Abstract
Fibroblast growth factors (FGFs) make up a large family of polypeptide growth factors that are found in organisms ranging from nematodes to humans. In vertebrates, the 22 members of the FGF family range in molecular mass from 17 to 34 kDa and share 13-71% amino acid identity. Between vertebrate species, FGFs are highly conserved in both gene structure and amino-acid sequence. FGFs have a high affinity for heparan sulfate proteoglycans and require heparan sulfate to activate one of four cell-surface FGF receptors. During embryonic development, FGFs have diverse roles in regulating cell proliferation, migration and differentiation. In the adult organism, FGFs are homeostatic factors and function in tissue repair and response to injury. When inappropriately expressed, some FGFs can contribute to the pathogenesis of cancer. A subset of the FGF family, expressed in adult tissue, is important for neuronal signal transduction in the central and peripheral nervous systems.
Gene organization and evolutionary history
Gene organization
The prototypical Fgf genes contain three coding exons (Figure 1), with exon 1 containing the initiation methionine, but several Fgf genes (for example, Fgf2 and Fgf3) have additional 5' transcribed sequence that initiates from upstream CUG codons [1,2]. The size of the coding portion of Fgf genes ranges from under 5 kb (in Fgf3 and Fgf4) to over 100 kb (in Fgf12). In several Fgf subfamilies, exon 1 is subdivided into between two and four alternatively spliced sub-exons (denoted 1A-1D in the case of Fgf8). In these Fgf genes, a single initiation codon (ATG) in exon 1A is used. This gene organization is conserved in humans, mouse and zebrafish, but its functional consequences are poorly understood. Other subfamilies of Fgfs (such as Fgf11-14) have alternative amino termini, which result from the use of alternative 5' exons. It is not known whether a common 5' untranslated exon splices to these exons or whether alternative promoter and regulatory sequences are used.
Figure 1.
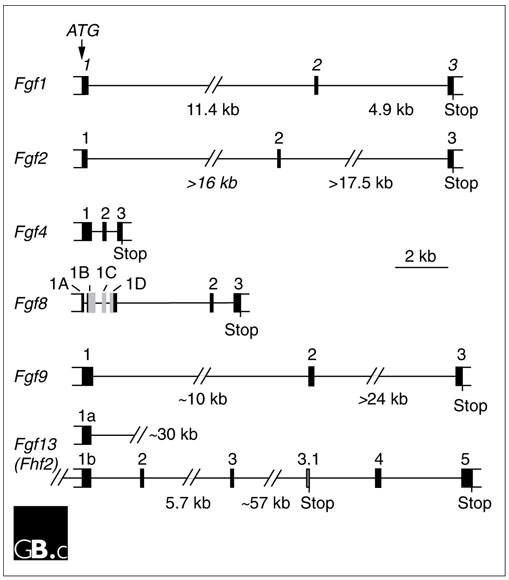
Gene structure of selected members of the Fgf family. Only the portion of each gene containing coding exons is shown. Constitutively expressed exons are in black; alternatively spliced exons are in gray. Fgfs1, 2, 4 and 9 contain the prototypic three-exon organization. For Fgf1, 5' untranslated exons are not shown; inclusion of these exons extends the gene by approximately 69 kb [78]. Fgf8 is an example of a gene with 5' alternative splicing, and Fgf13 demonstrates alternatively used 5' exons separated by over 30 kb. References: Fgf1 [78]; Fgf2 [79]; Fgf4 [80]; Fgf8 [52]; Fgf9 [81]; Fgf13 [76].
Most Fgf genes are found scattered throughout the genome. In human, 22 FGF genes have been identified and the chromosomal locations of all except FGF16 are known (Table 1) [3,4,5,6,7]. Several human FGF genes are clustered within the genome. FGF3, FGF4 and FGF19 are located on chromosome 11q13 and are separated by only 40 and 10 kb, respectively; FGF6 and FGF23 are located within 55 kb on chromosome 12p13; and FGF17 and FGF20 map to chromosome 8p21-p22. These gene locations indicate that the FGF gene family was generated both by gene and chromosomal duplication and translocation during evolution. Interestingly, a transcriptionally active portion of human FGF7, located on chromosome 15q13-q22, has been amplified to about 16 copies, which are dispersed throughout the human genome [8].
Table 1.
Chromosomal localizations of FGFs in human and mouse
| Human | Mouse | References | Accession numbers | |||
| Gene | Location | Gene | Location | Human | Mouse | |
| FGF1 | 5q31 | Fgf1 | 18 | [82,83] | X65778, E03692, E04557 | U67610, M30641 |
| FGF2 | 4q26-27 | Fgf2 | 3A2-B | [84,85] | E05628, M27968 | M30644, AF065903, AF065904, AF065905 |
| FGF3 | 11q13 | Fgf3 | 7F | [86,87,88] | X14445 | Y00848 |
| FGF4 | 11q13.3 | Fgf4 | 7F | [87,89] | E03343 | M30642 |
| FGF5 | 4q21 | Fgf5 | 5E1-F | [85,90] | M37825 | M30643 |
| FGF6 | 12p13 | Fgf6 | 6F3-G1 | [91,92] | X63454 | M92416 |
| FGF7 | 15q15-21.1 | Fgf7 | 2F-G | [93,94] | M60828 | Z22703 |
| FGF8 | 10q24 | Fgf8 | 19C3-D | [54,95] | U36223, U56978 | Z48746 |
| FGF9 | 13q11-q12 | Fgf9 | 14D | [81,96,97] | D14838 | U33535, D38258 |
| FGF10 | 5p12-p13 | Fgf10 | 13A3-A4 | [98,99] | AB002097 | D89080 |
| FGF11 | 17p13.1 | Fgf11 | - | [100] | U66199 | U66203 |
| (FHF3) | ||||||
| FGF12 | 3q28 | Fgf12 | 16B1-B3 | [31,100,101,102] | U66197 | U66201 |
| (FHF1) | ||||||
| FGF13 | Xq26 | Fgf13 | X | [31,76,103] | U66198 | U66202, AF020737 |
| (FHF2) | ||||||
| FGF14 | 13q34 | Fgf14 | 14 | [31] | U66200 | U66204 |
| (FHF4) | ||||||
| - | Fgf15* | 7F | (N.I., unpublished observations) | AF007268 | ||
| FGF16 | - | Fgf16 | - | AB009391 | AB049219 | |
| FGF17 | 8p21 | Fgf17 | 14 | [104] | AB009249 | AB009250 |
| FGF18 | 5q34 | Fgf18 | - | [105] | AB007422, AF075292 | AB004639, AF075291 |
| FGF19* | 11q13.1 | - | [106] | AB018122, AF110400 | ||
| FGF20 | 8p21.3-p22 | Fgf20 | - | [27,107] | AB030648, AB044277 | AB049218 |
| FGF21 | 19q13.1-qter | Fgf21 | - | [108] | AB021975 | AB025718 |
| FGF22 | 19p13.3 | Fgf22 | - | [109] | AB021925 | AB036765 |
| FGF23 | 12p13.3 | Fgf23 | 6F3-G1 | [7,75] (N.I., unpublished) | AB037973, AF263537 | AB037889, AF263536 |
*Human FGF19 and mouse Fgf15 may be orthologous genes.
In the mouse, there are at least 22 Fgf genes [3,9], and the locations of 16 have been identified (Table 1). Many of the mouse Fgf genes are scattered throughout the genome, but as in the human, Fgf3, Fgf4 and Fgf19 are closely linked (within 80 kb on chromosome 7F) and Fgf6 and Fgf23 are closely linked on chromosome 6F3-G1.
Evolutionary history
Fgfs have been identified in both invertebrates and vertebrates [3]. Interestingly, an Fgf-like gene is also encoded in the nuclear polyhedrosis virus genome [10]. Fgf-like sequences have not been found in unicellular organisms such as Escherichia coli and Saccharomyces cerevisiae. Although the Drosophila and Caenorhabditis elegans genomes have been sequenced, only one Fgf gene (branchless) has been identified in Drosophila [11] and two (egl-17 and let-756) have been identified in C. elegans [12,13], in contrast to the large number of Fgf genes identified in vertebrates. The evolutionary relationship between invertebrate and vertebrate Fgfs is shown in Figure 2a.
Figure 2.
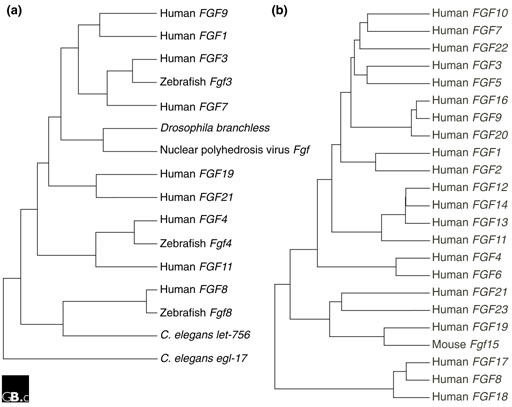
Evolutionary relationships within the FGF family. (a) Apparent evolutionary relationships between FGFs from vertebrates, invertebrates and a virus. Amino-acid sequences of nine representative FGFs were chosen from human and compared with FGFs from Drosophila, C. elegans, zebrafish and Autographa californica nuclear polyhedrosis virus. (b) Apparent evolutionary relationships of the 22 known human and murine FGFs. Sequences were aligned using Genetyxsequence analysis software and trees were constructed from the alignments using the neighbor-joining method.
The Fgf gene expansion has been hypothesized to be coincident with a phase of global gene duplications that took place during the period leading to the emergence of vertebrates [14]. Across species, most orthologous FGF proteins are highly conserved and share greater than 90% amino-acid sequence identity (except human FGF15 and mouse Fgf19; see below). To date, four Fgfs (Fgf3, 8, 17 and 18) have been identified in zebrafish, seven (Fgf3, Fgf(i), Fgf(ii), Fgf8, 9 and 20) in Xenopus (Fgf(i) and Fgf(ii) are most closely related to Fgf4 and Fgf6 [15]) and seven (Fgf2, 4, 8, 12, 14, 18 and 19) in chicken [3].
