Abstract
p63 is a p53-related transcription factor. Utilization of two different promoters and alternative splicing at the C terminus lead to generation of six isoforms. The α isoforms of TAp63 and ΔNp63 contain a transactivation-inhibitory (TI) domain at the C termini, which can bind to the transactivation (TA) domain and inhibit its transcriptional activity. Consequently, TAp63α can directly inhibit its activity through an intramolecular interaction; similarly, ΔNp63α can inhibit the activity of the active TAp63 isoforms through an intermolecular interaction. In this work, we demonstrate that after induction of apoptosis, the TI domain of the p63α isoforms is cleaved by activated caspases. Cleavage of ΔNp63α relieves its inhibitory effect on the transcriptionally active p63 proteins, and the cleavage of TAp63α results in production of a TAp63 protein with enhanced transcriptional activity. In agreement with these data, generation of the N-terminal TAp63 fragment has a role in apoptosis because stable cell lines expressing wild-type TAp63 are more sensitive to apoptosis compared with cells expressing the noncleavable mutant. We also used a model system in which TAp63 expression was induced by trichostatin-A treatment in HCT116 cells. Trichostatin-A sensitized these cells to apoptosis, and this sensitization was associated with cleavage of up-regulated p63.
Keywords: cancer, p53, transcription, histone deacetylase inhibitor, trichostatin A
TP63 belongs to the p53 family of transcription factors and shares high homology with the other family members (1). The p63 gene encodes six isoforms with different N and/or C termini [supporting information (SI) Fig. 6]. The transactivation (TA) domain containing isoforms are encoded from an upstream promoter and, because of the presence of an N-terminal transactivation domain, activate a set of target genes, including bax, mdm2, and p21, and can consequently induce cell cycle arrest and apoptosis (1–3). The ΔN isoforms are expressed from an intronic promoter and therefore lack the TA domain. However, the presence of a second TA domain between residues 410 and 512 (TA2) confers transcriptional activity on at least some ΔNp63 isoforms (4–7).
Alternative splicing of both TAp63 and ΔNp63 generates proteins with different C termini (TAp63α, β, and γ isoforms and ΔNp63α, β, and γ isoforms). The C terminus of the α isoforms contains a transactivation-inhibitory (TI) domain that can interact with the TA domain and mask the residues that are important for transactivation, thereby suppressing TAp63-mediated transactivation (1, 8).
Unlike p53, expression of p63 is highly tissue-specific. Although ΔNp63α is expressed predominantly in the regenerative compartment of epithelial tissues, TAp63 is the predominant isoform expressed in the nervous system and in oocytes (1, 9–13). p63 mutations are very rare in cancer, and the role of specific isoforms of p63 in cancer is not clear. It has recently been shown that in squamous cell carcinoma, TAp63 and ΔNp63 isoforms are both up-regulated at the mRNA level although only ΔNp63 protein could be detected (R. Cui, J. He, R. Mei, C. C. de Fromentel, G. Martel-Planche, P. Taniere, and P. Hainaut, unpublished data). TAp63 has been shown to be up-regulated in malignant lymphomas, and treatment of hepatocellular carcinoma cells with chemotherapeutic drugs results in a dramatic increase of TAp63α levels (2, 15).
TAp63 isoforms can transactivate many p53 target genes because they can bind DNA through p53-responsive elements. We have recently shown that TAp63α can activate major apoptosis pathways by triggering death receptor signaling and mitochondria and thus sensitizes hepatocellular carcinoma cell lines toward chemotherapy (2). TAp63α can directly induce the expression of death receptors such as CD95, TNF-R, and TRAIL-R, as well as proapoptotic Bcl-2 family members such as Bax. Furthermore, inhibition of TAp63 function results in chemoresistance.
Although both the TA and ΔNp63 isoforms can act as transcription factors when homodimerized, the ΔN isoforms can heterooligomerize with other p53 family members as well as with the TAp63 isoforms and can modify their activity in vitro (8, 16–18). ΔNp63α inhibits the function of p53 and TAp63 efficiently, whereas ΔNp63γ is incapable of inhibiting TAp63α but can still inhibit p53 in vitro (8).
Here we demonstrate that p63 is cleaved after an apoptotic stimulus by activated caspases. Cleavage of α isoforms resulted in the loss of the C-terminal TI domain. Although the transcriptional activity of ΔNp63α was unchanged after caspase cleavage, this modification relieved the inhibitory effect of ΔNp63α on the transcriptionally active TAp63 isoforms. In contrast, cleavage of TAp63α resulted in a marked increase in its transcriptional activity. Cleavage of the TI domain was biologically relevant because cell lines stably expressing the noncleavable mutant of TAp63α were more resistant to apoptosis. We also demonstrated that TAp63α expression was up-regulated by treatment of HCT116 cells with the histone deacetylase (HDAC) inhibitor trichostatin-A (TSA). After pretreatment with TSA, these cells became sensitized to apoptosis, which coincided with induction and cleavage of TAp63α.
