct nuclear structures that are linked to transcription regulation and apoptosis and contain the PML protein, retinoblastoma protein Rb, Sp100, and PIC1/SUMO-1 (7, 45). Speckles or interchromatin granules are highly enriched in splicing factors, although it is unclear if these structures are storage sites for splicing factors or whether they actively participate in mRNA maturation (34, 36).
Now we show that MVM NS1 specifically interacts with a component of Cajal bodies, the SMN protein, and that SMN and NS1 colocalize in Cajal bodies following transient expression of NS1. As suggested by earlier results, at early time points following MVM infection of synchronized cells, NS1 and SMN fail to colocalize. However, at later time points (20 to 30 h postinfection) NS1 and SMN colocalize in large nuclear bodies that are the active sites of virus replication and viral capsid assembly. Unlike transfection of NS1 alone, at these later time points, MVM infection induces a massive nuclear reorganization in which constituents from Cajal bodies, PODs, interchromatin granules, and APAR bodies accumulate in multiple large nuclear bodies, termed SMN-associated APAR bodies (SAABs). These results highlight the dynamic nature of nuclear bodies and identify a novel structure generated in response to parvovirus infection. They also suggest that some feature(s) of virus replication other than merely expression of the viral nonstructural proteins is required for the observed nuclear reorganization following parvovirus infection.
MATERIALS AND METHODS
Virus propagation.
Wild-type prototypic MVM (MVMp) was grown and titered on NB324K cells and used to infect A92L and NB324K cells at a multiplicity of infection of 10. Cells were highly synchronized by growth in isoleucine-deficient media for 40 h, followed by incubation in the presence of 36 μM aphidicolin and MVM for 12 to 16 h. The cells were then washed in phosphate-buffered saline and released in complete Dulbecco's modified Eagle's medium. Viral replication is detectable 5 to 6 h postrelease (44). Neuraminidase at a concentration of 0.05 U/ml was added 7 h after release to prevent reinfection. At various time points cells were labeled for 20 min with bromodeoxyuridine (BrdU) at a concentration of 10 μM.
Recombinant proteins.
NS1 cDNA was cloned into glutathione S-transferase (GST) expression vector pGEX3X (Pharmacia). Wild-type SMN cDNA was cloned into the pET32c vector (Novagen). Transformed bacteria were induced with 1 mM isopropyl-β-d-thiogalactopyranoside for 4 h at 37°C. Expressed recombinant proteins were purified from inclusion bodies by sequential extraction with increasing urea concentrations (2, 4, 6, and 8 M) in phosphate-buffered saline (PBS) as previously described (52).
Transient transfections.
Approximately 105 A92L cells were transiently transfected with 2 μg of pCI:HA-NS1 by using calcium phosphate (32).
Cellular extracts.
Total cellular extracts were derived from approximately 105 A92L cells infected with MVM at 0 and 30 h postrelease. Total extracts were normalized, and equivalent levels were used for biomolecular interaction analysis experiments.
BIA.
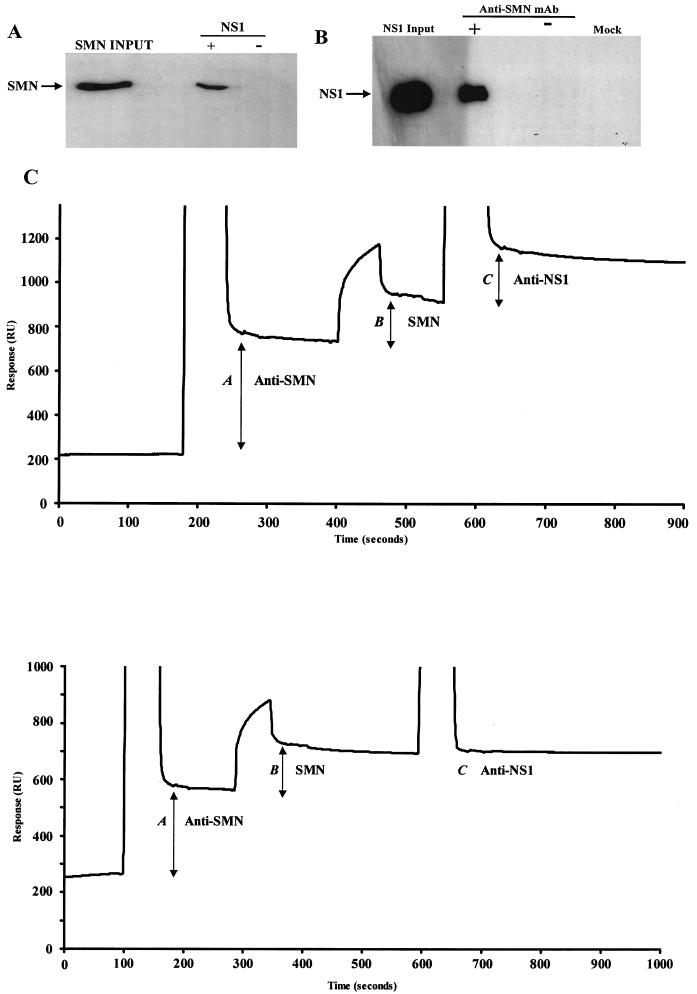
The BIAcore biosensor detects changes in total mass at the surface of a sensor chip by measuring variations of the critical angle needed to produce total internal refraction. The change in critical angle is proportional to the amount of bound protein and is expressed as resonance units (RU). In Fig. 1C, the shift in RU is plotted against time and is displayed as a sensorgram. Injected samples contain protein, urea, imidazole, and HEPES buffered saline (HBS) and therefore are denser than HBS running buffer. This increase in medium density during protein injections produces the large vertical spikes seen on the sensorgram (the initial vertical line indicates the start of an injection and the second vertical line indicates the end of an injection). The amount of bound protein following an injection is indicated by the increase in the horizontal baseline. A Cm5 sensor chip was covalently linked with rabbit anti-mouse immunoglobulin G by the amino-coupling protocol according to the manufacturer's instructions (BIAcore AB, Stevenage, Herts, United Kingdom). A BIAcore-X apparatus with an operating flow rate of 5 μl per min was used. All injected volumes were 10 μl. Concentrations of stock recombinant proteins for each reaction were determined prior to injection. Proteins were diluted in HBS to 10 μg/ml unless otherwise stated. During biomolecular interaction analysis (BIA) experiments, nonspecific interactions were ruled out by ensuring that neither SMN nor NS1 bound directly to the chip in the absence of specific antibodies or protein binding partners.
FIG. 1.
SMN and NS1 interact directly both in vitro and in vivo. (A) Western blot analysis of recombinant polyhistidine (six-His-)-tagged human SMN captured by GST-tagged NS1 immobilized on GST resin. The blot was developed with an anti-SMN monoclonal antibody. (B) Western blot analysis of NS1 coimmunoprecipitated with endogenous SMN from transfected A92L cells. Anti-SMN coprecipitated endogenous SMN, and anti-NS1 rabbit polyclonal sera detected NS1 complexed with SMN. NS1-SMN binding was not detected when the immunoprecipitating antibody (−) was omitted or when cells were not transfected with NS1 (mock). mAb, monoclonal antibody. (C) (Top) BIA of NS1 coimmunoprecipitated with endogenous SMN from MVM-infected double-blocked A92L cells. Total protein extracts from A92L cells 30 h postinfection were used. Anti-SMN (MANSMA3) coprecipitated SMN, and anti-NS1 rabbit polyclonal sera detected NS1 complexed with SMN. Levels of the captured anti-SMN antibody (A), SMN-NS1 complex (B), and anti-NS1 rabbit polyclonal antibody (C) are indicated. (Bottom) NS1 is not coprecipitated with SMN from double-blocked noninfected A92L cells. Total protein extracts from A92L cells 30 h after being released from the blocking process were used. Anti-SMN (MANSMA3) precipitated SMN, and anti-NS1 rabbit polyclonal sera detected no NS1 complexed with SMN. Levels the captured anti-SMN antibody (A), SMN (B), and anti-NS1 rabbit polyclonal antibody (C) are indicated.
