Abstract
Site-specific recombination catalyzed by bacteriophage λ integrase (Int) is essential for establishment and termination of the viral lysogenic life cycle. Int is the archetype of the tyrosine recombinase family whose members are responsible for DNA rearrangement in prokaryotes, eukaryotes and viruses. The mechanism regulating catalytic activity during recombination is incompletely understood. Studies of tyrosine recombinases bound to their target substrates suggest that the C-termini of the proteins are involved in protein–protein contacts that control the timing of DNA cleavage events during recombination. We investigated an Int truncation mutant (W350) that possesses enhanced topoisomerase activity but greater than 100-fold reduced recombination activity. Alanine scanning mutagenesis of the C-terminus indicates that two mutants, W350A and I353A, cannot perform site-specific recombination although their DNA binding, cleavage and ligation activities are at wild-type levels. Two other mutants, R346A and R348A, are deficient solely in the ability to cleave DNA. To explain these results, we have constructed a homology-threaded model of the Int structure using a Cre crystal structure. We propose that residues R346 and R348 are involved in orientation of the catalytic tyrosine that cleaves DNA, whereas W350 and I353 control and make intermolecular contacts with other Int proteins in the higher order recombination structures known as intasomes. These results suggest that Int and the other tyrosine recombinases have evolved regulatory contacts that coordinate site-specific recombination at the C-terminus.
INTRODUCTION
Site-specific recombination is a precise genetic exchange of DNA segments between sites of limited homology. Much of our knowledge about the mechanism of site-specific recombination in general derives from studies of the integration and excision reactions mediated by bacteriophage λ integrase (Int) (1). Int is a member of the tyrosine recombinase family, which contains diverse members including invertases, resolvases and transposases in addition to integrases (2,3).
Int catalyzes DNA cleavage and ligation reactions during site-specific recombination (Fig. 1). Integration of the bacteriophage genome into the host chromosome requires Int and the host-encoded integration host factor (IHF) protein. Int and IHF recombine regions of DNA called attachment (att) sites. The sites on the phage and host bacteria used for integration are called attP and attB, respectively. After integration, the resulting hybrid attachment sites on either side of the prophage are called attL and attR. These sites are substrates for site-specific excision that precisely restores the attP and attB sites. Excision of the prophage from the host chromosome requires Int, IHF and the λ-encoded excisionase (Xis). Both integration and excision are stimulated by the host-encoded factor for inversion stimulation (FIS) protein (4,5).
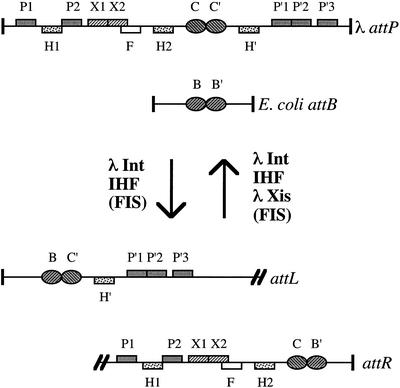
Figure 1.
Substrates and products of bacteriophage λ site-specific recombination. Relative positions of relevant DNA-binding sites are indicated. The attL and attR sites are separated by the λ chromosome (represented by double slash marks). Int arm binding sites, P1, P2, P′1, P′2 and P′3; Int core binding sites, B, B′, C and C′; IHF binding sites, H1, H2 and H′; Xis binding sites, X1 and X2; FIS binding site, F.
The tyrosine recombinases Cre and Flp have been trapped with DNA substrates as recombination intermediate crystal structures (6–9). These crystal structures predict that four recombinase monomers are required for recombination. Each monomer binds to core sites and cleaves and ligates one of the four DNA strands involved in the reaction. A similar structure is thought to be present during Int site-specific recombination (Fig. 2).
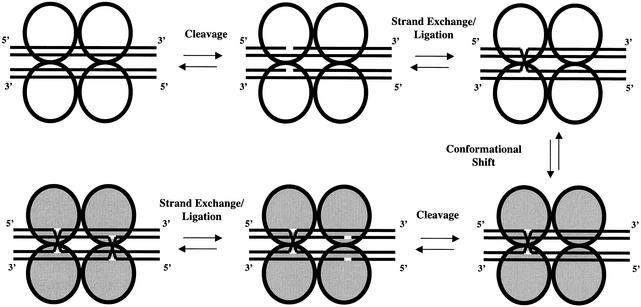
Figure 2.
A model of site-specific recombination. Only the integrase molecules (represented by circles) and their target DNA substrates (represented by lines) are shown. Two recombination substrates are recognized and bound by four integrases. Once the first series of cleavage, strand exchange and ligation events has occurred, changes in the DNA substrate alter the intasome conformation (indicated by gray circles). The site-specific recombination reaction then proceeds to completion.
During integration, the binding of IHF to the three sites on attP DNA facilitates the binding of Int molecules to both arm and core sites, thereby forming the attP ‘intasome’ complex. This complex synapses with naked attB core DNA (10) to assemble the higher order intasome complex that undergoes recombination. Once bound in the intasome complex, recombination proceeds stepwise: the first pair of DNA strands in attP and attB are cleaved by two Int molecules to form covalent 3′ phosphotyrosyl bonds with their respective Int Y342 residues. The free 5′ hydroxyl ends generated by the cleavage event migrate away from their original strand and attack the phosphotyrosyl bond on the other strand, ligating the two strands to form a Holliday junction. Formation of the Holliday junction likely allows a conformational shift in the intasome that prevents reversal of the reaction and permits cleavage, exchange and ligation of the second pair of DNA strands by two other Int molecules. Excisive recombination consists of a similar series of Int-mediated strand exchanges that take the integration products and regenerate the attP and attB sites.
Biochemical experiments define three functional domains in Int (11,12). The N-terminal domain, comprising residues 1–64, specifically binds the arm-type sites in the P and P′ arm regions. The central domain, comprising residues 65–169, specifically binds core-type sites. The C-terminal domain, comprising residues 170–356, possesses type I topoisomerase activity and performs the DNA cleavage, strand exchange and ligation steps in the recombination reaction. The carboxyl region also contains the conserved Arg-His-Arg-Tyr (RHRY) motif that defines the tyrosine recombinase family (13). The RHRY residues are essential for DNA cleavage, formation of the 3′ phosphorotyrosyl bonds with DNA and ligation of the cleaved DNA ends to their new partner strand (14,15).
The catalytic domains of the tyrosine recombinase family are similar to the type IB topoisomerases in both tertiary structure and catalytic mechanism. Both cleave DNA and create transient 3′ phosphotyrosyl bonds at their respective DNA target sites (16). The tyrosine recombinases are distinguished from the type IB topoisomerases in their ability to control precisely the timing of the cleavage, strand exchange and ligation events. Thus, four recombinases can cooperatively cleave, exchange and ligate four DNA strands to new DNA partner strands. How catalytic activity is regulated during Int-mediated recombination remains unclear.
Han et al. (14) isolated an Int mutant that lowered the efficiency of site-specific excision and increased topoisomerase I activity. This mutant, Int W350, is a nonsense mutant that deletes the last seven residues of Int. Han et al. (14) proposed that the deleted residues regulate the recombination reaction. In this report, we systematically changed residues 346–356 in the carboxyl region of Int by alanine-scanning mutagenesis. We assayed the topoisomerase, recombination, DNA binding, cleavage and ligation activities of the mutants. Our results allow us to identify those amino acids that are involved in and essential for the regulation of cleavage activity during the integrative and excisive recombination reactions.
MATERIALS AND METHODS
Media, chemicals and enzymes
Luria–Bertani (LB) and MacConkey growth media were supplied by Difco. Galactose, isopropyl β-d-thiogalactopyranoside (IPTG), ampicillin, kanamycin and spectinomycin were supplied by Sigma. Smith-Beecham Laboratories supplied Timetin. Gibco BRL or Fisher supplied Taq polymerase used in PCRs. New England Biolabs supplied all restriction and modification enzymes except T4 DNA ligase, which was supplied by Promega. Oligodeoxyribonucleotides (ODNs) and acrylamide gels were supplied by Genosys Laboratories and Invitrogen, respectively.
