Abstract
The Bacillus subtilis nprE gene lacking its own promoter sequence was inserted in the lactococcal expression vector pMG36e. Upon introduction of the recombinant plasmid into Lactococcus lactis subsp. lactis strain MG1363, neutral protease activity could be visualized by the appearance of large clearing zones around colonies grown on milk agar plates. By measuring the activities of the neutral protease and the intracellular enzyme lactate dehydrogenase in culture supernatants and cell fractions, it was demonstrated that the neutral protease was actively secreted into the growth medium. This was corroborated by using the Western blot (immunoblot) technique, which showed the presence of the mature form of the neutral protease in the culture supernatant. On the basis of these results, it is concluded that the B. subtilis neutral protease gene was expressed in L. lactis and that the gene product was secreted into the growth medium and was apparently correctly processed to produced a biologically active protein. The secretion of this particular enzyme may be helpful in achieving accelerated cheese ripening.
Full text
PDF

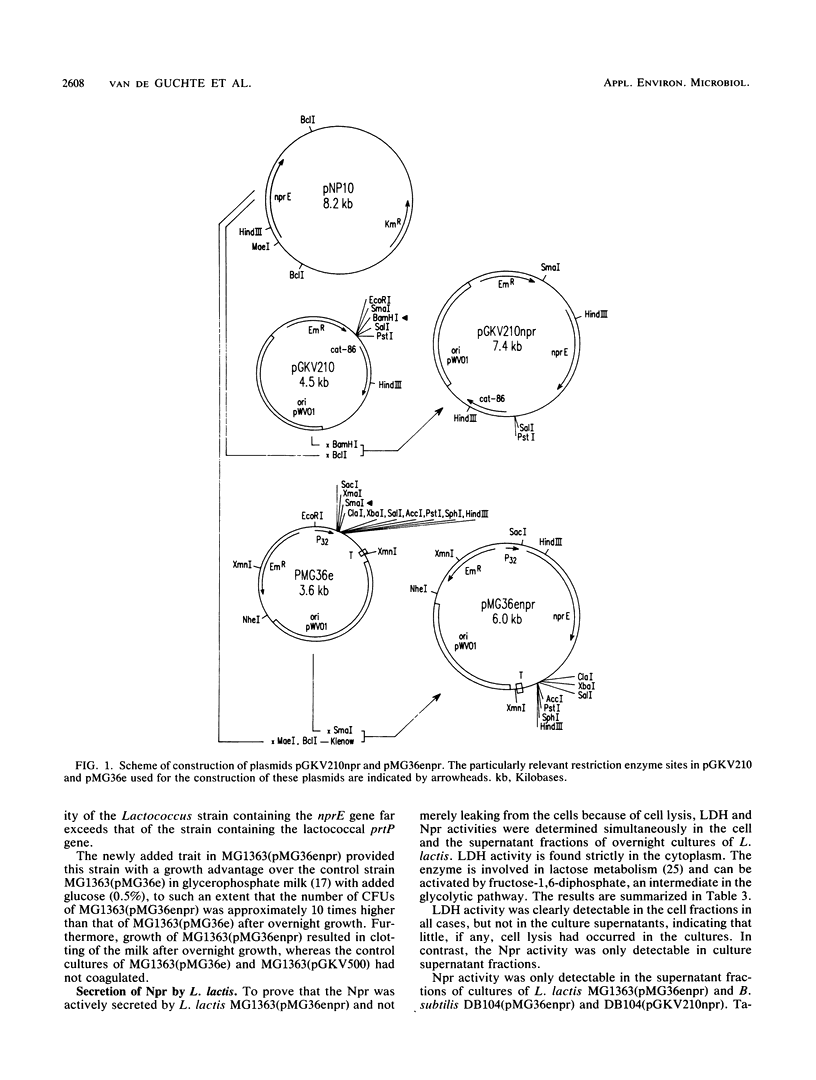



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1006/abio.1976.9999. [DOI] [PubMed] [Google Scholar]
- Chang S., Cohen S. N. High frequency transformation of Bacillus subtilis protoplasts by plasmid DNA. Mol Gen Genet. 1979 Jan 5;168(1):111–115. doi: 10.1007/BF00267940. [DOI] [PubMed] [Google Scholar]
- Gaur N. K., Dubnau E., Smith I. Characterization of a cloned Bacillus subtilis gene that inhibits sporulation in multiple copies. J Bacteriol. 1986 Nov;168(2):860–869. doi: 10.1128/jb.168.2.860-869.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haandrikman A. J., Kok J., Laan H., Soemitro S., Ledeboer A. M., Konings W. N., Venema G. Identification of a gene required for maturation of an extracellular lactococcal serine proteinase. J Bacteriol. 1989 May;171(5):2789–2794. doi: 10.1128/jb.171.5.2789-2794.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kok J., van Dijl J. M., van der Vossen J. M., Venema G. Cloning and expression of a Streptococcus cremoris proteinase in Bacillus subtilis and Streptococcus lactis. Appl Environ Microbiol. 1985 Jul;50(1):94–101. doi: 10.1128/aem.50.1.94-101.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagami Y., Tanaka T. Molecular cloning and nucleotide sequence of a DNA fragment from Bacillus natto that enhances production of extracellular proteases and levansucrase in Bacillus subtilis. J Bacteriol. 1986 Apr;166(1):20–28. doi: 10.1128/jb.166.1.20-28.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rottländer E., Trautner T. A. Genetic and transfection studies with B, subtilis phage SP 50. I. Phage mutants with restricted growth on B. subtilis strain 168. Mol Gen Genet. 1970;108(1):47–60. doi: 10.1007/BF00343184. [DOI] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. DNA sequence analysis with a modified bacteriophage T7 DNA polymerase. Proc Natl Acad Sci U S A. 1987 Jul;84(14):4767–4771. doi: 10.1073/pnas.84.14.4767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka T., Kawata M. Cloning and characterization of Bacillus subtilis iep, which has positive and negative effects on production of extracellular proteases. J Bacteriol. 