The apparent evolutionary relationships of the 22 known human FGFs are shown in Figure 2b. Vertebrate FGFs can be classified into several subgroups or subfamilies. Members of a subgroup of FGFs share increased sequence similarity and biochemical and developmental properties. For example, members of the FGF8 subfamily (FGF8, FGF17, and FGF18) have 70-80% amino acid sequence identity, similar receptor-binding properties and some overlapping sites of expression (for example, the midbrain-hindbrain junction) [16,17]. Members of FGF subgroups are not closely linked in the genome, however, indicating that the subfamilies were generated by gene-translocation or by genome-duplication events, not by local duplication events.
Human FGF15 and mouse Fgf19 have not been identified. Human FGF19 is evolutionarily most closely related to mouse Fgf15 (51% amino acid identity; Figure 2b) [18] and both the human FGF19 and mouse Fgf15 genes are closely linked to the human and mouse Fgf3 and Fgf4 genes on orthologous regions of human chromosome 11q13 and mouse chromosome 7F (N.I., unpublished observations). These findings indicate that human FGF19 may be the human ortholog of mouse Fgf15. Because all other Fgf orthologs share greater than 90% amino acid identity, it remains possible that the true orthologs of these genes have not been identified, have been lost or have diverged during vertebrate evolution.
Characteristic structural features
FGFs range in molecular weight from 17 to 34 kDa in vertebrates, whereas the Drosophila FGF is 84 kDa. Most FGFs share an internal core region of similarity, with 28 highly conserved and six identical amino-acid residues [19]. Ten of these highly conserved residues interact with the FGF receptor (FGFR) [20]. Structural studies on FGF1 and FGF2 identify 12 antiparallel β strands in the conserved core region of the protein (Figure 3) [21,22]. FGF1 and FGF2 have a β trefoil structure that contains four-stranded β sheets arranged in a triangular array (Figure 3b; reviewed in [23]). Two β strands (strands β10 and β11) contain several basic amino-acid residues that form the primary heparin-binding site on FGF2. Regions thought to be involved in receptor binding are distinct from regions that bind heparin (Figure 3) [21,22,23,24].
Figure 3.
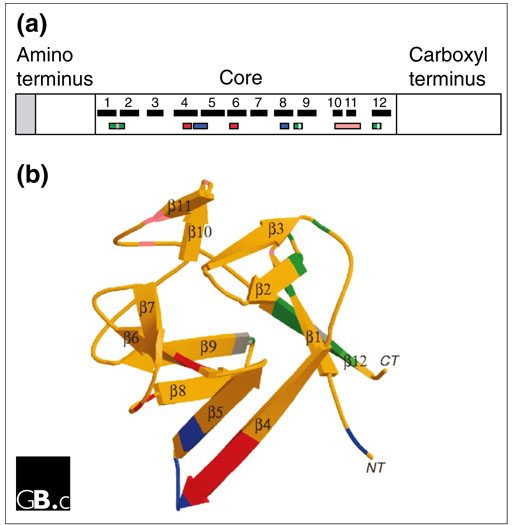
(a) Structural features of the FGF polypeptide. The amino terminus of some FGFs contains a signal sequence (shaded). All FGFs contain a core region that contains conserved amino-acid residues and conserved structural motifs. The locations of β strands within the core region are numbered and shown as black boxes. The heparin-binding region (pink) includes residues in the loop between β strands 1 and 2 and in β strands 10 and 11. Residues that contact the FGFR are shown in green (the region contacting Ig-domain 2 of the receptor), blue (contacting Ig-domain 3) and red (contacting the alternatively spliced region of Ig-domain 3). Amino-acid residues that contact the linker region are shown in gray [20]. (b) Three-dimensional structure of FGF2, a prototypical member of the FGF family. A ribbon diagram of FGF2 is shown; β strands are labeled 1-12 and regions of contact with the FGFR and heparin are color-coded as in (a) [22,24]. Image provided by M. Mohammadi.
Localization and function
Localization
Subcellular localization and secretion
Most FGFs (FGFs 3-8, 10, 15, 17-19, and 21-23) have amino-terminal signal peptides and are readily secreted from cells. FGFs 9, 16 and 20 lack an obvious amino-terminal signal peptide but are nevertheless secreted [25,26,27]. FGF1 and FGF2 also lack signal sequences, but, unlike FGF9, are not secreted; they can, however, befound on the cell surface and within the extracellular matrix. FGF1 and FGF2 may be released from damaged cells or could be released by an exocytotic mechanism that is independent of the endoplasmic-reticulum-Golgi pathway [28]. FGF9 has been shown to contain a non-cleaved amino-terminal hydrophobic sequence that is required for secretion [29,30]. A third subset of FGFs (FGF11-14) lack signal sequences and are thought to remain intracellular [31,32,33,34]. It is not known whether these FGFs interact with known FGFRs or function in a receptor-independent manner within the cell. FGF2 and FGF3 have high-molecular-weight forms that arise from initiation from upstream CUG codons [2,14,35]. The additional amino-terminal sequence in these proteins contains nuclear-localization signals, and the proteins can be found in the nucleus; the biological function of nuclear-localized FGF is unclear.
Developmental expression patterns and function
The 22 members of the mammalian FGF family are differentially expressed in many, if not all, tissues, but the patterns and timing of expression vary. Subfamilies of FGFs tend to have similar patterns of expression, although each FGF also appears to have unique sites of expression. Some FGFs are expressed exclusively during embryonic development (for example, Fgf3, 4, 8, 15, 17 and 19), whereas others are expressed in embryonic and adult tissues (for example, Fgf1, 2, 5-7, 9-14, 16, 18, and 20-23).
Function
The expression patterns of FGFs (see above) suggest that they have important roles in development. FGFs often signal directionally and reciprocally across epithelial-mesenchymal boundaries [36]. The integrity of these signaling pathways requires extremely tight regulation of FGF activity and receptor specificity. For example, in vertebrate limb development, mesenchymally expressed Fgf10 in the lateral-plate mesoderm induces the formation of the overlying apical ectodermal ridge; the ridge subsequently expresses Fgf8, which signals back to the underlying mesoderm [37]. This directional signaling initiates feedback loops and, along with other signaling molecules, regulates the outgrowth and patterning of the limb. Importantly, the differential expression of the alternative splice forms of the receptors in the apical ectodermal ridge and underlying mesoderm is such as to limit or prevent autocrine signaling within a given compartment.
Studies of the biochemical activities of FGFs have focused on the specificity of interactions between FGFs and FGFRs, on factors that affect the stability of FGFs and on the composition and mechanism of the active FGF-FGFR signaling complex.
Specificity of FGFs for FGF receptors
The FGFR tyrosine kinase receptors contain two or three immunoglobulin-like domains and a heparin-binding sequence [38,39,40]. Alternative mRNA splicing of the FGFR gene specifies the sequence of the carboxy-terminal half of immunoglobulin-domain III, resulting in either the IIIb or the IIIc isoform of the FGFR [41,42,43]. This alternative-splicing event is regulated in a tissue-specific manner and dramatically affects ligand-receptor binding specificity [44,45,46,47,48]. Exon IIIb is expressed in epithelial lineages and exon IIIc tends to be expressed in mesenchymal lineages [44,46,47,48]. In vitro patterns of binding specificity have been determined for each splice form of FGFR1-3 and for FGFR4, which is not alternatively spliced [49,50,51]. Ligands specific for these receptor splice forms are expressed in adjacent tissues, resulting in directional epithelial-mesenchymal signaling. For example, epithelially expressed FGFR2b (that is, FGFR2 IIIb isoform) can be activated by FGF7 and FGF10, ligands produced in mesenchymal tissue [49,50,51]. These ligands show no activity towards mesenchymally expressed FGFR2c. Conversely, FGF8 is expressed in epithelial tissue and activates FGFR2c but shows no activity towards FGFR2b ([49,52] and our unpublished observations). Notably, FGF8 expression is often restricted to epithelial tissue such as the apical ectodermal ridge of the developing limb bud [53,54].
Interaction with heparin or heparan sulfate proteoglycans
An important feature of FGF biology involves the interaction between FGF and heparin or heparan sulfate (HS) proteoglycan (HSPG) [19]. These interactions stabilize FGFs to thermal denaturation and proteolysis and may severely limit their diffusion and release into interstitial spaces [55,56]. FGFs must saturate nearby HS-binding sites before exerting an effect on tissue further away, or else must be mobilized by heparin/HS-degrading enzymes. The interaction between FGFs and HS results in the formation of dimers and higher-order oligomers [57,58,59]. Although the biologically active form of FGF is poorly defined, it has been established that heparin is required for FGF to effectively activate the FGFR in cells that are deficient in or unable to synthesize HSPG or in cells pretreated with heparin/HS-degrading enzymes or inhibitors of sulfation [60,61,62]. Genetic studies have also shown that mutations in enzymes involved in HS biosynthesis affect FGF signaling pathways during development [19,63]. Additional studies have shown that heparin and/or HS act to increase the affinity and half-life of the FGF-FGFR complex (reviewed in [40,64]).
A minimal complex containing one FGF molecule per FGFR can form in the absence of HS [24]. Structural studies suggest that HS may bridge FGF2 and the FGFR by binding to a groove formed by the heparan-binding sites of both the ligand and the receptor [24,65]. Binding studies with soluble chimeric FGFRs have identified a second potential FGF-binding site that, in some cases, can interact cooperatively with the primary FGF-binding site [66].
Important mutants
Many members of the Fgf family have been disrupted by homologous recombination in mice. The phenotypes range from very early embryonic lethality to subtle phenotypes in adult mice. The major phenotypes observed in Fgf knockout mice are shown in Table 2. Because FGFs within a subfamily have similar receptor-binding properties and overlapping patterns of expression, functional redundancy is likely to occur. This has been demonstrated for Fgf17 and Fgf8, which cooperate to regulate neuroepithelial proliferation in the midbrain-hindbrain junction [17]. In the case of Fgf knockouts resulting in early lethality, other functions later in development will need to be addressed by constructing conditional alleles that can be targeted at specific times and places in development. For example, Fgf8-/- mice die by embryonic day 9.5 [67]. A conditional allele for Fgf8 targeted to the apical ectodermal ridge has been used to demonstrate an essential role for Fgf8 in early limb development [68,69].