Bacterial strains and plasmids
Table 1 lists the bacterial strains, plasmids, bacteriophage and ODNs used in this study.
Table 1. Bacterial strains, plasmids, phage and oligonucleotides.
| Strain or plasmid | Relevant characteristics/sequence | Source or reference |
|---|---|---|
| Bacteria | ||
| Escherichia coli | ||
| DH1 | F– thi-1 supE44 hsdR17 endA1 recA1 gyr-96 λ– | |
| JG4117 | sup° galT::λ (int-G)Δ pYWH101 | (14) |
| BMS209 | araD139 Δ(ara-leu)7696 galE15 galK16 Δ(lac)X74 rpsL (StrR) hsdR2 (r–m+) mcrA mcrB1 zbh620::Tn10 Δλ::galETK from RW1817 | B. Swalla; (17) |
| Salmonella typhimurium | ||
| MS1868 | leuA414(am) Fels– (r–m+) endE40 hsdSB | M. Susskind |
| Plasmids | ||
| pBR322 | ||
| pCKR101 | pBR322 Plac–lacIq Ptac polylinker | (35) |
| pYWH7 | pCKR101 Plac-λint | (14) |
| pYWH101 | pSC101 Plac-xis | (14) |
| pXM122 | pCKR101 Plac-λint (W350) from pYWH7 | (14) |
| pCINT | pCKR101 Plac-λint | This study |
| pCINT Y342A | pCKR101 Plac-λint (Y342A) | This study |
| pCINT R346A | pCKR101 Plac-λint (R346A) | This study |
| pCINT R348A | pCKR101 Plac-λint (R348A) | This study |
| pCINT W350A | pCKR101 Plac-λint (W350A) | This study |
| pCINT D351A | pCKR101 Plac-λint (D351A) | This study |
| pCINT K352A | pCKR101 Plac-λint (K352A) | This study |
| pCINT I353A | pCKR101 Plac-λint (I353A) | This study |
| pCINT E354A | pCKR101 Plac-λint (E354A) | This study |
| pCINT I355A | pCKR101 Plac-λint (I355A) | This study |
| pCINT K356A | pCKR101 Plac-λint (K356A) | This study |
| Bacteriophage | ||
| P22-P′1 (II) | Pant P′1-arc-ant | (35) |
| P22-P′123 (I) | Pant P′123-arc-ant | (35) |
| P22-attL (I) | COC′-H′-P′321-arc-ant | (19) |
| Deoxyribonucleotides | ||
| IX-1 | d(GCCGGAATTCTTGCGTGTAATTGCGGAGACTTTGC) | |
| IX-2 | d(CGACGCATGCTCATTATTTGATTTCAATTTTGTCC) | |
| IX-2 INTY342A | d(CGACGCATGCTCATTATTTGATTTCAATTTTGTCCCACTCCCTGCC TCTGTCATCACGAGCCTGTGATGCC) | |
| IX-2 INTR346A | d(CGACGCATGCTCATTATTTGATTTCAATTTTGTCCCACTCCCTGCC TGCGTCATCACGA) | |
| IX-2 INTR348A | d(CGACGCATGCTCATTATTTGATTTCAATTTTGTCCCACTCCGCGCC TCTGTCA) | |
| IX-2 INTW350A | d(CGACGCATGCTCATTATTTGATTTCAATTTTGTCCGCCTCCCTGCCT) | |
| IX-2 INTD351A | d(CGACGCATGCTCATTATTTGATTTCAATTTTGGCCCACTCCCTG) | |
| IX-2 INTK352A | d(CGACGCATGCTCATTATTTGATTTCAATTGCGTCCCACTCC) | |
| IX-2 INTI353A | d(CGACGCATGCTCATTATTTGATTTCAGCTTTGTCCCAC) | |
| IX-2 INTE354A | d(CGACGCATGCTCATTATTTGATTGCAATTTTGTCC) | |
| IX-2 INTI355A | d(CGACGCATGCTCATTATTTGGCTTCAATTTTGTCC) | |
| IX-2 INTK356A | d(CGACGCATGCTCATTATGCGATTTCAATTTTGTCC) | |
| topattB2Ts5′Bio | Bio-d(CGTTGAAGCCTGCTTTTT) | |
| attBtop nosubstr | d(ATACTAACTTGAGC) | |
| attBBottom5′PO4 | d(pGCTCAAGTTAGTATAAAAAAGCAGGCTTCAACG) | |
| attB3′pNP | d(CGTTGAAGCCTGCTTT)-pNP | |
| attBLig2 | d(TTTATACTAACTTGAGC) | |
| oHN685′PO4 | d(pTCGCTCAAGTTAGTATAAAAAAGCAGGCTTCAACG) | |
| oHN68 | d(TCGCTCAAGTTAGTATAAAAAAGCAGGCTTCAACG) | |
| oHN69S | d(TCCGTTGAAGCCTGCTTT-ops-TTTATACTAACTTGAGC) |
Bio, biotin; pNP, p-nitrophenol; ops, 5′ bridging phosphorothiolate.
Construction of Int expression plasmids
Plasmids for producing mutant Int proteins were constructed by using ODNs that encoded the desired mutated int sequences and an engineered SphI restriction site downstream of the int translational stop codon. The latter ODNs contained nucleotide changes that produced the alanine substitutions and were used in PCRs with a partner ODN (IX-1) that hybridized upstream of int on a phage λ template. Successful PCR generated 1.3 kb products that contained int with the desired mutation. These products were digested with SphI and XmnI restriction enzymes and ligated into the pCKR101 vector, which had been digested with SphI and SnaBI restriction enzymes to place the int gene downstream of a Ptac promoter. The plasmids were transformed into competent Escherichia coli DH1 cells and plated on LB ampicillin (50 µg/ml) medium to select for the plasmid. Putative int clones were identified by electrophoresis of linearized plasmids in 1% agarose gels. The presence of the desired mutation and absence of secondary mutations in int was confirmed by DNA sequencing.
Site-specific recombination assays
Site-specific integration assays were performed using strain BWS209, a RW1817 derivative that has a galETK operon located in the chromosome flanked by the wild-type attP and attB sites (17). This strain forms red colonies on MacConkey plates that contain galactose. Recombination between the attP and attB sites deletes the galETK operon and leaves the attR site on the chromosome. Progeny of recombined cells can no longer use galactose as a carbon source and form white colonies on MacConkey galactose plates. Plasmids containing the int gene were transformed into electrocompetent BMS209 cells and incubated with aeration in 1 ml of LB at 37°C for 1 h. Dilutions were plated on MacConkey plates supplemented with galactose (1%) and Timetin (20 µg/ml). The plates were incubated for 24 h at 37°C. Integration efficiencies were calculated as the fraction of white colonies to total colonies. Results are reported as the average of at least two independent experiments.
Site-specific excision assays were performed using strain JG4117, which has a recombination-deficient λ prophage inserted into a secondary att site in the chromosomal galT gene (14). It also contains a plasmid (pYWH101) that supplies λ Xis. Int-induced excision of the prophage restores galT and allows the cell to use galactose as a carbon source. Plasmids containing the int gene were transformed into electrocompetent JG4117 cells and incubated with aeration in 1 ml of LB at 37°C for 1 h. Dilutions were plated on MacConkey plates supplemented with galactose (1%), Timetin (20 µg/ml), spectinomycin (100 µg/ml) and IPTG (0.5 mM). The plates were incubated for 48 h at 37°C. Recombination-proficient colony phenotypes ranged from red to white with numerous red papillae on excision medium; recombination-deficient colonies remained white. Excision efficiencies were calculated as the fraction of red and papillated colonies to total colonies. Results are reported as the average of at least two independent experiments.
