Abstract
Antimicrobial peptides (AMPs) are an important component of innate immunity and have generated considerable interest as a potential new class of antibiotic. The biological activity of AMPs is strongly influenced by peptide-membrane interactions; however, for many of these peptides the molecular details of how they disrupt and/or translocate across target membranes are not known. CM15 is a linear, synthetic hybrid AMP composed of the first seven residues of the cecropin A and residues 2–9 of the bee venom peptide mellitin. Previous studies have shown that upon membrane binding CM15 folds into an α-helix with its helical axis aligned parallel to the bilayer surface and have implicated the formation of 2.2–3.8 nm pores in its bactericidal activity. Here we report site-directed spin labeling electron paramagnetic resonance studies examining the behavior of CM15 analogs labeled with a methanethiosulfonate spin label (MTSL) and a brominated MTSL as a function of increasing peptide concentration and utilize phospholipid-analog spin labels to assess the effects of CM15 binding and accumulation on the physical properties of membrane lipids. We find that as the concentration of membrane-bound CM15 is increased the N-terminal domain of the peptide becomes more deeply immersed in the lipid bilayer. Only minimal changes are observed in the rotational dynamics of membrane lipids, and changes in lipid dynamics are confined primarily to near the membrane surface. However, the accumulation of membrane-bound CM15 dramatically increases accessibility of lipid-analog spin labels to the polar relaxation agent, nickel (II) ethylenediaminediacetate, suggesting an increased permeability of the membrane to polar solutes. These results are discussed in relation to the molecular mechanism of membrane disruption by CM15.
INTRODUCTION
Antimicrobial peptides (AMPs) are an essential part of innate immune defense against microbial infection. Naturally occurring AMPs are basic peptides composed of 12–50 amino acids that are ubiquitously distributed throughout all kingdoms of life (1–5) with over 800 peptides now listed in AMP databases (6). As a group, AMPs display extensive sequence heterogeneity; however they do share a number of common characteristics, including a net positive charge of ≥+2 (with +4 to +6 being most common), ∼50%–70% hydrophobic amino acids, and a propensity to fold into amphipathic conformations in the presence of membranes (2,7,8). AMPs display a broad spectrum of antimicrobial activity against both Gram-negative and Gram-positive bacteria, fungi, and enveloped viruses (5). Importantly, they retain activity against antibiotic-resistant strains and do not readily elicit resistance (5,9).
Although potential intracellular targets are receiving increasing attention, the primary target for most AMPs appears to be the bacterial cytoplasmic membrane (4,5). AMPs bind strongly to membranes, dissipate transmembrane ionic potentials, and cause leakage of liposome-entrapped solutes. Peptides synthesized with D-amino acids retain activity comparable to their corresponding L-amino acid enantiomer, indicating that the interaction with their biological target is nonstereospecific (10–13). The bacterial cell surface has a highly negative surface charge density due to the presence of lipopolysaccharides and lipoteichoic acids in Gram-negative and Gram-positive organisms, respectively, and it is believed that electrostatic interactions make a significant contribution to AMP selectivity.
A large and important subclass of AMPs is linear peptides that fold into amphipathic α-helices upon membrane binding. For several of these peptides it has now been demonstrated that initial binding occurs with the helical axis parallel to the membrane surface (14–19) followed by insertion into the bilayer and disruption of the membrane permeability barrier as the amount of bound peptide exceeds a critical concentration (20–22). Several models have been proposed to describe the molecular events involved in AMP-mediated membrane disruption, including the formation of barrel-stave peptide channels, induction of peptide-lipid toroidal pores, “sinking-raft” and “micellar aggregate” models, and a detergent-like carpet mechanism (2,4,5,19,20,23–25). It is likely that the precise mechanism of membrane disruption depends on the specific peptide as well as the composition of the target membrane (22,26,27).
Our studies have focused on a linear, synthetic hybrid AMP composed of the first seven residues of cecropin A and residues 2–9 of the bee venom peptide mellitin. This 15-residue peptide, designated CM15, retains the two-domain structure of native cecropins (28,29), with a highly cationic N-terminal region and a mostly hydrophobic C-terminal region. CM15 displays potent, broad-spectrum antimicrobial activity yet lacks the strong hemolytic activity of mellitin (30). We have previously shown that CM15 folds into an α-helix upon membrane binding (16,19) and that at low peptide/lipid (P/L) ratios (i.e., under initial binding conditions) the helical axis is positioned ∼5 Å below the hydrophobic interface of the membrane and aligned parallel to the bilayer surface (16). Osmoprotection studies with live bacteria indicate that cell killing by CM15 is mediated by the formation of membrane pores with a diameter of 2.2–3.8 nm (31); however, nothing is known about the intermediate stages between initial binding and pore formation.
In this study we have used site-directed spin labeling (SDSL) electron paramagnetic resonance (EPR) spectroscopy to investigate the behavior of a spin-labeled analog of CM15 as a function of increasing peptide concentration and utilized phospholipid-analog spin labels to examine the effects of CM15 binding and accumulation on physical properties of membrane lipids. We find that as the concentration of membrane-bound CM15 is increased, the N-terminal domain of the peptide becomes more deeply immersed in the lipid bilayer. Changes in the rotational dynamics of membrane lipids are minimal and confined primarily to near the membrane surface. However, peptide binding dramatically increases interaction of the lipid-analog spin labels with the polar relaxation agent NiEDDA (nickel (II) ethylenediaminediacetate), indicating that there are significant changes in the physical state of the lipid bilayer that are not readily detected by methods that examine motional dynamics. These results are discussed in relation to the molecular mechanism of membrane disruption by CM15.
