Abstract
microRNAs (miRNAs) are widespread among eukaryotes, and studies in several systems have revealed that miRNAs can regulate expression of specific genes. Primary miRNA transcripts are initially processed to ≈70-nucleotide (nt) stem–loop structures (pre-miRNAs), exported to the cytoplasm, further processed to yield ≈22-nt dsRNAs, and finally incorporated into ribonucleoprotein particles, which are thought to be the active species. Here we study nuclear export of pre-miRNAs and show that the process is saturable and thus carrier-mediated. Export is sensitive to depletion of nuclear RanGTP and, according to this criterion, mediated by a RanGTP-dependent exportin. An unbiased affinity chromatography approach with immobilized pre-miRNAs identified exportin 5 as the pre-miRNA-specific export carrier. We have cloned exportin 5 from Xenopus and demonstrate that antibodies raised against the Xenopus receptor specifically block pre-miRNA export from nuclei of Xenopus oocytes. We further show that exportin 5 interacts with double-stranded RNA in a sequence-independent manner.
Keywords: miRNA, tRNA, nuclear transport, exportin, translation regulation, silencing
INTRODUCTION
The nuclear envelope (NE) separates eukaryotic cells into a nuclear and a cytoplasmic compartment and thereby necessitates nucleocytoplasmic transport (Mattaj and Englmeier 1998; Görlich and Kutay 1999; Macara 2001; Weis 2002). Nuclei lack protein synthesis and therefore import their proteins from the cytoplasm. Conversely, ribosomal subunits, tRNAs, mRNAs, and other RNA species are generated in the nucleus and need to be exported to the cytoplasm. Nucleocytoplasmic exchange proceeds through nuclear pore complexes, which penetrate the NE and permit passage of material in two modes: passive diffusion and facilitated translocation. Passive diffusion is fast for small molecules, but becomes increasingly restricted as the size of the transported species approaches a size limit of 20–40 kD. In contrast, facilitated translocation allows passage of very large objects with a molecular mass of up to several million daltons. Facilitated translocation is typically receptor-mediated and coupled to an input of metabolic energy (active transport), allowing accumulation of cargoes even against gradients of chemical activity.
Most nuclear protein transport pathways are mediated by importins or exportins, which are all (at least distantly) related to importin β (Impβ; for reviews, see Mattaj and Englmeier 1998; Wozniak et al. 1998; Görlich and Kutay 1999; Macara 2001; Weis 2002). These receptors possess a RanGTP-binding site, circulate between nucleus and cytoplasm, and transfer cargo molecules from one side of the NE to the other. As an energy source, they use the RanGTP gradient across the NE, which regulates the interaction with their cargoes in a compartment-specific manner. Exportins bind their cargoes at high RanGTP levels in the nucleus (Fornerod et al. 1997; Kutay et al. 1997) and translocate as trimeric RanGTP • exportin • cargo complexes to the cytoplasm, where cargo and Ran are released upon GTP hydrolysis. The exportin can then return to the nuclear compartment and mediate another round of transport. The human genome codes for 20 members of the Impβ family, and seven of them are known to mediate nuclear export (Mattaj and Englmeier 1998; Görlich and Kutay 1999; Macara 2001; Weis 2002).
All characterized RNA transport pathways are mediated by proteinaceous factors. General mRNA export proceeds through a RanGTPase-independent mechanism, whereby NPC passage is facilitated by the yeast Mex67p/Mtr2p complex or its higher eukaryotic counterpart, the NXF (TAP)/p15 complex (Segref et al. 1997; Grüter et al. 1998), and metabolic energy is fed in by RNA-helicases such as Dbp5p (Snay-Hodge et al. 1998; Tseng et al. 1998). Smaller cellular RNAs, such as tRNA and U1 snRNA, are exported in an RanGTP-dependent manner (Izaurralde et al. 1997).
Two exportins function in tRNA export, namely, exportin-t (Arts et al. 1998a; Kutay et al. 1998) and exportin 5 (Bohnsack et al. 2002; Calado et al. 2002). However, there are significant differences between the two. First, both recognize different parts of the tRNA. The binding site on a tRNA for exportin 5 is complementary to that for eEF1A, and this circumstance permits Exp5 not only to export tRNA, but also to deplete aa-tRNA • eEF1A complexes from the nuclear compartment. In contrast, the binding sites on tRNA for eEF1A and exportin-t overlap. Second, the spectrum of tRNA recruited to exportin 5 differs from that preferred by exportin-t. Finally, exportin-t recognizes tRNA only, whereas exportin 5 appears to have a broader specificity and also exports the adenovirus VA1 RNA via a minihelix motif (Gwizdek et al. 2003).