Results
p63 Is a Target of Caspases.
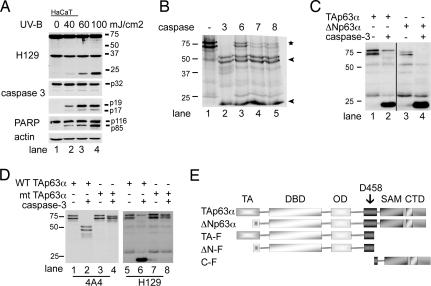
We have recently detected a change in the molecular weight of p63 protein after induction of apoptosis (Fig. 1A). To understand whether this change was a result of caspase cleavage, we in vitro translated TAp63α and incubated these proteins with recombinant active caspases (Fig. 1B). All of the caspases tested (caspase-3, -6, -7, and -8) cleaved p63 with different efficiencies in vitro, yielding two cleavage products. ΔNp63α was also cleaved when incubated with caspase-3 in vitro (Fig. 1C). By using two different p63 antibodies directed against either the N terminus (4A4) or the C terminus (H129) of p63, we identified the smaller molecular weight cleavage product as a C-terminal p63 fragment (Fig. 1D, lanes 1 and 2, and 5 and 6). Mutation of candidate aspartate residues by site-directed mutagenesis in the C-terminal region based on putative caspase cleavage sites revealed that p63 is cleaved at a single site, at aa 458 (Fig. 1D).
Fig. 1.
p63 is a caspase target. (A) HaCaT cells were treated with different UV-B doses, and cells were collected 24 h after treatment. Apoptosis was assessed by caspase-3 activation and PARP cleavage. H129 antibody was used to detect p63. (B) TAp63α protein, produced by in vitro translation in the presence of [35S] methionine, was incubated with 200 nM recombinant active caspase-3, -6, -7, or -8 for 2 h. All caspases cleaved p63 efficiently, yielding two cleavage products, indicated by the arrows. The star indicates the p63 isoform generated by an in-frame upstream methionine (see SI Fig. 6D). (C) TAp63α and ΔNp63α were generated by in vitro translation and cleaved by 200 nM active caspase-3. Both isoforms were cleaved by caspase-3. (D) The candidate caspase-3 cleavage site, Asp-458, was mutated by site-directed mutagenesis, and in vitro translated WT and mutant (mt) p63 proteins were incubated with 200 nM caspase-3 for 2 h. The cleavage products were detected by using two different antibodies. The 4A4 antibody (α N-ter) detected the high molecular weight p63 fragment (Left), whereas the H129 antibody (α C-ter) antibody detected the small molecular weight fragment (Right), which suggests that p63 has a single caspase-3 site. (E) Schematic representation of the p63 cleavage products. TA-F is the N-terminal fragment of TAp63, ΔN-F is the N-terminal fragment of ΔNp63, and C-F is the C-terminal fragment of p63α.
Cleavage of p63 by caspase-3 was efficient because 5 nM active caspase-3 was sufficient to cleave in vitro translated p63 (SI Fig. 6B), and 100 nM recombinant active caspase cleaved p63 within the first 5 min of incubation (SI Fig. 6C). Kinetics of caspase-8 cleavage was similar to cleavage by caspase-3 (data not shown).
To evaluate the biochemical and biological characteristics of the p63 cleavage fragments, we cloned them into pcDNA3.1 as the N-terminal fragment of TAp63 (TA-F), the N-terminal fragment of ΔNp63 (ΔN-F), and the C-terminal fragment of the α isoforms (C-F) (Fig. 1E).
N-Terminal p63 Cleavage Fragments Remain in the Nucleus.
When expressed in H1299 cells, both TA-F and ΔN-F localized to the nucleus, whereas the C-terminal fragment was detected outside the nucleus (Fig. 2A). The α isoforms of p63 contain a C-terminal TI domain that controls the activity of p63 by interacting with the TA domain. It has been suggested that this intramolecular interaction inhibits the transcriptional activity of TAp63α, therefore explaining why TAp63γ, which lacks the TI domain, is transcriptionally more active than the α isoform (8). Retention of both TA-F and ΔN-F in the nucleus after cleavage and the lack of the TI domain in both fragments raised the question as to whether these N-terminal fragments were transcriptionally active and if so, whether the loss of the TI domain resulted in enhanced transcriptional activity.
