Abstract
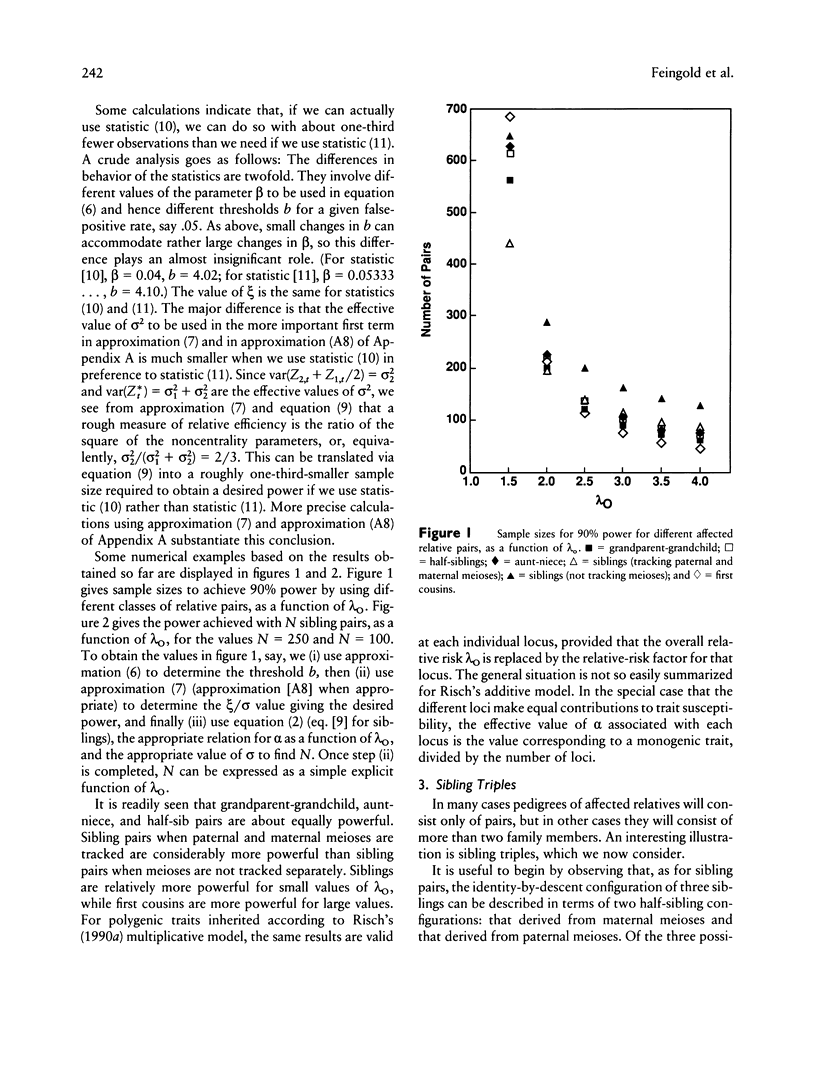
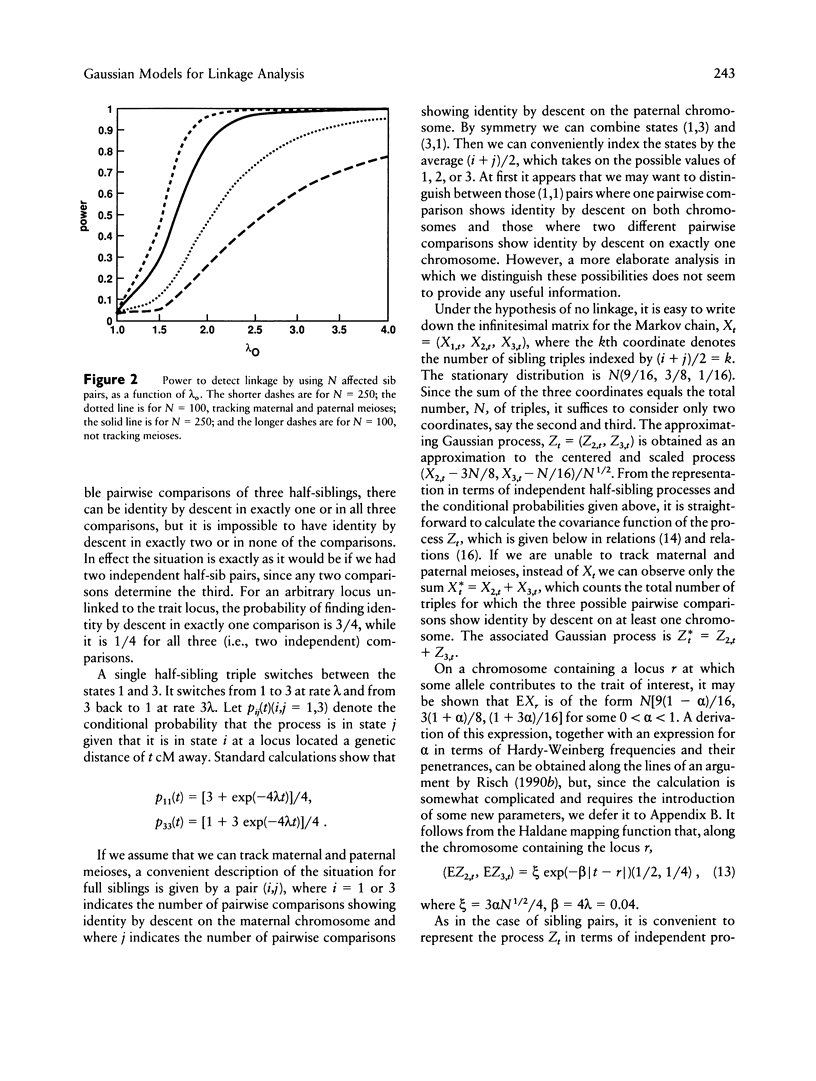
Gaussian-process models are developed to detect genetic linkage using complete high-resolution maps of identity by descent between affected relative pairs. Approximations are given for the significance level and power of the likelihood-ratio test of no linkage and for likelihood-ratio confidence regions for trait loci. The sample sizes required to detect linkage by using different classes of affected relative pairs are compared, and the problem of combining data from different classes of relatives is discussed.
Full text
PDF















Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Botstein D., White R. L., Skolnick M., Davis R. W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet. 1980 May;32(3):314–331. [PMC free article] [PubMed] [Google Scholar]
- James J. W. Frequency in relatives for an all-or-none trait. Ann Hum Genet. 1971 Jul;35(1):47–49. doi: 10.1111/j.1469-1809.1956.tb01377.x. [DOI] [PubMed] [Google Scholar]
- Risch N. Linkage strategies for genetically complex traits. I. Multilocus models. Am J Hum Genet. 1990 Feb;46(2):222–228. [PMC free article] [PubMed] [Google Scholar]