Immunohistochemistry.
Subconfluent A92L and NB324K cell lines were washed three times with PBS, fixed for 1 min with 50% acetone-50% methanol, and then washed an additional three times with PBS. Cells were then analyzed by double-label immunofluorescence. Images were captured with tetramethyl rhodamine isothiocyanate (TRITC) and fluorescein isothiocyanate (FITC) filter sets and a 100× oil immersion objective in conjunction with a Leica TCS-NT confocal microscope. All experiments were repeated in the absence of the primary antibodies to eliminate the possibility of FITC-TRITC cross-reaction.
Immunofluorescence experiments were repeated with various primary antibodies to eliminate epitope masking and cross-reaction during time courses. Western blotting was performed on total protein extracts from all infection time points to ensure that each NS1 and SMN antibody used was specific to the protein it was generated against. All other antibodies were obtained from commercial vendors and have been shown to have no nonspecific cross-reactivity. The antibodies used were as follows: anti-SMN antibodies MANSMA1 (exon 2 specific), MANSMA4 (exon 4 specific), and MANSMA3 (exon 5 specific); anti-NS1 (monoclonal antibody 3D9); anti-MVM VP1 (peptide polyclonal antibody); anti-capsid/assembled (monoclonal antibody D4H) (30); anti-SIP1 antibody (MANSIP1A); anti-p80 coilin antibody (5P10); anti-PML antibody (PG-M3; Santa Cruz Biotechnology); anti-cyclin A antibody (C-19; Santa Cruz Biotechnology); anti-cyclin E (N-20; Santa Cruz Biotechnology), anti-BrdU (Chemicon International); anti-SR (1H4; Zymed Laboratories); NS1 (rabbit polyclonal antibody M50); and anti-Sm (Y12; Neomarkers).
In vitro binding assays.
GST and His-binding resins (Novagen) were washed three times in binding buffer (500 mM NaCl, 20 mM Tris-HCl [pH 7.8], 0.2% NP-40) and incubated with recombinant proteins diluted to 100 μg/ml in binding buffer for 1 h at 4°C. The resins were washed three times with binding buffer and incubated with the respective binding proteins diluted to 100 μg/ml in binding buffer for 1 h at 4°C. Resins were washed three times with binding buffer, and bound fractions were resolved by sodium dodecyl sulfate-polyacrylamide gel electrophoresis.
Protein concentrations were confirmed by Coomassie blue staining
RESULTS
Identification of NS1 as an SMN-interacting protein.
The NS1 nonstructural protein of MVM and the bovine papillomavirus transactivator E2 protein interact with similar subsets of cellular transcription factors to activate viral gene expression, including TATA binding protein TBP and Sp1 (8, 32, 43, 48). Recently, E2 has been shown to interact with SMN, a protein that has reported functions in transcriptional activation and RNA-processing pathways (29, 48). Based on the previously observed similarities between NS1 and E2 the following experiments were devised to determine whether NS1 and SMN associate. Initially, a bacterially expressed and purified GST-NS1 fusion protein and polyhistidine (six-His-)-tagged SMN were incubated together to determine whether an NS1-SMN complex formed in vitro. GST-NS1-bound fractions were separated with glutathione-linked agarose beads. Specific NS1-SMN in vitro binding was observed by Western analysis with an anti-SMN monoclonal antibody (Fig. 1A). SMN did not nonspecifically interact with the agarose beads alone (Fig. 1A) or with certain NS1-mutant GST fusions (P. J. Young, K. T. Jensen, D. J. Pintel, and C. L. Lorson, unpublished data). Coimmunoprecipitations were then performed with extracts from murine A92L cells to determine whether the NS1-SMN complex formed in vivo. A92L cells were transfected with hemagglutinin (HA)-tagged NS1, and coimmunoprecipitation reactions were performed with a monoclonal antibody specific for the peptide encoded by SMN exon 5. Endogenous SMN coprecipitated the HA-tagged NS1 protein (Fig. 1B). NS1 was not detected when cells were not transfected or when the anti-SMN monoclonal antibody was omitted from the immunoprecipitation reactions (Fig. 1B).
BIA was performed to confirm and further characterize the NS1-SMN interaction in vivo. To determine whether SMN and NS1 form a complex during MVM infection, highly synchronized A92L cells (23) were harvested at 30 h postrelease. BIA was used to immunoprecipitate endogenous SMN with an anti-SMN monoclonal antibody. At 30 h postinfection, NS1 coprecipitated with endogenous SMN (Fig. 1C, top). NS1 is not coprecipitated with SMN from double-blocked noninfected A92L cells (Fig. 1C, bottom; compare the horizontal plateaus for peak C in the top and bottom sensorgrams). A thorough analysis of the SMN-NS1 interaction is the subject of a manuscript in preparation (P. J. Young, K. T. Jensen, D. J. Pintel, and C. L. Lorson, unpublished data). Taken together these results demonstrate that NS1 and SMN interact in vivo and in vitro and form a complex during viral infection.
NS1 localizes in SAABs in the later stages of MVM infection.
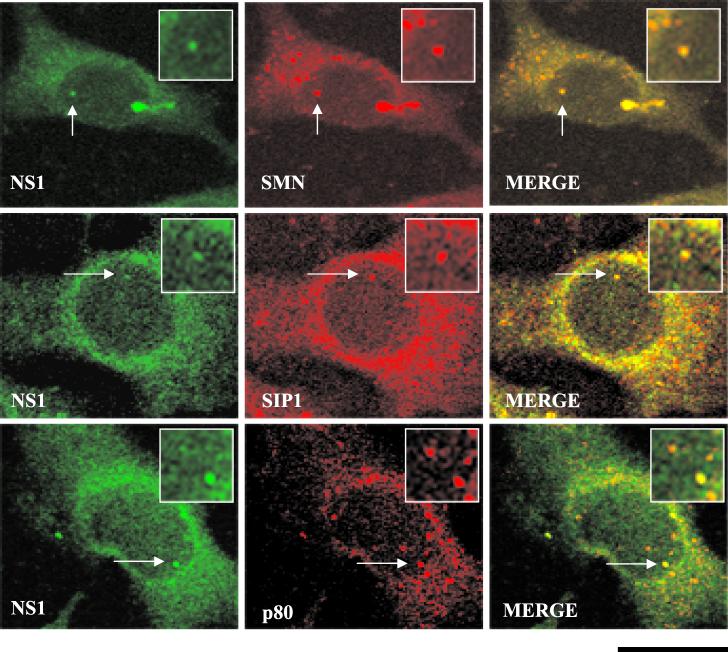
Nuclear SMN accumulates within Cajal bodies and the nucleolus (51, 52). At early time points (pre-15 h) after infection, however, NS1 has been reported to localize in APAR bodies, nuclear structures distinct from Cajal bodies. Since NS1 and SMN interact avidly in vitro and in vivo during infection, we performed the following series of indirect double-label immunofluorescence experiments to address this apparent discrepancy. First, HA epitope-tagged NS1 was transiently transfected into A92L cells. Cajal bodies were identified using antibodies specific for SMN, SIP1, and the Cajal body marker protein, p80 coilin. By 30 h after transient NS1 transfection, NS1 colocalized with all three proteins within Cajal bodies (Fig. 2). As expected, Cajal bodies were positively stained for SMN and SIP1 in mock-transfected cells.