Fig. 2.
TAp63 cleavage product has high transcriptional activity. (A) H1299 cells were plated on coverslips and transfected with TAp63α or p63 fragments by using Lipofectamine-2000. Twenty-four hours after transfection, cells were fixed, permeabilized, and probed with 4A4 antibody (for TAp63α, TA-F, and DN-F) or the H129 antibody (for C-F). Nuclei were stained with DAPI. Both TA-F and DN-F displayed a predominantly nuclear staining, whereas C-F localized mainly outside the nucleus. (Scale bars, 10 μm.) (B) TA-F has an enhanced transcriptional activity compared with TAp63α on p53-responsive promoters. H1299 cells were cotransfected with the indicated plasmids and promoters. Luciferase assay was performed 36 h after transfection. The results represent the mean ± SE for three independent experiments.
To answer this question, we transfected H1299 cells with these fragments and compared their transcriptional activity with TAp63, ΔNp63, and p53 on different promoters. After transfection, TAp63, ΔNp63, and the cleavage fragments were expressed at comparable levels (SI Fig. 7A). With all of the p53-responsive promoters tested (bax, PG13, mdm2, and cyclin G) TA-F showed a marked increase in transcriptional activity compared with uncleaved TAp63α (SI Fig. 7B, compare second and fourth lanes) as also verified at the protein level for mdm2 and cyclin G (SI Fig. 7B). Cleavage of ΔNp63 had no effect on its transcriptional activity on these promoters (Fig. 2B, compare third and fifth bars). We also tested the activities of these fragments on p63-responsive promoters of genes important in epithelial differentiation with similar results (SI Fig. 7C) (19, 20). These results demonstrated that caspases could regulate p63 activity by transforming the transcriptionally less active full-length α isoform into a more active truncated form. However, because TA-F transcriptional activity was more markedly enhanced over that of TAp63α on apoptosis-related promoters and overexpression of TA-F in H1299 cells resulted in 25% apoptosis after 3 days, whereas TAp63α caused only 14% apoptosis (SI Fig. 7D), we focused on the effects of p63 cleavage fragments on apoptosis in subsequent experiments.
Endogenous p63 Is Targeted by Caspases.
We next asked whether endogenous p63 is also modified by caspases after apoptotic stimuli. Treatment of immortalized human keratinocyte HaCaT cells with UV-B, staurosporine, or cisplatin resulted in cleavage of p63 in direct proportion to the apoptosis observed (Fig. 3A). As before, we detected the cleavage products by using two different p63 antibodies against the N terminus and the C terminus of p63. Apoptosis was assessed by phosphatidylserine externalization, processing of caspase-3, and poly(ADP-ribose) polymerase (PARP) cleavage. Generation of these cleavage products was caspase-dependent because it was inhibited by Z-Val-Ala-Asp-fluoromethyl ketone (z-VAD-fmk) (Fig. 3A, lanes 3, 5, and 7).
Fig. 3.
Endogenous p63 is cleaved after apoptotic stimuli. (A) Detection of endogenous p63 cleavage forms after apoptotic stimuli. HaCaT cells were treated with 60 mJ/cm2 UV-B, 500 nM staurosporine, or 30 μM cisplatin for 24 h in the presence or absence of 100 μM z-VAD-fmk. Apoptosis was measured by FACS analysis for annexin-V-positive cells. For the analysis of p63 processing, equal number of cells were sonicated in Laemmli buffer and subjected to Western blotting. Caspase-3 activation and PARP cleavage were also analyzed in the same preparations. Arrowheads indicate the full-length p63 and the p63 cleavage products detected by the 4A4 (N-terminal) and the H129 (C-terminal) antibodies, whose production was inhibited in z-VAD-fmk-treated cells. (B) To test the possibility that cleavage in apoptotic cells may be the result of calpain activation, HaCaT cells were treated with UV-B, and the C-terminal cleavage fragment was detected by H129 antibody. Calpeptin treatment did not inhibit the p63 cleavage, but cleavage was inhibited by z-VAD. (C) To demonstrate further the cleavage of endogenous p63, HT1376 were treated with 100 μM cisplatin, 75 μM etoposide, and 1 μM Taxol. Apoptosis was evaluated by caspase-3 activation PARP cleavage and annexin-V. All drugs produced p63 cleavage as detected by the H129 antibody, together with caspase-3 activation and PARP cleavage.
p53 and the other family member p73 are also known to be targeted and cleaved by calpains (21, 22). To exclude the possibility that cleavage of p63, after apoptotic stimuli, was through calpain activation, we treated HaCaT cells with UV-B in the presence or absence of either calpeptin or z-VAD-fmk. Although no cleavage product was observed in z-VAD-fmk-treated cells, calpeptin failed to protect cells against UV-induced p63 cleavage (Fig. 3B). This result demonstrated that generation of these p63-derived peptides was the result of caspase- and not calpain-mediated proteolysis.