microRNAs (miRNAs) are widespread among eukaryotes and represent key components of a conserved system of RNA-based gene regulation (Cullen 2002; Hutvagner and Zamore 2002b; Carrington and Ambros 2003; Cerutti 2003). The prototypic miRNAs function as negative, posttranscriptional regulators and likely inhibit the translation of target mRNAs. lin-4 and let-7, for example, play important roles in the development of Caenorhabditis elegans and inhibit translation of lin-28 and lin-41, respectively (Lee et al. 1993; Reinhart et al. 2000; Pasquinelli and Ruvkun 2002).
The biogenesis of miRNAs involves several processing steps and proceeds in animal cells according to the following scheme (Lee et al. 2002). pre-miRNAs represent an initial intermediate and form hairpins of 60–80 nucleotides (nt) that are excised by the nuclear RNase III Drosha (Lee et al. 2003) from the primary transcripts (pri-miRNAs). They are then exported to the cytoplasm, processed by a cytoplasmic RNase III, Dicer, to yield double-stranded RNA of ≈21 nt and can finally be detected as single-stranded RNAs in ribonucleoprotein particles (Mourelatos et al. 2002; Zeng and Cullen 2002). miRNAs can direct the degradation of mRNAs that contain perfectly complementary sequences (Hutvagner and Zamore 2002a; Doench et al. 2003; Zeng et al. 2003). Interestingly, mRNAs with imperfect complementarity are not degraded, but instead their translation is inhibited. It is not clear whether the same type of RNP accounts for mRNA cleavage and for degradation-independent translational silencing.
Here, we study nuclear export of pre-miRNAs and show that it is an exportin-mediated and RanGTPase-driven process. Affinity chromatography on immobilized pre-miRNAs identified exportin 5 as the export receptor. We have cloned Xenopus exportin 5 and demonstrate that antibodies directed against this receptor block pre-miRNA export. We further show that exportin 5 interacts with double-stranded RNA specifically, but in a sequence-independent fashion.
RESULTS AND DISCUSSION
microRNA precursors are actively exported from nuclei
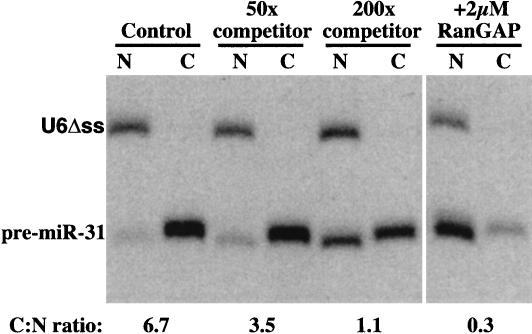
Nuclear export of pre-miRNA represents an essential step in the biogenesis of active miRNA. To study the process, we injected 32P-labeled pre-miR-31 into nuclei of Xenopus oocytes (Fig. 1 ▶). Export from the nuclei was rapid, and after 30 min of incubation, nearly 90% of pre-miR-31 had reached the cytoplasm, while the injection control U6Δss remained nuclear. Interestingly, the exported pre-miR-31 (Fig. 1 ▶) and other pre-miRNAs (data not shown) were fully stable in the cytoplasm of the Xenopus oocytes, suggesting that a Dicer-dependent maturation to miRNAs occurs only later in development.
FIGURE 1.
Microinjected pre-miR-31 is actively exported from nuclei of Xenopus laevis oocytes. Here, 15 nL of a mixture of 10 nM pre-miR-31 and U6Δss injection control were coinjected into nuclei of stage IV–V oocytes. If indicated, RNA mixtures also included 2 μM RanGAP or 500 nM or 2 μM unlabeled pre-miR-31 (competitor). Oocytes were separated into (N) nuclear and (C) cytoplasmic fractions 30 min after injection; RNA was extracted and analyzed on a 10% denaturing PAGE gel. The dried gel was quantified by a phoshorimager, and export was expressed as the cytoplasmic:nuclear (C:N) ratio.
Coinjection of an excess of unlabeled pre-miRNA-31 competed the export of labeled pre-miRNA significantly and in a dose-dependent manner (Fig. 1 ▶), indicating that a saturable export carrier mediates the process. Exportins require RanGTP for cargo binding; depletion of nuclear RanGTP by nuclear injection of RanGAP therefore blocks exportin function. This treatment also inhibited pre-miRNA export (Fig. 1 ▶), which in turn strongly suggests that an exportin of the Impβ family transfers pre-miRNAs to the cytoplasm.
