Abstract
Three new photoreactive tRNA derivatives have been synthesized for use as probes of the peptidyl transferase center of the ribosome. In two of these derivatives, the 3′ adenosine of yeast tRNAPhe has been replaced by either 2-azidodeoxyadenosine or 2-azido-2′-O-methyl adenosine, while in a third the 3′-terminal 2-azidodeoxyadenosine of the tRNA is joined to puromycin via a phosphoramidate linkage to generate a photoreactive transition-state analog. All three derivatives bind to the P site of 70S ribosomes with affinities similar to that of unmodified tRNAPhe and can be cross-linked to components of the 50S ribosomal subunit by irradiation with near-UV light. Characteristic differences in the cross-linking patterns suggest that these tRNA derivatives can be used to follow subtle changes in the position of the tRNA relative to the components of the peptidyl transferase center.
Keywords: peptidyl transferase center, photoreactive tRNA probes, ribosomes
INTRODUCTION
Dynamic interactions between the amino acid acceptor end of tRNA and the ribosome underlie the sequential synthesis of peptide bonds at the peptidyl transferase center of the 50S ribosomal subunit (Schmeing et al. 2005b). Photo-cross-linking of the 3′-terminal residue of tRNA—which is adjacent to the attached amino acid or peptide—to components of the 50S subunit has proven to be a sensitive means for identifying specific protein and RNA segments at the peptidyl transferase center. At the ribosomal P site, a derivative of yeast tRNAPhe with the photoreactive nucleoside 2-azidoadenosine in position 76, (2N3A76)tRNAPhe, cross-links to nucleotides U2506 and U2585 of the 23S rRNA as well as to the N-terminal sequence of 50S-subunit protein L27 (Wower et al. 2000; Maguire et al. 2005), indicating their close proximity to the site of peptide bond formation. The role of these components in transpeptidation is also supported by numerous genetic, footprinting, modification-interference, and structural studies (Moazed and Noller 1989; Porse et al. 1996; Bocchetta et al. 1998; Maguire et al. 2005; Schmeing et al. 2005b). To further probe the peptidyl transferase center (PTC), we have developed three new photoreactive derivatives of yeast tRNAPhe with modifications at position 76, one containing 2-azidodeoxyadenosine, (2N3dA76)tRNAPhe, a second containing 2-azido-2′-O-methyladenosine (2N32′OMeA76)tRNAPhe, and a third, (2N3dA76)tRNAPhe-p-Puro, based on a transition-state analog of peptidyl transferase designed by Yarus and coworkers (Welch et al. 1995).
The Yarus analog consists of a P-site moiety, CCdAp, joined to an A-site moiety, puromycin, via a phosphoramidate linkage between the 3′-phosphate of dA and the α-amino group of puromycin (Welch et al. 1995). This analog, CCdApPuro, binds to Escherichia coli ribosomes with a Kd in the submicromolar range and is a potent inhibitor of peptidyl transferase. Chemical footprinting experiments revealed that bound CCdApPuro affords protection to several nucleotides that are associated with the ribosomal P site. It is also a competitive inhibitor of the “fragment reaction” at the A site. This compound therefore appears to span the peptidyl transferase reaction center. Subsequent work has shown that the presence or absence of the 2′-OH group of the P-site A residue, equivalent to A76 in tRNAPhe, does not significantly affect either the affinity of similar transition-state analogs for the 50S subunit or their positioning at the PTC (Weinger et al. 2004; Schmeing et al. 2005a).
We have recently undertaken the synthesis of a photoreactive counterpart of CCdApPuro in which dA is replaced by 2-azidodeoxyadenosine (2N3dA) and the CC portion by an otherwise unmodified, full-length tRNAPhe molecule. We anticipate that (2N3dA76)tRNAPhe-p-Puro, like CCdApPuro, will bind simultaneously to the A and P sites and can therefore be used to monitor dynamic changes at the peptidyl transferase center that occur during peptide bond formation. There is in fact considerable evidence that peptide bond formation is accompanied by conformational rearrangements within the 23S rRNA (Polacek et al. 2000; Bayfield et al. 2001). In addition, significant differences in the position and orientation of nucleotides at the peptidyl transferase center have emerged from comparisons of 23S rRNA conformation within the 50S subunits of Haloarcula marismortui, Deinococcus radiodurans, and Thermus thermophilus (Ban et al. 2000; Harms et al. 2001: Yusupov et al. 2001; Schmeing et al. 2005b). Cross-linking of (2N3dA76)tRNAPhe-p-Puro to the ribosome can be used to track changes in the orientation of the 3′ end of P-site tRNA relative to ribosomal components during peptide bond formation at the peptidyl transferase center, as labeling will occur from the position that corresponds to the 3′ A residue of P-site tRNA. Cross-linking patterns arising from the use of (2N3dA76)tRNAPhe-p-Puro can therefore be compared directly with those of P-site bound (2N3A76)tRNAPhe.