To demonstrate further the cleavage of endogenous p63 after treatment with chemotherapeutic drugs, we exposed the bladder cancer cell line HT1376 to cisplatin, etoposide, and Taxol (Fig. 3C). As in HaCaT cells, all three drugs caused cleavage of p63 along with caspase-3 activation and PARP cleavage.
Our data based on overexpression of p63 cleavage fragments implied that the N-terminal p63 fragments were retained in the nucleus whereas the C-F was predominantly localized to the cytosol. To verify the localization of p63 fragments after cleavage of endogenous p63, we induced apoptosis in HT1376 cells by etoposide and prepared nuclear lysates from untreated and treated cells. Although we were readily able to detect the N-terminal cleavage product in the nuclear lysate, the C-terminal fragment was barely detectable in the nucleus (SI Fig. 8). This finding suggested that after cleavage of endogenous p63, the C-terminal fragment was released from the N terminus of the protein and excluded from the nucleus.
ΔNp63 Cleavage Reduces Its Inhibitory Effects on Transcriptionally Active TAp63 Isoforms.
Our results demonstrated that cleavage of ΔNp63 had no effect on the transcriptional activity of the protein (Fig. 2B). However, ΔNp63 isoforms can oligomerize with TAp63, and the interaction between the TA domain of TAp63 with the TI domain of ΔNp63α results in inhibition of transcriptional activity of the TA isoforms in vitro (1, 8). Therefore, we wanted to evaluate the effect of ΔNp63 cleavage on its ability to inhibit transcriptionally active TAp63 isoforms and also on TA-F because the activated caspases would also cleave TAp63α. We cotransfected TAp63α, TAp63γ isoforms, or TA-F, together with ΔNp63α or ΔN-F and tested their transcriptional activities on mdm2, bax, and PG13 promoters (Fig. 4). Cleavage of the ΔNp63α TI domain reduced the inhibitory effect of ΔNp63 on TAp63-mediated transactivation. However, this reduction in inhibitory activity was more evident when ΔN-F was coexpressed with TAp63γ or TA-F. Thus, expression of TAp63γ with ΔNp63α resulted in a 25–60% loss in the transcriptional activity of TAp63γ, whereas when coexpressed with ΔN-F this loss was reduced to 0–12% (Fig. 4B). The results were very similar when TA-F was coexpressed with ΔNp63α (40–60% loss) or ΔN-F (5–20% loss) (Fig. 4C). These results were also confirmed at protein level for mdm2, p21, and cyclin G expression (SI Fig. 9).
Fig. 4.
Cleavage of ΔNp63 relieves its inhibitory effect on TAp63α- or TAp63γ-mediated transcription. TAp63α (A), TAp63γ (B), or TA-F (C) was expressed in H1299 cells alone or together with ΔNp63α or ΔN-F. Twenty-four hours after transfection-relative activities on the indicated promoters were measured by luciferase assay. When coexpressed with ΔNp63α, the transcriptional activities of TAp63α, TAp63γ, and TA-F were reduced by ≈55–70%, 25–60%, and 40–60%, respectively. When coexpressed with ΔN-F, both TA isoforms retained most of their transcriptional activities (20–25% loss with TAp63α, 0–12% loss with TAp63γ, and 5–20% loss with TA-F).
TAp63 Cleavage Confers Sensitivity to Apoptosis.
To evaluate the biological significance of p63 cleavage after apoptotic stimuli, we generated cell lines stably expressing TAp63α or the noncleavable mutant of TAp63α (TAp63α-D458A) (SI Fig. 10A). We then exposed these cells to different doses of UV-B to induce apoptosis. As expected, although we were able to detect the cleavage fragment in the stable cell lines expressing wild-type (WT) p63, no cleavage product was detected in the stable cell lines containing the noncleavable mutant (Fig.5A). Before analysis of the effect of TAp63 cleavage on apoptosis in these stable cell lines, we tested the transcriptional activities of the WT and TAp63α-D458A mutant on bax, mdm2, and p21 promoters by luciferase assay. Both of the p63 forms, WT and the D458A noncleavable mutant, showed similar activities on these promoters (SI Fig. 10B). To compare the transcriptional activities of TAp63α and TAp63α-D458A under stress conditions, we transfected H1299 cells with these plasmids and induced cell death by a low concentration of staurosporine (SI Fig. 10C). Analysis of mdm2 and p21 expression levels after 6 and 12 h of treatment revealed that although TAp63α could rapidly induce mdm2 and p21 levels, the noncleavable mutant failed to up-regulate expression of these genes as efficiently as the WT isoform, providing evidence that the cleavage of TAp63α after an apoptotic stimuli enhances the transcriptional activity in vivo.