Identification of Exp5 as a potential export receptor for miRNA precursors
We then immobilized four different pre-miRNAs (precursors of human miR-31, miR-23, and miR-20, as well as Drosophila miR-2a-1) to agarose beads and used these affinity matrices to enrich export mediators from a HeLa cell extract, which had been depleted of competing, endogenous RNAs. Our microinjection experiments predicted the interaction between pre-miRNAs and corresponding export mediators to be RanGTP-dependent. We therefore performed the binding also in the presence of the GTPase-deficient RanQ69L mutant (GTP-form), which mimics a nuclear environment. Strikingly, analysis of the bound fractions revealed a single protein band that interacted with the miRNAs in the predicted RanGTP-dependent manner. Peptide mass fingerprinting (data not shown) and immunoblotting (Fig. 2 ▶) identified this putative export mediator as Exp5. We and others previously described Exp5 as an exportin for tRNA and eEF1A (Bohnsack et al. 2002; Calado et al. 2002). Figure 2 ▶ now demonstrates that Exp5 bound immobilized tRNA and pre-miRNA with similar efficiency. As a control, exportin-t bound tRNA only and showed no interaction with the pre-miRNAs.
FIGURE 2.
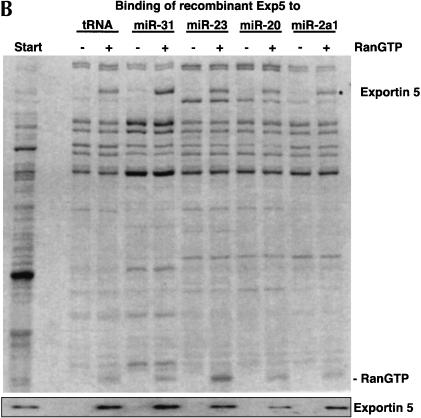
Exportin 5 forms export complexes with pre-miRNAs as well as with tRNA. (A) To identify a potential nuclear export receptor for pre-miRNAs, we first removed competing, endogenous RNAs from a HeLa cell lysate (see Materials and Methods) and bound the resulting extract (Start) to immobilized tRNA, indicated pre-miRNAs, or double-stranded DNA. Where indicated, 2 μM RanQ69L (GTP-form) had been included. Analysis was by SDS-PAGE, followed by Coomassie staining or immunoblotting with indicated antibodies. The load of bound material corresponds to 20 times the input. Note that Exp5 bound to tRNA and to any of the pre-miRNAs specifically and in an RanGTP-dependent manner. In contrast, exportin-t formed export complexes with tRNA only. None of the exportins bound immobilized dsDNA, which served as a control for nonspecific ionic interactions. (B) Escherichia coli lysate from bacteria expressing human Exp5 was depleted of endogenous RNA and subjected to binding to either immobilized tRNA or various pre-miRNAs in the absence or presence of RanQ69L GTP (2 μM). Binding assays were performed and analyzed as described in A.
The interaction between Exp5 and the pre-miRNAs in Figure 2A ▶ could have been direct or bridged by an additional export adaptor present in the HeLa extract. To distinguish between these possibilities, we expressed Exp5 in Escherichia coli (which lack nuclei and hence nuclear export components) and repeated the binding experiment with the corresponding Exp5-containing bacterial lysate (Fig. 2B ▶). Also under these conditions, specific and RanGTP-dependent binding of Exp5 to immobilized pre-miRNAs was observed. We can therefore conclude that Exp5 interacts with pre-miRNAs directly. This conclusion is further confirmed by the fact that a stable pre-miRNA • Exp5 • RanGTP complex efficiently forms also from purified components (data not shown).