FIG. 2.
Colocalization of SMN, p80 coilin, SIP1, and NS1 in transfected A92L cells. SMN (primary antibody: MANSMA1; secondary antibody: TRITC [red]), SIP1 (primary antibody: MANSIP1A; secondary antibody: anti-mouse TRITC conjugate [red]), p80 coilin (primary antibody: 5P10 monoclonal antibody; secondary: anti-mouse TRITC conjugate [red]), and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: anti-rabbit FITC conjugate [green]) are indicated. Arrows, Cajal bodies containing NS1. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm.
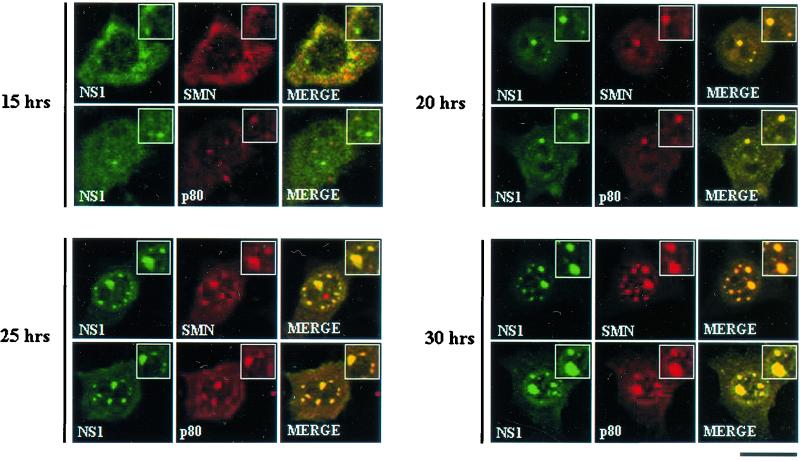
To determine whether this localization pattern was a consequence of NS1 transfection or whether similar distributions would be observed following viral infection, double-label immunofluorescence was performed on a series of highly synchronized A92L cells infected with MVM. Cells were harvested at multiple time points after release into S phase (0, 15, 20, and 30 h). Consistent with previous reports, at 15 h postrelease, NS1-positive APAR bodies and Cajal bodies were independent nuclear structures (Fig. 3). In contrast, at 20 h postrelease, SMN, p80 coilin (Fig. 3), SIP1 (data not shown), and NS1 (Fig. 3) colocalized in abnormally large nuclear structures. At subsequent time points (25 and 30 h), both the size and the number of these large nuclear bodies increased dramatically. Prior to release, nuclear structures displayed a normal distribution pattern.
FIG. 3.
SMN, p80 coilin, and NS1 colocalize in SAABs in MVM-infected A92L cells. SMN (primary antibody: anti-SMN monoclonal antibody MANSMA1; secondary antibody: anti-mouse TRITC conjugate [red]), p80 coilin (primary antibody: 5P10 monoclonal antibody; secondary antibody: anti-mouse TRITC conjugate [red]), and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: anti-rabbit FITC conjugate [green]) are indicated. Synchronized infections were obtained by performing an isoleucine-aphidicolin double block on A92L cells prior to infection. Immunofluorescence experiments were performed on infected cells 15, 20, 25, and 30 h after entry into S phase. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm.
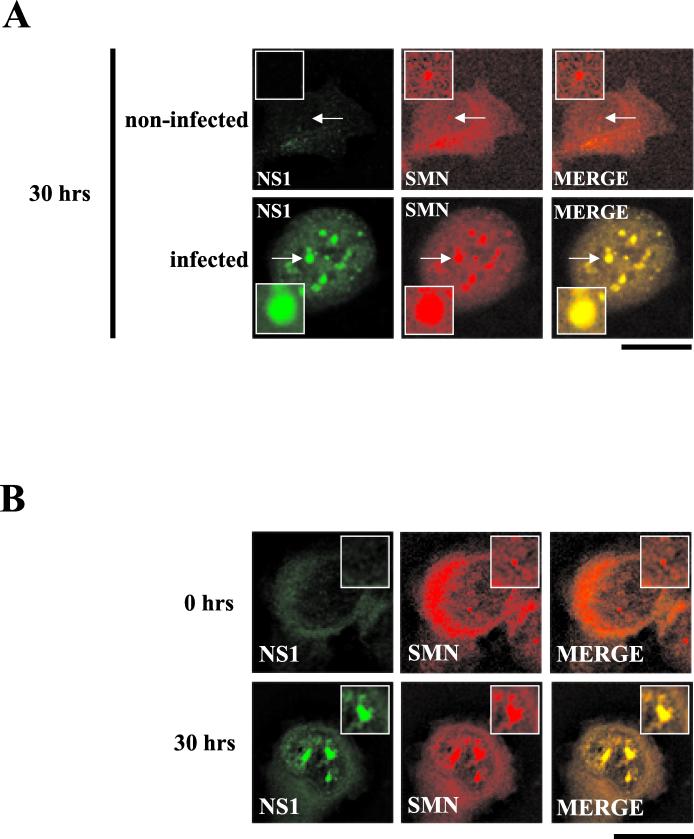
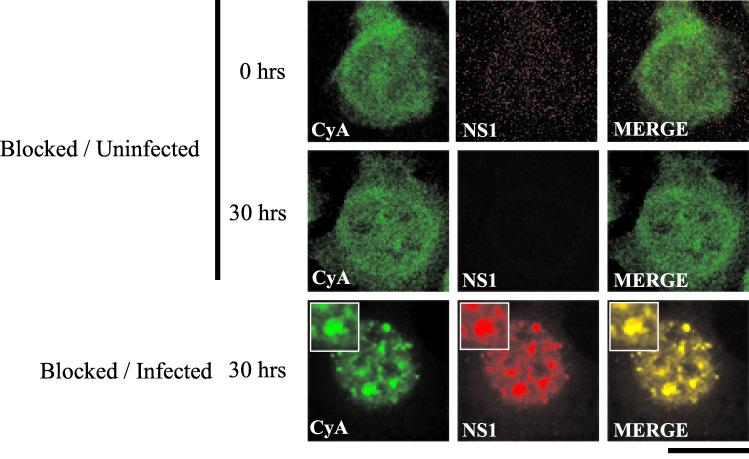
To confirm that the observed nuclear body formation was not a consequence of the synchronization procedure, immunofluorescence was performed in parallel with synchronized but uninfected A92L cells. In highly synchronized uninfected cells, SMN distribution was unaffected at 0 and 30 h postrelease, resulting in small, infrequent nuclear bodies, consistent with normal Cajal body staining and in contrast to the numerous large structures formed in A92L cells at 20 to 30 h postrelease (Fig. 4A). These large nuclear bodies were also observed at late time points following infection of unblocked A92L cells, confirming that the distribution pattern observed was not a direct consequence of the blocking process (data not shown). This novel localization pattern was not simply a consequence of a virus-induced overexpression of the various Cajal body components; BIA of total protein extracts from MVM-infected A92L cells 0 and 30 h after the cells entered S phase demonstrated that there was no significant increase in the levels of these proteins (data not shown).