Topoisomerase I activity assays
Topoisomerase I activity assays were performed following the protocol described by Kikuchi and Nash (18). CsCl-purified pBR322 DNA was used as the supercoiled plasmid substrate. Each sample contained either 100 ng of wild-type or mutant Int protein. Topoisomerase assays were performed in triplicate. Int concentrations were determined by electrophoresis of the crude extracts and 0.5–4.0 µg ovalbumin protein standards on 5%/12% discontinuous Tris–glycine– SDS gels. The gels were stained with Coomassie blue dye and scanned. The standards and Int were quantified using NIH Image software (Macintosh v.1.61). Int concentrations were determined from the linear portion of the ovalbumin standard curve.
Challenge phage assays
Challenge phage assays were performed as described by Han et al. (14) and MacWillams et al. (19). Challenge phages carrying the P′1, P′123 and attL sites were used in this study to measure the arm and core DNA binding affinities and cooperative interactions of wild-type and mutant Int proteins. P22-P′1 (II) contains the P′1 arm-type site and measures binding to a single arm site. P22-P′123 (I) measures cooperative binding to three consecutive arm sites. P22-attL (I) measures binding of core-type sites and formation of the IHF-dependent attL intasome complex. Relative binding affinities can be measured in vivo by comparing the frequency of challenge phage lysogeny induced by binding of wild-type or mutant Int to target sites at different levels of IPTG-controlled Int expression.
Purification of Int
Partially purified Int extracts were made for cleavage and ligation assays in vitro using the purification protocol described by Bankhead and Segall (20) except that no protease inhibitors were added after cell lysis and 0.4 M KCl was used in the initial extraction buffer.
Suicide attB cleavage assays
The attB cleavage assay is a variation of the suicide substrate assay originally described by Nunes-Duby et al. (21). The ODN for the cleavage assay [top strand, biotin-d(CGTTGAAGCCTGCTTTTT); bottom strand, d(pGCTCAAGTTAGTATAAAAAAGCAGGCTTCAACG)] were annealed at a top:bottom strand ratio of 1:10 to make a substrate containing the B core site. A 32P-labeled TMP was added to the 3′ end of the top strand by incubating 1 nmol attB substrate with 10 mM Tris–HCl (pH 7.5), 5 mM MgCl2, 7.5 mM dithiothreitol, 130 nM [α-32P]dTTP and 20 U exonuclease-minus large Klenow fragment (New England Biolabs) at 37°C for 60 min. Unincorporated [α-32P]dTTP was removed with a G50 Sephadex column. Cleavage assays were performed in 20 µl reactions containing Int extracts, 10 nM labeled attB substrate, 50 mM Tris–HCl (pH 8.0), 1 mM EDTA, 65 mM KCl, 10% glycerol and 200 µg/ml BSA. Extracts containing Int were added to the reaction and vortexed to start the reaction. Twenty microliter aliquots were removed at indicated time points and quenched by adding an equivalent volume of a solution containing 10 M urea and 0.05% SDS. Five microliters of MagnaBind™ streptavidin beads (Pierce) were added to the quenched reactions and incubated at room temperature for 5 min. Under these conditions the streptavidin beads bind >95% of the substrate. The reaction mixtures were then centrifuged at 14 000 g for 2 min and the supernatant and pellet moved to individual scintillation vials for quantification of cleavage events (represented by the movement of the 3′ label from the pellet to the supernatant). Background signal and cleavage efficiencies were calculated as the counts in the supernatant fraction divided by the total counts found in both supernatant and pellet. Subtracting the supernatant counts of a sample that lacked Int extract eliminated background signal that varied from 12 to 17% of the total radioactivity in a single reaction. Results are reported as the average of at least two independent experiments.
Phosphorothiolate cleavage assays
A phosphorothiolated ODN oHN69S [d(TCCGTTGAAG CCTGCTTT-ops-TTTATACTAACTTGAGC)] was synthesized as described (22). This ODN contains a 5′-bridging phosphorothiolate linkage (ops) at the site of Int-mediated strand cleavage and exchange. The phosphorothiolate cleavage substrate was prepared by mixing 1.5 nmol 5′ [α-32P]dATP-labeled oHN69S with 5.7 nmol oHN68 [d(TCG CTCAAGTTAGTATAAAAAAGCAGGCTTCAACG)] in 0.1 M KCl and 10 mM Tris–HCl (pH 8.0). The ODNs were annealed by heating to 70°C and cooling to 22°C at a rate of 1°C/min. The resulting substrate was stored at –20°C. Int-dependent cleavage reactions (50 µl) were performed as described in Burgin et al. (23) except that 2 or 4 µg Int extract (prepared as described above) was used per reaction. Cleavage was measured by removing 10 µl aliquots and mixing them with 10 µl of 10 M urea and 0.1% SDS at the specified times to stop the reaction. The quenched reactions were mixed with 20 µl of 90% formamide/1× TBE and heated to 100°C for 5 min to denature the DNA. Aliquots (10 µl) were immediately loaded onto 4–20% Tris–glycine gradient gels and subjected to electrophoresis at 180 V for 90 min at 37°C in 25 mM Tris base, 192 mM glycine and 0.1% SDS (pH 8.3). Gels were dried and exposed to phosphorimager plates. Cleavage was quantified using a Fuji FLA-3000 phosphorimager and Fujifilm Image Gauge software (Macintosh v.3.4). Results for each sample were calculated by dividing the total amount of counts present in the cleavage product band by the total amount of counts in the substrate and product bands. Results are reported as the average of at least two independent experiments.
p-Nitrophenol (pNP) attB ligation assays
A 3′ pNP DNA ODN [d(CGTTGAAGCCTGCTTT-pNP)] was synthesized as described (23). This ODN (attB3′pNP) was labeled with polynucleotide kinase and [α-32P]dATP at the 5′ end, then mixed with ODN attBLig2 [d(TTTATA CTAACTTGAGC)] and 5′ phosphorylated oHN68 in a 1:10:5 ratio. The mixture was heated to 70°C and cooled at a rate of 1°C/min to 22°C in 0.2 M NaCl and 1× TE. The annealed product (attB ligation substrate) was purified on a 20% acrylamide native gel, eluted from an excised gel slice in 1× TE, concentrated by chromatography on a DE52 column (1 cm) and resuspended in 1× TE buffer containing 0.2 M NaCl. The concentration of the attB ligation substrate was determined by spectrophotometry and additional attBLig2 ODN (350 pmol) was added in a ratio of 1:10 (attB ligation substrate:attBLig2) to compensate for loss of attBLig2 during purification. The attB ligation substrate was stored at –20°C. Assays were performed in 10 µl reactions containing 10 nM 32P-labeled attB ligation substrate, 50 mM Tris–HCl (pH 8.0), 1 mM EDTA, 65 mM KCl, 10% glycerol, 200 µg/ml BSA and Int extract. Reactions were started by the addition of Int protein and quenched by mixing reactants with equal volumes of 90% deionized formamide in 1× TBE. Samples were heated for 5 min at 100°C, loaded onto 15% TBE–urea gels and subjected to electrophoresis at 180 V for 90 min at 37°C in 89 mM Tris base, 89 mM boric acid and 2 mM EDTA (pH 8.3). Gels were dried and exposed to phosphorimager plates. Ligation levels were quantified using a Fuji FLA-3000 phosphorimager and Fujifilm Image Gauge software (Macintosh v.3.4). Results for each sample were calculated by dividing the total amount of counts present in the ligation product band by the total amount of counts in the substrate and product bands. Results are reported as the average of at least two independent experiments.
Comparative modeling of the Int catalytic domain and carboxyl tail region
To create a comparative three-dimensional model for the CAT domain of Int, residues 170–356 were modeled onto the 3crx atomic coordinates for Cre (24). The 3crx template structure was chosen because it facilitated examination of DNA binding and dimerization interactions when a protein dimer is complexed with a Holliday junction intermediate. According to the alignment published by Yang and Mizuuchi (25), the λ Int residues were manually threaded onto the 3crx structure using Deep View Swiss-PdbViewer client software (26). The sequence–structure alignment was then submitted to the Swiss-Model server (26) (http://www.expasy.org/swissmod/) for construction of a full-atom model. Graphic representations of the model were rendered with the free molecular graphics program PyMOL (27) (http://www.pymol.org/).