Fig. 5.
TAp63α-D458A expression confers resistance to apoptosis. (A) TAp63α-1 and TAp63α-D458A-1 cells were treated with two different doses of UV-B to verify the noncleavable phenotype of the TAp63α-D458A cells. No cleavage product was observed in either of the noncleavable cell lines with 120 or 400 mJ/cm2 UV-B. (B) TAp63α-1 and TAp63α-D458A-1 cells were treated with 100 nM staurosporine for 24 h, and p63 processing, caspase-3 activation, and PARP cleavage were assessed by Western blotting. Actin was used as an equal loading control. The p63 cleavage product in staurosporine-treated TAp63α-1 cells was produced in parallel with caspase-3 activation and PARP cleavage. Although staurosporine also induced caspase-3 activation in TAp63-D458A cells (lanes 11 and 12), there was no corresponding p63 cleavage. Arrows indicate the full-length p63 and the p63 cleavage products detected by H129 antibody. (C) Three different stable clones of TAp63α and TAp63α-D458A were treated as in B to analyze apoptosis by FACS analysis. Although 33–40% of TAp63α cells were apoptotic at 24 h, only 12–18% TAp63α-D458A cells were in the sub-G1 population at this time. (D) Treatment of HCT116 cells with TSA resulted in induction of TAp63α expression. HCT116 cells were treated with 5 μM TSA, and p63 expression was analyzed by RT-PCR after 24 h. (E) TSA-induced TAp63α is cleaved after apoptotic stimuli. HCT116 cells were treated with 250 nM TSA for 24 h. Fresh medium was added to cells containing the indicated drugs, and cells were incubated with these drugs for 12 h. With the drugs tested, up-regulated TAp63α was cleaved efficiently. (F) Up-regulation and cleavage of TAp63α coincide with sensitization of cells to apoptosis. HCT116 cells were treated with 250 nM TSA, 100 μM etoposide (etop; for 12 h), or 100 nM staurosporine (sts; for 12 h) alone or treated with etoposide or staurosporine after pretreatment of cells with 250 nM TSA. With both of the drugs, pretreatment with TSA sensitized these cells to apoptosis significantly.
To evaluate the contribution of TA-F production to apoptosis, we treated the stable cell lines with a low concentration of staurosporine for 24 h and analyzed p63 processing and caspase-3 activation by Western blotting (Fig. 5B). As expected, staurosporine treatment induced cleavage of TAp63α but not of TAp63α-D458A. Caspase-3 activation was observed at an earlier time point in cells expressing WT TAp63α compared with the cells expressing mutant TAp63α or cells transfected with empty vector, suggesting that cells containing the noncleavable mutant of TAp63 were more resistant to apoptosis. A fraction of cells from the same experiment was also used to measure apoptosis at different time points by sub-G1 analysis (Fig. 5C). In accordance with the data obtained by Western blotting, 32–40% of cells were detected in the sub-G1 population at the end of the treatment in cells expressing WT TAp63α. However, cells expressing the noncleavable TAp63α-D458A mutant were resistant to apoptosis (13–18% sub-G1).
These data suggested that TAp63α can induce apoptosis efficiently and this induction relies predominantly on the cleavage of C-terminal domain of the protein by activated caspases. Therefore, we thought it would be clinically relevant to identify a pharmacological agent that would specifically induce TAp63 in cancer cells. For this purpose, we treated HCT116 colon cancer cells with various drugs, including etoposide, cisplatin, TSA, Taxol, doxorubicin, and staurosporine for 24 h. Among these compounds only the HDAC inhibitor TSA produced a marked increase in p63 expression, as detected by the anti-p63 antibody (Fig. 5D). By using TAp63- or ΔNp63-specific primers, we showed that the p63 isoform induced by TSA is TAp63 (Fig. 5D).