FIG. 4.
SAAB formation is neither a consequence of the blocking process nor restricted to murine cell lines. (A) Double-label experiments were performed on infected and uninfected blocked A92L cells at 30 h postrelease. SMN (primary antibody: anti-SMN monoclonal antibody MANSMA1; secondary antibody: anti-mouse TRITC conjugate) and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: anti-rabbit FITC conjugate) are indicated. Arrows indicate SMN-positive nuclear bodies. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm. (B) SMN (primary antibody: anti-SMN monoclonal antibody MANSMA1; secondary antibody: TRITC) and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: FITC) are indicated. Experiments were performed on NB324K cells 0 and 30 h postinfection. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm.
To determine whether these large nuclear bodies were specific to murine cells, an MVM-permissive human cell line (NB324K) was also examined following MVM infection. At 0 h postinfection, discrete Cajal bodies were observed (Fig. 4B). At 30 h postinfection, multiple large nuclear bodies formed, demonstrating that these large structures were not cell type specific but rather were a consequence of viral infection. Similar to murine cells, the uninfected NB324K cells did not exhibit these large nuclear structures (data not shown).
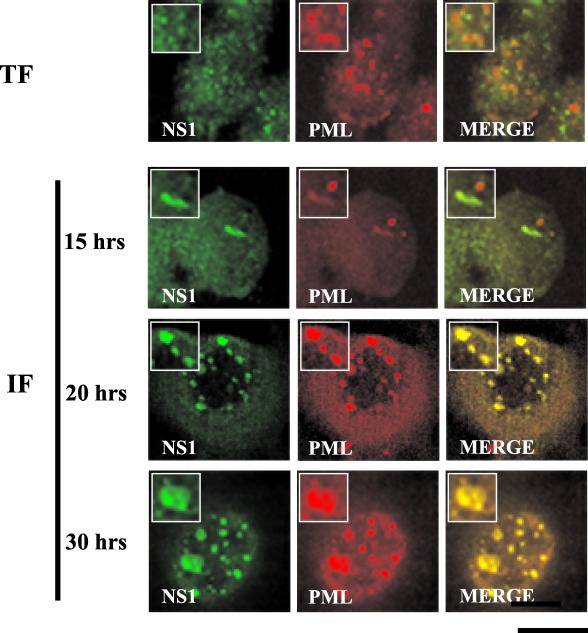
PODs are nuclear structures originally identified in oncovirus infection which are distinct from Cajal bodies and other nuclear bodies such as interchromatin granules and gems. To determine the relationship between PODs and NS1-SMN-positive bodies double-label immunofluorescence experiments were performed on both HA-NS1-transfected and highly synchronized, MVM-infected A92L cells. In transfected A92L cells (30 h posttransfection) and in MVM-infected cells 15 h after release into S phase, PODs and NS1 bodies were identified as completely independent structures (Fig. 5). However, at 20 and 30 h postrelease, PML was present in the enlarged NS1-containing nuclear bodies (Fig. 5). Noninfected synchronized cells exhibited normal POD nuclear localization patterns (data not shown). To distinguish the previously described APAR bodies from the larger, POD-NS1-SMN-inclusive structures found later in infection, the latter have been termed SAABs. As with SMN and p80 coilin, BIA was used to determine whether the appearance of SAABs was accompanied by an increase in PML expression levels. Total protein extracts from A92L cells at 0 and 30 h postrelease were collected and assayed on a sensor chip containing a bound anti-PML monoclonal antibody. There was no difference in PML protein level between the two time points (data not shown), suggesting that our observations were due to relocalization rather than increased expression of PML.
FIG. 5.
PML colocalizes with NS1 in SAABs at 20 h postinfection but not in A92L cells transiently expressing HA-tagged NS1. Double-label experiments were performed on A92L cells transfected with HA-tagged NS1 (TF) and blocked A92L cells 15, 20, and 30 h after infection with MVM (IF). PML (primary antibody: anti-PML monoclonal antibody; secondary antibody: anti-mouse TRITC conjugate) and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: anti-rabbit FITC conjugate) are indicated. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm.
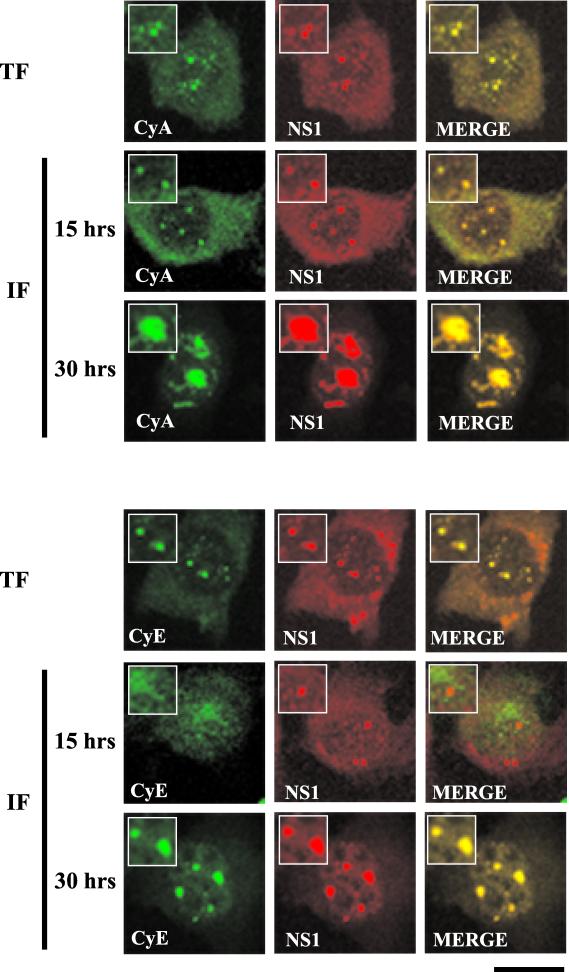
Cyclin A and cyclin E accumulate in SAABs 30 h postinfection. APAR bodies contain cyclin A but not cyclin E (5). These proteins are involved in the G1/S phase transition of the cell cycle (27). A role for cyclins in the viral life cycle has been suggested (4). Similar to previous observations (4), at 15 h postinfection, we found that APAR bodies contained cyclin A but not cyclin E (Fig. 6). Following NS1 transfection, NS1 colocalized with cyclin A and cyclin E in small NS1-positive nuclear bodies. However, at 30 h after release of MVM infection both cyclin A and E accumulated within SAABs (Fig. 6). The presence of SMN and p80 coilin within these SAABs was confirmed in parallel experiments (data not shown). Nuclear redistribution of cyclin A (Fig. 7) and cyclin E (data not shown) was not observed in blocked, noninfected A92L cells at all time points tested.
FIG. 6.