RESULTS
The topoisomerase I activity of Int performs strand cleavage and ligation reactions during site-specific recombination (18,28). An Int mutant, W350, eliminates the last seven residues of the protein and possesses increased topoisomerase I activity relative to the wild-type protein (14). Int W350 binds arm-type and core-type sites and forms the attL intasome complex as well as the wild-type protein (14). These observations suggest that the mutant phenotype results from altered catalytic activity or regulation of the cleavage and ligation reactions. We reasoned that other mutants with alanine substitutions in residues 350–356 could provide insight into the regulation of the strand cleavage or ligation reactions. We made single alanine substitutions in residue 342 (the position of the conserved catalytic tyrosine) and in residues 346–356 by directed PCR mutagenesis. These mutants were analyzed for topoisomerase, recombination, DNA binding, cleavage and ligation activities.
Recombination and topoisomerase I assays
Analysis of the recombination data and topoisomerase enzyme assays (Table 2) allowed classification of the mutants into two classes. The first class contains mutants that affect both topoisomerase and recombination activity (Y342A, R346A, R348A and K352A). As expected, the Y342A substitution mutant, which changes the catalytic tyrosine to alanine, lacked both topoisomerase and recombination activity. The R346A mutant lost all topoisomerase and recombination activity. The R348A mutant was partially deficient in topoisomerase activity and showed 10- and 3-fold lower site-specific integration and excision activity, respectively. The K352A mutant showed decreased topoisomerase activity and approximately 10-fold lowered recombination activity.
Table 2. Characterization of Int topoisomerase and recombination activity of alanine substitution mutants.
| Int mutationa | Recombination efficiencyb | Topoisomerase I activityc | |
|---|---|---|---|
| Integration (%) | Excision (%) | ||
| Wild-type Int | 98.0 | 100.0 | +++ |
| Y342A | 0.0 | 0.0 | – |
| R346A | 0.0 | 0.0 | – |
| R348A | 11.1 | 33.8 | ++ |
| W350 | <0.5 | <0.5 | ++++ |
| W350A | 6.0 | 5.6 | ++++ |
| D351A | 98.0 | 100.0 | +++ |
| K352A | 13.3 | 9.4 | ++ |
| I353A | <0.5 | <0.5 | +++ |
| E354A | 96.7 | 100.0 | +++ |
| I355A | 96.5 | 100.0 | +++ |
| K356A | 100.0 | 100.0 | +++ |
| Vector control | 0.0 | 0.0 | – |
aMutants were tested using the pCINT and pXM plasmids listed in Table 1. pCINT was used as the wild-type Int construct and pCKR101 was used as the parental vector control. W350 is a nonsense mutant.
bRecombination efficiencies were calculated as described in Materials and Methods.
cQualitative results are described. +++, topoisomerase I activity equivalent to wild-type Int incubated with supercoiled plasmid; ++++, enhanced topoisomerase activity; ++, reduced topoisomerase activity; –, no detectable relaxation. Topoisomerase assays were performed as described in Materials and Methods.
The other class of mutants disrupts recombination but does not inhibit topoisomerase activity (W350, W350A and I353A). Mutants W350 and W350A showed enhanced topoisomerase activity but greatly reduced recombination. Recombination of the W350 truncation mutant was reduced over 100-fold and recombination of the W350A mutant was reduced over 10-fold. Interestingly, Int W350A topoisomerase activity was enhanced yet reproducibly less than that of the W350 mutation. The I353A mutant possessed wild-type topoisomerase activity yet was more than 100-fold deficient in recombination. The remaining mutants with alanine substitutions at residues 351, 354, 355 and 356 exhibited wild-type recombination and topoisomerase activities.
DNA binding assays
P22-P′1 (II), P22-P′123 (I) and P22-attL (I) challenge phages were used to determine if the alanine substitutions affected DNA binding of the Int mutants in vivo. The P22-P′1 (II) challenge phage contains the P′1 arm site and is designed to test binding affinity to a single arm-type site. The P22-P′123 (I) challenge phage contains the contiguous P′123 sites located in attP and attL. This phage tests the ability of Int to bind the three arm-type sites cooperatively. Binding to the P22-attL (I) challenge phage is dependent on IHF binding to its wild-type H′ site and simultaneous binding of a single Int molecule to both an arm and core site; this cooperative interaction assembles the attL intasome (29,30).
All of the mutants repressed P22-P′1 (II), P22-P′123 (I) and P22-attL (I) challenge phages (Fig. 3). The substitution mutants had slightly better repression efficiencies than wild-type Int. The P22-P′123 and P22-attL challenge phage results demonstrated efficient binding with wild-type Int and all of the substitution mutants. These findings suggest that the recombination defects observed are not due to dramatic changes in DNA binding affinity, cooperative binding to arm-type sites or in the protein–protein interactions necessary for repressing attL challenge phage. These results also suggest that the recombination and topoisomerase phenotypes of the alanine substitution mutants are due to specific disruptions in the catalytic activity of the site-specific recombination pathway or in the regulation of such activity after DNA binding.
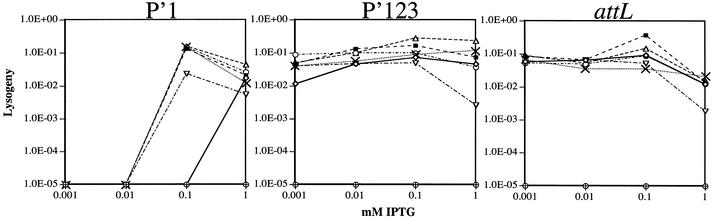
Figure 3.
Challenge phage assays. Results of P′1, P′123 and attL challenge phage assays with wild-type Int (open diamonds), no Int (open crossed circles) and Int mutants Y342A (solid squares), R346A (open circles), R348A (open triangles), W350A (crosses) and I353A (open inverted triangles). The curves show the frequency of lysogeny as a function of total cells infected by the phage at each IPTG concentration.
Cleavage and ligation assays
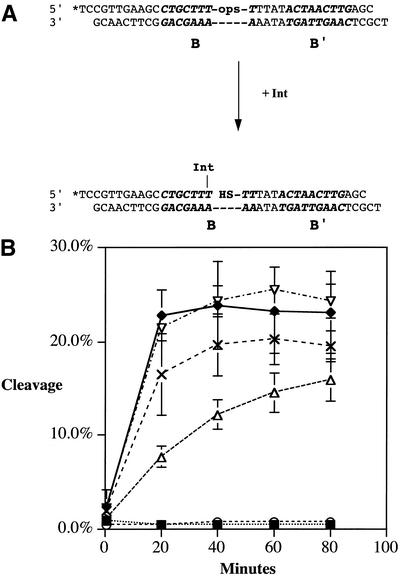
The topoisomerase I assays indicated that some of the Int mutants had topoisomerase defects. Since Int relaxes DNA by successive cleavage and ligation events, the mutant phenotypes could arise from alterations in either of these activities. Quantitative assays were developed to independently measure the rates of cleavage and ligation when Int binds to the B core-type site of an attB substrate. To determine the maximal cleavage rates of mutant and wild-type Int, a suicide attB cleavage assay was developed. This assay is a modification of the suicide cleavage assay developed by Nunes-Duby et al. (21). The substrate has a nick in the DNA backbone positioned three bases after the initial strand Int cleavage site; consequently, Int-mediated cleavage results in the production of a trinucleotide that dissociates. Removal of this segment of DNA covalently traps the Int on the DNA substrate in a phosphotyrosyl linkage (Fig. 4A). We measured the release of the 3′-labeled DNA product after Int cleavage at the B core site as described in Materials and Methods. To assay the maximal cleavage rates without possible artifacts caused by differences in protein stability or core-type DNA binding defects, the concentration of Int in the suicide attB cleavage reactions was increased until cleavage rates were maximal. At excess concentrations of Int, the observed cleavage rates should not be affected by changes in substrate binding affinity and therefore represent the maximal rate of cleavage under saturating Int concentrations.