tRNA and pre-miRNAs compete with each other for exportin 5 binding
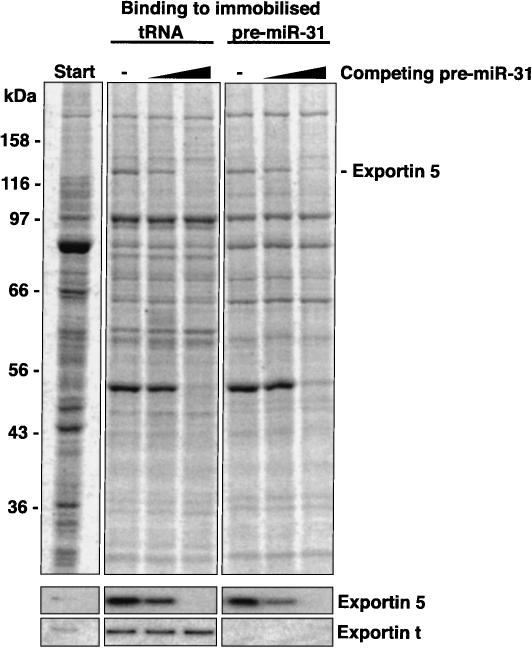
A key criterion for a specific binding to an affinity matrix is competition of the interaction with the free ligand. Indeed, addition of increasing amounts of pre-miRNA to the RNA-free HeLa extract selectively inhibited the recruitment of Exp5 to either immobilized pre-miRNA or immobilized tRNA, but left background bands and the interaction of exportin-t with tRNA largely unaffected (Fig. 3 ▶). We can thus conclude that Exp5 binds pre-miRNAs not only in immobilized form but also free in solution. The experiment also indicates that a given Exp5 molecule cannot simultaneously bind a tRNA and a pre-miRNA molecule.
FIGURE 3.
tRNA and pre-miRNA appear to bind to overlapping sites on Exp5. Binding of Exp5 from HeLa extract to immobilized tRNA and pre-miRNA was performed in the presence of RanQ69L (GTP) as described in Figure 2A ▶. Where indicated, 0.4 μM or 4 μM nonimmobilized pre-miR-31 had been added. The soluble pre-miRNA competed not only binding of Exp5 to immobilized pre-miRNA, but also to tRNA, which points to overlapping binding sites for tRNA and pre-miRNA on Exp5. The interaction between Exp-t and tRNA was not competed by pre-miRNA.
Exportin 5 binds double-stranded RNA in a sequence-independent manner
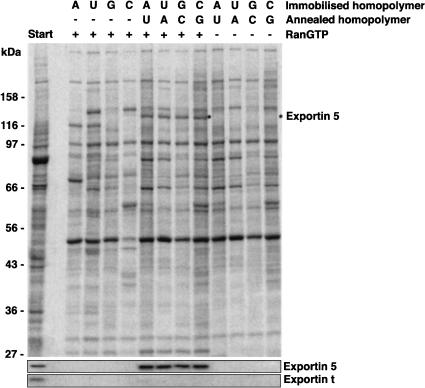
Gwizdek et al. (2003) reported that Exp5 recognizes a double-stranded minihelix motif in adenovirus VA1 RNA and mediates export of this RNA. pre-miRNAs are also predicted to form stem–loop structures, which can contain mismatches. There is no obvious sequence homology among any of the four pre-miRNAs tested in this study and the VA1 RNA, raising the possibility that Exp5 binds double-stranded RNA in a sequence-independent manner. We decided to subject this assumption to a rigorous test and immobilized RNA homopolymers (A30, C30, G30, and U30) to agarose. These homopolymers are not only single stranded, but also highly unlikely to form any secondary structures. As seen from Figure 4 ▶, Exp5 failed to bind these single-stranded RNAs even in the presence of RanQ69L (GTP). When, however, the complementary strands were annealed to form poly(A):poly(U) or poly(C):poly(G) double strands, Exp5 bound efficiently and in an RanGTP-dependent manner. As Exp5 binds the homopolymer-based dsRNAs and the four unrelated pre-miRNAs with comparable efficiency, the data demonstrate that Exp5 interacts with double-stranded RNA in an entirely sequence-independent fashion.
FIGURE 4.
Exportin 5 binds double-stranded RNA in a sequence-independent manner. Immobilized RNA homopolymers (30 nt) were used as ssRNA or annealed with the complementary homopolymer (dsRNA) to retrieve interacting proteins from cytosolic HeLa extract depleted of endogenous RNA. HeLa extract was fractionated and binding assays were performed as described for Figure 2 ▶. Formation of export complexes was analyzed by SDS-PAGE, followed by Coomassie staining. Western blotting confirmed that Exp5 (but not Exp-t) could be retrieved with dsRNA independent of nucleotide composition and sequence, provided RanQ69L GTP had been present.
Numerous factors from eukaryotic, prokaryotic, and viral origin have been identified that recognize dsRNA independently of sequence, examples being Staufen, the dsRNA-dependent protein kinase PKR, and Dicer (Saunders and Barber 2003). Most of them share a common, conserved structural motif, the dsRNA-binding domain (DRBD). No such motif is evident in Exp5, and therefore it will be very interesting to learn how the dsRNA-binding site of Exp5 is structured.