Here we show that the three new probes (1) bind efficiently to the P site of E. coli 70S ribosomes with affinities similar to unmodified tRNA, (2) can be competed from the P site by unmodified tRNA, and (3) become cross-linked to protein and RNA components of the 50S subunit when irradiated with near-UV light. Characteristic differences in the cross-linking patterns suggest that these tRNA derivatives can be used to follow subtle changes in the position of the tRNA relative to the components of the peptidyl transferase center at different stages of peptide bond formation.
RESULTS AND DISCUSSION
The photoreactive nucleoside bisphosphates, p2N3dAp and p2N32′OMeAp, were synthesized and incorporated into the 3′-terminal position of yeast tRNAPhe in place of A76 to yield the tRNA derivatives (2N3dA76)tRNAPhe and (2N32′OMeA76)tRNAPhe (Fig. 1). In addition, we prepared a new, photoreactive transition-state analog, based on the Yarus analog, CCdApPuro (Welch et al. 1995), which consists of a P-site (2N3dA76)tRNAPhe moiety joined to an A-site puromycin moiety by a 3′ phosphoramidate linkage. Together, the photoreactive tRNA derivatives and the photoreactive transition-state analog were used to probe the PTC, and their cross-linking patterns were compared.
FIGURE 1.
Synthetic schemes and structures of photoreactive nucleotides and modified tRNA ligands. (A) Synthesis of 2-azido-2′-O-methyladenosine 3′,5′-bisphosphate (p2N32′OMeAp). (B) Synthesis of 2-azido-2′-deoxyadenosine 3′,5′-bisphosphate (p2N3dAp) and its final incorporation into the photoreactive transition-state analog (2N3dA76)tRNAPhe-p-Puro.
Synthesis of tRNA derivatives
p2N3Ap, p2N3dAp, and p2N32′OMeAp were labeled with 32P prior to ligation to the tRNA via an exchange reaction catalyzed by 3′-phosphatase-free polynucleotide kinase (PNK). The labeling efficiency of p2N32′OMeAp was significantly lower than that of the other two nucleotides. The incorporation of 32P into all three nucleoside bisphosphates decreased with increasing amounts of ATP, most likely due to pyrophosphate and phosphate impurities present in the ATP stock that inhibited PNK. The addition of PEG-8000 to the exchange reaction significantly improved the labeling (Harrison and Zimmerman 1986), as did both PPase and PNP. PPase is presumed to stimulate the reaction by hydrolyzing inhibitory pyrophosphate (Lillehaug and Kleppe 1977), while PNP was used to convert inorganic phosphate into the noninhibitory ribose-1-phosphate, in the presence of inosine (Nixon et al. 1998). The bisphosphates were next ligated to the 3′ end of yeast tRNAPhe from which A76 had been removed. PPase was also used to improve the efficiency of this reaction by removing pyrophosphate byproducts (Sabatini and Hajduk 1995). In the preparation of (2N3dA76)tRNAPhe and (2N32′OMeA76)tRNAPhe, the 3′-terminal phosphate was removed by CIAP.
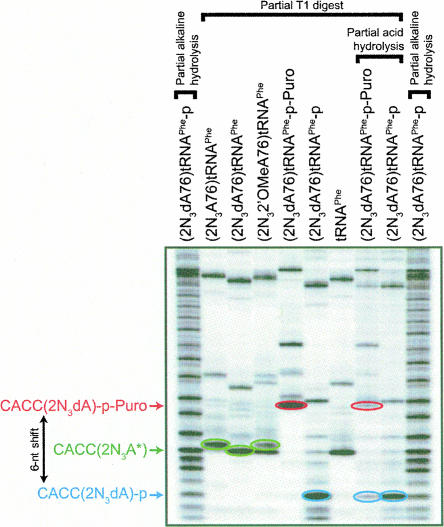
In the synthesis of the transition-state analog, 3′-phosphorylated (2N3dA76)tRNA was condensed with puromycin to form (2N3dA76)tRNA-p-Puro. To verify the presence of puromycin, the product was digested with RNase T1 and the resulting oligonucleotides were subjected to electrophoresis on a denaturing 25% polyacrylamide gel alongside a T1 digest of (2N3dA76)tRNA-p. The shortest 32P-labeled digestion fragments derived from (2N3dA76)tRNA-p-Puro and (2N3dA76)tRNA-p correspond to CACCdA-p-Puro and CACCdA-p, respectively. Figure 2 shows that the presence of puromycin leads to a characteristic electrophoretic mobility shift equivalent to ∼6 nucleotides (nt), confirming that puromycin is joined to the 3′ end of the tRNA. This mobility shift was eliminated upon treatment of CACCdA-p-Puro with 1% TFA for 30 min, consistent with hydrolysis of the acid-labile phosphoramidate bond that joins puromycin to tRNA (Benkovic and Sampson 1971; Welch et al. 1995).
FIGURE 2.