We used TSA treatment of HCT116 cells as a model system to understand the consequences of up-regulation of TAp63 and to see whether cleavage of endogenous TAp63 has any influence on apoptosis. First we identified a sublethal dose of TSA that will induce TAp63 expression in the absence of apoptosis (SI Fig. 10E). We identified 250 nM TSA as the most efficient concentration. After pretreatment of HCT116 cells with TSA, we incubated these cells with sublethal concentrations of various agents. After TSA treatment, the drugs tested produced efficient TAp63 cleavage (Fig. 5E). Moreover, apoptosis was greater when TSA was used in combination with etoposide or staurosporine (Fig. 5F).
Discussion
Our results demonstrate that p63 is a caspase target, and in vitro, low concentrations of recombinant active caspases are sufficient to cleave p63. The cleavage occurs at aa 458, which results in loss of the TI domain at the C terminus of the p63α isoforms. This cleavage site is not a classical caspase cleavage consensus (SI Fig. 6E) as also described recently for the p53 caspase cleavage site (23). Cleavage of TAp63α results in enhanced transcriptional activity as a result of loss of the TI domain. Cleavage of ΔNp63α does not affect its transactivational ability but abrogates its inhibitory effects on TAp63 isoforms, again after loss of the TI domain. Treatment of cells with various DNA-damaging agents also resulted in cleavage of endogenous p63, and cells expressing a noncleavable mutant showed reduced apoptosis after DNA damage. Moreover, increased p63 expression induced by TSA (and consequently, increased expression of the N-terminal cleavage form) conferred enhanced chemosensitivity.
A previous study has also demonstrated caspase-mediated cleavage of a p63 fragment, p40 (24). However, p40 fragment does not contain the Asp-458 residue, which is shown to be the caspase target in the present work. Moreover, p40 cleavage was mediated by caspase-1 but not by other caspases. Therefore, because we were interested primarily in the effects of p63 cleavage on apoptosis rather than the inflammatory response, we have focused on the Asp-458 cleavage mediated by apoptotic caspases.
We have recently shown that p53 is also a caspase target and that p53 cleavage results in enhanced apoptosis. However, some p53 cleavage products localize to mitochondria, and the increased apoptosis is at least partly the result of mitochondrial damage. Although the C-terminal p63 fragment exits the nucleus and becomes predominantly cytosolic, there is no clear evidence for its mitochondrial localization. Therefore, the increased apoptosis seen after p63 cleavage is unlikely to be caused by mitochondrial injury but to result from two factors. First, cleavage of the TI domain from TAp63α relieves its intramolecular interaction with the N-terminal TA domain and results in increased transcriptional activity on proapoptotic genes. Second, removal of the TI domain from ΔNp63 isoforms abrogates their ability to inhibit the transcriptional activity of TA isoforms in an intermolecular manner, again resulting in an overall enhancement of transcription. Therefore, the effects of caspase cleavage of p63 are nuclear and transcriptional, whereas the effects of caspase cleavage of p53 are mitochondrial and nontranscriptional.
Chemoresistance is a major problem in cancer therapy, particularly in the 50% or so of tumors where p53 is either absent or present only as a mutant inactive form. TAp63 is important for DNA damage-induced oocyte apoptosis (13), and TAp63 can activate both death receptor- and mitochondrial-mediated apoptotic pathways. Inhibition of TAp63 function leads to enhanced chemoresistance (2). p63 is an epithelial protein, and epithelial tumors (e.g., breast, prostate, lung) form a large percentage of total tumor incidence. Therefore, strategies to enhance the apoptotic effects of p63 in epithelial cancers may offer therapeutic benefits. In our experiments, we have shown that DNA damage-induced apoptosis is in proportion to the degree of p63 cleavage induced by the DNA-damaging agent. Moreover, increasing p63 expression with an HDAC inhibitor enhances chemosensitivity to DNA-damaging agents (which cause caspase activation) in parallel with increased expression of p63 cleavage products, which may be an additional mechanism whereby HDAC inhibitors exert their clinical effects. Therefore, therapeutic approaches that enhance both p63 expression and its cleavage are likely to improve the management of chemoresistant tumors, especially those of epithelial origin.
Materials and Methods
Cell culture and drug treatments are discussed in SI Materials and Methods.
Plasmids and Luciferase Assay.
To generate the noncleavable p63 mutants or different p63 cleavage fragments, a plasmid containing human WT p63 with an N-terminal HA tag was used. The QuikChange site-directed mutagenesis kit was used to mutate candidate aspartate residues (Stratagene, La Jolla, CA). p63 fragments were cloned in pcDNA3.1 with V5-His tag using a TOPO cloning kit (Invitrogen, Carlsbad, CA). Luciferase assays were performed as described previously (23).