The most straightforward explanation for the competition experiment (Fig. 3 ▶) is that tRNA and dsRNA bind to a similar or even identical site on exportin 5. This could indicate that exportin 5 also recognizes tRNA via a dsRNA motif. The mapping of the VA1 RNA had indicated that a double-stranded region of ≈17 nt is needed for Exp5 binding (Gwizdek et al. 2001), which is longer than any stem–loop in a typical tRNA. We would therefore assume that exportin 5 contacts at least two of the stems of a tRNA molecule.
Exportin 5 is essential for pre-miRNA export from nuclei of Xenopus oocytes
The abovementioned binding and microinjection experiments indicate that exportin 5 mediates pre-miRNA export and that Xenopus oocytes contain an Exp5-like activity. To test these assumptions, we retrieved ESTs of the putative Xenopus tropicalis ortholog from the database of the Sanger center and identified a clone that contained the 5′-end of the cDNA. Sequencing of the entire cDNA insert revealed that this clone comprised the entire coding region. X. tropicalis exportin 5 has a predicted mass of 135 kD and is at the amino acid level 57% identical to the human factor. Binding assays from egg extract confirmed that Xenopus exportin 5 recognizes tRNA and dsRNA with virtually the same specificity as the human one (data not shown), indicating that the two are not only related in sequence, but also functionally equivalent.
We then expressed a C-terminal fragment of Xenopus laevis exportin 5 in bacteria and used the protein to raise antibodies, which were then affinity-purified on the immobilized antigen. Strikingly, injection of this antibody into Xenopus oocytes blocked the export of pre-miRNA-31 completely (Fig. 5A ▶). The effect was highly specific as judged by two sets of controls: First, anti-Drosophila-exportin-5 antibodies (“control”), which do not cross-react with the Xenopus protein, had no effect (Fig. 5A ▶). Second, the anti-Xenopus-exportin-5 antibodies did not block export of tRNAPhe (Fig. 5B ▶), which largely relies on exportin-t for its own export (Arts et al. 1998b), and also had no effect on the export of U1 snRNA (Fig. 5B ▶), which is mediated by the CRM1 • PHAX • CBC complex (Ohno et al. 2000). We can thus conclude that pre-miRNA export proceeds along the exportin 5-dependent pathway and that no other transport receptor can substitute for exportin 5 in this process.
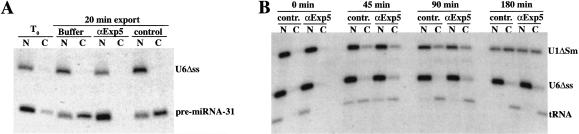
FIGURE 5.
Antibodies against exportin 5 block export of pre-miR-31 from nuclei of Xenopus oocytes. (A) Here, 15 nL of a mixture of radiolabeled 10 nM pre-miR-31 and U6Δss nuclear injection control were coinjected into nuclei of stage IV–V oocytes. Both RNA mixtures contained 1 μM unlabeled pre-miR-31, 1 Unit/μL Superasin (Ambion), and either 2 mg/mL anti-Xenopus-exportin-5 antibody (αExp5) or the same concentration of a control antibody (anti-Drosophila-exportin-5, which does not cross-react with the Xenopus protein). Oocytes were separated into (N) nuclear and (C) cytoplasmic fractions 30 min after injection; RNA was extracted and analyzed on a 10% denaturing PAGE gel. (B) In this panel, 15 nL of a mixture of radiolabeled 10 nM U6Δss injection control, tRNAPhe, and 7 nM U1ΔSm were coinjected into nuclei of stage IV–V oocytes. Injection mixtures contained 1 μM unlabeled tRNAPhe, 1 Unit/μL Superasin (Ambion), and either 2 mg/mL affinity-purified anti-Xenopus-exportin-5 or control antibodies as indicated. Oocytes were separated into nuclear and cytoplasmic fractions either immediately (0 min), 45, 90, or 180 min after injection and processed as in A. Each timepoint in both panels represents the average of five oocytes.
Exportin 5 also exists in plants (Bollman et al. 2003) and Drosophila (Bohnsack et al. 2002). Our preliminary data suggest that at least the insect protein also recognizes dsRNA in an RanGTP-dependent manner. We would therefore assume that it also functions in pre-miRNA export and would predict that it is essential for miRNA-dependent regulation of gene expression. In contrast, C. elegans lacks a recognizable exportin 5 ortholog, and it will therefore be very interesting to learn how nematodes export their microRNA precursors to the cytoplasm.