MATERIALS AND METHODS
Materials
Phospholipids POPE (1-palmitoyl-2-oleoylphosphatidylethanolamine), POPG (1-palmitoyl-2-oleoylphosphatidylglycerol), tetraoleoyl-cardiolipin (CL), and n-PCSL (1-(n-doxylpalmitoyl)-2-stearoylphosphatidylcholine; n = 5, 7, 12) spin labels were obtained from Avanti Polar Lipids (Alabaster, AL). NiEDDA was synthesized according to a protocol provided by Dr. Christian Altenbach (Jules Stein Eye Institute, UCLA School of Medicine, Los Angeles, CA). The methanethiosulfonate spin label, MTSL (1-oxy-2,2,5,5-tetramethylpyrroline-3-methyl methanethiosulfonate), and its brominated derivative, BrMTSL (4-bromo-(1-oxy-2,2,5,5-tetramethylpyrroline-3-methyl) methanethiosulfonate) (Fig. 1), were obtained from Toronto Research Chemicals (North York, ON, Canada).
FIGURE 1.
Structures of the MTSL and BrMTSL spin labels and the resulting side chains produced by reaction with the peptide cysteine residue.
Peptide synthesis and spin labeling
Peptides used in this study included the “wild-type” cecropin-mellitin hybrid peptide CM15 with N-terminal acetylation and C-terminal amidation (Ac-KWKLFKKIGAVLKVL-amide) and an analog containing a leucine-to-cysteine substitution at position 4 (Ac-KWKCFKKIGAVLKVL-amide, designated C4). Peptides were synthesized by standard n-(9-fluororenyl)methoxycarbonyl (Fmoc) solid phase synthesis methods on Rink amide p-methylbenzhydrylamine (MBHA) resin as previously described (16). Crude peptides were purified by reverse-phase semipreparative high performance liquid chromatography (HPLC) on a 10-μm, 1.0 × 2.5-cm C8 column (Vydac, Hesperia, CA) using a linear gradient of 10%–80% acetonitrile/0.1% trifluoroacetic acid (TFA) in water/0.1% TFA over 56 min. Upon elution the peptides were lyophilized and stored at −20°C until use. For spin labeling, the C4 peptide was resuspended in 50 mM MOPS (3-(N-morpholino)propanesulfonic acid) pH 6.8 and reacted with a fivefold molar excess of spin label (MTSL or BrMTSL) for 3 h at room temperature. To remove excess spin label the peptide was bound to a 2-ml column of SP Sepharose (Amersham, Buckinghamshire, UK), washed with 50 ml of 50 mM MOPS, pH 6.8, and eluted with 6 M guanidine hydrochloride in the same buffer. The spin labeled peptide was then repurified by HPLC as described above, lyophilized, and stored at −20°C. Final purity was confirmed by matrix-assisted laser desorption ionization-time of flight mass spectrometry at the Medical College of Wisconsin Protein and Nucleic Acid facility. Peptides were rehydrated in 50 mM MOPS, pH 6.8 at time of use, and peptide concentrations were determined based on the absorbance of the single tryptophan residue using an extinction coefficient of 5.69 mM−1 cm−1 (32).
Liposome preparation
Liposomes were composed of POPE/POPG/CL (molar ratio of 70:25:5), in agreement with the composition of bacterial inner membrane lipids (33). Lipids in the desired molar ratio were dried down from chloroform stock solutions under a stream of nitrogen gas and then dried overnight under vacuum. The resulting lipid film was hydrated by the addition of 50 mM MOPS, pH 6.8 to give a concentration of 50 mM phospholipid. Large unilamellar vesicles (LUVs) were prepared by freeze-thawing this lipid suspension five times, followed by extrusion through 200-nm polycarbonate membrane filters using a miniextruder syringe device (Avanti Polar Lipids). Final lipid concentration was measured by the method of Stewart (34). LUVs containing 1 mol % of 5-, 7-, or 12PCSL were prepared as described above.
Circular dichroism
Far ultraviolet circular dichroism (CD) spectra were obtained at room temperature on a Jasco (Tokyo, Japan) J-710 spectropolarimeter at a scan rate of 50 nm/min, 0.5 s response time, 1 nm pitch, and 1 nm bandwidth. Peptides were at a concentration of 0.1 mM (∼0.2 μg/ml) in 5 mM phosphate buffer pH 7 (5P7), 50% TFE in 5P7, or in the presence of 10 mM LUVs in 5P7. Spectra taken in the presence of liposomes were truncated at 200 nm due to high background scattering and saturation of the detector below this wavelength. Secondary structure content was estimated via the DICHROWEB server (35) using the Selcon3 and K2D analysis programs.
EPR spectroscopy
Conventional continuous-wave (CW) EPR spectra were recorded on a Bruker (Billerica, MA) Elexsys E-500 X-band spectrometer equipped with a super-high-Q cavity. Room temperature spectra were recorded at a microwave power of 10 mW, using a 100 kHz, 1.0 G field modulation. Spectra taken at 200 K were recorded at a microwave power of 2 mW. For peptide-binding assays a constant amount of C4MTSL or C4BrMTSL peptide was mixed with various concentrations of LUVs to give the desired lipid/peptide (L/P) ratios and incubated at room temperature overnight. Final peptide concentration was ∼50 μM, and the final sample volume was 30 μl. The fraction of peptide remaining free in solution was calculated based on the peak-to-peak amplitude of the high-field (MI = −1) line as previously described (16). From the fraction of bound peptide and known total peptide concentration, the concentrations of membrane-bound peptide (Cb), peptide remaining free in solution (Cf), and the molar ratio of bound peptide/lipid (Cb/L) were calculated. A plot of Cb/L against Cf yields the apparent partition coefficient, Kp, where Kp [L] = Cb/Cf.
To assess the rotational mobility of 12PCSL, the apparent rotational correlation time, τc, was determined according to (36):
 |
where ΔH0 is the peak-to-peak width of the center line in gauss, A0 is the amplitude of the center line, and A−1 is the amplitude of the high field line (see Fig. 7). The rotational correlation time is inversely related to the motional rate, i.e., an increase in τ indicates slower motion.
FIGURE 7.
EPR spectra of (A) 5PCSL, (B) 7PCSL, and (C) 12PCSL in PE/PG/CL liposomes. In each set of spectra the upper spectrum is in the absence of peptide and the lower spectrum is in the presence of wild-type CM15 at an L/P ratio of 25:1. The increase in 2T// observed for 5- and 7PCSL indicate a decrease in rotational mobility. For 12PCSL there is no measurable difference in the width of the center line (ΔH0) or in the relative line amplitudes (A0 and A−1). Scan widths are 100 G for 5- and 7PCSL, and 80 G for 12PCSL.