Analysis of modified photoreactive tRNA ligands. (2N3A76)tRNAPhe, (2N3dA76)tRNAPhe, (2N32′OMeA76)tRNAPhe, (2N3dA76)tRNAPhe-p-Puro, (2N3dA76)tRNAPhe-p, and unmodified tRNAPhe, labeled with 32P as described in Materials and Methods, were subjected to digestion with RNase T1 under partial conditions to avoid overdigestion. In addition, (2N3dA76)]tRNAPhe-p-Puro and (2N3dA76)tRNAPhe-p were subjected to partial acid hydrolysis (60 min in 1% trifluoroacetic acid at 23°C) after T1 digestion. Partial alkaline digestion of (2N3dA76)tRNAPhe-p was used to generate a reference ladder. The samples were resolved by denaturing 25% PAGE and visualized by autoradiography. Labeled products released from the 3′ ends of the photoreactive tRNA ligands are CACC(2N3dA), CACC(2N32′OMeA), and CACC(2N3A) (collectively designated as CACC[2N3A*] in the figure), CACC(2N3dA)-p, and CACC(2N3dA)-p-Puro. A characteristic mobility shift of about 6 nt in the 3′ T1 RNase product is observed when (2N3dA76)tRNAPhe-p is condensed with puromycin. The observed 6-nt shift in (2N3dA76)tRNAPhe-p-Puro is reversed upon treatment with acid, consistent with hydrolysis of phosphoramidate linkage between (2N3dA76)tRNAPhe-p and puromycin (Benkovic and Sampson 1971; Welch et al. 1995).
Since (2N3dA76)tRNA-p-Puro has a full-length tRNA as its P-site moiety, it resembles the transition state of peptidyl transfer more closely than the shorter analogs reported previously (Welch et al. 1995; Weinger et al. 2004; Schmeing et al. 2005a). It had been proposed earlier that interactions between the tRNA and the ribosome at sites remote from the PTC make an important contribution to precise substrate alignment (Bashan et al. 2003; Agmon et al. 2004).
Binding of tRNA derivatives to 70S ribosomes
The interaction of the photoreactive tRNA derivatives and the transition-state analog with 70S tight-couple ribosomes was assessed by heterologous competition assays. 32P-labeled (2N3A76)tRNA, (2N3dA76)tRNA, (2N32′OMeA76)tRNA, and (2N3dA76)tRNA-p-Puro were bound to poly(U)-programmed ribosomes in P-site buffer in the presence of unlabeled yeast tRNAPhe (Fig. 3). Homologous competition with unmodified tRNAPhe was also performed. The amount of radiolabeled tRNA associated with the ribosomes at various competitor concentrations was measured by a nitrocellulose filter assay. In all cases, the unlabeled competitor, unmodified tRNAPhe, was able to fully displace the radioactive ligands, indicating competition for the same ribosomal binding site. Due to the very slow dissociation rate of tRNA from the P site at 4°C (half-life ≥ 10 h; Lill et al. 1986), the radiolabeled and competitor ligands were mixed together before adding ribosomes, and the incubation was continued for about five dissociation half-lives in order to reach equilibrium. Relative affinity values for the ligands (IC50) were determined from the competition data. The results show that all of the ligands have similar affinities for the ribosome.
FIGURE 3.
Competition binding assays. Radiolabeled (A) unmodified tRNAPhe, (B) (2N3A76)tRNAPhe, (C) (2N32′OMeA76)tRNAPhe, (D) (2N3dA76)tRNAPhe, and (E) (2N3dA76)tRNAPhe-p-Puro (∼100 nM) were first mixed with various concentrations of unlabeled, unmodified tRNAPhe and then mixed with 70S ribosomes (10 nM) in the presence of polyU and 20 mM MgCl2. The binding reactions were incubated at 4°C for 48 h to reach equilibrium and then analyzed by the nitrocellulose filter binding assay. Error bars represent the standard deviation of triplicate samples of each reaction. The concentrations of competing unlabeled, unmodified tRNAPhe that displace 50% of the specifically bound radiolabeled ligand (IC50) were calculated using the nonlinear regression function of GraphPad Prism software. In these assays, radiolabeled ligands with higher affinities for the ribosome lead to higher IC50 values.
Cross-linking of tRNA analogs to the peptidyl transferase center
Noncovalent complexes between the photoreactive ligands and poly(U)-programmed ribosomes were irradiated with 300-nm UV light. The irradiated ribosomes were then dissociated into subunits and fractionated by sucrose-gradient centrifugation. Only the 50S subunits of the irradiated samples contained covalently bound, radioactive ligands (Fig. 4).
FIGURE 4.
Sucrose-gradient profiles of dissociated ribosomal subunits after cross-linking. (A) unmodified tRNAPhe, (B) (2N3A76)tRNAPhe, (C) (2N3dA76)tRNAPhe, (D) (2N32′OMeA76)tRNAPhe, and (E) (2N3dA76)]tRNAPhe-p-Puro. Both UV absorbance (solid line) and radioactivity (dashed line with data points) were monitored as the gradients were fractionated. With the exception of unmodified tRNAPhe, the ligands cross-linked exclusively to the 50S subunit, as shown by the comigration of radioactivity with the 50S peak.