Cyclin A and cyclin E colocalize in SAABs. Double-label experiments were performed on A92L cells transfected with HA-tagged NS1 (TF) and blocked infected A92L cells 15 and 30 h after entry into S phase (IF). (Top) Cyclin A (CyA) colocalizes with NS1 in transfected and infected cells. Cyclin A (primary antibody: goat anti-cyclin A polyclonal; secondary antibody: anti-goat FITC conjugate) and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: anti-rabbit TRITC conjugate) are indicated. (Bottom) Cyclin E (CyE) accumulates in SAABs at around 20 h postinfection. Cyclin E (primary antibody: goat anti-cyclin E polyclonal; secondary antibody: anti-goat FITC conjugate) and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: anti-rabbit TRITC conjugate) are indicated. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm.
FIG. 7.
Cyclin A nuclear distribution is not altered by the blocking process. Double-label experiments were performed on noninfected A92L cells 0 and 30 h after release from the blocking process. Cyclin A (CyA; primary antibody: goat anti-cyclin A polyclonal; secondary antibody: anti-goat FITC conjugate) and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: anti-rabbit TRITC conjugate) are indicated. In comparison, double-label experiments performed on blocked A92L cells 30 h postinfection are shown. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm.
SAABs are likely sites of virus replication.
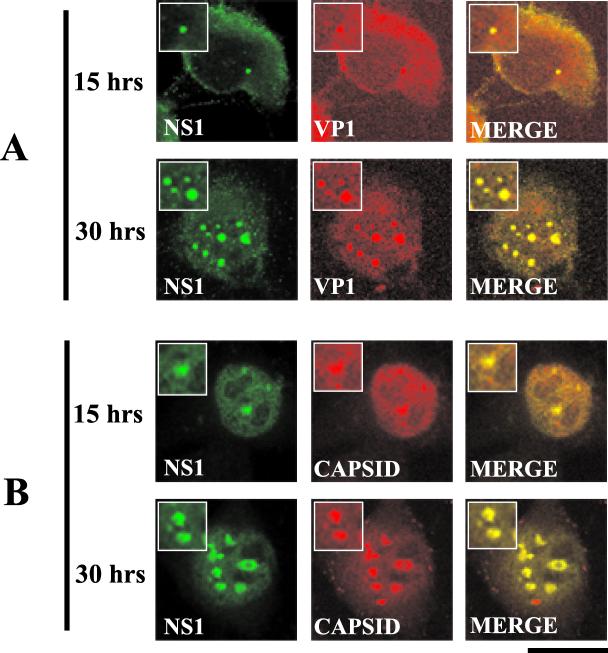
Double-label immunofluorescence experiments were performed on synchronized A92L MVM-infected cells to determine whether the capsid proteins accumulate within SAABs. The smaller APAR bodies identified at 15 h postinfection and the larger SAABs identified at 30 h postinfection contained VP1, VP1-containing intermediates, and fully assembled MVM capsids (Fig. 8). The capsids are likely due to de novo synthesis since capsids are not detected at earlier time points (data not shown). However, there were differences in the overall cellular distribution of assembled capsids compared to those of VP1 and VP1-containing capsid assembly intermediates at 15 and 30 h postrelease. At 15 h postrelease, assembled capsids were diffusely distributed throughout the nucleoplasm, as were the capsid-positive APAR bodies (Fig. 8B). In contrast, preassembled VP1 capsid protein intermediates were localized mainly within the cytoplasm at this time, with lower levels colocalized within nuclear APAR bodies (Fig. 8A). Both assembled and nonassembled capsids were exclusively confined to the larger SAABs at 30 h, with reduced nucleoplasmic and cytoplasmic staining.
FIG. 8.
Viral capsid components (VP1) and the assembled MVM capsid accumulate within SAABs. Double-label experiments were performed on blocked infected A9 cells 15 and 30 h postrelease. (A) NS1 and VP1 colocalize at the 15- and 30-h time points. NS1 (primary antibody: anti-NS1 polyclonal; secondary antibody: anti-rabbit FITC conjugate) and VP1 (primary antibody: anti-VP1 polyclonal peptide antibody; secondary antibody: anti-mouse TRITC conjugate) are indicated. (B) NS1 and the assembled MVM capsid colocalize at the 15- and 30-h time points. NS1 (primary antibody: anti-NS1 polyclonal; secondary antibody: anti-rabbit FITC conjugate) and the capsid (primary antibody: anticapsid monoclonal antibody [30]; secondary antibody: anti-mouse TRITC conjugate) are indicated. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm.
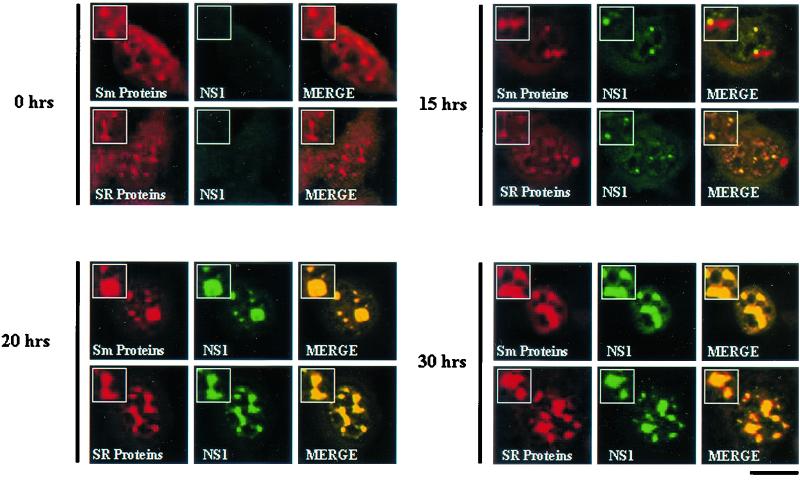
Interchromatin granules or speckled domains are distinct nuclear bodies that contain a variety of RNA splicing factors, including U snRNPs and SR proteins (36, 47). Antibodies against the SR proteins and the Sm core proteins were used in double-label experiments with MVM-infected synchronized A92L cells to determine whether SAABs contain factors essential for RNA splicing. As expected, in synchronized uninfected A92L cells (data not shown) and at 0 h postrelease, the Sm and SR proteins localized within interchromatin granules (Fig. 9). However, following MVM infection, SR and Sm core proteins colocalized with NS1 in the small nuclear bodies at 15 h postrelease and in SAABs at 20 and 30 h postrelease (Fig. 9).
FIG. 9.
RNA splicing factors accumulate within SAABs. Double-label analysis was performed on blocked infected A92L cells 0, 15, 20, and 30 h after entry into S phase. Sm proteins (primary antibody: Y12; secondary antibody: anti-mouse TRITC conjugate), SR proteins (primary antibody: anti-SR monoclonal antibody; secondary antibody: anti-mouse TRITC conjugate), and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: anti-rabbit FITC conjugate) are indicated. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm.