The accessibility of spin-labeled peptides and PCSLs to the relaxation agents O2 and NiEDDA was determined by CW power saturation. Liposomes containing either 5-, 7-, or 12PCSL and native CM15 or spin labeled C4 and unlabeled LUVs were mixed to obtain the desired L/P ratios. Samples (final volume 10 μl) with or without 20 mM NiEDDA were incubated at room temperature overnight and then placed into gas-permeable TPX capillaries (Molecular Specialties, Milwaukee, WI). EPR spectra for power saturation studies were obtained on a Varian E-102 Century series spectrometer equipped with an X-band two-loop one-gap resonator (Molecular Specialties). The saturation parameter P1/2 was determined under various conditions (under N2, saturated with air (20% O2) and under N2 with samples containing 20 mM NiEDDA) by measuring the amplitude of the center line at a series of microwave powers from 0.25 to 64 mW as described previously (37,38). The change in the saturation parameter, ΔP1/2, in the presence of O2 or NiEDDA is directly proportional to the bimolecular collision rate with the respective paramagnetic relaxation agent (37). The depth parameter, Φ, where
 |
was calculated as a measure of the bilayer immersion depth of the spin label side chain (16,38,39).
RESULTS
Secondary structure of the spin-labeled peptides
One conserved characteristic of linear cationic AMPs is their ability to fold into amphipathic secondary structures upon membrane binding (1–5). We used CD to evaluate secondary structure formation by C4MTSL and C4BrMTSL. Previous CD studies have shown that wild-type CM15 adopts an α-helical secondary structure upon membrane binding (19), and a nitroxide-scanning study of CM15 was also consistent with the formation of a continuous α-helix in the membrane-bound state (16). CD spectra of C4BrMTSL in aqueous solution, in the helix-promoting solvent trifluoroethanol (TFE), and in the presence of LUVs are shown in Fig. 2. C4BrMTSL exhibits little or no secondary structure in solution but adopts a significant degree of α-helical structure in the presence of 50% (v/v) TFE or when bound to LUVs (helix content estimated at 86% and 58%, respectively). CD spectra of C4MTSL were quite similar in appearance (data not shown) although estimates of α-helical content were consistently lower (68% and 52% in TFE and bound to LUVs, respectively). Helix content for wild-type CM15 in 50% TFE and bound to LUVs was 65% and 58%, respectively (H. Sato and J. B. Feix, unpublished data). These results indicate that the spin-labeled C4 peptides adopt an α-helical secondary structure upon membrane binding that is similar to wild-type CM15 and suggest that the BrMTSL side chain may facilitate or stabilize helix formation.
FIGURE 2.
CD spectra of C4BrMTSL in 5 mM phosphate buffer (crosses) in 50% TFE (squares) and in the presence of LUVs at an L/P ratio of 100:1 (triangles). The peptide concentration was 0.1 mM (∼0.2 mg/ml) in each sample.
Binding and mobility of the spin-labeled peptides
To further examine peptide-membrane interactions by EPR spectroscopy, we utilized the C4MTSL and C4BrMTSL analogs of CM15. Since the spin label side chain is relatively hydrophobic (40), spin labeling at this site (normally a leucine) is a conservative substitution. EPR spectra of C4MTSL in aqueous solution, in 50% TFE, and bound to liposomes at various L/P ratios are shown in Fig. 3. In solution, the EPR spectrum of C4MTSL consists of three narrow lines, typical of an unstructured peptide with few constraints on the motion of the spin label side chain and consistent with the observed random coil CD spectrum. Addition of 50% TFE to induce α-helix formation produced only a slight decrease in spin label motion (Fig. 3 B). However, upon addition of liposomes a dramatic broadening of the spectrum was observed (Fig. 3, C–E), indicating a significant reduction in spin label mobility upon membrane binding. Similar experiments were carried out using C4 labeled with BrMTSL (Fig. 4). In aqueous solution the EPR spectrum of C4BrMTSL was quite similar to that observed for C4MTSL, with only a slightly longer rotational correlation time, τc (0.41 ns for C4MTSL and 0.56 ns for C4BrMTSL). Induced α-helix formation by the addition of 50% TFE moderately decreased the motional freedom of the spin label side chain (Fig. 4 B) and to a somewhat greater degree than for the nonbrominated spin label (τc values 0.77 ns and 1.17 ns for C4MTSL and C4BrMTSL, respectively). As with C4MTSL, binding of C4BrMTSL to LUVs resulted in a significant reduction in spin label mobility (Fig. 4, C–E). Previous studies have shown that for labeling sites in α-helices, reduced mobility of MTSL and its analogs results from interaction between the substituent at the 4′ position of the nitroxide ring and a Cα hydrogen of the peptide backbone (41,42). Replacing the 4′-hydrogen of MTSL with the large bromine atom substantially increases this interaction (43). Thus, even in these small peptides, BrMTSL can be used to reduce the flexibility of the spin label side chain, tethering it more closely to the peptide backbone relative to MTSL under the same conditions (compare Figs. 3 and 4). The relatively low mobility of C4BrMTSL in the presence of LUVs also indicates that the peptide itself has little flexibility when membrane bound.
FIGURE 3.
EPR spectra of C4-MTSL in (A) 50 mM MOPS buffer, (B) in 50% TFE, and in the presence of PE/PG/CL (70:25:5) LUVs at L/P ratios of (C) 250:1, (D) 50:1, and (E) 25:1. Spectra for the membrane-bound peptide are presented at a 10-fold higher gain. The scan width is 100 G.
FIGURE 4.