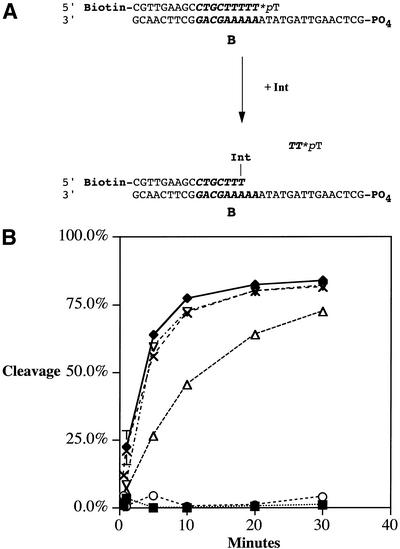
Figure 4.
Suicide attB cleavage assays. (A) Int binds to the attB core-type site and cleaves the top strand. Unreacted substrate is attached to streptavidin-coated beads and removed from the reaction volume by centrifugation; cleavage levels are determined by scintillation counting of the label present in the supernatant. Biotin, 5′ attached biotinyl group; *pT, terminal thymidylate labeled with 32P at the last phosphodiester bond; B, B core-type binding site (indicated in bold italicized sequence); PO4, 5′ phosphate group. (B) Results of the suicide attB cleavage assay with wild-type Int (solid diamonds) and Int mutants Y342A (solid squares), R346A (open circles), R348A (open triangles), W350A (crosses) and I353A (open inverted triangles). The curves show the release of radiolabeled product over time.
The results of these assays are shown in Table 3 and illustrated in Figure 4B. Mutants Y342A and R346A lacked cleavage activity, which was expected given the topoisomerase and recombination phenotypes. The R348A mutant depressed cleavage activity. Int mutants W350 and W350A cleaved the B site as well as the wild-type protein. The K352A, I353A and remaining substitution mutants D351A, E354A, I355A and K356A (data not shown) cleaved at wild-type levels.
Table 3. Cleavage and ligation activities of alanine substitution mutants.
| Int mutationa | 3′ attB cleavage assayb | Phosphorothiolate cleavage assayb | p-Nitrophenol ligation assayb |
|---|---|---|---|
| Wild-type Int | +++ | +++ | +++ |
| Y342A | – | – | +++ |
| R346A | – | – | +++ |
| R348A | ++ | ++ | +++ |
| W350 | +++c | +++c | |
| W350A | +++ | +++ | +++ |
| D351A | +++ | +++ | |
| K352A | +++ | +++ | |
| I353A | +++ | +++ | +++ |
| E354A | +++ | +++ | |
| I355A | +++ | +++ | |
| K356A | +++ | +++ | |
| Vector control | – | – | – |
aMutants used are described in Table 2.
bAssays performed with purified extracts of indicated proteins as described in Materials and Methods. +++, wild-type activity; ++, reduced activity; –, no activity.
cCleavage assays were performed with wild-type and W350 crude extracts.
Phosphorothiolate cleavage assays. Phosphorothiolate cleavage assays were performed as described in Materials and Methods and illustrated in Figure 5A. This assay is a modification of the cleavage assay developed by Burgin et al. (23). It uses an attB DNA substrate with a bridging phosphorothiolate at the Int cleavage site on the top strand. The presence of the phosphorothiolate at the cleavage site allows cleavage but prevents the reverse ligation reaction. This assay supplied a second, independent method for measuring cleavage without using a nicked DNA substrate.
Figure 5.
Phosphorothiolate attB cleavage assays. (A) Int cleaves the top strand of attB. Cleavage is measured by separation of the labeled DNA–protein product on 4–20% non-denaturing polyacrylamide gradient gels and quantifying the complex on a Fuji FLA-3000 phosphorimager. *, 5′ 32P label; B and B′, core-type binding site (indicated in bold italicized sequence). The -ops- in the top sequence indicates the phosphorothiolate linkage; the dashes in the sequence indicate a continuous DNA backbone. (B) Results of the phosphorothiolate attB cleavage assay with wild-type Int (solid diamonds) and Int mutants Y342A (solid squares), R346A (open circles), R348A (open triangles), W350A (crosses) and I353A (open inverted triangles). The curves show accumulation of attB–Int complexes over time.
The results of these assays are shown in Table 3 and illustrated in Figure 5B. As seen in the attB suicide cleavage assay, we detected no significant cleavage with the Y342A and R346A mutant Int proteins using the phosphorothiolated substrate. Similarly, the R348A protein exhibited partial cleavage activity with both the suicide and phosphorothiolated substrate. The wild-type and I353A Int proteins had equivalent cleavage activity with both substrates. The cleavage activity of the W350A mutant displayed wild-type cleavage levels when tested with the attB suicide substrate but displayed slightly yet reproducibly lower cleavage activity using the phosphorothiolated substrate. The other substitution mutants tested (D351A, K352A, E354A, I355A and K356A) had cleavage activity equivalent to wild-type Int (data not shown).
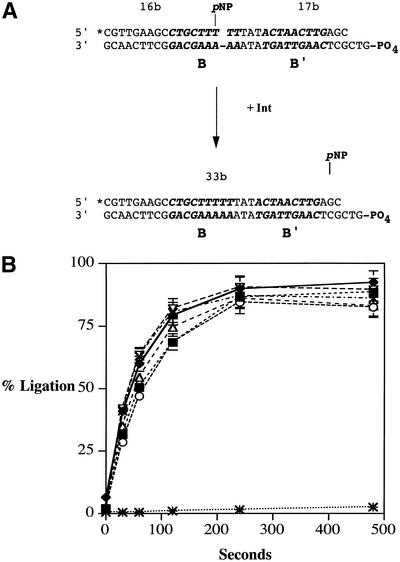
attB ligation assays. To determine the maximal ligation rates of mutant and wild-type Int, a pNP ligation assay was developed. This assay is a modification of the pNP ligation assay developed by Burgin and co-workers (22) and represents the first use of this ligation assay with λ Int. As described in Figure 6A and Materials and Methods, ligation is measured by quantifying the amount of 33 bp ligation product formed after incubation of the activated attB ligation substrate with Int. The concentration of Int in the pNP ligation assays was increased until ligation rates did not change. Results from the pNP ligation assay are shown in Table 3 and illustrated in Figure 6B. All of the alanine substitution mutants from residues 346–356 and the W350 truncation mutant had ligation activity like that of wild-type Int. Since the substitution and truncation mutants have no effect on attB ligation, the enhanced topoisomerase activity exhibited by the W350 and W350A mutants is not due to inherent defects in cleavage or ligation by the protein; rather, it likely arises from events that coordinate these activities. This conclusion may also apply to the I353A substitution mutant, whose topoisomerase activity is greatly enhanced relative to its recombination activity, yet displays wild-type cleavage and ligation activity.
Figure 6.
attB p-nitrophenol ligation assays. (A) Int ligates the attB top strand using a pNP-activated substrate. Ligation rates were measured by separating the 33 bp product on 15% denaturing polyacrylamide gels; products were quantified using a Fuji FLA-3000 phosphorimager. *, 5′ 32P label; 16b, 17b and 33b, bases; B and B′, core-type binding site (indicated in bold italicized sequence); pNP, p-nitrophenol; PO4, 5′ phosphate group. The gap in the top sequence indicates a nick; dashes in the sequence indicate a continuous DNA backbone. (B) Results of the attB pNP ligation assay with wild-type Int (solid diamonds), no Int (asterisks) and Int mutants Y342A (solid squares), R346A (open circles), R348A (open triangles), W350A (crosses) and I353A (open inverted triangles). The curves show the accumulation of ligated attB product over time.
DISCUSSION
Int catalyzes the λ site-specific recombination reaction in conjunction with accessory factors. Site-specific recombination reactions require recombinases with the ability to specifically bind, cleave, exchange and ligate their DNA targets. The topoisomerase I activity exhibited by Int is responsible for the cleavage and ligation activity occurring during site-specific recombination.