Table 2.
FGF knockout mice
| Gene | Survival of null mutant* | Phenotype | References |
| Fgf1 | Viable | None identified | [110] |
| Fgf2 | Viable | Mild cardiovascular, skeletal, neuronal | [110,111,112,113,114] |
| Fgf3 | Viable | Mild inner ear, skeletal (tail) | [115] |
| Fgf4 | Lethal, E4-5 | Inner cell mass proliferation | [116] |
| Fgf5 | Viable | Long hair, angora mutation | [72] |
| Fgf6 | Viable | Subtle, muscle regeneration | [117,118,119] |
| Fgf7 | Viable | Hair follicle growth, ureteric bud growth | [120,121] |
| Fgf8 | Lethal, E7 | Gastrulation defect, CNS development, limb development | [67,70,122,123] |
| Fgf9 | Lethal, P0 | Lung mesenchyme, XY sex reversal | [124]; (J.S. Colvin et al., personal communication) |
| Fgf10 | Lethal, P0 | Development of multiple organs, including limb, lung, thymus, pituitary | [125,126,127] |
| Fgf12 (Fhf1) | Viable | Neuromuscular phenotype | (J. Schoorlemmer and M. Goldfarb, personal communication) |
| Fgf14 (Fhf4) | Viable | Neurological phenotypes | (Q. Wang, personal communication) |
| Fgf15 | Lethal, E9.5 | Not clear | (J.R. McWhirter, personal communication) |
| Fgf17 | Viable | Cerebellar development | [17] |
| Fgf18 | Lethal, P0 | Skeletal development | (N. Ohbayashi, Z. Liu, personal communication) |
*E, embryonic day; P, postnatal day.
Several mutations in Fgf genes have been identified in C. elegans, Drosophila, zebrafish, mouse and human. The C. elegans gene egl-17 is required for sex myoblast migration [12], and a null allele of let-756 causes developmental arrest of the early larva [13]. The Drosophila branchless gene is required for tracheal branching and cell migration [11]. In zebrafish, acerebellar (ace) embryos lack the cerebellum and the midbrain-hindbrain boundary organizer. The ace gene encodes the zebrafish homolog of Fgf8 [70]. Interestingly, zebrafish aussicht mutant embryos, which overexpress Fgf8, also have defects in development of the central nervous system [71].
In the mouse, the angora mutation, which affects hair growth, was found to be allelic with Fgf5 [72]. A mouse mutant with a Crouzon-syndrome-like craniofacial dysmorphology phenotype was found to result from an insertional mutation in the Fgf3/Fgf4 locus [73]. Recently, positional cloning of the autosomal dominant hypophosphataemic rickets gene identified missense mutations in human FGF23 [74]. A recent paper demonstrates that this disease is caused by a gain-of-function mutation [75]. The chromosomal location (Xq26) and tissue-specific expression pattern of Fgf13 (also called Fhf2) suggests that it may be a candidate gene for Borjeson-Forssman-Lehmann syndrome, an X-linked mental retardation syndrome [76].
Frontiers
Issues most studied
FGFs have been intensely studied for nearly 30 years. Most of the early work focused on the mechanisms that regulate stability, secretion, export and interactions with heparin and on the mechanisms and consequences of signal transduction in various types of cells. More recent work has focused on the mechanisms regulating receptor specificity and receptor activation, the structure of the FGF-FGFR-HS complex, and the identification of new members of the FGF family. Functional studies have begun to address the role of FGFs in cell biology, development and physiology. Initial studies focused on the regulation of cell proliferation, migration and differentiation; more recent work has addressed the negative effect of FGFs and FGFRs on proliferation of some cell types, which was surprising as FGFs were thought to promote proliferation. In vitro studies have now been complemented by gene targeting in mice. The knockout approach has been fairly successful in identifying primary phenotypes but will be challenged by the need to address redundancy amongst the 22 FGFs and to study their developmental and physiological functions after the point of lethality of the null allele.
Unresolved questions
A major unresolved question concerns the mechanism(s) regulating FGF activity in vivo in the presence of cell-surface and extracellular-matrix HSPG. Current hypotheses predict that tissue-specific heparan fragments of defined sequence (and particularly of defined sulfation pattern) will differentially regulate FGFs by controlling their diffusion in the extracellular matrix and their ability to activate specific receptors [77]. These issues will be resolved by determining the sequence of tissue-specific HS and by demonstrating whether specific HS sequences can modulate the binding specificity of FGFs beyond that determined by the specific FGFR and its alternative splice form in the presence of heparin.
A second area of research will aim to elucidate the developmental roles of all the FGFs, first alone and then in various combinations. This will include determining whether a single FGF with a defined developmental function interacts with one or multiple FGFRs. A third major frontier will be to elucidate the physiological roles of FGFs that are expressed in adult tissues. This will again involve testing combinations of FGFs in cases in which knockouts are viable and designing conditional alleles in cases of embryonic lethality. Major areas being considered include neuronal and cardiovascular physiology, neuronal regeneration and homeostasis and tissue repair.
The last major frontier will be to elucidate the primary roles of FGFs in genetic diseases and cancer. Several FGFs were initially cloned from human and animal tumors. Future work will be required to determine whether FGF activation is itself an etiological agent in primary human tumors or whether it is a progression factor in the pathogenesis of cancer. As functional roles for FGFs are elucidated in embryonic development, it is expected that various human birth defects and genetic diseases will be attributed to mutations in Fgf genes. These studies will probably lead to the development of pharmacogenetic agents to treat these diseases. Because a large number of skeletal diseases are caused by mutations in Fgfr genes, it is anticipated that mutations in some Fgf genes will also be involved in skeletal pathology.
Acknowledgments
Acknowledgements
This work was supported by NIH grants CA60673 and HD35692 and by a grant from the American Heart Association (to D.M.O.) and by the Grant-in-Aid for Scientific Research from the Ministry of Education, Science and Culture, Japan and a grant from the Human Frontier Science Program, France (to N.I.).
References
- Kiefer P, Acland P, Pappin D, Peters G, Dickson C. Competition between nuclear localization and secretory signals determines the subcellular fate of a single CUG-initiated form of FGF3. EMBO J. 1994;13:4126–4136. doi: 10.1002/j.1460-2075.1994.tb06730.x. This is one of first papers to show that an FGF can initiate from an upstream CUG codon and can be localized in the nucleus. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arnaud E, Touriol C, Boutonnet C, Gensac MC, Vagner S, Prats H, Prats AC. A new 34-kilodalton isoform of human fibroblast growth factor 2 is cap dependently synthesized by using a non-AUG start codon and behaves as a survival factor. Mol Cell Biol. 1999;19:505–514. doi: 10.1128/mcb.19.1.505. Identification of a high-molecular-weight form of FGF2 that initiates from an upsteam CUG codon and is localized in the nucleus. [DOI] [PMC free article] [PubMed] [Google Scholar]
- GenBank http://www.ncbi.nlm.nih.gov/Genbank/index.html DNA sequences of FGFs listed in Table 1 can be accessed through this website.
- HUGO Gene Nomenclature Database http://www.gene.ucl.ac.uk/nomenclature/ This database contains nomenclature and mapping information for human genes.
- Cytokine Family cDNA Database http://cytokine.medic.kumamoto-u.ac.jp/CFC/FGF/FGF.html This database contains nomenclature and mapping information for the FGF family.
- LocusLink http://ncbi.nlm.nih.gov/LocusLink/ This database contains updated mapping information for the FGF family with links to a variety of other databases.
- Yamashita T, Yoshioka M, Itoh N. Identification of a novel fibroblast growth factor, FGF-23, preferentially expressed in the ventrolateral thalamic nucleus of the brain. Biochem Biophys Res Commun. 2000;277:494–498. doi: 10.1006/bbrc.2000.3696. FGF23 is the most recent reported member of the FGF family. This paper summarizes the chromosomal localization and apparent evolutionary relationships of human FGFs. [DOI] [PubMed] [Google Scholar]
- Kelley MJ, Pech M, Seuanez HN, Rubin JS, O'Brien SJ, Aaronson SA. Emergence of the keratinocyte growth factor multigene family during the great ape radiation. Proc Natl Acad Sci USA. 1992;89:9287–9291. doi: 10.1073/pnas.89.19.9287. This paper identifies a human multigene family composed of amplified and dispersed FGF7 gene copies and suggests models for the evolution of the great apes and humans. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mouse Genome Informatics http://www.informatics.jax.org This website provides information on mouse gene sequences, map position and function.