In Vitro Transcription–Translation of p63 and in Vitro Caspase Cleavage Assay.
WT and alanine-substituted p63 plasmids were in vitro translated and [35S]methionine labeled by the TnT-T7-coupled reticulocyte lysate system (Promega, Madison, WI) according to the manufacturer's instructions. For in vitro cleavage, proteins were incubated with either 200 nM active caspase-3, -6, -7, or -8 or 20–400 nM recombinant caspase-3.
Antibodies and Immunoblot Analysis.
Equal numbers of cells were sonicated in 2× Laemmli buffer and boiled at 95°C for 5 min. HA tag, actin, and anti-p63 antibodies 4A4 (sc-8431) and H129 (sc-8344), cyclin G antibody (sc-7865), anti-p21 antibody (sc-756), and anti-mdm2 antibody (sc-13161) were purchased from Santa Cruz Biotechnology (Santa Cruz, CA). Anti-PARP antibody was from Alexis Biomedicals (San Diego, CA). Nuclear lysates were prepared by using nuclei isolation kit (Sigma, St. Louis, MO).
Immunofluorescence.
H1299 cells (105) were plated on coverslips and transfected with expression vectors containing p63, TA-F, ΔN-F, or C-F. Immunofluorescence was done as described before (23) and analyzed with a confocal laser microscope (LSM 510; Zeiss, Thornwood, NY).
Analysis of Apoptosis.
For sub-G1 cell population, cells were treated and collected at the indicated time points for analysis. Briefly, cells collected and fixed, and treated cells were treated with RNase and propidium iodide. Annexin-V/propidium iodide treatment was performed as described previously (23). Apoptosis was measured by using a flow cytometer (FACSCalibur; BD Bioscience, San Jose, CA).
PCR.
RNA extraction and cDNA synthesis were done as described before (14).
Supplementary Material
Acknowledgments
We thank Dr. Xiao-Ming Sun (MRC Toxicology) for the active caspases and Dr. G. Brown for critical reading of the manuscript. This work was supported by the Medical Research Council and Telethon Grant GGPO4110 (to G.M.); Associazione Italiana per la Ricerca sul Cancro Grant 2743, European Union Grants LSGBH-2005-019067-Epistem, Blandino and LSHC-CT-2004-503576-Active p53, FIRB-2001-RBNE01KJHT_004 (Marino), and RBNE01NWCH_008 (Rotilio), and MIUR, MinSan (to G.M.); a Kimmel Scholar Award (to R.I.A.); and National Cancer Institute/National Institutes of Health grant (to C.M.C.).
Abbreviations
- HDAC
histone deacetylase
- PARP
poly(ADP-ribose) polymerase
- TA domain
transactivation domain
- TI domain
transactivation-inhibitory domain
- TSA
trichostatin-A
- z-VAD-fmk
Z-Val-Ala-Asp-fluoromethyl ketone.
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/cgi/content/full/0700761104/DC1.
References
- 1.Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dotsch V, Andrews NC, Caput D, McKeon F. Mol Cell. 1998;2:305–316. doi: 10.1016/s1097-2765(00)80275-0. [DOI] [PubMed] [Google Scholar]
- 2.Gressner O, Schilling T, Lorenz K, Schulze Schleithoff E, Koch A, Schulze-Bergkamen H, Maria Lena A, Candi E, Terrinoni A, et al. EMBO J. 2005;24:2458–2471. doi: 10.1038/sj.emboj.7600708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Melino G, Lu X, Gasco M, Crook T, Knight RA. Trends Biochem Sci. 2003;28:663–670. doi: 10.1016/j.tibs.2003.10.004. [DOI] [PubMed] [Google Scholar]
- 4.Duijf PH, Vanmolkot KR, Propping P, Friedl W, Krieger E, McKeon F, Dotsch V, Brunner HG, van Bokhoven H. Hum Mol Genet. 2002;11:799–804. doi: 10.1093/hmg/11.7.799. [DOI] [PubMed] [Google Scholar]