Cross-linked material was extracted from the isolated 50S subunits and analyzed by PAGE. As shown in Figure 5, 32P-labeled (2N3A76)tRNA and (2N3dA76)tRNA cross-linked to 23S rRNA and to 50S-subunit proteins in a ratio of about 50:50, while the corresponding ratio for (2N32′OMeA76)tRNA was 60:40 and that for (2N3dA76)tRNA-p-Puro was 70:30, suggesting different orientations of the 3′ end of the P-site ligands relative to components of the PTC. Analysis of the cross-linked proteins by one- and two-dimensional PAGE revealed that (2N3A76)tRNA, (2N3dA76)tRNA, and (2N32′OMeA76)tRNA labeled mainly L27, with a small amount of incorporation into L33 owing to the spillover of nonaminoacylated tRNA derivatives into the E-site (Fig. 6; see Maguire et al. 2005). In contrast, (2N3dA76)tRNA-p-Puro cross-linked exclusively to L27, since the transition-state analog spans the P and A sites and cannot spill into the E site.
FIGURE 5.
Analysis of the distribution of cross-links between 23S rRNA and 50S-subunit proteins. (A) 32P-labeled material isolated from 50S subunits was resuspended in 6 M urea, resolved by 4%–20% gradient SDS-PAGE, and visualized by autoradiography. With the exception of unmodified tRNAPhe, the ligands cross-linked to both 23S rRNA and to proteins. (B) Bar graph showing the relative distribution of cross-linked 23S rRNA and protein estimated by PhosphorImager analysis of the gels. Error bars represent the standard deviation in measurements obtained from three separate SDS-PAGE runs.
FIGURE 6.
Identification of ribosomal proteins cross-linked by each photoreactive ligand. 50S-subunit components were solubilized in 6 M urea and incubated with ribonuclease S1, which entirely digested the RNA, leaving a radioactively labeled 5′-phosphate and its modified A76 nucleotide covalently attached to a ribosomal protein. The digested material was resolved by (A) 10% SDS-PAGE and by (B) two-dimensional PAGE, and then visualized by autoradiography. In panel A the labeled bands contain different derivatives of A76, collectively designated as A*. In panel B, the samples were spiked with nonradioactive proteins that were acid-extracted from purified 70S ribosomes to provide reference markers. The proteins in the two-dimensional (2D) gel were stained with bromophenol blue and the image was overlaid with the autoradiogram of the same gel. The positions of (p2NA)-L27, (p2NA)-L33, and (2NdA-p-Puro)-L27 are displaced from those of L27 and L33 owing to the presence of the negatively charged nucleotide moieties (Maguire et al. 2005).
The three new tRNA-derived ligands described in this report—all with photoreactive nucleotides at their 3′ termini—should prove useful in characterizing the interaction of tRNA with the PTC. They have been shown to bind to 70S tight-couple ribosomes with affinities similar to that of unmodified tRNA and all can be cross-linked to RNA and protein components of the 50S ribosomal subunit upon irradiation with UV light. Furthermore, we find that the nature and extent of the protein cross-linking by the photoreactive transition-state analog differs from that of the tRNA analogs. Identification of the cross-linked nucleotide(s) within the 23S rRNA, currently underway, should elucidate the extent to which the position of A76 in the P-site tRNA changes relative to the rRNA in the transition state.
MATERIALS AND METHODS
Materials
Yeast tRNAPhe, poly(U), puromycin, yeast pyrophosphatase (PPase), yeast purine nucleoside phosphorylase (PNP), inosine, ADP, linear polyacrylamide (GeneElute), bovine serum albumin (BSA), and polyethylene glycol (PEG-8000) were from Sigma. T4 RNA ligase was from Epicentre Technologies; calf intestinal alkaline phosphatase (CIAP) and 3′ phosphatase-free T4 polynucleotide kinase (PNK) were from Roche. 1-ethyl-3-[3-(dimethylamino)propyl]carbodiimide (EDC) was from Avocado Organics. 2-chloro-2′-deoxyadenosine (cladribine) was a product of General Intermediates of Canada. Partially purified [γ-32P]ATP (6000 Ci/mmol) was obtained from Perkin Elmer Life Sciences. Forty percent polyacrylamide mix (acrylamide/bis-acrylamide, 19:1) was from Amresco, 96-well filter plates (MultiScreen-HA) were from Millipore, Nucleobond AX20 columns were from Machery & Nagel, DEAE-Sephadex A-25 was from GE Healthcare, and precast 4%–20% polyacrylamide gradient gels were from Gradipore. MRE600 cells were from the University of Alabama Fermentation Facility. Plasmid pQECCA, which encodes His-tagged yeast nucleotidyl transferase under lac regulation, was the generous gift of Dr. D.L. Thurlow, Clark University, Worcester, MA.
Preparation of 70S tight-couple ribosomes
For the isolation of tight-couple 70S ribosomes, ∼10 g of MRE600 cells were broken by grinding with alumina and further processed as described (Rheinberger et al. 1988; Maguire et al. 2005).
Synthesis of photoreactive nucleotides
2-Azidoadenosine-3′,5′-bisphosphate (p2N3Ap)
p2N3Ap was prepared according to Sylvers et al. (1989).