- Ayres MD, Howard SC, Kuzio J, Lopez-Ferber M, Possee RD. The complete DNA sequence of Autographa californica nuclear polyhedrosis virus. Virology. 1994;202:586–605. doi: 10.1006/viro.1994.1380. The complete nucleotide sequence of the genome of the baculovirus Autographa californica nuclear polyhedrosis virus (AcNPV) revealed that this genome encodes an FGF-like protein. [DOI] [PubMed] [Google Scholar]
- Sutherland D, Samakovlis C, Krasnow MA. branchless encodes a Drosophila FGF homolog that controls tracheal cell migration and the pattern of branching. Cell. 1996;87:1091–1101. doi: 10.1016/s0092-8674(00)81803-6. This paper identifies a Drosophila homolog (branchless, bnl) of mammalian FGFs, which activates the breathless FGF receptor tyrosine kinase. This FGF pathway regulates tracheal branching patterns by guiding tracheal cell migration during primary branch formation and by activating cytoplasmic branching at the ends of primary branches. [DOI] [PubMed] [Google Scholar]
- Burdine RD, Chen EB, Kwok SF, Stern MJ. egl-17 encodes an invertebrate fibroblast growth factor family member required specifically for sex myoblast migration in Caenorhabditis elegans. Proc Natl Acad Sci USA. 1997;94:2433–2437. doi: 10.1073/pnas.94.6.2433. The C. elegans gene egl-17 is a member of the Fgf family and one of the first known functional invertebrate FGFs. EGL-17 acts as a ligand for EGL-15 (FGFR) to regulate sex myoblast migration. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roubin R, Naert K, Popovici C, Vatcher G, Coulier F, Thierry-Mieg J, Pontarotti P, Birnbaum D, Baillie D, Thierry-Mieg D. let-756, a C. elegans Fgf essential for worm development. Oncogene. 1999;18:6741–6747. doi: 10.1038/sj/onc/1203074. Null mutations in C. elegansl et-756 lead to developmental arrest in early larval stages; let-756 is thus an essential gene for C. elegans development. [DOI] [PubMed] [Google Scholar]
- Coulier F, Pontarotti P, Roubin R, Hartung H, Goldfarb M, Birnbaum D. Of worms and men: an evolutionary perspective on the fibroblast growth factor (FGF) and FGF receptor families. J Mol Evol. 1997;44:43–56. doi: 10.1007/pl00006120. This review compares FGF and FGF receptor sequences in vertebrates and nonvertebrates and concludes that the FGF and FGF receptor families have evolved through phases of gene duplications, one of which may have coincided with the emergence of vertebrates. The review includes all four Fgfrs and 11 Fgfs. [DOI] [PubMed] [Google Scholar]
- Isaacs HV, Tannahill D, Slack JM. Expression of a novel FGF in the Xenopus embryo. A new candidate inducing factor for mesoderm formation and anteroposterior specification. Development. 1992;114:711–720. doi: 10.1242/dev.114.3.711. Identification of novel Xenopus FGFs. [DOI] [PubMed] [Google Scholar]
- Maruoka Y, Ohbayashi N, Hoshikawa M, Itoh N, Hogan BLM, Furuta Y. Comparison of the expression of three highly related genes, Fgf8, Fgf17 and Fgf18, in the mouse embryo. Mech Dev. 1998;74:175–177. doi: 10.1016/s0925-4773(98)00061-6. This paper compares the expression pattern of the Fgf8 subfamily which includes Fgf8, 17 and 18. Expression is examined during early development and in the developing brain and limbs. [DOI] [PubMed] [Google Scholar]
- Xu JS, Liu ZH, Ornitz DM. Temporal and spatial gradients of Fgf8 and Fgf17 regulate proliferation and differentiation of midline cerebellar structures. Development. 2000;127:1833–1843. doi: 10.1242/dev.127.9.1833. This is the first paper to demonstrate cooperation between two Fgfs during vertebrate development. [DOI] [PubMed] [Google Scholar]
- Nishimura T, Utsunomiya Y, Hoshikawa M, Ohuchi H, Itoh N. Structure and expression of a novel human FGF, FGF-19, expressed in the fetal brain. Biochim Biophys Acta. 1999;1444:148–151. doi: 10.1016/s0167-4781(98)00255-3. Identification of human FGF19 as a possible homolog of mouse Fgf15. Both of these genes are most closely related to FGFs 21 and 23. [DOI] [PubMed] [Google Scholar]
- Ornitz DM. FGFs, heparan sulfate and FGFRs: complex interactions essential for development. BioEssays. 2000;22:108–112. doi: 10.1002/(SICI)1521-1878(200002)22:2<108::AID-BIES2>3.0.CO;2-M. This review article addresses the developmental implications of the interactions between FGFs, FGFRs and heparan sulfate proteoglycans. The paper by Lin et al. [63], which demonstrates that heparan sulfate is essential for FGF function during Drosophila development, is highlighted. [DOI] [PubMed] [Google Scholar]
- Plotnikov AN, Hubbard SR, Schlessinger J, Mohammadi M. Crystal structures of two FGF-FGFR complexes reveal the determinants of ligand-receptor specificity. Cell. 2000;101:413–424. doi: 10.1016/s0092-8674(00)80851-x. The analysis of two different ligand-receptor crystal structures is used to determine elements that regulate ligand-binding specificity. [DOI] [PubMed] [Google Scholar]
- Zhu X, Komiya H, Chirino A, Faham S, Fox GM, Arakawa T, Hsu BT, Rees DC. Three-dimensional structures of acidic and basic fibroblast growth factors. Science. 1991;251:90–93. doi: 10.1126/science.1702556. This paper presents one of the first structures of FGF1 and FGF2. [DOI] [PubMed] [Google Scholar]
- Eriksson AE, Cousens LS, Weaver LH, Matthews BW. Three-dimensional structure of human basic fibroblast growth factor. Proc Natl Acad Sci USA. 1991;88:3441–3445. doi: 10.1073/pnas.88.8.3441. This paper presents one of the first structures of FGF2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Faham S, Linhardt RJ, Rees DC. Diversity does make a difference: fibroblast growth factor-heparin interactions. Curr Opin Struct Biol. 1998;8:578–586. doi: 10.1016/s0959-440x(98)80147-4. A review that models the formation of an FGF-FGFR-proteoglycan complex, based on the structure of FGF in complex with fragments of heparin. [DOI] [PubMed] [Google Scholar]
- Plotnikov AN, Schlessinger J, Hubbard SR, Mohammadi M. Structural basis for FGF receptor dimerization and activation. Cell. 1999;98:641–650. doi: 10.1016/s0092-8674(00)80051-3. Crystal structure of FGF2 complexed with FGFR1 showing extensive interaction between immunoglobulin domains 2 and 3 and the interdomain linker of the FGFR. [DOI] [PubMed] [Google Scholar]
- Miyake A, Konishi M, Martin FH, Hernday NA, Ozaki K, Yamamoto S, Mikami T, Arakawa T, Itoh N. Structure and expression of a novel member, FGF-16, of the fibroblast growth factor family. Biochem Biophys Res Commun. 1998;243:148–152. doi: 10.1006/bbrc.1998.8073. This paper identifies Fgf16 as a homolog of Fgf9. [DOI] [PubMed] [Google Scholar]
- Miyamoto M, Naruo K, Seko C, Matsumoto S, Kondo T, Kurokawa T. Molecular cloning of a novel cytokine cDNA encoding the ninth member of the fibroblast growth factor family, which has a unique secretion property. Mol Cell Biol. 1993;13:4251–4259. doi: 10.1128/mcb.13.7.4251. This paper identifies FGF9 as a new member of the FGF family without a classical signal peptide. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohmachi S, Watanabe Y, Mikami T, Kusu N, Ibi T, Akaike A, Itoh N. FGF-20, a novel neurotrophic factor, preferentially expressed in the substantia nigra pars compacta of rat brain. Biochem Biophys Res Commun. 2000;277:355–360. doi: 10.1006/bbrc.2000.3675. This paper identifies Fgf20 as a homolog of Fgf9 and Fgf16. [DOI] [PubMed] [Google Scholar]
- Mignatti P, Morimoto T, Rifkin DB. Basic fibroblast growth factor, a protein devoid of secretory signal sequence, is released by cells via a pathway independent of the endoplasmic reticulum-Golgi complex. J Cell Physiol. 1992;151:81–93. doi: 10.1002/jcp.1041510113. FGF2 lacks a signal peptide but is known to act extracellularly. This study shows that FGF2 can be released via an exocytotic mechanism independent of the endoplasmic reticulum-Golgi pathway. [DOI] [PubMed] [Google Scholar]
- Miyakawa K, Hatsuzawa K, Kurokawa T, Asada M, Kuroiwa T, Imamura T. A hydrophobic region locating at the center of fibroblast growth factor-9 is crucial for its secretion. J Biol Chem. 1999;274:29352–29357. doi: 10.1074/jbc.274.41.29352. FGF9 lacks an amino-terminal signal sequence but is secreted from expressing cells. This study shows that nascent FGF9 polypeptides translocate into endoplasmic reticulum without peptide cleavage via a co-translational pathway in which both an amino terminus and a central hydrophobic domain is important. [DOI] [PubMed] [Google Scholar]
- Revest JM, DeMoerlooze L, Dickson C. Fibroblast growth factor 9 secretion is mediated by a non-cleaved amino-terminal signal sequence. J Biol Chem. 2000;275:8083–8090. doi: 10.1074/jbc.275.11.8083. FGF9 lacks a recognizable amino-terminal signal sequence, although it is efficiently secreted. This study shows that FGF9 enters the endoplasmic reticulum and traverses the Golgi complex in a similar manner to other constitutively secreted proteins. They demonstrate that the first 28 amino acids of FGF9 can function as an efficient non-cleaved signal peptide. [DOI] [PubMed] [Google Scholar]