EPR spectra of C4-BrMTSL in (A) 50 mM MOPS buffer, (B) in 50% TFE, and in the presence of PE/PG/CL (70:25:5) LUVs at L/P ratios of (C) 250:1, (D) 50:1, and (E) 25:1. Scan width 100 G. Spectra for the membrane-bound peptide are presented at a 10-fold higher gain. Binding to the LUVs results in a pronounced reduction in spin label mobility, and the BrMTSL side chain is significantly more immobilized than the nonbrominated MTSL side chain in the membrane-bound state (compare with Fig. 3).
Both C4MTSL and C4BrMTSL bound to POPE/POPG/CL (70:25:5) liposomes with high affinity. Binding isotherms, determined as described previously (16), yielded partition coefficients (Kp) in 50 mM MOPS buffer of 1.9 × 105 M−1 and 2.3 × 105 M−1 for C4MTSL and C4BrMTSL, respectively. These values are significantly higher than previously reported for POPE/POPG (80:20) liposomes in 50 mM MOPS, 0.1 M KCl (16), consistent with the higher content of anionic lipids and lower ionic strength in this system. As indicated by the high Kp, the peptide was essentially fully bound even at high P/L ratios (e.g., at 2.5 mM lipid and 100 μM peptide, L/P = 25:1, the calculated ratio of bound/free peptide is 575:1).
There were no evident changes in line width or normalized amplitude for either C4MTSL or C4BrMTSL as the concentration of membrane-bound peptide increased (decreasing the L/P ratio from 250:1 to 25:1, Figs. 3 and 4 and data not shown), indicating an absence of spin-spin interactions. As noted above, both C4MTSL and C4BrMTSL were essentially fully bound even at an L/P of 25:1, as evidenced by the lack of a sharp component in the EPR spectrum due to unbound peptide (Figs. 3 and 4). We also examined EPR spectra for the membrane-bound peptides as a function of L/P ratio in frozen samples. Freezing should eliminate residual rotational motion as well as stabilize the association of any peptide oligomers that might exist, enhancing the ability to observe dipole-dipole interactions. However, no changes in line shape or relative line widths were observed over a wide range of L/P ratios (Fig. 5). The lack of observed dipolar coupling indicates that spin labels are separated by at least 20 Å at all L/P ratios examined. Alternatively, a dynamic equilibrium between oligomeric channels and monomers could limit our ability to detect spin-spin interactions, especially if the oligomeric species represents a minor population of the total peptide. Motional parameters (i.e., central line width, outer hyperfine splittings) also remained unchanged as a function of bound peptide concentration (data not shown), indicating that rotational motion was not altered by the accumulation of bound peptide. The absence of line broadening and lack of motional restriction at increasing peptide concentrations argue against direct peptide-peptide contact, at least in the vicinity of the spin label side chain.
FIGURE 5.
EPR spectra of C4MTSL bound to membranes in the frozen state. C4MTSL was mixed with LUVs at final L/P ratios of (A) 160:1, (B) 80:1, (C) 40:1, and (D) 20:1 and allowed to equilibrate at room temperature for 1 h. The samples were then placed in the EPR cavity and brought to 200 K. Scan widths are 160 G. No changes in line widths or relative amplitudes were observed as a function of the L/P ratio.
Accessibility of the spin labeled peptide
To determine the accessibility of the spin labeled side chain of the peptides, we used power saturation EPR to examine interaction with the paramagnetic relaxation agents O2 and NiEDDA. NiEDDA is an uncharged, polar reagent that penetrates only weakly into phospholipid bilayers, with diminishing concentrations at increasing bilayer depths (39). The NiEDDA accessibility parameter, ΔP1/2(NiEDDA), is proportional to the bimolecular collision rate between the spin label and the relaxation agent and, therefore, a direct reflection of the local NiEDDA concentration. Conversely, molecular oxygen is nonpolar and partitions favorably into lipid bilayers, with O2 concentration increasing as a function of bilayer depth (39,44). As seen in Fig. 6 A, interaction with NiEDDA gradually diminished with increasing concentrations of bound peptide for both C4-MTSL and C4-BrMTSL. The decrease in ΔP1/2(NiEDDA) was approximately linear up to an L/P ratio of 30:1, followed by a sharp decrease at L/P = 25:1. Both spin-labeled peptides also showed a trend of increasing O2 accessibility with increasing concentrations of bound peptide (Fig. 6 B). Interaction with O2 was much greater than with NiEDDA under all conditions (note the different vertical scales in Fig. 6, A and B), confirming the hydrophobic localization of the MTSL and BrMTSL side chains (16). The ratio of accessibility parameters for NiEDDA and O2 can be used to calculate an EPR depth parameter, Φ (see Materials and Methods). Although significant changes in the accessibilities of standard lipid-analog spin labels (discussed below) precluded actual depth calculations, depth parameters for both spin-labeled peptide analogs indicated a gradually increasing immersion depth at increasing peptide concentrations, again with a sharp transition between L/P ratios of 30:1 and 25:1 (Fig. 6 C). Except at the highest concentration of bound peptide, C4BrMTSL consistently exhibited lower accessibility to NiEDDA, greater O2 accessibility, and an increased depth parameter compared to C4MTSL, consistent with the greater hydrophobicity of the brominated MTSL side chain.
FIGURE 6.
Accessibilities of C4-MTSL (squares) and C4-BrMTSL (triangles) as a function of the L/P ratio. (A) The change in the EPR saturation parameter, P1/2, in the presence of 20 mM NiEDDA, (B) the change in P1/2 upon equilibration with air (20% O2), and (C) the EPR depth parameter, calculated as described in the text. Error bars indicate standard deviations from at least three separate measurements.