2-Azido-2′-deoxyadenosine-3′,5′-bisphosphate (p2N3dAp)
2-Azido-2′-deoxyadenosine was synthesized from 2-chloro-2′-deoxyadenosine (cladribine) as described (Lioux et al. 2003). Fifty-four milligrams (0.18 mmol) of 2-azido-2′-deoxyadenosine, 83 mg (1.2 mmol) of 1,2,4-triazole, 42 μL (0.3 mmol) of pyrophosphoryl chloride, and 121 mg (1.2 mmol) of triethylamine were reacted in 6 mL of pyridine following the procedure of Folkman et al. (1990). The crude product obtained by quenching the reaction with ice-cold 1 M triethylammonium bicarbonate solution (pH 7) was chromatographed on a 2 × 40-cm DEAE Sephadex A-25 column using a 1200-mL linear gradient of 0.05–2 M triethylammonium buffer (pH 7.5) to give 2-azido-2′-deoxyadenosine-3′,5′-bisphosphate in 5% yield. The compound was identified by its mass spectrum (electrospray, negative ion mode, m/z 451.2 [M − 1]) and by changes in its ultraviolet spectrum as a function of pH, which were similar to those reported for 2-azidoadenosine (MacFarlane et al. 1982).
2-Azido-2′-O-methyladenosine-3′,5′-bisphosphate (p2N32′OMeAp)
The method used to prepare 2-azido-2′-O-methyladenosine was very similar to that for 2′-O-methyladenosine (Yano et al. 1980) except that a greater than fourfold molar excess of sodium hydride was used since equimolar amounts of base and nucleoside appeared to give mostly the N-6-methylated compound. One hundred twenty milligrams (0.388 mmol) of vacuum-dried 2-azidoadenosine were dissolved in 9 mL of anhydrous DMF and cooled to −20°C, after which 40.8 mg (1.70 mmol) of sodium hydride were added. Next, a solution containing 62 μL (0.463 mmol) of methyl iodide in 0.5 mL of anhydrous DMF was added dropwise over a 1-h period. The mixture was stirred for 2 h at 25°C and then quenched by the addition of 131.2 mg (1.702 mmol) of ammonium acetate. The crude product was purified by flash column chromatography to give 2-azido-2′-O-methyladenosine in 25% yield. The compound was identified by its mass spectrum (electrospray, positive ion mode, m/z 323.5 [M + 1]) and by 1H NMR. The compound was then phosphorylated and purified exactly as 2-azidoadenosine-3′,5′-bisphosphate (Sylvers et al. 1989) to give 2-azido-2′-O-methyladenosine-3′,5′-bisphosphate in 50% yield. The compound was identified by its mass spectrum (electrospray, negative ion mode, m/z 481.5 [M − 1]) and by its 31P NMR spectrum (D2O, pH 10; 85% H3PO4 external standard), which showed signals for the 3′-phosphates of the azide and tetrazole tautomers at δ 4.34 and 4.35 and for the 5′-phosphates at δ 3.90 and 3.95 (Barrio et al. 1978).
Labeling of photoreactive nucleotides with 32P
Nucleoside-3′,5′-bisphosphates were labeled with 32P at the 5′ position using a phosphate exchange reaction essentially as described by Sylvers et al. (1989). In brief, the nucleotide-3′,5′-bisphosphate (0.25 mM) was incubated for 30 min at 37°C in a buffer containing 50 mM imidazole (pH 6.6), 0.25 mM ADP, 10 mM MgCl2, 0.1 mM EDTA, 10 mM inosine, 12.5 μM [γ-32P]ATP (6000 Ci/mmol), and PEG-8000 (5% w/v) in the presence of yeast PPase (25 U/mL) and yeast PNP (25 U/mL). T4 polynucleotide kinase, 3′-phosphatase free (500 U/mL), was then added and the reaction continued for another 30 min, after which it was quenched by heating for 1 min at 95°C. The 32P-labeled nucleoside bisphosphates were ligated to tRNA without further purification (Barrio et al. 1978; England and Uhlenbeck 1978).
Preparation of tRNA
Yeast nucleotidyl transferase was expressed from plasmid pQECCA in E. coli JM109 and purified as described (Maguire et al. 2005). The 3′ terminus of yeast tRNAPhe was regenerated using purified yeast nucleotidyl transferase, and the regenerated tRNA was subjected to a single cycle of the Whitfeld degradation to remove the 3′-terminal A76 (Wower et al. 1988). 32P-labeled nucleoside-3′,5′-bisphosphates were ligated to the 3′ end of tRNA lacking A76 (Sylvers et al. 1989). Ligation mixtures, which consisted of 2 μM tRNA (or 1 μM of tRNA for the ligation of p2N3Ap) and 4 μM nucleoside-3′,5′-bisphosphate in 50 mM HEPES (pH 7.5), 15 mM MgCl2, 10 μM ATP, 15% (v/v) DMSO, and 10 μg/mL BSA, and contained 10 units/mL of yeast PPase and 320 units/mL of T4 RNA ligase, were incubated at 4°C for 19 h. The resulting 32P-labeled tRNA-p was diluted twofold with 0.1 M Tris acetate (pH 6.3) and 15% ethanol, and then applied to a Nucleobond AX20 column. The column was washed with 4 mL of 0.1 M Tris acetate (pH 6.3), 0.2 M NaCl, and 15% ethanol to remove unincorporated nucleotides and enzymes used in the previous steps, followed by 4 mL of 30% ethanol. The tRNA-p sample was eluted with 1.0 mL of 0.6 M NaOAc and 30% ethanol and precipitated by adding ethanol to 70% and decreasing the pH to about 5.2 with glacial acetic acid. Linear polyacrylamide (GeneElute) was used as a coprecipitant. When necessary, the tRNA-p was treated with CIAP to remove the 3′ phosphate. The resulting tRNAs were further purified by denaturing polyacrylamide gel electrophoresis (PAGE) and extracted as described (Maguire et al. 2005).