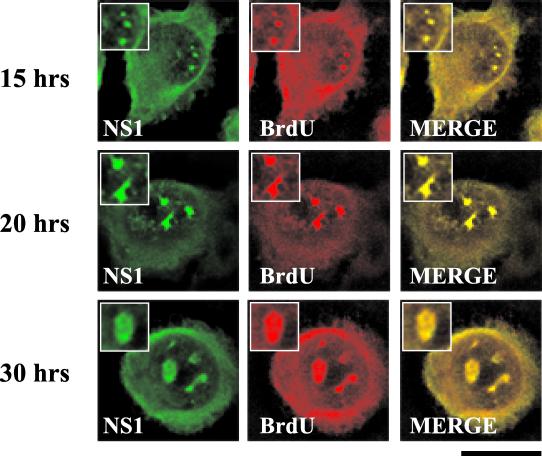
APAR bodies have been shown to be the sites of ongoing viral DNA replication (14). To determine whether SAABs are the sites of viral replication during the later stages of viral infection, similar experiments were performed on infected cells previously labeled with BrdU. In this procedure, the lack of a denaturation step ensures that, while replicating viral single-stranded DNA is detected, cellular chromosomal DNA is not (50). BrdU was identified with an anti-BrdU monoclonal antibody. BrdU colocalized with NS1 in both the smaller nuclear bodies identified at 15 h postinfection (Fig. 10) and the larger SAABs identified at 20 and 30 h postinfection (Fig. 10).
FIG. 10.
SAABs are the sites of viral DNA replication. Double-label experiments were performed on BrdU-treated blocked infected A92L cells 15, 20, and 30 h after entry into S phase. Cells were labeled for 20 min with BrdU at a concentration of 10 μM. BrdU (primary antibody: anti-BrdU monoclonal antibody; secondary antibody: anti-mouse TRITC conjugate) and NS1 (primary antibody: rabbit anti-NS1 polyclonal; secondary antibody: anti-rabbit FITC conjugate) are indicated. Insets show nuclear regions of interest magnified by an additional factor of 2. Bar ≈ 30 μm.
Taken together, these results identify a novel subnuclear structure that is induced upon viral infection. Furthermore, this nuclear reorganization sequesters the components of several previously distinct nuclear bodies, suggesting that SAABs are the active site of viral replication and capsid assembly.
DISCUSSION
The eukaryotic nucleus is a highly organized organelle that is composed of a number of discrete nuclear bodies (34). Interest in the various nuclear bodies has increased recently as it has become clear that these structures are not simply storage sites but rather are highly dynamic structures that can change in size, number, and constituents in response to internal and external stimuli (14, 31, 34, 45). Here we identify the SMN protein as an MVM NS1-interacting protein. This was initially surprising since previous reports suggested that NS1 accumulated in nuclear structures that were independent of SMN-positive nuclear bodies (14). Nuclear SMN is normally a component of Cajal bodies and typically colocalizes with the Cajal body marker protein p80 coilin (9, 28, 51, 52). To address this apparent discrepancy, we investigated the distribution of NS1 after transfection and following the progression of a highly synchronized infection. Transiently expressed NS1 colocalized with SMN in Cajal bodies; however, nuclear bodies such as PODs and speckles remained separate. Consistent with the previous reports of APAR bodies, at time points before 15 h postinfection APAR bodies were SMN-independent structures and were separate from PODs. However, this was in stark contrast to the later time points, at which MVM infection induced dramatic nuclear changes, resulting in the accumulation of Cajal body, POD, speckle, and APAR components within large SAABs. Similar structures were also seen at late time points in infected, unsynchronized cells (data not shown). This dramatic reorganization of the nuclear bodies is likely a specific response to viral infection since the transient expression of NS1 alone (Fig. 2) or NS2 alone or NS1 and NS2 combined (data not shown) did not result in SAAB formation. Additionally, in NS1-transfected cells infected with wild-type MVM, transiently expressed NS1 and virus-derived NS1 are present in SAABs later than 15 h after release h after release (data not shown). Although the detailed kinetics of viral replication is incomplete, significant amounts of DNA replication, capsid production, and encapsidation appear to occur at these late times in infection when the cells have become arrested (49).
APAR bodies are previously described nuclear bodies that appear immediately upon infection and that do not spatially colocalize with other known nuclear bodies such as Cajal bodies, PODs, and speckles (14). Localization in APAR bodies is likely the initial step in an MVM infection. At early time points, a fraction of NS1 was found in APAR bodies but there was also considerable diffuse cytoplasmic staining. As the infection progressed, however, essentially all detectable NS1 was concentrated within SAABs. Since NS1 is essential for several phases of the viral life cycle, including replication and transcriptional regulation of the capsid promoter, these results suggest that SAABs may play a role in the viral life cycle. The status of viral expression and replication past 20 h during synchronous infection, however, has not been well characterized. Alternatively, this massive nuclear aggregation may represent the beginnings of NS1-mediated cytotoxicity or a virus-induced cell death pathway.
The requirement for the infection process in the formation of SAABs suggests that nuclear reorganization occurs in response to a step within the viral life cycle rather than simply in response to the expression of an individual protein. Whether this is a consequence of an early stimulus, such as the entering capsid, viral single-stranded DNA, or the hairpin structures, or whether the stimulus comes at some later time point is not known. There are mutations within the MVM NS2 protein and capsid genes that result in infections by viruses that are deficient in various stages of the viral life cycle, for example, the production of the monomer replicative form or progeny single-stranded template (10, 11, 38). Experiments to determine which of these steps is required for SAAB formation within the viral life cycle are ongoing.
PODs/PML bodies have been associated with transcription, cell growth, and antiviral responses (17, 19, 26). DNA and RNA viruses also frequently target PODs, presumably to facilitate the early stages of transcription and replication (1, 15, 18, 20, 50). PODs can also be targeted for reorganization following viral infection (20). For example, adenovirus protein E4-ORF3 localizes to PODs/PML bodies, thereby causing a physical restructuring of the bodies from spherical to extended fibril-like structures termed nuclear tracks (20). Additionally, many viral infections induce interferon expression, and these infections or treatment with exogenous interferon can increase the size and number of PODs (21, 26). Although parvovirus infection does not induce an interferon response, a dramatic relocalization of PODs is seen late in MVM infection.
The functional significance of the interaction between SMN and NS1 is still unknown. However, as SMN has been implicated in RNA transcription and processing, cellular transportation, and apoptosis (6, 9, 22, 25, 29), NS1 may sequester SMN to facilitate viral RNA processing or to inhibit host cell apoptosis, thus enabling efficient viral turnover. Consistent with the SMN-NS1 complex performing a role during the viral life cycle is the recent finding that novel heterogeneous nuclear ribonucleoprotein-like protein NS1-associated protein 1 (NSAP1) interacts with SMN and NS1 (24, 37). The functional significance of these interactions has not been determined. SMN is not required for the maintenance of nuclear structures such as Cajal bodies (51, 52); therefore it is unlikely that SMN is required for SAAB formation. It is possible, however, that SMN plays a role in targeting or retaining NS1 in SAABs. Alternatively, since SMN is involved in snRNP biogenesis and transcriptional regulation, the NS1-SMN interaction may facilitate an NS1 activity that is required at later time points in infection.
We have identified SAABs, a novel nuclear structure required for parvovirus infection, that contain factors typically associated with other distinct nuclear bodies. The site of this dramatic nuclear reorganization is also a site of viral replication and contains the viral capsids, demonstrating that SAABs may play a role in the viral life cycle. SAABs also contain cyclin A and cyclin E, suggesting that viral infection and viral cycle progression may be dependent on the ability to control specific stages of the cell cycle. Although it has been suggested that cyclin A is sequestered by APAR bodies early in viral infection to arrest the cell in the G1 phase, thus enabling efficient expression of NS1, the exact function of cyclin A and cyclin E in the latter stages of infection within SAABs has yet to be determined. These results highlight the fluidity and dynamic nature of nuclear structures and their contents and help address the functional significance of nuclear compartmentalization. In addition, while this study has no direct link with spinal muscular atrophy, identifying the role SMN plays in facilitating the course of viral infection may shed light on the cellular function of SMN.