Effect of CM15 binding on membrane lipids
To assess the effects of CM15 binding on the lipid phase of the bilayer, we examined the motion and accessibility parameters for phosphatidylcholine spin labels (PCSLs) with the nitroxide moiety positioned at various depths along the alkyl chain. CW-EPR spectra for 5-, 7-, and 12PCSL in POPE/POPG/CL (70:25:5) LUVs with and without bound wild-type CM15 peptide are shown in Fig. 7. For 5PCSL, a plot of the motional parameter 2T// as a function of the L/P ratio suggests a slight decrease in motion (i.e., an increase in 2T//) with increasing concentration of bound peptide (Fig. 8, Table 1). A similar trend is observed for 7PCSL, although changes are smaller than for 5PCSL (Fig. 8, Table 1). Because 12PCSL undergoes more rapid motion than 5- or 7PCSL, the 2T// parameter is less sensitive to changes in mobility than the width of the center line (ΔH0) or relative line amplitudes, which can be used to calculate an empirical rotational correlation time, τc (see Materials and Methods). For 12PCSL, increasing concentrations of bound peptide produced no significant changes in either ΔH0 or τc (Table 2). These results indicate that perturbation of bilayer lipid motional dynamics upon accumulation of bound CM15 is observed only near the membrane surface, and even those effects are minimal.
FIGURE 8.
Effect of wild-type CM15 binding on the motion of PCSLs. CM15 was added to PE/PG/CL (70:25:5) LUVs containing 1 mol % PCSL and equilibrated at room temperature overnight before recording the EPR spectrum. The motion parameter 2T// for 5PCSL (squares) and 7PCSL (triangles) is plotted as a function of the lipid ratio (with ∞ corresponding to control values in the absence of peptide). An increase in 2T// indicates decreased mobility of the spin label. Error bars indicate the standard deviation from at least three separate experiments.
TABLE 1.
Motional parameters for 5PCSL and 7PCSL
| 2T// (G)
|
||
|---|---|---|
| Lipid/peptide | 5PCSL | 7PCSL |
| No peptide | 54.2 ± 0.2 | 52.0 ± 0.1 |
| 250:1 | 54.9 ± 0.3 | 53.0 ± 0.3 |
| 100:1 | 54.7 ± 0.1 | 52.4 ± 0.2 |
| 80:1 | 55.4 ± 0.7 | 52.3 ± 0.1 |
| 50:1 | 55.0 ± 0.9 | 53.2 ± 0.6 |
| 40:1 | 55.2 ± 0.5 | 53.0 ± 0.9 |
| 30:1 | 55.5 ± 0.8 | 52.8 ± 0.7 |
| 25:1 | 56.1 ± 1.2 | 52.8 ± 0.6 |
Motional parameters for 5- and 7PCSL as a function of L/P molar ratio. Mean ± SD of 2T// are from at least three independent experiments. All values are in gauss (G).
TABLE 2.
Motional parameters for 12PCSL
| Lipid/peptide | ΔH0 (G) | τ (nsec) |
|---|---|---|
| No peptide | 3.72 ± 0.5 | 3.63 ± 0.04 |
| 250:1 | 3.71 ± 0.5 | 3.52 ± 0.04 |
| 100:1 | 3.74 ± 0.7 | 3.55 ± 0.06 |
| 80:1 | 3.66 ± 0.4 | 3.50 ± 0.03 |
| 50:1 | 3.72 ± 0.5 | 3.53 ± 0.04 |
| 40:1 | 3.88 ± 0.8 | 3.60 ± 0.07 |
| 30:1 | 3.44 ± 0.5 | 3.63 ± 0.04 |
| 25:1 | 3.49 ± 0.5 | 3.63 ± 0.04 |
Motional parameters for 12PCSL as a function of L/P molar ratio. Mean ± SE of the peak-peak width of the center line (ΔH0) and rotational correlation time (τ) are from two independent experiments.
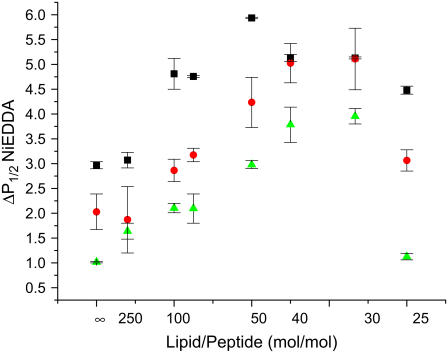
The binding of CM15 induced far more pronounced changes in PCSL accessibilities, significantly altering the accessibility of all three PCSLs to NiEDDA (Fig. 9). In the absence of peptide, NiEDDA accessibility parameters exhibit the expected profile, with the accessibility of 5PCSL > 7PCSL > 12PCSL (Fig. 9). At concentrations of CM15 giving P/L ratios in the range 0–0.02 (L/P = ∞ to 50:1), an increase in NiEDDA interaction is observed for all three of the PCSLs, indicating an increase in NiEDDA concentration at all depths of the bilayer. At L/P ratios in the range 40:1–30:1, interactions between NiEDDA and the PCSLs plateau, with no difference observed in the accessibilities of 5- and 7PCSL. These results are consistent with a pronounced disruption of the membrane permeability barrier, particularly near the membrane surface, and may also indicate a thinning of the bilayer (see Discussion). Remarkably, as the peptide concentration is increased even further, to an L/P of 25:1, NiEDDA accessibility of the PCSLs reverts to a profile similar to that observed in the absence of peptide.
FIGURE 9.
Effect of wild-type CM15 binding on the accessibility of PCSLs to NiEDDA. Large unilamellar liposomes (PE/PG/CL, molar ratio 70:25:5) containing 1 mol % of 5PCSL (squares), 7PCSL (circles), or 12PCSL (triangles) were mixed with CM15 to give the desired L/P ratio, incubated overnight at room temperature and the CW saturation parameter P1/2 determined under N2. Final NiEDDA concentration was 20 mM.