MATERIALS AND METHODS
Cloning of microRNA precursors
Constructs for in vitro transcription of human pre-miR-20 and Drosophila pre-miR-2a-1 have been the kind gift of E. Izaurralde (EMBL, Heidelberg, Germany). The precursor of human miR-23 (Mourelatos et al. 2002) was cloned as an NotI/ClaI construct into the pSilencer vector (Ambion) for in vitro transcription, and the construct for in vitro transcription of pre-miR-31 (Lagos-Quintana et al. 2001) was inserted into a pQE80 derivative (QIAGEN). microRNAs were named according to Ambros et al. (2003).
Immobilization of RNA
tRNA from bakers yeast (Roche) and in vitro translated microRNA precursors were purified by anion exchange chromatography on a Mono Q column (Amersham Biosciences). RNA homopolymers, purified tRNA, or pre-miRNA were oxidized at their 3′-ribose with sodium periodate; residual periodate was quenched; and the RNA was precipitated and finally immobilized at ~1 mg/mL to polyacryl hydrazido-agarose (Sigma). The covalent bond was stabilized by reduction with NaBH3CN. The matrix with immobilized dsDNA was purchased from Sigma (#D8515).
Binding assays
A cytosolic HeLa extract was prepared as described (Kutay et al. 1998). The extract was fractionated by anion exchange chromatography on a Q-Sepharose FF column to deplete it of endogenous RNA, and fractions of the eluate containing protein but no RNA were pooled. Buffer was exchanged to 50 mM Tris-HCl (pH 7.5), 50 mM NaCl, and 5 mM Mg(OAc)2. Aliquots of the protein eluate equivalent to 200 μL of HeLa extract or E. coli lysate containing Exp5 depleted of endogenous RNA (as described for the HeLa extract) were subjected to binding to 20 μL of immobilized RNA, DNA, or empty matrix in the absence or presence of RanQ69L (GTPase-deficient Ran mutant). After binding, beads were extensively washed, and immobilized proteins were eluted with SDS-containing buffer. Binding was analyzed by SDS-PAGE followed by either Coomassie staining or Western blot analysis with anti-exportin antibodies.
Cloning of Xenopus exportin 5
The EST TEgg076k22 (Sanger Institute) containing the ORF of X. tropicalis exportin 5 was obtained from MRC Geneservice, and the cDNA insert was sequenced on both strands. Exp5 was cloned at BspHI–HindIII into a derivative of pQE80 (QIAGEN). The C terminus of X. laevis Exp5 was cloned from cDNA, expressed in E. coli, and used for the production of antibodies.
Antibodies
Antibodies against human and Drosophila exportin 5 (Bohnsack et al. 2002) as well as against human exportin-t (Kutay et al. 1998) have been described previously. Antibodies against Xenopus exportin 5 were raised against a fragment comprising the 368 C-terminal amino acids. All antibodies were used after affinity purification on the appropriate antigen columns.
Oocyte injection and analysis
Collagenase-treated X. laevis oocytes were prepared and injected according to standard procedures as previously described (Jarmolowski et al. 1994). RNAs were double labeled with [α-32P]GTP and [α-32P]UTP by in vitro transcription from U6Δss, U1Δsm (Jarmolowski et al. 1994), tRNAPhe (Arts et al. 1998b), and the pre-miR-31 construct (this study).
Accession number
The nucleotide sequence of X. tropicalis exportin 5 will be listed under the accession number AJ585214.
Acknowledgments
We thank E. Izaurralde for clones of the precursors of miRNAs miR-20 (human) and miR-2a-1 (Drosophila) as well as the DFG (SFB 352 and Graduiertenkolleg Molekulare Zellbiologie) and the Alfried Krupp-Foundation for financial support. K.C. is grateful to Iain Mattaj for support during the execution of this work.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Article and publication are at http://www.rnajournal.org/cgi/doi/10.1261/rna.5167604.