Acknowledgments
P.J. Young and K.T. Jensen contributed equally to this work.
We thank Glenn Morris, Angus Lamond, and Colin Parish for antibodies, Kori Wallace for technical assistance, and the W. M. Keck BioImaging Laboratory for confocal imaging.
P.J.Y. was supported by a fellowship from Families of SMA, and K.T.J. was supported by a postdoctoral fellowship from the University of Missouri Life Science Mission Enhancement Program. Funding for these studies was provided by Families of SMA (C.L.L.), the Muscular Dystrophy Association (C.L.L), and the National Institutes of Health (C.L.L., RO1 NS41584-01; D.J.P., RO1 AI21302 and RO1 AI46458).
REFERENCES
- 1.Ahn, J. H., and G. S. Hayward. 2000. Disruption of PML-associated nuclear bodies by IE1 correlates with efficient early stages of viral gene expression and DNA replication in human cytomegalovirus infection. Virology 274:39-55. [DOI] [PubMed] [Google Scholar]
- 2.Ahn, J. K., Z. W. Pitluk, and D. C. Ward. 1992. The GC box and TATA transcription control elements in the P38 promoter of the minute virus of mice are necessary and sufficient for transactivation by the nonstructural protein NS1. J. Virol. 66:3776-3783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Anouja, F., R. Wattiez, S. Mousset, and P. Caillet-Fauquet. 1997. The cytotoxicity of the parvovirus minute virus of mice nonstructural protein NS1 is related to changes in the synthesis and phosphorylation of cell proteins. J. Virol. 71:4671-4678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bashir, T., R. Horlein, J. Rommelaere, and K. Willwand. 2000. Cyclin A activates the DNA polymerase delta-dependent elongation machinery in vitro: a parvovirus DNA replication model. Proc. Natl. Acad. Sci. USA 97:5522-5527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bashir, T., J. Rommelaere, and C. Cziepluch. 2001. In vivo accumulation of cyclin A and cellular replication factors in autonomous parvovirus minute virus of mice-associated replication bodies. J. Virol. 75:4394-4398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bechade, C., P. Rostaing, C. Cisterni, R. Kalisch, V. La Bella, B. Pettmann, and A. Triller. 1999. Subcellular distribution of survival motor neuron (SMN) protein: possible involvement in nucleocytoplasmic and dendritic transport. Eur. J. Neurosci. 11:293-304. [DOI] [PubMed] [Google Scholar]
- 7.Boisvert, F. M., M. J. Hendzel, and D. P. Bazett-Jones. 2000. Promyelocytic leukemia (PML) nuclear bodies are protein structures that do not accumulate RNA. J. Cell Biol. 148:283-292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Breiding, D. E., M. J. Grossel, and E. J. Androphy. 1996. Genetic analysis of the bovine papillomavirus E2 transcriptional activation domain. Virology 221:34-43. [DOI] [PubMed] [Google Scholar]
- 9.Carvalho, T., F. Almeida, A. Calapez, M. Lafarga, M. T. Berciano, and M. Carmo-Fonseca. 1999. The spinal muscular atrophy disease gene product, SMN: a link between snRNP biogenesis and the Cajal (coiled) body. J. Cell Biol. 147:715-728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cater, J. E., and D. J. Pintel. 1992. The small non-structural protein NS2 of the autonomous parvovirus minute virus of mice is required for virus growth in murine cells. J. Gen. Virol. 73:1839-1843. [DOI] [PubMed] [Google Scholar]
- 11.Cotmore, S. F., A. M. D'Abramo, Jr., L. F. Carbonell, J. Bratton, and P. Tattersall. 1997. The NS2 polypeptide of parvovirus MVM is required for capsid assembly in murine cells. Virology 231:267-280. [DOI] [PubMed] [Google Scholar]
- 12.Cotmore, S. F., and P. Tattersall. 1987. The autonomously replicating parvoviruses of vertebrates. Adv. Virus Res. 33:91-174. [DOI] [PubMed] [Google Scholar]
- 13.Cotmore, S. F., and P. Tattersall. 1986. Organization of nonstructural genes of the autonomous parvovirus minute virus of mice. J. Virol. 58:724-732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cziepluch, C., S. Lampel, A. Grewenig, C. Grund, P. Lichter, and J. Rommelaere. 2000. H-1 parvovirus-associated replication bodies: a distinct virus-induced nuclear structure. J. Virol. 74:4807-4815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Day, P. M., R. B. Roden, D. R. Lowy, and J. T. Schiller. 1998. The papillomavirus minor capsid protein, L2, induces localization of the major capsid protein, L1, and the viral transcription/replication protein, E2, to PML oncogenic domains. J. Virol. 72:142-150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Deleu, L., A. Pujol, S. Faisst, and J. Rommelaere. 1999. Activation of promoter P4 of the autonomous parvovirus minute virus of mice at early S phase is required for productive infection. J. Virol. 73:3877-3885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Doucas, V. 2000. The promyelocytic (PML) nuclear compartment and transcription control. Biochem. Pharmacol. 60:1197-1201. [DOI] [PubMed] [Google Scholar]
- 18.Doucas, V., A. M. Ishov, A. Romo, H. Juguilon, M. D. Weitzman, R. M. Evans, and G. G. Maul. 1996. Adenovirus replication is coupled with the dynamic properties of the PML nuclear structure. Genes Dev. 10:196-207. [DOI] [PubMed] [Google Scholar]
- 19.Duprez, E., A. J. Saurin, J. M. Desterro, V. Lallemand-Breitenbach, K. Howe, M. N. Boddy, E. Solomon, H. de The, R. T. Hay, and P. S. Freemont. 1999. SUMO-1 modification of the acute promyelocytic leukaemia protein PML: implications for nuclear localisation. J. Cell Sci. 112:381-393. [DOI] [PubMed] [Google Scholar]
- 20.Everett, R. D., and G. G. Maul. 1994. HSV-1 IE protein Vmw110 causes redistribution of PML. EMBO J. 13:5062-5069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Fabunmi, R. P., W. C. Wigley, P. J. Thomas, and G. N. DeMartino. 2001. Interferon gamma regulates accumulation of the proteasome activator PA28 and immunoproteasomes at nuclear PML bodies. J. Cell Sci. 114:29-36. [DOI] [PubMed] [Google Scholar]
- 22.Fischer, U., Q. Liu, and G. Dreyfuss. 1997. The SMN-SIP1 complex has an essential role in spliceosomal snRNP biogenesis. Cell 90:1023-1029. [DOI] [PubMed] [Google Scholar]
- 23.Hardt, N., C. Dinsart, S. Spadari, G. Pedrali-Noy, and J. Rommelaere. 1983. Interrelation between viral and cellular DNA synthesis in mouse cells infected with the parvovirus minute virus of mice. J. Gen. Virol. 64:1991-1998. [DOI] [PubMed] [Google Scholar]