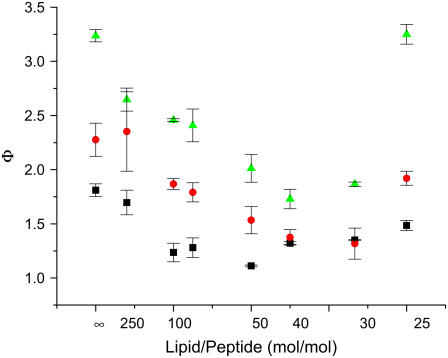
Peptide binding had little effect on oxygen accessibility parameters for the PCSLs (data not shown), so that a plot of Φ against the L/P ratio is approximately the inverse of the dependence seen with NiEDDA (Fig. 10). In the absence of peptide we observed the expected gradient in the depth parameter, with 12PCSL > 7PCSL > 5PCSL and significant differences between the Φ values of each of the labels. Depth parameters decreased (reflecting increased accessibility to NiEDDA) for all three of the PCSLs up to an L/P ratio of 50:1 and then plateaued at L/P ratios between 50:1 and 30:1. In the L/P range of 40:1–30:1, the difference in Φ values between 12PCSL and the other PCSLs was much less than in the absence of peptide, and no difference in Φ was observed between 5- and 7PCSL. Again, as the L/P ratio was further decreased to 25:1, the PCSL depth profile reverted to a pattern similar to that observed in the absence of peptide.
FIGURE 10.
Effect of wild-type CM15 binding on the EPR depth parameter. The depth parameter for 12PCSL (triangles), 7PCSL (circles), and 5PCSL (squares) is plotted against the L/P molar ratio. The spin labels were at a concentration of 1 mol % in PE/PG/CL LUVs. Error bars represent the standard deviation from at least three independent experiments.
DISCUSSION
AMPs interact with membranes by a variety of mechanisms that can result in disruption of bilayer structure and loss of the differential permeability barrier. Several studies have indicated that for most linear, α-helical AMPs, including CM15, initial interactions (i.e., at low concentrations of membrane-bound peptide) occur primarily near the membrane surface, with the peptide aligned parallel to the plane of the bilayer (14–19). Peptide insertion near the hydrophobic-hydrophilic interface of the membrane is postulated to result in expansion of the outer leaflet of the bilayer, with continued accumulation of bound peptide leading to membrane thinning as a prelude to pore formation or detergent-like disintegration of the bilayer (21,45–47). The goals of this study were to examine peptide-induced changes in the lipid phase of the membrane and to explore potential changes in peptide localization and aggregation state at increasing concentrations of membrane-bound CM15.
Using phosphatidylcholine-analog spin labels, we find that CM15-induced changes in lipid motional dynamics are minimal and occur primarily near the membrane surface, even at relatively high concentrations of membrane-bound peptide (i.e., up to 4 mol %). For 5PCSL, spin label mobility gradually decreased across the entire range of P/L ratios examined, as might be expected if peptide insertion increased the membrane lateral pressure near the interfacial region. This result is consistent with a previous spin labeling study of a 34-residue cecropin B analog in which decreased mobility of PCSLs upon peptide binding to liposomes was also observed (48). In contrast to CM15 however, the most active cecropin B analog caused decreased lipid mobility (i.e., increased order) at all depths of the bilayer (48), which may reflect a fundamental difference in membrane interactions between the shortened 15-residue peptide studied here and full-length cecropins. The observation that lipid perturbation by CM15 is restricted to near the membrane surface is consistent with a toroidal-pore model for permeabilization, as proposed for other linear α-helical AMPs such as the magainins (49), magainin derivatives (50), and LL-37 (15,51). In this model, peptides remain associated with lipid headgroups even as they transition from an orientation parallel to the membrane surface to an alignment along the bilayer normal (22,23).
In contrast to the minimal effects on lipid motion, peptide binding significantly increased accessibility to the polar relaxation agent NiEDDA at all depths of the membrane. For example, PCSL accessibility to NiEDDA increased by 1.5–2-fold at an L/P ratio of 100:1, with 3–4-fold increases observed at an L/P of 30:1 (Fig. 9). Accessibility, as measured by changes in P1/2, is a reflection of the diffusion-concentration product of the relaxation agent. Given the absence of large changes in membrane fluidity, it seems unlikely that the diffusion coefficient of NiEDDA has significantly increased. Consequently the observed increases in accessibility most likely indicate an increase in NiEDDA concentration within the bilayer, reflecting a disruption of the permeability barrier to polar solutes. The observed changes in accessibility to the polar solute NiEDDA clearly indicate that interaction of CM15 with the membrane alters the physical properties of the lipid phase of the bilayer. Such changes are not readily apparent using methods (e.g., 31P- and 2H-NMR and spin label motion analysis) that examine lipid dynamics.
Differences in NiEDDA accessibility at different bilayer depths (5 > 7 > 12PCSL) were maintained up to an L/P ratio of 50:1. However, at concentrations of bound peptide with L/P ratios in the range of 40:1–30:1 differences between 12PCSL and the other two PC spin labels were diminished, and no difference was observed between 5- and 7PCSL. We speculate that this may be a reflection of membrane thinning. X-ray diffraction (21,45,46) and atomic force microscopy (47) studies have provided strong evidence that peptide accumulation leads to membrane thinning before peptide reorientation and insertion into the bilayer. Our data showing diminished differences in the NiEDDA accessibilities of 5-, 7-, and 12PCSL may be an indication of membrane thinning by CM15 (thus reducing differences in the apparent depths of the PCSL isomers), although further studies under conditions where thinning of the bilayer is known to occur are needed to support this conclusion. Our data also indicate that CM15-induced changes in bilayer structure undergo an abrupt transition at L/P ratios of 30:1–25:1. This is consistent with a two-state mechanism proposed by Huang and co-workers for the interaction of several AMPs with lipid bilayers, based primarily on oriented CD studies in which peptides associate parallel to the membrane surface up to a given critical concentration, at which point they reorient along the bilayer normal and insert into the membrane (20,22). Observed threshold concentrations for various AMPs depend on both the peptide and the composition of the target membrane (22). In this work, reversion of the NiEDDA accessibility profile for PCSL spin labels at an L/P of 25:1 to one similar to that observed in the absence of peptide suggests that as this critical concentration of bound peptide is reached CM15 may become sequestered (i.e., into localized pores), leaving the remaining bulk lipid phase relatively unperturbed.