Preparation of (2N3dA76)tRNA-p-Puro
(2N3dA76)tRNA-p was condensed with puromycin by a method similar to that reported by Welch et al. (1995). The reaction was carried out overnight at room temperature in a 50 μL reaction mixture containing 100–150 nmol of (2N3dA76)tRNA-p, 75 mM neutralized puromycin, 0.4 M EDC, and 0.4 M MES (pH 5.6). The reactants were then diluted by addition of 250 μL of RNA extraction buffer (0.5 M NH3+ acetate, 10 mM Mg++ acetate, 1 mM EDTA, 1% SDS), extracted with phenol and chloroform, and precipitated. The sample was then resuspended in 15 μL of RNA loading dye (90 mM Tris-90 mM boric acid at pH 8.3, 2 mM EDTA, 6 M urea, 0.03% xylene cyanol, 0.03% bromophenol blue), resolved by 43-cm denaturing 8% PAGE, and extracted (Maguire et al. 2005). Typical reaction yields were over 80%. The purity and homogeneity of the resulting (2N3dA76)tRNA-p-Puro were routinely checked by RNase T1 digestion, followed by analysis of the digested fragments by 25% denaturing PAGE.
Competitive binding of tRNA ligands to ribosomes
Heterologous competition binding of tRNA ligands to the P site of the 70S tight-couple ribosomes was performed in 15 μL reaction mixtures under conditions similar to those of Wower et al. (1988). P-site buffer consisted of 50 mM Tris-HCl (pH 7.5), 100 mM NH3Cl, and 20 mM MgCl2. Various concentrations of unlabeled competitor yeast tRNAPhe, whose 3′ terminus had been regenerated by yeast nucleotidyl transferase, were used. The unlabeled and radiolabeled ligands were mixed and, after the addition of ribosomes, the mixtures were incubated for 48 h at 4°C to reach equilibrium. Following incubation, 180 μL of binding buffer were added to each mixture. The mixtures were immediately filtered and washed twice with 200 μL of the same buffer using a 96-well nitrocellulose filter plate. As the hydrophobicity of (2N3dA76)tRNA-p-Puro results in an unusually high background in this assay, 5% DMSO was added to the buffer with which the filters were washed after ligand deposition. The addition of DMSO dramatically reduces nonspecific binding of the ligand to the filter but has no significant effect on the binding of the ligand to ribosomes (data not shown). The filter plates were next disassembled and exposed to an imaging screen that was scanned using a Fuji FLA-5000 Imaging System; the radioactivity was quantified with the aid of Fuji Multi Gage software. Each competition binding reaction was performed in triplicate; relative affinity values (IC50) were determined by fitting displacement binding inhibition values by nonlinear regression using Prism data software (GraphPad, Inc.).
Cross-linking of tRNA to ribosomes and analysis of cross-links
Cross-linking of tRNA ligands to 70S tight-couple ribosomes was performed in 100–1000 μL reaction mixtures according to Wower et al. (1988). Sample mixtures were spotted onto a Petri dish that was placed on ice and irradiated for 5 min at 300 nm in a Rayonet RPR-100 photochemical reactor (Southern New England Ultra Violet Company) with six RPR-3000 lamps installed. The emitted light was filtered through a polystyrene Petri dish cover and a 0.025-mm cellulose triacetate sheet to significantly reduce the transmission of light below 300 nm. The sample was then precipitated with two volumes of ethanol and processed as described (Maguire et al. 2005). In brief, the 50S subunits were separated by centrifugation through 15%–30% (w/v) sucrose gradients and recovered using an ISCO density gradient fractionator. To estimate the proportion of tRNA that was covalently bound to 23S ribosomal RNA or to ribosomal proteins, a portion of the cross-linked sample was subjected to SDS-PAGE using a 4%–20% (w/v) gradient gel. The gel was dried and exposed to an imaging screen and radioactivity was quantified using a Fuji FLA-5000 imager. Cross-linked tRNA–protein adducts were isolated and analyzed by one- and two-dimensional polyacrylamide gel electrophoresis as described (Maguire et al. 2005).
ACKNOWLEDGMENTS
We are grateful to Dr. Charles Dickenson for assistance with the 31P NMR measurements. This work was supported by grant GM 67267 From the National Institutes of Health.
Footnotes
Article published online ahead of print. Article and publication date are at http://www.rnajournal.org/cgi/doi/10.1261/rna.425907.