- 24.Harris, C. E., R. A. Boden, and C. R. Astell. 1999. A novel heterogeneous nuclear ribonucleoprotein-like protein interacts with NS1 of the minute virus of mice. J. Virol. 73:72-80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kerr, D. A., J. P. Nery, R. J. Traystman, B. N. Chau, and J. M. Hardwick. 2000. Survival motor neuron protein modulates neuron-specific apoptosis. Proc. Natl. Acad. Sci. USA 97:13312-13317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lavau, C., A. Marchio, M. Fagioli, J. Jansen, B. Falini, P. Lebon, F. Grosveld, P. P. Pandolfi, P. G. Pelicci, and A. Dejean. 1995. The acute promyelocytic leukaemia-associated PML gene is induced by interferon. Oncogene 11:871-876. [PubMed] [Google Scholar]
- 27.Liu, J., M. D. Hebert, Y. Ye, D. J. Templeton, H. Kung, and A. G. Matera. 2000. Cell cycle-dependent localization of the CDK2-cyclin E complex in Cajal (coiled) bodies. J. Cell Sci. 113:1543-1552. [DOI] [PubMed] [Google Scholar]
- 28.Liu, Q., and G. Dreyfuss. 1996. A novel nuclear structure containing the survival of motor neurons protein. EMBO J. 15:3555-3565. [PMC free article] [PubMed] [Google Scholar]
- 29.Liu, Q., U. Fischer, F. Wang, and G. Dreyfuss. 1997. The spinal muscular atrophy disease gene product, SMN, and its associated protein SIP1 are in a complex with spliceosomal snRNP proteins. Cell 90:1013-1021. [DOI] [PubMed] [Google Scholar]
- 30.Lombardo, E., J. C. Ramirez, M. Agbandje-McKenna, and J. M. Almendral. 2000. A beta-stranded motif drives capsid protein oligomers of the parvovirus minute virus of mice into the nucleus for viral assembly. J. Virol. 74:3804-3814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lorenz, P., B. F. Baker, C. F. Bennett, and D. L. Spector. 1998. Phosphorothioate antisense oligonucleotides induce the formation of nuclear bodies. Mol. Biol. Cell 9:1007-1023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lorson, C., J. Pearson, L. Burger, and D. J. Pintel. 1998. An Sp1-binding site and TATA element are sufficient to support full transactivation by proximally bound NS1 protein of minute virus of mice. Virology 240:326-337. [DOI] [PubMed] [Google Scholar]
- 33.Lorson, C., and D. J. Pintel. 1997. Characterization of the minute virus of mice P38 core promoter elements. J. Virol. 71:6568-6575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Matera, A. G. 1999. Nuclear bodies: multifaceted subdomains of the interchromatin space. Trends Cell Biol. 9:302-309. [DOI] [PubMed] [Google Scholar]
- 35.Matera, A. G. 1998. Of coiled bodies, gems, and salmon. J. Cell. Biochem. 70:181-192. [PubMed] [Google Scholar]
- 36.Mintz, P. J., and D. L. Spector. 2000. Compartmentalization of RNA processing factors within nuclear speckles. J. Struct. Biol. 129:241-251. [DOI] [PubMed] [Google Scholar]
- 37.Mourelatos, Z., L. Abel, J. Yong, N. Kataoka, and G. Dreyfuss. 2001. SMN interacts with a novel family of hnRNP and spliceosomal proteins. EMBO J. 20:5443-5452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Naeger, L. K., N. Salome, and D. J. Pintel. 1993. NS2 is required for efficient translation of viral mRNA in minute virus of mice-infected murine cells. J. Virol. 67:1034-1043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Oleksiewicz, M. B., and S. Alexandersen. 1997. S-phase-dependent cell cycle disturbances caused by Aleutian mink disease parvovirus. J. Virol. 71:1386-1396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Op De Beeck, A., F. Anouja, S. Mousset, J. Rommelaere, and P. Caillet-Fauquet. 1995. The nonstructural proteins of the autonomous parvovirus minute virus of mice interfere with the cell cycle, inducing accumulation in G2. Cell Growth Differ. 6:781-787. [PubMed] [Google Scholar]
- 41.Op De Beeck, A., and P. Caillet-Fauquet. 1997. The NS1 protein of the autonomous parvovirus minute virus of mice blocks cellular DNA replication: a consequence of lesions to the chromatin? J. Virol. 71:5323-5329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Op De Beeck, A., J. Sobczak-Thepot, H. Sirma, F. Bourgain, C. Brechot, and P. Caillet-Fauquet. 2001. NS1- and minute virus of mice-induced cell cycle arrest: involvement of p53 and p21(cip1). J. Virol. 75:11071-11078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Rank, N. M., and P. F. Lambert. 1995. Bovine papillomavirus type 1 E2 transcriptional regulators directly bind two cellular transcription factors, TFIID and TFIIB. J. Virol. 69:6323-6334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Schoborg, R. V., and D. J. Pintel. 1991. Accumulation of MVM gene products is differentially regulated by transcription initiation, RNA processing and protein stability. Virology 181:22-34. [DOI] [PubMed] [Google Scholar]
- 45.Seeler, J. S., and A. Dejean. 1999. The PML nuclear bodies: actors or extras? Curr. Opin. Genet. Dev. 9:362-367. [DOI] [PubMed] [Google Scholar]
- 46.Sleeman, J., C. E. Lyon, M. Platani, J. P. Kreivi, and A. I. Lamond. 1998. Dynamic interactions between splicing snRNPs, coiled bodies and nucleoli revealed using snRNP protein fusions to the green fluorescent protein Exp. Cell Res. 243:290-304. [DOI] [PubMed] [Google Scholar]
- 47.Sleeman, J. E., and A. I. Lamond. 1999. Newly assembled snRNPs associate with coiled bodies before speckles, suggesting a nuclear snRNP maturation pathway. Curr. Biol. 9:1065-1074. [DOI] [PubMed] [Google Scholar]
- 48.Strasswimmer, J., C. L. Lorson, D. E. Breiding, J. J. Chen, T. Le, A. H. Burghes, and E. J. Androphy. 1999. Identification of survival motor neuron as a transcriptional activator-binding protein. Hum. Mol. Genet. 8:1219-1226. [DOI] [PubMed] [Google Scholar]
- 49.Tullis, G., R. V. Schoborg, and D. J. Pintel. 1994. Characterization of the temporal accumulation of minute virus of mice replicative intermediates. J. Gen. Virol. 75:1633-1646. [DOI] [PubMed] [Google Scholar]
- 50.Wu, F. Y., J. H. Ahn, D. J. Alcendor, W. J. Jang, J. Xiao, S. D. Hayward, and G. S. Hayward. 2001. Origin-independent assembly of Kaposi's sarcoma-associated herpesvirus DNA replication compartments in transient cotransfection assays and association with the ORF-K8 protein and cellular PML. J. Virol. 75:1487-1506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Young, P. J., T. T. Le, M. Dunckley, T. M. Nguyen, A. H. Burghes, and G. E. Morris. 2001. Nuclear gems and Cajal (coiled) bodies in fetal tissues: nucleolar distribution of the spinal muscular atrophy protein, SMN. Exp. Cell Res. 265:252-261. [DOI] [PubMed] [Google Scholar]
- 52.Young, P. J., T. T. Le, N. thi Man, A. H. Burghes, and G. E. Morris. 2000. The relationship between SMN, the spinal muscular atrophy protein, and nuclear coiled bodies in differentiated tissues and cultured cells. Exp. Cell Res. 256:365-374. [DOI] [PubMed] [Google Scholar]