- Smallwood PM, Munoz-sanjuan I, Tong P, Macke JP, Hendry SH, Gilbert DJ, Copeland NG, Jenkins NA, Nathans J. Fibroblast growth factor (FGF) homologous factors: new members of the FGF family implicated in nervous system development. Proc Natl Acad Sci USA. 1996;93:9850–9857. doi: 10.1073/pnas.93.18.9850. This study identifies a novel FGF-related subfamily, termed fibroblast growth factor homologous factors (FHF). FHFs lack a classical signal sequence and contain nuclear localization signals. FHFs are expressed in the developing and adult nervous systems, suggesting a role in nervous system development and function. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto S, Mikami T, Ohbayashi N, Ohta M, Itoh N. Structure and expression of a novel isoform of mouse FGF homologous factor (FHF)-4. Biochim Biophys Acta. 1998;1398:38–41. doi: 10.1016/s0167-4781(98)00050-5. Identification of alternative amino terminal coding exon for FHF4/FGF14. [DOI] [PubMed] [Google Scholar]
- Munoz-Sanjuan I, Smallwood PM, Nathans J. Isoform diversity among fibroblast growth factor homologous factors is generated by alternative promoter usage and differential splicing. J Biol Chem. 2000;275:2589–2597. doi: 10.1074/jbc.275.4.2589. Characterization of multiple isoforms of Fhfs1-4/Fgfs 11-14, generated through the use of alternative 5' exons. Isoforms show different subcellular distributions and distinct expression patterns in developing and adult mouse tissues. [DOI] [PubMed] [Google Scholar]
- Wang Q, McEwen DG, Ornitz DM. Subcellular and developmental expression of alternatively spliced forms of fibroblast growth factor 14. Mech Dev. 2000;90:283–287. doi: 10.1016/s0925-4773(99)00241-5. Identification of isoforms of Fhf4/Fgf14 that result from the alternative usage of two different first exons. Isoforms show different subcellular localization and distinct expression patterns in developing and adult mouse tissues. Expression was observed in migrating and post migratory neurons. [DOI] [PubMed] [Google Scholar]
- Antoine M, Reimers K, Dickson C, Kiefer P. Fibroblast growth factor 3, a protein with dual subcellular localization, is targeted to the nucleus and nucleolus by the concerted action of two nuclear localization signals and a nucleolar retention signal. J Biol Chem. 1997;272:29475–29481. doi: 10.1074/jbc.272.47.29475. Identification of an FGF3 CUG initiation site. This amino-terminal extended product is localized in both the nucleus/nucleolus and secretory pathway. The amino terminus contains both a signal sequence and a nuclear localization signal. [DOI] [PubMed] [Google Scholar]
- Hogan BL. Morphogenesis. Cell. 1999;96:225–233. doi: 10.1016/s0092-8674(00)80562-0. Review article highlighting the roles of multigene families, such as Fgfs, Bmps, Hedgehogs, Wnts and Egfs in morphogenesis. [DOI] [PubMed] [Google Scholar]
- Martin GR. The roles of FGFs in the early development of vertebrate limbs. Genes Dev. 1998;12:1571–1586. doi: 10.1101/gad.12.11.1571. Review article summarizing data on the role of FGFs in vertebrate limb development. Demonstrates that FGFs play essential roles in signaling centers that control establishment, outgrowth and patterning of the limb bud. [DOI] [PubMed] [Google Scholar]
- Lee PL, Johnson DE, Cousens LS, Fried VA, Williams LT. Purification and complementary DNA cloning of a receptor for basic fibroblast growth factor. Science. 1989;245:57–60. doi: 10.1126/science.2544996. This is the first cloning of an FGF receptor. [DOI] [PubMed] [Google Scholar]
- Johnson DE, Lee PL, Lu J, Williams LT. Diverse forms of a receptor for acidic and basic fibroblast growth factors. Mol Cell Biol. 1990;10:4728–4736. doi: 10.1128/mcb.10.9.4728. Identification of FGF receptor 1 variants which contain either two or three immunoglobulin-like domains. Identification of putative secreted forms of FGF receptor 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKeehan WL, Wang F, Kan M. The heparan sulfate-fibroblast growth factor family: diversity of structure and function. Prog Nucleic Acid Res Mol Biol. 1998;59:135–176. doi: 10.1016/s0079-6603(08)61031-4. This review presents models for FGF receptor activation by FGF and heparin. Also discussed are developmental roles for FGFs in liver and prostate development. [DOI] [PubMed] [Google Scholar]
- Miki T, Bottaro DP, Fleming TP, Smith CL, Burgess WH, Chan AM, Aaronson SA. Determination of ligand-binding specificity by alternative splicing: two distinct growth factor receptors encoded by a single gene. Proc Natl Acad Sci USA. 1992;89:246–250. doi: 10.1073/pnas.89.1.246. Demonstration that FGFR alternative splicing can regulate ligand binding specificity. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chellaiah AT, McEwen DG, Werner S, Xu J, Ornitz DM. Fibroblast growth factor receptor (FGFR) 3. Alternative splicing in immunoglobulin-like domain III creates a receptor highly specific for acidic FGF/FGF-1. J Biol Chem. 1994;269:11620–11627. Identification of an alternative splice form of FGFR3 with very restricted ligand binding properties. [PubMed] [Google Scholar]
- Naski MC, Ornitz DM. FGF signaling in skeletal development. Front Biosci. 1998;3:D781–D794. doi: 10.2741/a321. Review article highlighting the functions of the Fgfrs with a focus on roles in skeletal development. [DOI] [PubMed] [Google Scholar]
- Yan G, Fukabori Y, McBride G, Nikolaropolous S, McKeehan WL. Exon switching and activation of stromal and embryonic fibroblast growth factor (FGF)-FGF receptor genes in prostate epithelial cells accompany stromal independence and malignancy. Mol Cell Biol. 1993;13:4513–4522. doi: 10.1128/mcb.13.8.4513. Demonstration that tumor progression can be accompanied by a change in Fgfr alternative splicing with consequent change in ligand binding specificity. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orr-Urtreger A, Bedford MT, Burakova T, Arman E, Zimmer Y, Yayon A, Givol D, Lonai P. Developmental localization of the splicing alternatives of fibroblast growth factor receptor-2 (FGFR2). Dev Biol. 1993;158:475–486. doi: 10.1006/dbio.1993.1205. Demonstration that alternative Fgfr splice forms are expressed preferentially in epithelial and mesenchymal lineages. [DOI] [PubMed] [Google Scholar]
- Gilbert E, Del Gatto F, Champion-Arnaud P, Gesnel MC, Breathnach R. Control of BEK and K-SAM splice sites in alternative splicing of the fibroblast growth factor receptor 2 pre-mRNA. Mol Cell Biol. 1993;13:5461–5468. doi: 10.1128/mcb.13.9.5461. Identification of the mechanism regulating tissue-specific usage of alternative splice forms of Fgfr2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Avivi A, Yayon A, Givol D. A novel form of FGF receptor-3 using an alternative exon in the immunoglobulin domain III. FEBS Lett. 1993;330:249–252. doi: 10.1016/0014-5793(93)80882-u. Identification of alternative splice forms of Fgfr3. [DOI] [PubMed] [Google Scholar]
- Scotet E, Houssaint E. The choice between alternative IIIb and IIIc exons of the FGFR-3 gene is not strictly tissue-specific. Biochim Biophys Acta. 1995;1264:238–242. doi: 10.1016/0167-4781(95)00156-b. The less stringent alternative splicing choice between exons IIIb and IIIc of FGFR3 is in contrast with the very tissue-specific alternative splicing observed for FGFR2 in which epithelial cells use only the IIIb exon and fibroblasts use only the IIIc exon. [DOI] [PubMed] [Google Scholar]
- Ornitz DM, Xu J, Colvin JS, McEwen DG, MacArthur CA, Coulier F, Gao G, Goldfarb M. Receptor specificity of the fibroblast growth factor family. J Biol Chem. 1996;271:15292–15297. doi: 10.1074/jbc.271.25.15292. The first comparison of the relative mitogenic activity of a large number of FGFs on cells expressing a single alternatively spliced FGFR 1-3 or FGFR4. [DOI] [PubMed] [Google Scholar]
- Igarashi M, Finch PW, Aaronson SA. Characterization of recombinant human fibroblast growth factor (Fgf-10) reveals functional similarities with keratinocyte growth factor (Fgf-7). J Biol Chem. 1998;273:13230–13235. doi: 10.1074/jbc.273.21.13230. Demonstrates that the receptor-binding specificity of FGF10 is similar to that of the closely related FGF7. [DOI] [PubMed] [Google Scholar]
- Miki T, Fleming TP, Bottaro DP, Rubin JS, Ron D, Aaronson SA. Expression cDNA cloning of the KGF receptor by creation of a transforming autocrine loop. Science. 1991;251:72–75. doi: 10.1126/science.1846048. First cloning of a receptor for FGF7/KGF. [DOI] [PubMed] [Google Scholar]
- MacArthur CA, Lawshé A, Xu J, Santos-Ocampo S, Heikinheimo M, Chellaiah AT, Ornitz DM. FGF-8 isoforms activate receptor splice forms that are expressed in mesenchymal regions of mouse development. Development. 1995;121:3603–3613. doi: 10.1242/dev.121.11.3603. Identification of alternatively spliced forms of FGF8 and demonstration that these molecules bind the mesenchymal splice forms of Fgfr2. [DOI] [PubMed] [Google Scholar]
- Heikinheimo M, Lawshé A, Shackleford GM, Wilson DB, MacArthur CA. Fgf-8 expression in the post-gastrulation mouse suggests roles in the development of the face, limbs and central nervous system. Mech Dev. 1994;48:129–138. doi: 10.1016/0925-4773(94)90022-1. Identification of temporal and spatial patterns of Fgf8 expression in the developing mouse. [DOI] [PubMed] [Google Scholar]
- Crossley PH, Martin GR. The mouse Fgf8 gene encodes a family of polypeptides and is expressed in regions that direct outgrowth and patterning in the developing embryo. Development. 1995;121:439–451. doi: 10.1242/dev.121.2.439. Identification of alternative splice forms of Fgf8 and Fgf8 expression in the apical ectodermal ridge. [DOI] [PubMed] [Google Scholar]