REFERENCES
- Agmon, I., Amit, I., Auerbach, T., Bashan, A., Baram, D., Bartels, H., Berisio, R., Greenberg, I., Harms, J., Hansen, H.A.S., et al. Ribosomal crystallography: A flexible nucleotide anchoring tRNA translocation, facilitates peptide-bond formation, chirality discrimination, and antibiotics synergism. FEBS Lett. 2004;567:20–26. doi: 10.1016/j.febslet.2004.03.065. [DOI] [PubMed] [Google Scholar]
- Ban, N., Nissen, P., Hansen, J., Moore, P.B., Steitz, T.A. The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution. Science. 2000;289:905–920. doi: 10.1126/science.289.5481.905. [DOI] [PubMed] [Google Scholar]
- Barrio, J.R., Barrio, M.C., Leonard, N.J., England, T.E., Uhlenbeck, O.C. Synthesis of modified nucleoside 3′,5′-bisphosphates and their incorporation into oligoribonucleotides with T4 RNA ligase. Biochemistry. 1978;17:2077–2081. doi: 10.1021/bi00604a009. [DOI] [PubMed] [Google Scholar]
- Bashan, A., Agmon, I., Zarivach, R., Schluenzen, F., Harms, J., Berisio, R., Bartels, H., Franceschi, F., Auerbach, T., Hansen, H.A.S., et al. Structural basis of the ribosomal machinery for peptide bond formation, translocation, and nascent chain progression. Mol. Cell. 2003;11:91–102. doi: 10.1016/s1097-2765(03)00009-1. [DOI] [PubMed] [Google Scholar]
- Bayfield, M.A., Dahlberg, A.E., Schulmeister, U., Dorner, S., Barta, A. A conformational change in the ribosomal peptidyl transferase center upon active/inactive transition. Proc. Natl. Acad. Sci. USA. 2001;98:10096–10101. doi: 10.1073/pnas.171319598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benkovic, S.J., Sampson, E.J. Structure-reactivity correlation for the hydrolysis of phosphoramidate monoanions. J. Am. Chem. Soc. 1971;93:4009–4016. [Google Scholar]
- Bocchetta, M., Xiong, L., Mankin, A.S. 23S rRNA positions essential for tRNA binding in ribosomal functional sites. Proc. Natl. Acad. Sci. 1998;95:3525–3530. doi: 10.1073/pnas.95.7.3525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- England, T.E., Uhlenbeck, O.C. Enzymatic oligoribonucleotide synthesis with T4 RNA ligase. Biochemistry. 1978;17:2069–2076. doi: 10.1021/bi00604a008. [DOI] [PubMed] [Google Scholar]
- Folkman, W., Kusmierek, J.T., Singer, B. A new one-step method for the preparation of 3′,5′-bisphosphates of acid-labile deoxynucleosides. Chem. Res. Toxicol. 1990;3:536–539. doi: 10.1021/tx00018a007. [DOI] [PubMed] [Google Scholar]
- Harms, J., Schluenzen, F., Zarivach, R., Bashan, A., Gat, S., Agmon, I., Bartels, H., Franceschi, F., Yonath, Y. High-resolution structure of the large ribosomal subunit from a mesophilic eubacterium. Cell. 2001;107:679–688. doi: 10.1016/s0092-8674(01)00546-3. [DOI] [PubMed] [Google Scholar]
- Harrison, B., Zimmerman, S.B. Stabilization of T4 polynucleotide kinase by macromolecular crowding. Nucleic Acids Res. 1986;14:1863–1870. doi: 10.1093/nar/14.4.1863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lill, R., Robertson, J.M., Wintermeyer, W. Affinities of tRNA binding sites of ribosomes from Escherichia coli. Biochemistry. 1986;25:3245–3255. doi: 10.1021/bi00359a025. [DOI] [PubMed] [Google Scholar]
- Lillehaug, J.R., Kleppe, K. Phosphorylation of tRNA by T4 polynucleotide kinase. Nucleic Acids Res. 1977;4:373–380. doi: 10.1093/nar/4.2.373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lioux, T., Gosselin, G., Mathé, C. Azido/tetrazole tautomerism in 2-azidoadenine β-D-pentofuranonucleoside derivatives. Eur. J. Org. Chem. 2003;2003:3997–4002. [Google Scholar]
- MacFarlane, D.E., Mills, D.C.B., Srivastava, P.C. Binding of 2-azidoadenosine [α-32P]diphosphate to the receptor on intact human blood platelets which inhibits adenylate cyclase. Biochemistry. 1982;21:544–549. doi: 10.1021/bi00532a020. [DOI] [PubMed] [Google Scholar]
- Maguire, B.A., Beniaminov, A.D., Ramu, H., Mankin, A.S., Zimmermann, R.A. A protein component at the heart of an RNA machine: The importance of protein L27 for the function of the bacterial ribosome. Mol. Cell. 2005;20:427–435. doi: 10.1016/j.molcel.2005.09.009. [DOI] [PubMed] [Google Scholar]