The Int W350 mutant, which is missing the last seven amino acids present in the wild-type, is defective in both integrative and excisive recombination but has increased topoisomerase activity. The Int W350 protein displays wild-type activities in DNA binding and protein–protein interactions leading to the formation of intasome complexes (14). These findings suggest that its phenotype could be due to changes in the inherent cleavage, strand exchange, or ligation activities. Alternatively, the phenotype could be due to changes in regulation of these functions during recombination. Han et al. (14) suggested that loss of the C-terminus in the W350 Int mutant might have uncovered the catalytic pocket, resulting in the enhanced cleavage and topoisomerase activity. Recent structural and genetic studies propose that the C-termini of the tyrosine recombinases function as active site switches that regulate the activity of recombinases in the intasome (6,31).
We constructed alanine substitutions of Int residues 342 and of several other positions between residues 346 and 356 to help understand the role of the Int C-terminus in site-specific recombination. We analyzed mutants for altered topoisomerase and recombination activities, since changes in topoisomerase activity likely reflect specific alterations in the site-specific recombination mechanism. Topoisomerase and recombination assays of the alanine substitution mutants in residues 346–356 demonstrate that only the W350A residue substitution mimics the enhanced topoisomerase phenotype displayed by the original W350 mutant. The enhanced topoisomerase I activity mutants W350 and W350A have greater than 100- and 10-fold recombination deficiencies, respectively. The R348A and K352A mutants are partially deficient in both topoisomerase and recombination activities, whereas the Y342A and R346A mutants displayed no detectable topoisomerase or recombination activity. An exception to the apparent connection between topoisomerase and recombination activity was the I353A substitution mutant that displayed wild-type topoisomerase activity but was more than 100-fold deficient in recombination activity. The remaining mutants had both wild-type topoisomerase and recombination activities.
These recombination defects cannot be explained by dramatic alterations in DNA binding affinities. The challenge phage assays, which measured relative binding affinities for the arm sites as well as formation of the attL intasome structure in vivo, demonstrated that the mutants bind their target arm and form attL intasomes like wild-type Int. In contrast, all other recombination-defective Int substitution mutants isolated that contain amino acid substitutions in the catalytic domain ranging from residues 175 to 332 have significant defects in forming the attL intasome (14). These results suggest that the residues in the region 346–356 are not essential for DNA binding affinity or assembly of an attL intasome complex. Therefore, we conclude that the most likely causes of the recombination deficiencies in the substitution mutants were alterations in cleavage, strand transfer, or ligation activities.
To test this hypothesis, we developed suicide substrate assays and ligation assays to quantify the inherent cleavage and ligation activities of Int at the B core site. Cleavage and ligation assays were performed under saturating concentrations of enzyme. The data should therefore represent cleavage and ligation activity independent of protein instability or relative changes in core site affinity. Results from the suicide attB and phosphorothiolate cleavage assays indicated that the Y342A, R346A and R348A mutants demonstrated moderate to severe disruption in the ability to cleave DNA, which would explain their topoisomerase and recombination phenotypes. The W350A mutant had slightly lower cleavage activity in the phosphorothiolate cleavage assay yet wild-type cleavage activity in the attB suicide cleavage assay. Neither the W350 truncation mutant nor the alanine substitution mutants at residues 351–356 showed significant changes in inherent cleavage activity.
Recently, work done by Tekle et al. (32) showed that a W350 mutant that lacked the first 64 amino acid residues of Int possessed enhanced cleavage activity when assayed on suicide substrates that contain only the B core site. We cannot explain the difference between our results and theirs with the W350 mutant. Differences in experimental procedures included the structure of the proteins assayed, the structure of the suicide substrates and the assay conditions.
Ligation assays using the activated pNP attB ligation substrate demonstrated that the Y342A mutant and all the mutants in the 346–356 region could perform ligation as well as wild-type Int. Therefore, we conclude that the altered topoisomerase and recombination activities of the mutants from 350 to 356 are likely not due to disruption of the inherent ability of Int proteins to cleave or ligate at the wild-type target site. One model that explains these seemingly paradoxical results is that deletion of the last seven residues or substitution of the W350 residue to alanine alters activities that control the efficiency of cleavage or ligation in complex reactions involving topoisomerase and site-specific recombination substrates.
Mutations within the 346–356 residues cause disruption of active sites or alteration of Int-specific multimer contacts, leading to recombination defects
The W350 and W350A mutants form attL intasomes like the wild-type protein (this work; 19) and productive recombination can be observed using these mutants in vivo. Thus, it is likely that intasomes composed of these Int monomers can synapse with partner sites. However, productive recombination is a rare event, possibly because intra- or intermolecular contacts that control the order or velocities of strand cleavage and exchange are altered. This could, in turn, disrupt the coordination of strand exchanges so that the yield of recombinant products is drastically decreased.
The loss of the structure formed by the last seven amino acids of Int or alteration of the protein structure in the W350A mutant could disrupt a regulatory, intramolecular contact on their own catalytic domain. This hypothesis is consistent with the model derived from the crystal structure of the Int Y342F catalytic domain (CAT domain) presented by Tirumalai et al. (12) (Fig. 7). In the crystal structure, residues 350–356 are a loose coil structure on the surface of the catalytic domain. It is possible that loss of this domain in the W350 mutant disrupts the ability of Int to control cleavage or ligation activity in its own active site, either through contacts to the surface of the catalytic domain or by indirect alteration of the conformation of the catalytic tyrosine. However, we failed to detect either enhancement in cleavage activity or reduction of ligation activity in our assays of mutants possessing enhanced topoisomerase activity.
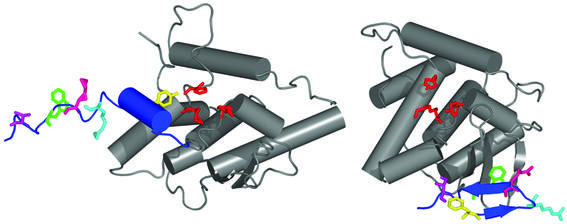
Figure 7.
Cre-threaded integrase model and Int carboxyl domain crystal structure model. An S173–K356 integrase monomer homology threaded onto the Cre Holliday junction structure model (left) and the crystal structure model of the inactive integrase catalytic domain (12) (right). A smoothed line represents the peptide backbone; cylinders and arrows represent predicted α-helices and β-sheets, respectively. The secondary structures of the two models are conserved (gray); changes between the models are from residues 334 to 356 (blue). Residues 334–341 are not defined in the Int crystal structure. The conserved Arg-His-Arg side chains are shown in red; the conserved catalytic tyrosine (phenylalanine in the Int crystal structure) is shown in yellow. Other labeled residues include R346 (light blue), R348 (pink), W350 (green) and I353 (purple). [Figure prepared with the program PyMOL (27).]
An alternative explanation for the W350 and W350A mutant phenotypes is disruption of intermolecular contacts necessary for both the topoisomerase and recombination reactions. During the topoisomerase assay, where intasomes are presumably not formed, the W350 and W350A proteins may be unable to regulate cleavage and/or ligation activity, thus producing increased topoisomerase activity. This suggests that the intermolecular interactions required for a productive recombination reaction may also be required for wild-type topoisomerase activity. If true, two or more Int monomers bound to DNA would be required for the properly regulated topoisomerase reaction.
An intermolecular model of topoisomerase and site-specific recombination regulation
Tyrosine recombinases make intermolecular contacts in recombination structures (6–9,33). The crystal structure prediction of four Cre monomers bound to a Holliday junction recombination intermediate showed that the C-terminal α-helices establish Cre–Cre contacts in a cyclic fashion (7,9,24). Two types of Cre–Cre contacts are observed in this recombination intermediate. One set of contacts are in a conformation that allows the catalytic tyrosine to cleave DNA substrate whereas the other contacts bind to and stabilize the active site of Cre in a conformation that prevents DNA cleavage by the receptor Cre proteins. From these structural data, Van Duyne (34) and Martin et al. (9) proposed that conformational alterations of the intasome structure suppress the cleavage activity of two Cre proteins at a given time, thus regulating the order of strand cleavage during recombination.