- 5.Ghioni P, Bolognese F, Duijf PH, van Bokhoven H, Mantovani R, Guerrini L. Mol Cell Biol. 2002;22:8659–8668. doi: 10.1128/MCB.22.24.8659-8668.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ponassi R, Terrinoni A, Chikh A, Rufini A, Lena AM, Sayan BS, Melino G, Candi E. Biochem Pharmacol. 2006;72:1417–1422. doi: 10.1016/j.bcp.2006.07.031. [DOI] [PubMed] [Google Scholar]
- 7.Wu G, Nomoto S, Hoque MO, Dracheva T, Osada M, Lee CC, Dong SM, Guo Z, Benoit N, Cohen Y, et al. Cancer Res. 2003;63:2351–2357. [PubMed] [Google Scholar]
- 8.Serber Z, Lai HC, Yang A, Ou HD, Sigal MS, Kelly AE, Darimont BD, Duijf PH, van Bokhoven H, McKeon F, Dotsch V. Mol Cell Biol. 2002;22:8601–8611. doi: 10.1128/MCB.22.24.8601-8611.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Candi E, Rufini A, Terrinoni A, Dinsdale D, Ranalli M, Paradisi A, De Laurenzi V, Spagnoli LG, Catani MV, Ramadan S, et al. Cell Death Differ. 2006;13:1037–1047. doi: 10.1038/sj.cdd.4401926. [DOI] [PubMed] [Google Scholar]
- 10.Jacobs WB, Govoni G, Ho D, Atwal JK, Barnabe-Heider F, Keyes WM, Mills AA, Miller FD, Kaplan DR. Neuron. 2005;48:743–756. doi: 10.1016/j.neuron.2005.10.027. [DOI] [PubMed] [Google Scholar]
- 11.Jacobs WB, Kaplan DR, Miller FD. J Neurochem. 2006;97:1571–1584. doi: 10.1111/j.1471-4159.2006.03980.x. [DOI] [PubMed] [Google Scholar]
- 12.Laurikkala J, Mikkola ML, James M, Tummers M, Mills AA, Thesleff I. Development (Cambridge, UK) 2006;133:1553–1563. doi: 10.1242/dev.02325. [DOI] [PubMed] [Google Scholar]
- 13.Suh EK, Yang A, Kettenbach A, Bamberger C, Michaelis AH, Zhu Z, Elvin JA, Bronson RT, Crum CP, McKeon F. Nature. 2006;444:624–628. doi: 10.1038/nature05337. [DOI] [PubMed] [Google Scholar]
- 14.Sayan AE, Sayan BS, Findikli N, Ozturk M. Oncogene. 2001;20:5111–5117. doi: 10.1038/sj.onc.1204669. [DOI] [PubMed] [Google Scholar]
- 15.Pruneri G, Fabris S, Dell'Orto P, Biasi MO, Valentini S, Del Curto B, Laszlo D, Cattaneo L, Fasani R, Rossini L, et al. J Pathol. 2005;206:337–345. doi: 10.1002/path.1787. [DOI] [PubMed] [Google Scholar]
- 16.Chan WM, Siu WY, Lau A, Poon RY. Mol Cell Biol. 2004;24:3536–3551. doi: 10.1128/MCB.24.8.3536-3551.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Choi HR, Batsakis JG, Zhan F, Sturgis E, Luna MA, El-Naggar AK. Hum Pathol. 2002;33:158–164. doi: 10.1053/hupa.2002.30722. [DOI] [PubMed] [Google Scholar]
- 18.Hibi K, Trink B, Patturajan M, Westra WH, Caballero OL, Hill DE, Ratovitski EA, Jen J, Sidransky D. Proc Natl Acad Sci USA. 2000;97:5462–5467. doi: 10.1073/pnas.97.10.5462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Rufini A, Weil M, McKeon F, Barlattani A, Melino G, Candi E. Biochem Biophys Res Commun. 2006;340:737–741. doi: 10.1016/j.bbrc.2005.12.065. [DOI] [PubMed] [Google Scholar]
- 20.Candi E, Terrinoni A, Rufini A, Chikh A, Lena AM, Suzuki Y, Sayan BS, Knight RA, Melino G. J Cell Sci. 2006;119:4617–4622. doi: 10.1242/jcs.03265. [DOI] [PubMed] [Google Scholar]
- 21.Kubbutat MH, Vousden KH. Mol Cell Biol. 1997;17:460–468. doi: 10.1128/mcb.17.1.460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Munarriz E, Bano D, Sayan AE, Rossi M, Melino G, Nicotera P. Biochem Biophys Res Commun. 2005;333:954–960. doi: 10.1016/j.bbrc.2005.05.188. [DOI] [PubMed] [Google Scholar]
- 23.Sayan BS, Sayan AE, Knight RA, Melino G, Cohen GM. J Biol Chem. 2006;281:13566–13573. doi: 10.1074/jbc.M512467200. [DOI] [PubMed] [Google Scholar]
- 24.Ratovitski EA, Patturajan M, Hibi K, Trink B, Yamaguchi K, Sidransky D. Proc Natl Acad Sci USA. 2001;98:1817–1822. doi: 10.1073/pnas.98.4.1817. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.