- Moazed, D., Noller, H.F. Interaction of tRNA with 23S rRNA in the ribosomal A, P, and E sites. Cell. 1989;57:585–597. doi: 10.1016/0092-8674(89)90128-1. [DOI] [PubMed] [Google Scholar]
- Nixon, A.E., Hunter, J.L., Bonifacio, G., Eccleston, J.F., Webb, M.R. Purine nucleoside phosphorylase: Its use in a spectroscopic assay for inorganic phosphate and for removing inorganic phosphate with the aid of phosphodeoxyribomutase. Anal. Biochem. 1998;265:299–307. doi: 10.1006/abio.1998.2916. [DOI] [PubMed] [Google Scholar]
- Polacek, N., Patzke, S., Nierhaus, K.H., Barta, A. Periodic conformational changes in rRNA: Monitoring the dynamics of translating ribosomes. Mol. Cell. 2000;6:159–171. [PubMed] [Google Scholar]
- Porse, B.T., Thi-Ngoc, H.P., Garrett, R.A. The donor substrate site within the peptidyl transferase loop of 23S rRNA and its putative interactions with the CCA-end of N-blocked aminoacyl-tRNA tRNAPhe. J. Mol. Biol. 1996;264:472–483. doi: 10.1006/jmbi.1996.0655. [DOI] [PubMed] [Google Scholar]
- Rheinberger, H.J., Geigenmuller, U., Wedde, M., Nierhaus, K.H. Parameters for the preparation of Escherichia coli ribosomes and ribosomal subunits active in tRNA binding. Methods Enzymol. 1988;164:658–670. doi: 10.1016/s0076-6879(88)64076-6. [DOI] [PubMed] [Google Scholar]
- Sabatini, R., Hajduk, S.L. RNA ligase and its involvement in guide RNA/mRNA chimera formation. J. Biol. Chem. 1995;270:7233–7240. doi: 10.1074/jbc.270.13.7233. [DOI] [PubMed] [Google Scholar]
- Schmeing, T.M., Huang, K.S., Kitchen, D.E., Strobel, S.A., Steitz, T.A. Structural insights into the roles of water and the 2′-hydoxyl of the P-site tRNA in the peptidyl transferase reaction. Mol. Cell. 2005a;20:437–438. doi: 10.1016/j.molcel.2005.09.006. [DOI] [PubMed] [Google Scholar]
- Schmeing, T.M., Huang, K.S., Strobel, S.A., Steitz, T.A. An induced-fit mechanism to promote peptide bond formation and exclude hydrolysis of peptidyl-tRNA. Nature. 2005b;438:520–524. doi: 10.1038/nature04152. [DOI] [PubMed] [Google Scholar]
- Sylvers, L.A., Wower, J., Hixson, S.S., Zimmermann, R.A. Preparation of 2-azidoadenosine 3′,5′-[5′-32P]bisphosphate for incorporation into transfer RNA. Photoaffinity labeling of Escherichia coli ribosomes. FEBS Lett. 1989;245:9–13. doi: 10.1016/0014-5793(89)80180-2. [DOI] [PubMed] [Google Scholar]
- Weinger, J.S., Kitchen, D., Scaringe, S.A., Strobel, S.A., Muth, G.W. Solid phase synthesis and binding affinity of peptidyl transferase transition state mimics containing 2′-OH at P-site position A76. Nucleic Acids Res. 2004;32:1502–1511. doi: 10.1093/nar/gkh311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Welch, M., Chastang, J., Yarus, M. An inhibitor of ribosomal peptidyl transferase using transition-state analogy. Biochemistry. 1995;34:387–390. doi: 10.1021/bi00002a001. [DOI] [PubMed] [Google Scholar]
- Wower, J., Hixson, S.S., Zimmermann, R.A. Photochemical cross-linking of yeast tRNAPhe containing 8-azidoadenosine at positions 73 and 76 to the Escherichia coli ribosome. Biochemistry. 1988;27:8114–8121. doi: 10.1021/bi00421a021. [DOI] [PubMed] [Google Scholar]
- Wower, J., Kirillov, S.V., Wower, I.K., Guven, S., Hixson, S.S., Zimmermann, R.A. Transit of tRNA through the Escherichia coli ribosome. Cross-linking of the 3′ end of tRNA to specific nucleotides of the 23 S ribosomal RNA at the A, P, and E sites. J. Biol. Chem. 2000;275:37887–37894. doi: 10.1074/jbc.M005031200. [DOI] [PubMed] [Google Scholar]
- Yano, J., Kan, L.S., Ts'o, P.O. A simple method of the preparation of 2′-O-methyladenosine. Methylation of adenosine with methyl iodide in anhydrous alkaline medium. Biochim. Biophys. Acta. 1980;629:178–183. doi: 10.1016/0304-4165(80)90276-7. [DOI] [PubMed] [Google Scholar]
- Yusupov, M.M., Yusupova, G.Zh., Baucom, A., Lieberman, K., Earnest, T.N., Cate, J.H.D., Noller, H.F. Crystal structure of the ribosome at 5.5 Å resolution. Science. 2001;292:883–896. doi: 10.1126/science.1060089. [DOI] [PubMed] [Google Scholar]