- Moscatelli D. High and low affinity binding sites for basic fibroblast growth factor on cultured cells: absence of a role for low affinity binding in the stimulation of plasminogen activator production by bovine capillary endothelial cells. J Cell Physiol. 1987;131:123–130. doi: 10.1002/jcp.1041310118. This paper challenges the role of cell surface heparan sulfate in FGFR activation but establishes a role for cell surface heparan sulfate in binding FGF. [DOI] [PubMed] [Google Scholar]
- Flaumenhaft R, Moscatelli D, Rifkin DB. Heparin and heparan sulfate increase the radius of diffusion and action of basic fibroblast growth factor. J Cell Biol. 1990;111:1651–1659. doi: 10.1083/jcb.111.4.1651. This paper demonstrates that the extracellular matrix can store FGF and limit its diffusion in tissues. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mach H, Volkin DB, Burke CJ, Middaugh CR, Linhardt RJ, Fromm JR, Loganathan D, Mattsson L. Nature of the interaction of heparin with acidic fibroblast growth factor. Biochemistry. 1993;32:5480–5489. doi: 10.1021/bi00071a026. This paper establishes that FGF1 binds heparin at high density (one molecule every 4-5 saccharide units) and with high affinity (50-140 nM). [DOI] [PubMed] [Google Scholar]
- Herr AB, Ornitz DM, Sasisekharan R, Venkataraman G, Waksman G. Heparin-induced self-association of fibroblast growth factor-2. Evidence for two oligomerization processes. J Biol Chem. 1997;272:16382–16389. doi: 10.1074/jbc.272.26.16382. Presents analytical data that indicates that biologically active heparin octasaccharides can induce a monomer-dimer-tetramer assembly of FGF2. [DOI] [PubMed] [Google Scholar]
- Moy FJ, Safran M, Seddon AP, Kitchen D, Bohlen P, Aviezer D, Yayon A, Powers R. Properly oriented heparin-decasaccharide-induced dimers are the biologically active form of basic fibroblast growth factor. Biochemistry. 1997;36:4782–4791. doi: 10.1021/bi9625455. Evidence that a cis-oriented FGF2 dimer is the minimal biologically active structural unit of FGF2 in the presence of a decasaccharide. [DOI] [PubMed] [Google Scholar]
- Ornitz DM, Yayon A, Flanagan JG, Svahn CM, Levi E, Leder P. Heparin is required for cell-free binding of basic fibroblast growth factor to a soluble receptor and for mitogenesis in whole cells. Mol Cell Biol. 1992;12:240–247. doi: 10.1128/mcb.12.1.240. First study to show that heparin, FGF and an FGF receptor binding domain form a trimolecular complex in vitro. This paper also shows that this complex is essential for receptor activation on cells lacking heparan sulfate proteoglycan. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rapraeger AC, Krufka A, Olwin BB. Requirement of heparan sulfate for bFGF-mediated fibroblast growth and myoblast differentiation. Science. 1991;252:1705–1708. doi: 10.1126/science.1646484. The first study to demonstrate that cell surface heparan sulfate is directly involved in FGF cell signaling. [DOI] [PubMed] [Google Scholar]
- Yayon A, Klagsbrun M, Esko JD, Leder P, Ornitz DM. Cell surface, heparin-like molecules are required for binding of basic fibroblast growth factor to its high affinity receptor. Cell. 1991;64:841–848. doi: 10.1016/0092-8674(91)90512-w. The first study to demonstrate that heparin is required for FGF to bind to an FGF receptor. [DOI] [PubMed] [Google Scholar]
- Lin XH, Buff EM, Perrimon N, Michelson AM. Heparan sulfate proteoglycans are essential for FGF receptor signaling during Drosophila embryonic development. Development. 1999;126:3715–3723. doi: 10.1242/dev.126.17.3715. The first genetic evidence that heparan sulfate glycosaminoglycans are essential for fibroblast growth factor receptor signaling in a well defined developmental context. These data support the model in which heparan sulfate facilitates FGF and/or FGF-FGFR oligomerization. [DOI] [PubMed] [Google Scholar]
- Szebenyi G, Fallon JF. Fibroblast growth factors as multifunctional signaling factors. Int Rev Cytol. 1999;185:45–106. doi: 10.1016/s0074-7696(08)60149-7. This is a comprehensive review that highlights all aspects of FGF biology including cell signaling, receptor binding and activation and roles for FGFs in embryonic development. [DOI] [PubMed] [Google Scholar]
- Schlessinger J, Plotnikov AN, Ibrahimi OA, Eliseenkova AV, Yeh BK, Yayon A, Linhardt RJ, Mohammadi M. Crystal structure of a ternary FGF-FGFR-heparin complex reveals a dual role for heparin in FGFR binding and dimerization. Mol Cell. 2000;6:743–750. doi: 10.1016/s1097-2765(00)00073-3. This paper presents a diffusion-based crystal structure which shows heparan fragments bound to both FGF and the FGFR in a head-to-head orientation. [DOI] [PubMed] [Google Scholar]
- Chellaiah A, Yuan W, Chellaiah M, Ornitz DM. Mapping ligand binding domains in chimeric fibroblast growth factor receptor molecules. Multiple regions determine ligand binding specificity. J Biol Chem. 1999;274:34785–34794. doi: 10.1074/jbc.274.49.34785. Identification of a second ligand-binding region on the FGF receptor. [DOI] [PubMed] [Google Scholar]
- Meyers EN, Lewandoski M, Martin GR. An Fgf8 mutant allelic series generated by Cre- and Flp-mediated recombination. Nat Genet. 1998;18:136–141. doi: 10.1038/ng0298-136. First null, hypomorphic and conditional allele created for an FGF. This study reveals requirements for Fgf8 during gastrulation, cardiac, craniofacial, forebrain, midbrain and cerebellar development. [DOI] [PubMed] [Google Scholar]
- Moon AM, Capecchi MR. Fgf8 is required for outgrowth and patterning of the limbs. Nat Genet. 2000;26:455–459. doi: 10.1038/82601. Demonstrates an essential role for Fgf8 in limb bud development. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewandoski M, Sun X, Martin GR. Fgf8 signalling from the AER is essential for normal limb development. Nat Genet. 2000;26:460–463. doi: 10.1038/82609. Demonstrates an essential role for Fgf8 in limb bud development. [DOI] [PubMed] [Google Scholar]
- Reifers F, Bohli H, Walsh EC, Crossley PH, Stainier DY, Brand M. Fgf8 is mutated in zebrafish acerebellar (ace) mutants and is required for maintenance of midbrain-hindbrain boundary development and somitogenesis. Development. 1998;125:2381–2395. doi: 10.1242/dev.125.13.2381. Demonstrates a role for Fgf8 in midbrain hindbrain development in zebrafish by demonstrating genetic linkage between Fgf8 and the acerebellar (ace) mutation. Demonstrates that ace is probably a null mutation in Fgf8. [DOI] [PubMed] [Google Scholar]
- Heisenberg CP, Brennan C, Wilson SW. Zebrafish aussicht mutant embryos exhibit widespread overexpression of ace (fgf8) and coincident defects in CNS development. Development. 1999;126:2129–2140. doi: 10.1242/dev.126.10.2129. Identification of a genetic locus that may regulate Fgf8 (ace) expression in zebrafish. [DOI] [PubMed] [Google Scholar]
- Hebert JM, Rosenquist T, Gotz J, Martin GR. FGF5 as a regulator of the hair growth cycle: evidence from targeted and spontaneous mutations. Cell. 1994;78:1017–1025. doi: 10.1016/0092-8674(94)90276-3. Demonstrates that a null allele in the Fgf5 gene is allelic with the mouse angora mutation. Identifies an essential role for FGF5 in regulating the hair growth cycle. [DOI] [PubMed] [Google Scholar]
- Carlton MBL, Colledge WH, Evans MJ. Crouzon-like craniofacial dysmorphology in the mouse is caused by an insertional mutation at the Fgf3/Fgf4 locus. Dev Dyn. 1998;212:242–249. doi: 10.1002/(SICI)1097-0177(199806)212:2<242::AID-AJA8>3.3.CO;2-Z. Identification of a retroviral integration in the intragenic region between Fgf3 and Fgf4 in the Bulgy-eye (Bey) mutant mouse. Expression of both Fgf3 and Fgf4 is up-regulated in the cranial sutures of Bey mice. Phenocopies some features of Crouzon syndrome. [DOI] [PubMed] [Google Scholar]
- White KE, Evans WE, O'Riordan JL, Speer MC, Econs MJ, Lorenz-Depiereux B, Grabowski M, Meitinger T, Strom TM. Autosomal dominant hypophosphataemic rickets is associated with mutations in FGF23. Nat Genet. 2000;26:345–348. doi: 10.1038/81664. First identification of a missense mutation in an FGF ligand in a human genetic disease. Autosomal dominant hypophosphataemic rickets (ADHR) is characterized by low serum phosphorus concentrations, rickets, osteomalacia, lower extremity deformities, short stature, bone pain and dental abscesses. [DOI] [PubMed] [Google Scholar]
- White KE, Jonsson KB, Carn G, Hampson G, Spector TD, Mannstadt M, Lorenz-Depiereux B, Miyauchi A, Yang IM, Ljunggren O, et al. The autosomal dominant hypophosphatemic rickets (ADHR) gene is a secreted polypeptide overexpressed by tumors that cause phosphate wasting. J Clin Endocrinol Metab. 2001;86:497–500. doi: 10.1210/jcem.86.2.7408. FGF-23, the gene mutated in ADHR, is a secreted protein and its mRNA is abundantly expressed by several different oncogenic hypophosphatemic osteomalacia tumors. [DOI] [PubMed] [Google Scholar]
- Gecz J, Baker E, Donnelly A, Ming JE, McDonald-McGinn DM, Spinner NB, Zackai EH, Sutherland GR, Mulley JC. Fibroblast growth factor homologous factor 2 (FHF2): gene structure, expression and mapping to the Borjeson-Forssman-Lehmann syndrome region in Xq26 delineated by a duplication breakpoint in a BFLS-like patient. Hum Genet. 1999;104:56–63. doi: 10.1007/s004390050910. Identification of linkage between FGF12/FHF2 and Borjeson-Forssman-Lehmann syndrome. [DOI] [PubMed] [Google Scholar]
- Chang Z, Meyer K, Rapraeger AC, Friedl A. Differential ability of heparan sulfate proteoglycans to assemble the fibroblast growth factor receptor complex in situ. FASEB J. 2000;14:137–144. doi: 10.1096/fasebj.14.1.137. First paper to demonstrate that tissue-specific heparan sulfate molecules can affect patterns of FGF and FGF receptor binding in situ. [DOI] [PubMed] [Google Scholar]