With regard to the effects of CM15 accumulation on peptide localization, our data indicate that the spin label side chain at position 4 of the CM15-C4 analogs becomes more deeply immersed in the membrane as the concentration of bound peptide is increased. Decreased interaction with NiEDDA and a concomitant increased interaction with O2 occurred gradually across the full range of bound peptide concentrations, again followed by an abrupt transition between L/P ratios of 30:1 and 25:1. The decrease in NiEDDA accessibility is even more remarkable given that the overall bulk lipid concentration of this polar relaxation agent increases with peptide binding at all depths of the bilayer, as indicated by the PCSL data. The decreased interaction of the C4-spin label side chain with NiEDDA may reflect a “sinking” of the entire peptide (25) or a reorientation of the peptide so that the N-terminal domain is more deeply buried. Alternatively, a localized thinning of the bilayer in the region of the peptide might allow the C4 side chain, positioned on the nonpolar face of the amphiphathic helix, to reach a more hydrophobic region of the bilayer while maintaining surface exposure of the hydrophilic face of the helix. Such localized effects on bilayer structure have recently been postulated (22). The formation of peptide aggregates, sequestering the spin label away from NiEDDA, could also produce a similar effect, but this seems unlikely given a) the lack of any observed changes in spin label mobility, b) the absence of spin-spin interaction, and c) the fact that interaction with O2 increases with the accumulation of bound peptide.
One of the goals of this study was to investigate association between membrane-bound peptides in the formation of transmembrane pores. Previous studies using osmoprotectants have provided strong evidence for pore formation by CM15 in intact Escherichia coli cells, with an estimated pore diameter in the range 2.2–3.8 nm (31). However, no spin-spin interactions were observed even at the highest concentrations of membrane-bound peptide. In addition, there were no evident changes in line width as a function of P/L ratio in the frozen state, again indicating a lack of dipole-dipole interaction. These results suggest that no stable aggregates are formed in which spin labels are within the ∼20 Å range of dipolar coupling for CW EPR (43,52) and would seem to rule out the existence of stable “barrel-stave”-type channels, which should place labels within this distance. This observed absence of dipolar broadening is best explained by formation of a toroidal pore-type structure in which peptides are separated by intervening phospholipids. However, we cannot rule out the formation of transient pores or channel formation by only a small fraction of membrane-bound peptides. In addition, it may be that the labeling site employed in this study—positioned on the nonpolar face of the peptide—may not be favorably located. The leucine 4 to Cys-MTSL substitution was chosen as a conservative (hydrophobic-hydrophobic) replacement; however CM15 analogs spin labeled at sites on the more polar face of the peptide (e.g., at Ala-10) do retain biological activity (H. Sato and J. B. Feix, unpublished data) and may be better suited for studies of channel formation. Further studies using pulsed EPR methods, which extend sensitivity to dipolar interactions up to 50 Å or more (53,54), and/or the use of other labeling sites (i.e., those on the hydrophilic face of the peptide) may allow us to further characterize the nature of the putative transmembrane pore.
CONCLUSION
In summary, these studies provide further insights into the disruptive effects of AMPs on membrane bilayers. We have shown that as the concentration of membrane-bound CM15 increases, the nonpolar face of the peptide samples an increasingly hydrophobic environment. There was no evidence of peptide-peptide association, suggesting that transmembrane pores formed by the peptide are either transient or that individual peptide monomers are separated by >20 Å. Binding of CM15 to model membranes with a lipid composition mimicking the bacterial inner membrane caused only slight perturbations in membrane lipid dynamics, and changes in lipid dynamics were observed only for 5PCSL, suggesting that CM15 remains in a region of the bilayer near the hydrophilic interface even as the concentration of bound peptide is increased. In contrast, CM15 binding significantly increased the accessibility of lipid-analog spin labels to the polar solute NiEDDA. Accessibility studies with both spin-labeled peptide and lipid-analog spin labels indicated an abrupt structural change at an L/P ratio of ∼25:1. Overall, these results are most consistent with the toroidal pore model as the mechanism of bilayer disruption by CM15.
Acknowledgments
We thank Dr. Candice Klug and Dr. Hiromi Sato for critically reading the manuscript.
This work was supported by National Institutes of Health grant GM068829. The National Biomedical EPR Center is supported by National Institutes of Health grant EB001980.
Editor: Lukas K. Tamm.
References
- 1.Boman, H. G. 1995. Peptide antibiotics and their role in innate immunity. Annu. Rev. Immunol. 13:61–92. [DOI] [PubMed] [Google Scholar]
- 2.Hancock, R. E. W., and D. S. Chapple. 1999. Peptide antibiotics. Antimicrob. Agents Chemother. 43:1317–1323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Zasloff, M. 2002. Antimicrobial peptides of multicellular organisms. Nature. 415:389–395. [DOI] [PubMed] [Google Scholar]
- 4.Brogden, K. A. 2005. Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria? Nat. Rev. Microbiol. 3:238–250. [DOI] [PubMed] [Google Scholar]
- 5.Jenssen, H., P. Hamill, and R. E. Hancock. 2006. Peptide antimicrobial agents. Clin. Microbiol. Rev. 19:491–511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Tossi, A. http://www.bbcm.univ.trieste.it/∼tossi/pag2.htm.