1988 Aug;170(8):3593–3600. doi: 10.1128/jb.170.8.3593-3600.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terzaghi B. E., Sandine W. E. Improved medium for lactic streptococci and their bacteriophages. Appl Microbiol. 1975 Jun;29(6):807–813. doi: 10.1128/am.29.6.807-813.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas T. D. Regulation of lactose fermentation in group N streptococci. Appl Environ Microbiol. 1976 Oct;32(4):474–478. doi: 10.1128/aem.32.4.474-478.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas T. D., Turner K. W., Crow V. L. Galactose fermentation by Streptococcus lactis and Streptococcus cremoris: pathways, products, and regulation. J Bacteriol. 1980 Nov;144(2):672–682. doi: 10.1128/jb.144.2.672-682.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toma S., Campagnoli S., De Gregoriis E., Gianna R., Margarit I., Zamai M., Grandi G. Effect of Glu-143 and His-231 substitutions on the catalytic activity and secretion of Bacillus subtilis neutral protease. Protein Eng. 1989 Jan;2(5):359–364. doi: 10.1093/protein/2.5.359. [DOI] [PubMed] [Google Scholar]
- Toma S., Del Bue M., Pirola A., Grandi G. nprR1 and nprR2 regulatory regions for neutral protease expression in Bacillus subtilis. J Bacteriol. 1986 Aug;167(2):740–743. doi: 10.1128/jb.167.2.740-743.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Den Burg B., Eijsink V. G., Stulp B. K., Venema G. One-step affinity purification of Bacillus neutral proteases using bacitracin-silica. J Biochem Biophys Methods. 1989 May;18(3):209–219. doi: 10.1016/0165-022x(89)90005-5. [DOI] [PubMed] [Google Scholar]
- Yang M. Y., Ferrari E., Henner D. J. Cloning of the neutral protease gene of Bacillus subtilis and the use of the cloned gene to create an in vitro-derived deletion mutation. J Bacteriol. 1984 Oct;160(1):15–21. doi: 10.1128/jb.160.1.15-21.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang M., Ferrari E., Chen E., Henner D. J. Identification of the pleiotropic sacQ gene of Bacillus subtilis. J Bacteriol. 1986 Apr;166(1):113–119. doi: 10.1128/jb.166.1.113-119.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van de Guchte M., van der Vossen J. M., Kok J., Venema G. Construction of a lactococcal expression vector: expression of hen egg white lysozyme in Lactococcus lactis subsp. lactis. Appl Environ Microbiol. 1989 Jan;55(1):224–228. doi: 10.1128/aem.55.1.224-228.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Lelie D., van der Vossen J. M., Venema G. Effect of Plasmid Incompatibility on DNA Transfer to Streptococcus cremoris. Appl Environ Microbiol. 1988 Apr;54(4):865–871. doi: 10.1128/aem.54.4.865-871.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Vossen J. M., Kok J., Venema G. Construction of cloning, promoter-screening, and terminator-screening shuttle vectors for Bacillus subtilis and Streptococcus lactis. Appl Environ Microbiol. 1985 Aug;50(2):540–542. doi: 10.1128/aem.50.2.540-542.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Vossen J. M., van der Lelie D., Venema G. Isolation and characterization of Streptococcus cremoris Wg2-specific promoters. Appl Environ Microbiol. 1987 Oct;53(10):2452–2457. doi: 10.1128/aem.53.10.2452-2457.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]