The ability of λ Int to adopt an intermolecular, Cre-like structure at the C-terminus appeared unlikely because of the length of the Int carboxyl tail compared to that of Cre (14 residues from the catalytic tyrosine in Int as opposed to 19 in Cre). In addition, there is a lack of sequence conservation between Int and Cre in the tail region and the crystal structure of the Y342F Int CAT domain lacks Int–Int intermolecular contacts at the C-terminus (12). However, the Cre crystal structure was solved when bound to its DNA substrate, whereas the crystal structure of the Int CAT domain was solved in the absence of DNA. The lack of a DNA substrate could allow crystallization of a different structure.
To examine whether Int could form an intermolecular structure similar to that of the Cre Holliday junction intasome structure, we threaded the Int sequence onto the crystal structure model of the DNA-bound Cre Holliday junction as described in Materials and Methods. We observed that Int folded in the form of Cre in the DNA-bound dimer, Holliday junction conformations appeared to be stable, and that residues 348–356 may form a loose α-helix that makes an intermolecular contact with another Int molecule (Fig. 8). We also compared the model of the threaded, DNA-bound Int structure to the crystal structure model of the Int CAT domain from Tirumalai et al. (12) (Fig. 7). This model repositions the catalytic tyrosine side chain residue proximal to the conserved catalytic triad in contrast to its distal position in the catalytic domain crystal structure.
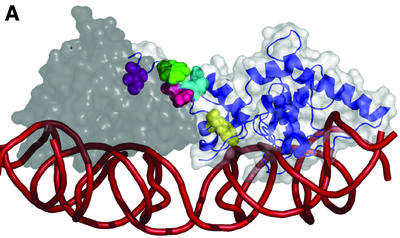
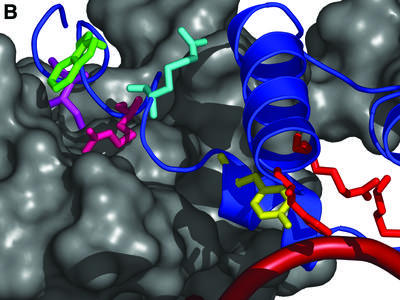
Figure 8.
Carboxyl tail-mediated intermolecular communication model. (A) Two integrase monomers homology threaded onto the Cre Holliday junction crystal structure. The S173–K356 catalytic domain is shown to illustrate the conformation of the C-terminus. The donor integrase (white space-filling structure with blue ribbon structure) is shown making an intermolecular contact to the acceptor integrase (gray) with the Int C-terminus. The Y342 (yellow), R346 (light blue), R348 (pink), W350 (blue) and I353 (purple) residues are highlighted on the donor Int molecule. The smoothed line (red) represents the DNA backbone. (B) A close-up of the donor Int carboxyl tail (blue ribbon structure) inserting into the acceptor Int groove (gray space-filled structure). Residues illustrated in (A) are shown. [Figure prepared with the program PyMOL (27).]
The threaded Int structure provides reasonable explanations for the changes in topoisomerase and recombination activity displayed by our substitution and truncation mutants. As proposed above, if two Int molecules are required for properly regulated relaxation of supercoiled DNA, it would be reasonable that the modeled carboxyl tail contact could form and suppress cleavage activity in one of the two Int monomers. Loss of the last seven residues of Int would remove this contact without disrupting the functional catalytic structure of the catalytic domain; as a result, cleavage activity would be unattenuated in both Int monomers and result in increased topoisomerase activity. Without the carboxyl tail to attenuate and regulate the order of strand cleavage, productive recombination events would be difficult to catalyze, thereby explaining the low frequency of productive recombination in vivo. Substitution of the large aromatic tryptophan side chain by alanine at residue 350 led to a phenotype similar to, but less extreme than, the W350 mutant. The carboxyl tail of this mutant may be destabilized and have more freedom to move in and out of the groove in the acceptor Int, resulting in decreased cleavage attenuation and frequency of productive recombination events.
Another mechanism that could rationalize the phenotypes of the W350 and W350A mutants in the threaded model would be incorrect positioning of the phosphotyrosyl intermediate during ligation. Loss or alteration of protein contacts could enhance mobility of the phosphotyrosyl bond within the active site. This could reduce the ligation efficiency during the topoisomerase assay and thereby increase relaxation of supercoiled substrates. Similarly, the movement of the phosphotyrosyl bond into a configuration sub-optimal for reaction could lower the efficiency of ligation during recombination. We would not observe lower ligation activities in our assays with the mutant enzymes in this scenario because the activated pNP substrate is a mimic of the true phosphotyrosyl intermediate in the ligation reaction and would not be affected by movement of a wild-type phosphotyrosyl bond.
The R346A and R348A substitution mutants resulted in complete or partial disruption of topoisomerase, recombination and cleavage activity. The threaded Int model predicts that the R346 and R348 side chains make the final contacts with the donor Int surface before extension of the carboxyl tail to make intermolecular contacts (Fig. 7). Substitution of these side chains might not only change the conformation of the carboxyl tail but also lead to destabilization of the Int active site. The localization of the R346 and R348 residues near the catalytic tyrosine could lead to movement of the catalytic tyrosine into alternative, non-catalytic conformations. These changes would be predicted to disrupt cleavage but not ligation, since substitution of the catalytic tyrosine does not affect the ability of Int to ligate a pNP-activated attB substrate in our assays (Y342A, Fig. 6B).
The CAT domain crystal structure model does not suggest an obvious reason for R346A-mediated disruption of cleavage, because the arginine side chain at residue 346 has been predicted to extend out into solution (12) (Fig. 7). Further evidence that the R346 residue plays an important role in the Int protein was revealed when a conservative R346K substitution had the same cleavage phenotype as the R346A substitution when tested with the attB suicide substrate (data not shown). Intolerance of the conservative arginine to lysine substitution suggests that the R346 residue either plays a role in a tightly coordinated function such as DNA cleavage, possibly by coordination of the catalytic tyrosine, or that it is essential for correct folding of the catalytic domain. However, we consider the latter possibility unlikely since R346A makes the arm and core site contacts necessary to form attL intasomes in vivo as well as binding and ligating activated DNA substrates.
The I353A substitution mutant is completely defective in both integrative and excisive site-specific recombination reactions yet it relaxes supercoiled DNA and binds, cleaves and ligates DNA like wild-type Int. Substitution of this residue appears to disrupt an activity unique to site-specific recombination. Both the threaded model (Fig. 8) and the crystal structure (Fig. 7) predict that the isoleucine residue at position 353 makes contacts with the catalytic domain, either in trans or in cis, respectively. It is possible that the I353A mutant is defective only when it is part of the intasome complex, where the mutation may cause improperly timed cleavage or ligation events that lead to unproductive recombination intermediates. Alternatively, the substitution mutant might encourage the formation of unproductive intasome conformations, thereby disrupting recombination activity. Further work will be required to understand, in detail, the specific functions of the I353 residue.
SUMMARY
Han et al. (14) described the W350 mutation as unique in its site-specific recombination and topoisomerase phenotypes yet wild-type DNA binding phenotype. It was predicted that the region from 350 to 356 specifically modulated the cleavage or ligation activity of Int. We isolated alanine substitution mutants of residues 346–356 and investigated their topoisomerase, recombination, cleavage and ligation activities. Based on our results, we constructed the Cre-threaded Int structure as a reasonable model of the Int catalytic domain during topoisomerase activity and site-specific recombination. We propose that the R346 and R348 residues are important for correct positioning of the catalytic tyrosine at residue Y342 and that controlled attenuation of DNA cleavage, mediated by the 350–356 Int terminus, is required for wild-type topoisomerase activity and efficient site-specific recombination.
Acknowledgments
ACKNOWLEDGEMENTS
We thank Ross Fasano for technical work. We also thank the members of the Gardner and Gumport laboratories for constructive discussions. This work was supported by NIH grant GM28717.
