Abstract
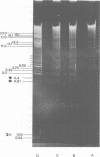
It has been shown earlier that the reiterated regions TRS and IRS bracketing the Us segment of herpes simplex virus type 1 Angelotti DNA are heterogeneous in size by stepwise insertion of one to six copies of a 550-base-pair nucleotide sequence. Considerably higher amplification of this sequence was observed in defective viral DNA: up to 14 copies were detected to be inserted in the repeat units of a major class of defective herpes simplex virus type 1 Angelotti DNA, dDNA1, which originated from noncontiguous sites located in UL and the inverted repeats of the S component of the parental genome. Physical maps were established for the cleavage sites of KpnI, PstI, XhoI, and BamHI restriction endonucleases on the repeats of dDNA1. The map position of the insertion sequence was determined. It was demonstrated that the amplified inserts were not distributed at random among or within the repeats. A given total population of dDNA1 molecules consisted of different homopolymers, each of which contained a constant number of inserts in all of its repeats. Assuming that a rolling-circle mechanism is involved in the generation of full-length defective herpes simplex virus type 1 Angelotti DNA from single repeat units, these data suggest that the 550-base-pair sequence is amplified in the repeats before the replication process.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ben-Porat T., Demarchi J. M., Kaplan A. S. Characterization of defective interfering viral particles present in a population of pseudorabies virions. Virology. 1974 Sep;61(1):29–37. doi: 10.1016/0042-6822(74)90239-6. [DOI] [PubMed] [Google Scholar]
- Bronson D. L., Dreesman G. R., Biswal N., Benyesh-Melnick M. Defective virions of herpes simplex viruses. Intervirology. 1973;1(3):141–153. doi: 10.1159/000148841. [DOI] [PubMed] [Google Scholar]
- Campbell D. E., Kemp M. C., Perdue M. L., Randall C. C., Gentry G. A. Equine herpesvirus in vivo: cyclic production of a DNA density variant with repetitive sequences. Virology. 1976 Feb;69(2):737–750. doi: 10.1016/0042-6822(76)90502-x. [DOI] [PubMed] [Google Scholar]
- Fleckenstein B., Bornkamm G. W., Ludwig H. Repetitive sequences in complete and defective genomes of Herpesvirus saimiri. J Virol. 1975 Feb;15(2):398–406. doi: 10.1128/jvi.15.2.398-406.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frenkel N., Jacob R. J., Honess R. W., Hayward G. S., Locker H., Roizman B. Anatomy of herpes simplex virus DNA. III. Characterization of defective DNA molecules and biological properties of virus populations containing them. J Virol. 1975 Jul;16(1):153–167. doi: 10.1128/jvi.16.1.153-167.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frenkeĺ N., Locker H., Batterson W., Hayward G. S., Roizman B. Anatomy of herpes simplex virus DNA. VI. Defective DNA originates from the S component. J Virol. 1976 Nov;20(2):527–531. doi: 10.1128/jvi.20.2.527-531.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graham B. J., Bengali Z., Vande Woude G. F. Physical map of the origin of defective DNA in herpes simplex virus type 1 DNA. J Virol. 1978 Mar;25(3):878–887. doi: 10.1128/jvi.25.3.878-887.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacob R. J., Morse L. S., Roizman B. Anatomy of herpes simplex virus DNA. XII. Accumulation of head-to-tail concatemers in nuclei of infected cells and their role in the generation of the four isomeric arrangements of viral DNA. J Virol. 1979 Feb;29(2):448–457. doi: 10.1128/jvi.29.2.448-457.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaerner H. C., Maichle I. B., Ott A., Schröder C. H. Origin of two different classes of defective HSV-1 Angelotti DNA. Nucleic Acids Res. 1979 Apr;6(4):1467–1478. doi: 10.1093/nar/6.4.1467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee A. S., Sinsheimer R. L. A cleavage map of bacteriophage phiX174 genome. Proc Natl Acad Sci U S A. 1974 Jul;71(7):2882–2886. doi: 10.1073/pnas.71.7.2882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Locker H., Frenkel N. BamI, KpnI, and SalI restriction enzyme maps of the DNAs of herpes simplex virus strains Justin and F: occurrence of heterogeneities in defined regions of the viral DNA. J Virol. 1979 Nov;32(2):429–441. doi: 10.1128/jvi.32.2.429-441.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Locker H., Frenkel N. Structure and origin of defective genomes contained in serially passaged herpes simplex virus type 1 (Justin). J Virol. 1979 Mar;29(3):1065–1077. doi: 10.1128/jvi.29.3.1065-1077.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murray B. K., Biswal N., Bookout J. B., Lanford R. E., Courtney R. J., Melnick J. L. Cyclic appearance of defective interfering particles of herpes simplex virus and the concomitant accumulation of early polypeptide VP175. Intervirology. 1975;5(3-4):173–184. doi: 10.1159/000149894. [DOI] [PubMed] [Google Scholar]
- Ott A., Föhring B., Kaerner H. C. Herpes simplex virus type 1 Angelotti and a defective viral genotype: analysis of genome structures and genetic relatedness by DNA-DNA reassociation kinetics. J Virol. 1979 Feb;29(2):423–430. doi: 10.1128/jvi.29.2.423-430.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reddy V. B., Thimmappaya B., Dhar R., Subramanian K. N., Zain B. S., Pan J., Ghosh P. K., Celma M. L., Weissman S. M. The genome of simian virus 40. Science. 1978 May 5;200(4341):494–502. doi: 10.1126/science.205947. [DOI] [PubMed] [Google Scholar]
- Roizman B. The structure and isomerization of herpes simplex virus genomes. Cell. 1979 Mar;16(3):481–494. doi: 10.1016/0092-8674(79)90023-0. [DOI] [PubMed] [Google Scholar]
- Schröder C. H., Stegmann B., Lauppe H. F., Kaerner H. C. An unusual defective genotype derived from herpes simplex virus strain ANG. Intervirology. 1975;6(4-5):270–284. doi: 10.1159/000149481. [DOI] [PubMed] [Google Scholar]
- Stegmann B., Zentgraf H., Ott A., Schröder C. H. Synthesis and packaging of herpes simplex virus DNA in the course of virus passages at high multiplicity. Intervirology. 1978;10(4):228–240. doi: 10.1159/000148986. [DOI] [PubMed] [Google Scholar]
- Thomas M., Davis R. W. Studies on the cleavage of bacteriophage lambda DNA with EcoRI Restriction endonuclease. J Mol Biol. 1975 Jan 25;91(3):315–328. doi: 10.1016/0022-2836(75)90383-6. [DOI] [PubMed] [Google Scholar]
- Wagner M. J., Summers W. C. Structure of the joint region and the termini of the DNA of herpes simplex virus type 1. J Virol. 1978 Aug;27(2):374–387. doi: 10.1128/jvi.27.2.374-387.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wagner M., Skare J., Summers W. C. Analysis of DNA of defective herpes simplex virus type 1 by restriction endonuclease cleavage and nucleic acid hybridization. Cold Spring Harb Symp Quant Biol. 1975;39(Pt 2):683–686. doi: 10.1101/sqb.1974.039.01.082. [DOI] [PubMed] [Google Scholar]