REFERENCES
- Ambros, V., Bartel, B., Bartel, D.P., Burge, C.B., Carrington, J.C., Chen, X., Dreyfuss, G., Eddy, S.R., Griffiths-Jones, S., Marshall, M., et al. 2003. A uniform system for microRNA annotation. RNA 9: 277–279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arts, G.J., Fornerod, M., and Mattaj, I.W. 1998a. Identification of a nuclear export receptor for tRNA. Curr. Biol. 8: 305–314. [DOI] [PubMed] [Google Scholar]
- Arts, G.J., Kuersten, S., Romby, P., Ehresmann, B., and Mattaj, I.W. 1998b. The role of exportin-t in selective nuclear export of mature tRNAs. EMBO J. 17: 7430–7441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bohnsack, M.T., Regener, K., Schwappach, B., Saffrich, R., Paraskeva, E., Hartmann, E., and Görlich, D. 2002. Exp5 exports eEF1A via tRNA from nuclei and synergizes with other transport pathways to confine translation to the cytoplasm. EMBO J. 21: 6205–6215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bollman, K.M., Aukerman, M.J., Park, M.Y., Hunter, C., Berardini, T.Z., and Poethig, R.S. 2003. HASTY, the Arabidopsis ortholog of exportin 5/MSN5, regulates phase change and morphogenesis. Development 130: 1493–1504. [DOI] [PubMed] [Google Scholar]
- Calado, A., Treichel, N., Muller, E.C., Otto, A., and Kutay, U. 2002. Exportin-5-mediated nuclear export of eukaryotic elongation factor 1A and tRNA. EMBO J. 21: 6216–6224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carrington, J.C. and Ambros, V. 2003. Role of microRNAs in plant and animal development. Science 301: 336–338. [DOI] [PubMed] [Google Scholar]
- Cerutti, H. 2003. RNA interference: Traveling in the cell and gaining functions? Trends Genet. 19: 39–46. [DOI] [PubMed] [Google Scholar]
- Cullen, B.R. 2002. RNA interference: Antiviral defense and genetic tool. Nat. Immunol. 3: 597–599. [DOI] [PubMed] [Google Scholar]
- Doench, J.G., Petersen, C.P., and Sharp, P.A. 2003. siRNAs can function as miRNAs. Genes & Dev. 17: 438–442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fornerod, M., Ohno, M., Yoshida, M., and Mattaj, I.W. 1997. Crm1 is an export receptor for leucine rich nuclear export signals. Cell 90: 1051–1060. [DOI] [PubMed] [Google Scholar]
- Görlich, D. and Kutay, U. 1999. Transport between the cell nucleus and the cytoplasm. Annu. Rev. Cell Dev. Biol. 15: 607–660. [DOI] [PubMed] [Google Scholar]
- Grüter, P., Tabernero, C., von Kobbe, C., Schmitt, C., Saavedra, C., Bachi, A., Wilm, M., Felber, B.K., and Izaurralde, E. 1998. TAP, the human homolog of Mex67p, mediates CTE-dependent RNA export from the nucleus. Mol. Cell 1: 649–659. [DOI] [PubMed] [Google Scholar]
- Gwizdek, C., Bertrand, E., Dargemont, C., Lefebvre, J.C., Blanchard, J.M., Singer, R.H., and Doglio, A. 2001. Terminal minihelix, a novel RNA motif that directs polymerase III transcripts to the cell cytoplasm. Terminal minihelix and RNA export. J. Biol. Chem. 276: 25910–25918. [DOI] [PubMed] [Google Scholar]
- Gwizdek, C., Ossareh-Nazari, B., Brownawell, A.M., Doglio, A., Bertrand, E., Macara, I.G., and Dargemont, C. 2003. Exportin-5 mediates nuclear export of minihelix-containing RNAs. J. Biol. Chem. 278: 5505–5508. [DOI] [PubMed] [Google Scholar]
- Hutvagner, G. and Zamore, P.D. 2002a. A microRNA in a multiple-turnover RNAi enzyme complex. Science 297: 2056–2060. [DOI] [PubMed] [Google Scholar]
- ———. 2002b. RNAi: Nature abhors a double-strand. Curr. Opin. Genet. Dev. 12: 225–232. [DOI] [PubMed] [Google Scholar]
- Izaurralde, E., Kutay, U., von Kobbe, C., Mattaj, I.W., and Görlich, D. 1997. The asymmetric distribution of the constituents of the Ran system is essential for transport into and out of the nucleus. EMBO J. 16: 6535–6547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarmolowski, A., Boelens, W.C., Izaurralde, E., and Mattaj, I.W. 1994. Nuclear export of different classes of RNA is mediated by specific factors. J. Cell Biol. 124: 627–635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawasaki, H. and Taira, K. 2003. Hes1 is a target of microRNA-23 during retinoic-acid-induced neuronal differentiation of NT2 cells. Nature 423: 838–842. [DOI] [PubMed] [Google Scholar]