REFERENCES
- 1.Landy A. (1989) Dynamic, structural and regulatory aspects of lambda site-specific recombination. Annu. Rev. Biochem., 58, 913–949. [DOI] [PubMed] [Google Scholar]
- 2.Esposito D. and Scocca,J.J. (1997) The integrase family of tyrosine recombinases: evolution of a conserved active site domain. Nucleic Acids Res., 25, 3605–3614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Nunes-Duby S.E., Kwon,H.J., Tirumalai,R.S., Ellenberger,T. and Landy,A. (1998) Similarities and differences among 105 members of the Int family of site-specific recombinases. Nucleic Acids Res., 26, 391–406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Thompson J.F., Moitoso de Vargas,L., Koch,C., Kahmann,R. and Landy,A. (1987) Cellular factors couple recombination with growth phase: characterization of a new component in the lambda site-specific recombination pathway. Cell, 50, 901–908. [DOI] [PubMed] [Google Scholar]
- 5.Ball C.A. and Johnson,R.C. (1991) Multiple effects of Fis on integration and the control of lysogeny in phage lambda. J. Bacteriol., 173, 4032–4038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Guo F., Gopaul,D.N. and van Duyne,G.D. (1997) Structure of Cre recombinase complexed with DNA in a site-specific recombination synapse. Nature, 389, 40–46. [DOI] [PubMed] [Google Scholar]
- 7.Guo F., Gopaul,D.N. and Van Duyne,G.D. (1999) Asymmetric DNA bending in the Cre-loxP site-specific recombination synapse. Proc. Natl Acad. Sci. USA, 96, 7143–7148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chen Y., Narendra,U., Iype,L.E., Cox,M.M. and Rice,P.A. (2000) Crystal structure of a Flp recombinase-Holliday junction complex: assembly of an active oligomer by helix swapping. Mol. Cell, 6, 885–897. [PubMed] [Google Scholar]
- 9.Martin S.S., Pulido,E., Chu,V.C., Lechner,T.S. and Baldwin,E.P. (2002) The order of strand exchanges in Cre-LoxP recombination and its basis suggested by the crystal structure of a Cre-LoxP Holliday junction complex. J. Mol. Biol., 319, 107–127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Richet E., Abcarian,P. and Nash,H.A. (1988) Synapsis of attachment sites during lambda integrative recombination involves capture of a naked DNA by a protein–DNA complex. Cell, 52, 9–17. [DOI] [PubMed] [Google Scholar]
- 11.Moitoso de Vargas L., Pargellis,C.A., Hasan,N.M., Bushman,E.W. and Landy,A. (1988) Autonomous DNA binding domains of lambda integrase recognize two different sequence families. Cell, 54, 923–929. [DOI] [PubMed] [Google Scholar]
- 12.Tirumalai R.S., Healey,E. and Landy,A. (1997) The catalytic domain of lambda site-specific recombinase. Proc. Natl Acad. Sci. USA, 94, 6104–6109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Argos P., Landy,A., Abremski,K., Egan,J.B., Haggard-Ljungquist,E., Hoess,R.H., Kahn,M.L., Kalionis,B., Narayana,S.V., Pierson,L.S.,III et al. (1986) The integrase family of site-specific recombinases: regional similarities and global diversity. EMBO J., 5, 433–440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Han Y.W., Gumport,R.I. and Gardner,J.F. (1994) Mapping the functional domains of bacteriophage lambda integrase protein. J. Mol. Biol., 235, 908–925. [DOI] [PubMed] [Google Scholar]
- 15.Pargellis C.A., Nunes-Duby,S.E., de Vargas,L.M. and Landy,A. (1988) Suicide recombination substrates yield covalent lambda integrase-DNA complexes and lead to identification of the active site tyrosine. J. Biol. Chem., 263, 7678–7685. [PubMed] [Google Scholar]
- 16.Champoux J.J. (2001) DNA topoisomerases: structure, function and mechanism. Annu. Rev. Biochem., 70, 369–413. [DOI] [PubMed] [Google Scholar]
- 17.Yagil E., Dorgai,L. and Weisberg,R.A. (1995) Identifying determinants of recombination specificity: construction and characterization of chimeric bacteriophage integrases. J. Mol. Biol., 252, 163–177. [DOI] [PubMed] [Google Scholar]
- 18.Kikuchi Y. and Nash,H.A. (1979) Nicking-closing activity associated with bacteriophage lambda int gene product. Proc. Natl Acad. Sci. USA, 76, 3760–3764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.MacWilliams M.P., Gumport,R.I. and Gardner,J.F. (1996) Genetic analysis of the bacteriophage lambda attL nucleoprotein complex. Genetics, 143, 1069–1079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bankhead T. and Segall,A.M. (2000) Characterization of a mutation of bacteriophage lambda integrase. Putative role in core binding and strand exchange for a conserved residue. J. Biol. Chem., 275, 36949–36956. [DOI] [PubMed] [Google Scholar]
- 21.Nunes-Duby S.E., Matsumoto,L. and Landy,A. (1987) Site-specific recombination intermediates trapped with suicide substrates. Cell, 50, 779–788. [DOI] [PubMed] [Google Scholar]
- 22.Woodfield G., Cheng,C., Shuman,S. and Burgin,A.B. (2000) Vaccinia topoisomerase and Cre recombinase catalyze direct ligation of activated DNA substrates containing a 3′-para-nitrophenyl phosphate ester. Nucleic Acids Res., 28, 3323–3331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Burgin A.B. Jr, Huizenga,B.N. and Nash,H.A. (1995) A novel suicide substrate for DNA topoisomerases and site-specific recombinases. Nucleic Acids Res., 23, 2973–2979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Gopaul D.N., Guo,F. and Van Duyne,G.D. (1998) Structure of the Holliday junction intermediate in Cre-loxP site-specific recombination. EMBO J., 17, 4175–4187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yang W. and Mizuuchi,K. (1997) Site-specific recombination in plane view. Structure, 5, 1401–1406. [DOI] [PubMed] [Google Scholar]
- 26.Guex N. and Peitsch,M.C. (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis, 18, 2714–2723. [DOI] [PubMed] [Google Scholar]
- 27.DeLano W.L. (2001) The PyMOL Molecular Graphics System. DeLano Scientific, San Carlos, CA.
- 28.Nash H.A. and Pollock,T.J. (1983) Site-specific recombination of bacteriophage lambda. The change in topological linking number associated with exchange of DNA strands. J. Mol. Biol., 170, 19–38. [DOI] [PubMed] [Google Scholar]
- 29.Kim S., Moitoso de Vargas,L., Nunes-Duby,S.E. and Landy,A. (1990) Mapping of a higher order protein–DNA complex: two kinds of long-range interactions in lambda attL. Cell, 63, 773–781. [DOI] [PubMed] [Google Scholar]
- 30.MacWilliams M., Gumport,R.I. and Gardner,J.F. (1997) Mutational analysis of protein binding sites involved in formation of the bacteriophage lambda attL complex. J. Bacteriol., 179, 1059–1067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hallet B., Arciszewska,L.K. and Sherratt,D.J. (1999) Reciprocal control of catalysis by the tyrosine recombinases XerC and XerD: an enzymatic switch in site-specific recombination. Mol. Cell, 4, 949–959. [DOI] [PubMed] [Google Scholar]
- 32.Tekle M., Warren,D.J., Biswas,T., Ellenberger,T., Landy,A. and Nunes-Duby,S.E. (2002) Attenuating functions of the C-terminus of λ integrase. J. Mol. Biol., in press. [DOI] [PubMed] [Google Scholar]
- 33.Jessop L., Bankhead,T., Wong,D. and Segall,A.M. (2000) The amino terminus of bacteriophage lambda integrase is involved in protein–protein interactions during recombination. J. Bacteriol., 182, 1024–1034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Van Duyne G.D. (2002) In Craig,N.L., Craigie,R., Gellert,M. and Lambowitz,A.M. (eds), Mobile DNA II. ASM Press, Washington, DC.
- 35.Lee E.C., Gumport,R.I. and Gardner,J.F. (1990) Genetic analysis of bacteriophage lambda integrase interactions with arm-type attachment site sequences. J. Bacteriol., 172, 1529–1538. [DOI] [PMC free article] [PubMed] [Google Scholar]