- Madiai F, Hackshaw KV, Chiu IM. Characterization of the entire transcription unit of the mouse fibroblast growth factor 1 (FGF-1) gene. Tissue-specific expression of the FGF-1.A mRNA. J Biol Chem. 1999;274:11937–11944. doi: 10.1074/jbc.274.17.11937. This paper defines the entire transcriptional unit of the mouse Fgf1 gene and shows that heart is the most abundant source of Fgf1.A mRNA. [DOI] [PubMed] [Google Scholar]
- Abraham JA, Whang JL, Tumolo A, Mergia A, Friedman J, Gospodarowicz D, Fiddes JC. Human basic fibroblast growth factor: nucleotide sequence and genomic organization. EMBO J. 1986;5:2523–2528. doi: 10.1002/j.1460-2075.1986.tb04530.x. First cloned member of the FGF family. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brookes S, Smith R, Thurlow J, Dickson C, Peters G. The mouse homologue of hst/k-FGF: sequence, genome organization and location relative to int-2. Nucleic Acids Res. 1989;17:4037–4045. doi: 10.1093/nar/17.11.4037. This paper idenitifies the mouse Fgf4 gene and shows that it is within 20 kb of the Fgf3 gene on chromosome 7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colvin JS, Feldman B, Nadeau JH, Goldfarb M, Ornitz DM. Genomic organization and embryonic expression of the mouse fibroblast growth factor 9 gene. Dev Dyn. 1999;216:72–88. doi: 10.1002/(SICI)1097-0177(199909)216:1<72::AID-DVDY9>3.3.CO;2-0. Contains a detailed analysis of the embryonic expression patterns of Fgf9. [DOI] [PubMed] [Google Scholar]
- Chiu IM, Liu Y, Payson RA. Isolation of yeast artificial chromosomes containing the entire transcriptional unit of the human FGF1 gene: a 720-kb contig spanning human chromosome 5q31.3->q32. Cancer Genet Cytogenet. 1998;106:1–10. doi: 10.1016/s0165-4608(98)00031-4. Localization of the Fgf1 gene to the q31-q33 region of chromosome 5, a regions that includes a number of genes encoding growth factors, growth factor receptors, hormone/neurotransmitter receptors. [DOI] [PubMed] [Google Scholar]
- Cox RD, Copeland NG, Jenkins NA, Lehrach H. Interspersed repetitive element polymerase chain reaction product mapping using a mouse interspecific backcross. Genomics. 1991;10:375–384. doi: 10.1016/0888-7543(91)90322-6. A rapid method to generate and map inter-repeat polymerase chain reaction products using DNA from interspecific backcross mice. [DOI] [PubMed] [Google Scholar]
- Lafage-Pochitaloff M, Galland F, Simonetti J, Prats H, Mattei MG, Birnbaum D. The human basic fibroblast growth factor gene is located on the long arm of chromosome 4 at bands q26-q27. Oncogene Res. 1990;5:241–244. Chromosomal localization of human FGF2. [PubMed] [Google Scholar]
- Mattei MG, Pebusque MJ, Birnbaum D. Chromosomal localizations of mouse Fgf2 and Fgf5 genes. Mamm Genome. 1992;2:135–137. doi: 10.1007/BF00353862. Mapping of Fgf2 and Fgf5 on mouse chromosome 4. [DOI] [PubMed] [Google Scholar]
- Kim HS, Crow TJ. Human proto-oncogene Int-2/FGF-3 map position 11q13.3-q13.4. Chromosome Res. 1998;6:579. Chromosomal localization of human FGF3. [PubMed] [Google Scholar]
- Yoshida CM, Wada M, Satoh H, Yoshida T, Sakamoto H, Miyagawa K, Yokota J, Koda T, Kakinuma M, Sugimura T, et al. Human HST1 (HSTF1) gene maps to chromosome band 11q13 and coamplifies with the Int2 gene in human cancer. Proc Natl Acad Sci USA. 1988;85:4861–4864. doi: 10.1073/pnas.85.13.4861. Mapping of human FGF4 to chromosome 11q13. Both FGF3 and FGF4 were co-amplified in DNA from a stomach cancer and a vulvar carcinoma cell line. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peters G, Kozak C, Dickson C. Mouse mammary tumor virus integration regions int-1 and int-2 map on different mouse chromosomes. Mol Cell Biol. 1984;4:375–378. doi: 10.1128/mcb.4.2.375. Demonstration that MMTV can activate expression of FGF3 to initiate oncogenesis in mouse mammary epithelial cells. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peters G, Brookes S, Smith R, Placzek M, Dickson C. The mouse homolog of the hst/k-FGF gene is adjacent to int-2 and is activated by proviral insertion in some virally induced mammary tumors. Proc Natl Acad Sci USA. 1989;86:5678–5682. doi: 10.1073/pnas.86.15.5678. Identification of a close linkage between Fgf3 and Fgf4 on mouse chromosome 7. Demonstration that either or both Fgf3 and Fgf4 can be activated by mouse mammary tumor virus integration. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nguyen C, Roux D, Mattei MG, de Lapeyriere O, Goldfarb M, Birnbaum D, Jordan BR. The FGF-related oncogenes hst and int.2, and the bcl.1 locus are contained within one megabase in band q13 of chromosome 11, while the fgf.5 oncogene maps to 4q21. Oncogene. 1988;3:703–708. Identification of physical linkage between Fgf3 and Fgf4 on human chromosome 11. [PubMed] [Google Scholar]
- deLapeyriere O, Rosnet O, Benharroch D, Raybaud F, Marchetto S, Planche J, Galland F, Mattei MG, Copeland NG, Jenkins NA, et al. Structure, chromosome mapping and expression of the murine Fgf-6 gene. Oncogene. 1990;5:823–831. Localization of Fgf6 on mouse chromosome 6. [PubMed] [Google Scholar]
- Marics I, Adelaide J, Raybaud F, Mattei MG, Coulier F, Planche J, de Lapeyriere O, Birnbaum D. Characterization of the HST-related FGF.6 gene, a new member of the fibroblast growth factor gene family. Oncogene. 1989;4:335–340. Cloning of FGF6 based on sequence similarity with FGF4. FGF6 was mapped to chromosome 12p13. [PubMed] [Google Scholar]
- Zimonjic DB, Kelley MJ, Rubin JS, Aaronson SA, Popescu NC. Fluorescence in situ hybridization analysis of keratinocyte growth factor gene amplification and dispersion in evolution of great apes and humans. Proc Natl Acad Sci USA. 1997;94:11461–11465. doi: 10.1073/pnas.94.21.11461. Chromosomal localization of Fgf7 sequences in human and great ape genomes indicates that amplification and dispersion occurred in multiple discrete steps during evolution. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mattei MG, deLapeyriere O, Bresnick J, Dickson C, Birnbaum D, Mason I. Mouse Fgf7 (fibroblast growth factor 7) and Fgf8 (fibroblast growth factor 8) genes map to chromosomes 2 and 19 respectively. Mamm Genome. 1995;6:196–197. doi: 10.1007/BF00293012. Mouse Fgf7 and Fgf8 genes were mapped to chromosomes 2F-G and 19C3-D, respectively. [DOI] [PubMed] [Google Scholar]
- Payson RA, Wu J, Liu Y, Chiu IM. The human Fgf-8 gene localizes on chromosome 10q24 and is subjected to induction by androgen in breast cancer cells. Oncogene. 1996;13:47–53. Mapping human FGF8 to chromosome 1Oq24. [PubMed] [Google Scholar]
- Mattei MG, Penault-Llorca F, Coulier F, Birnbaum D. The human FGF9 gene maps to chromosomal region 13q11-q12. Genomics. 1995;29:811–812. doi: 10.1006/geno.1995.9926. Chromosomal localization of human FGF9 to 13q11-q12. [DOI] [PubMed] [Google Scholar]
- Mattei MG, De Moerlooze L, Lovec H, Coulier F, Birnbaum D, Dickson C. Mouse fgf9 (fibroblast growth factor 9) is localized on chromosome 14. Mamm Genome. 1997;8:617–618. doi: 10.1007/s003359900516. [DOI] [PubMed] [Google Scholar]
- Emoto H, Tagashira S, Mattei MG, Yamasaki M, Hashimoto G, Katsumata T, Negoro T, Nakatsuka M, Birnbaum D, Coulier F, et al. Structure and expression of human fibroblast growth factor-10. J Biol Chem. 1997;272:23191–23194. doi: 10.1074/jbc.272.37.23191. Cloning and mapping of human FGF10 to 5p12-p13. Demonstration that FGF10 is similar to FGF7 in both sequence and biological activity. [DOI] [PubMed] [Google Scholar]
- Crackower MA, Heng HH, Tsui LC. Assignment of mouse fibroblast growth factor 10 (Fgf10) gene to the telomeric region of chromosome 13. Genomics. 1998;53:247–248. doi: 10.1006/geno.1998.5506. Mapping Fgf10 to mouse chromosome 13. [DOI] [PubMed] [Google Scholar]
- Verdier AS, Mattei MG, Lovec H, Hartung H, Goldfarb M, Birnbaum D, Coulier F. Chromosomal mapping of two novel human FGF genes, FGF11 and FGF12. Genomics. 1997;40:151–154. doi: 10.1006/geno.1996.4492. Mapping human FGF11 and FGF12 to chromosome 17p12-p13 and 3q28, respectively. [DOI] [PubMed] [Google Scholar]
- Hartung H, Lovec H, Verdier AS, Mattei MG, Coulier F, Goldfarb M, Birnbaum D. Assignment of Fgf12 to mouse chromosome bands 16B1->B3 by in situ hybridization. Cytogenet Cell Genet. 1997;76:185–186. doi: 10.1159/000134544. Mapping mouse Fgf12 to chromosome 13B1-3. [DOI] [PubMed] [Google Scholar]
- Liu Y, Chiu IM. Assignment of Fgf12, the human Fgf homologous factor 1 gene, to chromosome 3q29->3qter by fluorescence in situ hybridization. Cytogenet Cell Genet. 1997;78:48–49. doi: 10.1159/000134625. Mapping human FGF12 to chromosome 3q29-3qter. [DOI] [PubMed] [Google Scholar]
- Lovec H, Hartung H, Verdier AS, Mattei MG, Birnbaum D, Goldfarb M, Coulier F. Assignment of FGF13 to human chromosome band Xq21 by in situ hybridization. Cytogenet Cell Genet. 1997;76:183–184. doi: 10.1159/000134543. Mapping FGF13 to human chromosome Xq21. [DOI] [PubMed] [Google Scholar]
- Xu J, Lawshe A, MacArthur CA, Ornitz DM. Genomic structure, mapping, activity and expression of fibroblast growth factor 17. Mech Dev. 1999;83:165–78. doi: 10.1016/s0925-4773(99)00034-9. Expression patterns of Fgf17 and Fgf8 and demonstration that Fgf17 has similar receptor specificity, genomic organization and alternative splicing to Fgf8. [DOI] [PubMed] [Google Scholar]