- Kutay, U., Bischoff, F.R., Kostka, S., Kraft, R., and Görlich, D. 1997. Export of importin α from the nucleus is mediated by a specific nuclear transport factor. Cell 90: 1061–1071. [DOI] [PubMed] [Google Scholar]
- Kutay, U., Lipowsky, G., Izaurralde, E., Bischoff, F.R., Schwarzmaier, P., Hartmann, E., and Görlich, D. 1998. Identification of a tRNA-specific nuclear export receptor. Mol. Cell 1: 359–369. [DOI] [PubMed] [Google Scholar]
- Lagos-Quintana, M., Rauhut, R., Lendeckel, W., and Tuschl, T. 2001. Identification of novel genes coding for small expressed RNAs. Science 294: 853–858. [DOI] [PubMed] [Google Scholar]
- Lee, R.C., Feinbaum, R.L., and Ambros, V. 1993. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75: 843–854. [DOI] [PubMed] [Google Scholar]
- Lee, Y., Jeon, K., Lee, J.T., Kim, S., and Kim, V.N. 2002. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21: 4663–4670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee, Y., Ahn, C., Han, J., Choi, H., Kim, J., Yim, J., Lee, J., Provost, P., Radmark, O., Kim, S., et al. 2003. The nuclear RNase III Drosha initiates microRNA processing. Nature 425: 415–419. [DOI] [PubMed] [Google Scholar]
- Macara, I.G. 2001. Transport into and out of the nucleus. Microbiol. Mol. Biol. Rev. 65: 570–594, table of contents. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mattaj, I.W. and Englmeier, L. 1998. Nucleocytoplasmic transport: the soluble phase. Annu. Rev. Biochem. 67: 265–306. [DOI] [PubMed] [Google Scholar]
- Mourelatos, Z., Dostie, J., Paushkin, S., Sharma, A., Charroux, B., Abel, L., Rappsilber, J., Mann, M., and Dreyfuss, G. 2002. miRNPs: A novel class of ribonucleoproteins containing numerous microRNAs. Genes & Dev. 16: 720–728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohno, M., Segref, A., Bachi, A., Wilm, M., and Mattaj, I.W. 2000. PHAX, a mediator of U snRNA nuclear export whose activity is regulated by phosphorylation. Cell 101: 187–198. [DOI] [PubMed] [Google Scholar]
- Pasquinelli, A.E. and Ruvkun, G. 2002. Control of developmental timing by microRNAs and their targets. Annu. Rev. Cell Dev. Biol. 18: 495–513. [DOI] [PubMed] [Google Scholar]
- Reinhart, B.J., Slack, F.J., Basson, M., Pasquinelli, A.E., Bettinger, J.C., Rougvie, A.E., Horvitz, H.R., and Ruvkun, G. 2000. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403: 901–906. [DOI] [PubMed] [Google Scholar]
- Saunders, L.R. and Barber, G.N. 2003. The dsRNA binding protein family: Critical roles, diverse cellular functions. FASEB J. 17: 961–983. [DOI] [PubMed] [Google Scholar]
- Segref, A., Sharma, K., Doye, V., Hellwig, A., Huber, J., Lührmann, R., and Hurt, E. 1997. Mex67p, a novel factor for nuclear mRNA export, binds to both poly(A)+ RNA and nuclear pores. EMBO J. 16: 3256–3271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Snay-Hodge, C.A., Colot, H.V., Goldstein, A.L., and Cole, C.N. 1998. Dbp5p/Rat8p is a yeast nuclear pore-associated DEAD-box protein essential for RNA export. EMBO J. 17: 2663–2676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tseng, S.S., Weaver, P.L., Liu, Y., Hitomi, M., Tartakoff, A.M., and Chang, T.H. 1998. Dbp5p, a cytosolic RNA helicase, is required for poly(A)+ RNA export. EMBO J. 17: 2651–2662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weis, K. 2002. Nucleocytoplasmic transport: Cargo trafficking across the border. Curr. Opin. Cell Biol. 14: 328–335. [DOI] [PubMed] [Google Scholar]
- Wozniak, R.W., Rout, M.P., and Aitchison, J.D. 1998. Karyopherins and kissing cousins. Trends Cell Biol. 8: 184–188. [DOI] [PubMed] [Google Scholar]
- Zeng, Y. and Cullen, B.R. 2002. RNA interference in human cells is restricted to the cytoplasm. RNA 8: 855–860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeng, Y., Yi, R., and Cullen, B.R. 2003. MicroRNAs and small interfering RNAs can inhibit mRNA expression by similar mechanisms. Proc. Natl. Acad. Sci. 5: 5. [DOI] [PMC free article] [PubMed] [Google Scholar]