- Whitmore TE, Maurer MF, Sexson S, Raymond F, Conklin D, Deisher TA. Assignment of fibroblast growth factor 18 (FGF18) to human chromosome 5q34 by use of radiation hybrid mapping and fluorescence in situ hybridization. Cytogenet Cell Genet. 2000;90:231–233. doi: 10.1159/000056775. Mapping FGF18 to human chromosome 5q34. [DOI] [PubMed] [Google Scholar]
- Xie MH, Holcomb I, Deuel B, Dowd P, Huang A, Vagts A, Foster J, Liang J, Brush J, Gu Q, et al. FGF-19, a novel fibroblast growth factor with unique specificity for FGFR4. Cytokine. 1999;11:729–735. doi: 10.1006/cyto.1999.0485. Identification of human Fgf19 and demonstration of in vitro biological activity. [DOI] [PubMed] [Google Scholar]
- Kirikoshi H, Sagara N, Saitoh T, Tanaka K, Sekihara H, Shiokawa K, Katoh M. Molecular cloning and characterization of human FGF-20 on chromosome 8p21.3-p22. Biochem Biophys Res Commun. 2000;274:337–343. doi: 10.1006/bbrc.2000.3142. Cloning of human FGF20 and mapping to chromosome 8p21.3-p22. FGF20 is most closely related to FGF9 and FGF16. FGF20 mRNA was detected in a colon cancer cell line and at lower levels in human fetal tissues and primary tumors. [DOI] [PubMed] [Google Scholar]
- Nishimura T, Nakatake Y, Konishi M, Itoh N. Identification of a novel FGF, FGF-21, preferentially expressed in the liver. Biochim Biophys Acta. 2000;1492:203–206. doi: 10.1016/s0167-4781(00)00067-1. Cloning of Fgf21 an Fgf most closely related to Fgf19 and Fgf23. [DOI] [PubMed] [Google Scholar]
- Nakatake Y, Hoshikawa M, Asaki T, Kassai Y, Itoh N. Identification of a novel fibroblast growth factor, FGF-22, preferentially expressed in the inner root sheath of the hair follicle. Biochim Biophys Acta. 2001;1517:460–463. doi: 10.1016/s0167-4781(00)00302-x. Identification of Fgf22, a homolog of Fgf7 and Fgf10. [DOI] [PubMed] [Google Scholar]
- Miller DL, Ortega S, Bashayan O, Basch R, Basilico C. Compensation by fibroblast growth factor 1 (FGF1) does not account for the mild phenotypic defects observed in FGF2 null mice. Mol Cell Biol. 2000;20:2260–2268. doi: 10.1128/mcb.20.6.2260-2268.2000. The relatively mild phenotypic defects associated with loss of FGF2 led to the hypothesis that other FGFs partially compensate. However, FGF1-FGF2 double-knockout mice are viable and fertile and do not display any gross phenotypic defects.Thus Fgf1 null mice do not have a detectable phenotype and lack redundancy with Fgf2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tobe T, Ortega S, Luna JD, Ozaki H, Okamoto N, Derevjanik NL, Vinores SA, Basilico C, Campochiaro PA. Targeted disruption of the FGF2 gene does not prevent choroidal neovascularization in a murine model. Am J Pathol. 1998;153:1641–1646. doi: 10.1016/S0002-9440(10)65753-7. This study demonstrates that FGF2 is not required for the development of choroidal neovascularization. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ortega S, Ittmann M, Tsang SH, Ehrlich M, Basilico C. Neuronal defects and delayed wound healing in mice lacking fibroblast growth factor 2. Proc Natl Acad Sci, USA. 1998;95:5672–5677. doi: 10.1073/pnas.95.10.5672. Mice lacking Fgf2 are viable, fertile and phenotypically indistinguishable from wild type. Although not essential for embryonic development, loss of FGF2 leads to defects in cortical neurogenesis and skin wound healing in mice. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou M, Sutliff RL, Paul RJ, Lorenz JN, Hoying JB, Haudenschild CC, Yin M, Coffin JD, Kong L, Kranias EG, et al. Fibroblast growth factor 2 control of vascular tone. Nat Med. 1998;4:201–207. doi: 10.1038/nm0298-201. Mice lacking Fgf2 are morphologically normal but display decreased vascular smooth muscle contractility, low blood pressure and thrombocytosis. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultz JE, Witt SA, Nieman ML, Reiser PJ, Engle SJ, Zhou M, Pawlowski SA, Lorenz JN, Kimball TR, Doetschman T. Fibroblast growth factor-2 mediates pressure-induced hypertrophic response. J Clin Invest. 1999;104:709–719. doi: 10.1172/JCI7315. Identification of a role for FGF2 in the pathogenesis of cardiac hypertrophy. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mansour SL. Targeted disruption of int-2 (fgf-3) causes developmental defects in the tail and inner ear. Mol Reprod Dev. 1994;39:62–67. doi: 10.1002/mrd.1080390111. The first member of the Fgf family to be knocked out in mice. Developmental defects were discovered in the inner ear and in outgrowth of the tail. [DOI] [PubMed] [Google Scholar]
- Feldman B, Poueymirou W, Papaioannou VE, DeChiara TM, Goldfarb M. Requirement of FGF-4 for postimplantation mouse development. Science. 1995;267:246–249. doi: 10.1126/science.7809630. Fgf4 null embryos implant but die shortly thereafter. Fgf4 null embryos cultured in vitro displayed severely impaired proliferation of the inner cell mass. [DOI] [PubMed] [Google Scholar]
- Fiore F, Sebille A, Birnbaum D. Skeletal muscle regeneration is not impaired in Fgf6-/- mutant mice. Biochem Biophys Res Commun. 2000;272:138–143. doi: 10.1006/bbrc.2000.2703. Mice lacking Fgf6 have normal muscle regeneration. This is in contrast to the relatively severe phenotype noted by Floss et al. [118]. [DOI] [PubMed] [Google Scholar]
- Floss T, Arnold HH, Braun T. A role for Fgf-6 in skeletal muscle regeneration. Genes Dev. 1997;11:2040–2051. doi: 10.1101/gad.11.16.2040. Mice lacking Fgf6 show a severe muscle regeneration defect with fibrosis and myotube degeneration. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fiore F, Planche J, Gibier P, Sebille A, Delapeyriere O, Birnbaum D. Apparent normal phenotype of Fgf6-/- mice. Int J Dev Biol. 1997;41:639–642. Mice lacking Fgf6 are phenotypically normal. [PubMed] [Google Scholar]
- Guo L, Degenstein L, Fuchs E. Keratinocyte growth factor is required for hair development but not for wound healing. Genes Dev. 1996;10:165–175. doi: 10.1101/gad.10.2.165. Mice lacking Fgf7 have a unexpectedly subtle phenotype affecting hair growth. [DOI] [PubMed] [Google Scholar]
- Qiao J, Uzzo R, Obara-Ishihara T, Degenstein L, Fuchs E, Herzlinger D. FGF-7 modulates ureteric bud growth and nephron number in the developing kidney. Development. 1999;126:547–554. doi: 10.1242/dev.126.3.547. Identification of a second phenotype in mice lacking Fgf7. The developing ureteric bud and mature collecting system of Fgf7-null kidneys is markedly smaller than wild type and have 30% fewer nephrons than wild-type kidneys. [DOI] [PubMed] [Google Scholar]
- Sun X, Meyers EN, Lewandoski M, Martin GR. Targeted disruption of Fgf8 causes failure of cell migration in the gastrulating mouse embryo. Genes Dev. 1999;13:1834–1846. doi: 10.1101/gad.13.14.1834. This study identifies Fgf8 as a gene essential for gastrulation and shows that signaling via FGF8 and/or FGF4 is required for cell migration away from the primitive streak. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shanmugalingam S, Houart C, Picker A, Reifers F, Macdonald R, Barth A, Griffin K, Brand M, Wilson SW. Ace/Fgf8 is required for forebrain commissure formation and patterning of the telencephalon. Development. 2000;127:2549–2561. doi: 10.1242/dev.127.12.2549. Demonstration that Fgf8/Ace is required for the development of midline structures in the zebrafish forebrain. [DOI] [PubMed] [Google Scholar]
- Colvin JS, Green RP, Schmahl J, Capel B, Ornitz DM. Male to female sex reversal in mice lacking fibroblast growth factor 9. Cell. 2001. This study demonstrates that FGF9 is essential for testicular mesenchymal growth, cord formation, and Sertoli and Leydig cell differentiation. FGF9 may also regulate mesonephric cell migration into the male gonad. [DOI] [PubMed]
- Ohuchi H, Hori Y, Yamasaki M, Harada H, Sekine K, Kato S, Itoh N. FGF10 acts as a major ligand for FGF receptor 2 IIIb in mouse multi-organ development. Biochem Biophys Res Commun. 2000;277:643–649. doi: 10.1006/bbrc.2000.3721. Diverse phenotypes of mice lacking Fgf10 closely resemble those of mice lacking FGFR2b. Major phenotypes include the absence of thyroid, pituitary, and salivary glands in addition to the previously described absence of limbs and lungs. Minor defects were observed in the formation of the teeth, kidneys, hair follicles, and digestive organs. [DOI] [PubMed] [Google Scholar]
- Min H, Danilenko DM, Scully SA, Bolon B, Ring BD, Tarpley JE, DeRose M, Simonet WS. Fgf-10 is required for both limb and lung development and exhibits striking functional similarity to Drosophila branchless. Genes Dev. 1998;12:3156–3161. doi: 10.1101/gad.12.20.3156. Mice lacking Fgf10 exhibited perinatal lethality associated with complete absence of lungs. Fgf10 null mice also lack limbs. There is some disagreement with Sekine et al. [127] as to whether the apical ectodermal ridge transiently forms. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sekine K, Ohuchi H, Fujiwara M, Yamasaki M, Yoshizawa T, Sato T, Yagishita N, Matsui D, Koga Y, Itoh N, Kato S. Fgf10 is essential for limb and lung formation. Nat Genet. 1999;21:138–141. doi: 10.1038/5096. Mice lacking Fgf10 die at birth due to the lack of lung development beyond the trachea. Mice lacking Fgf10 also had complete truncation of the forelmbs and hindlimbs. In Fgf10 null embryos, limb bud formation was initiated but outgrowth of the limb buds did not occur. [DOI] [PubMed] [Google Scholar]