- 7.Giangaspero, A., L. Sandri, and A. Tossi. 2001. Amphipathic alpha-helical antimicrobial peptides. Eur. J. Biochem. 268:5589–5600. [DOI] [PubMed] [Google Scholar]
- 8.Zelezetsky, I., and A. Tossi. 2006. Alpha-helical antimicrobial peptides—using a sequence template to guide structure-activity relationship studies. Biochim. Biophys. Acta. 1758:1436–1449. [DOI] [PubMed] [Google Scholar]
- 9.Zhang, L., J. Parente, S. M. Harris, D. E. Woods, R. E. Hancock, and T. J. Falla. 2005. Antimicrobial peptide therapeutics for cystic fibrosis. Antimicrob. Agents Chemother. 49:2921–2927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wade, D., A. Boman, B. Wahlin, C. M. Drain, D. Andreu, H. G. Boman, and R. B. Merrifield. 1990. All-D amino acid-containing channel-forming antibiotic peptides. Proc. Natl. Acad. Sci. USA. 87:4761–4765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bessalle, R., A. Kapitkovsky, A. Gorea, I. Shalit, and M. Fridkin. 1990. All-D-magainin: chirality, antimicrobial activity and proteolytic resistance. FEBS Lett. 274:151–155. [DOI] [PubMed] [Google Scholar]
- 12.Bland, J. M., A. J. De Lucca, T. J. Jacks, and C. B. Vigo. 2001. All-D-cecropin B: synthesis, conformation, lipopolysaccharide binding, and antibacterial activity. Mol. Cell. Biochem. 218:105–111. [DOI] [PubMed] [Google Scholar]
- 13.Merrifield, R. B., P. Juvvadi, D. Andreu, J. Ubach, A. Boman, and H. G. Boman. 1995. Retro and retroenantio analogs of cecropin-melittin hybrids. Proc. Natl. Acad. Sci. USA. 92:3449–3453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Silvestro, L., and P. H. Axelsen. 2000. Membrane-induced folding of cecropin A. Biophys. J. 79:1465–1477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Henzler-Wildman, K. A., D. K. Lee, and A. Ramamoorthy. 2003. Mechanism of lipid bilayer disruption by the human antimicrobial peptide, LL-37. Biochemistry. 42:6545–6558. [DOI] [PubMed] [Google Scholar]
- 16.Bhargava, K., and J. B. Feix. 2004. Membrane binding, structure, and localization of cecropin-mellitin hybrid peptides: a site-directed spin-labeling study. Biophys. J. 86:329–336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bechinger, B. 1999. The structure, dynamics and orientation of antimicrobial peptides in membranes by multidimensional solid-state NMR spectroscopy. Biochim. Biophys. Acta. 1462:157–183. [DOI] [PubMed] [Google Scholar]
- 18.Ramamoorthy, A., S. Thennarasu, D. K. Lee, A. Tan, and L. Maloy. 2006. Solid-state NMR investigation of the membrane-disrupting mechanism of antimicrobial peptides MSI-78 and MSI-594 derived from magainin 2 and melittin. Biophys. J. 91:206–216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sato, H., and J. B. Feix. 2006. Peptide-membrane interactions and mechanisms of membrane destruction by amphipathic alpha-helical antimicrobial peptides. Biochim. Biophys. Acta. 1758:1245–1256. [DOI] [PubMed] [Google Scholar]
- 20.Huang, H. W. 2000. Action of antimicrobial peptides: two-state model. Biochemistry. 39:8347–8352. [DOI] [PubMed] [Google Scholar]
- 21.Lee, M. T., F. Y. Chen, and H. W. Huang. 2004. Energetics of pore formation induced by membrane active peptides. Biochemistry. 43:3590–3599. [DOI] [PubMed] [Google Scholar]
- 22.Huang, H. W. 2006. Molecular mechanism of antimicrobial peptides: the origin of cooperativity. Biochim. Biophys. Acta. 1758:1292–1302. [DOI] [PubMed] [Google Scholar]
- 23.Matsuzaki, K., O. Murase, N. Fujii, and K. Miyajima. 1996. An antimicrobial peptide, magainin 2, induced rapid flip-flop of phospholipids coupled with pore formation and peptide translocation. Biochemistry. 35:11361–11368. [DOI] [PubMed] [Google Scholar]
- 24.Papo, N., and Y. Shai. 2003. Can we predict biological activity of antimicrobial peptides from their interactions with model phospholipid membranes? Peptides. 24:1693–1703. [DOI] [PubMed] [Google Scholar]
- 25.Yandek, L. E., A. Pokorny, A. Floren, K. Knoelke, U. Langel, and P. F. F. Almeida. Mechanism of the cell-penetrating peptide transportan 10 permeation of lipid bilayers. Biophys. J. 92:2434–2444. [DOI] [PMC free article] [PubMed]
- 26.Blondelle, S. E., K. Lohner, and M. Aguilar. 1999. Lipid-induced conformation and lipid-binding properties of cytolytic and antimicrobial peptides: determination and biological specificity. Biochim. Biophys. Acta. 1462:89–108. [DOI] [PubMed] [Google Scholar]
- 27.Epand, R. F., M. A. Schmitt, S. H. Gellman, and R. M. Epand. 2006. Role of membrane lipids in the mechanism of bacterial species selective toxicity by two alpha/beta-antimicrobial peptides. Biochim. Biophys. Acta. 1758:1343–1350. [DOI] [PubMed] [Google Scholar]
- 28.Fink, J., R. B. Merrifield, A. Boman, and H. G. Boman. 1989. The chemical synthesis of cecropin D and an analog with enhanced antibacterial activity. J. Biol. Chem. 264:6260–6267. [PubMed] [Google Scholar]
- 29.Andreu, D., R. B. Merrifield, H. Steiner, and H. G. Boman. 1985. N-terminal analogues of cecropin A: synthesis, antibacterial activity, and conformational properties. Biochemistry. 24:1683–1688. [DOI] [PubMed] [Google Scholar]
- 30.Andreu, D., J. Ubach, A. Boman, B. Wahlin, D. Wade, R. B. Merrifield, and H. G. Boman. 1992. Shortened cecropin A-melittin hybrids. Significant size reduction retains potent antibiotic activity. FEBS Lett. 296:190–194. [DOI] [PubMed] [Google Scholar]
- 31.Sato, H., and J. B. Feix. 2006. Osmoprotection of bacterial cells from toxicity caused by antimicrobial hybrid peptide CM15. Biochemistry. 45:9997–10007. [DOI] [PubMed] [Google Scholar]
- 32.Pace, C. N., F. Vajdos, L. Fee, G. Grimsley, and T. Gray. 1995. How to measure and predict the molar absorption coefficient of a protein. Protein Sci. 4:2411–2423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Gennis, R. B. 1989. Biomembranes Molecular Structure and Function. Springer-Verlag, New York.
- 34.Stewart, J. C. M. 1980. Colorimetric determination of phospholipids with ammonium ferrothiocyanate. Anal. Biochem. 104:10–14. [DOI] [PubMed] [Google Scholar]
- 35.Lobley, A., L. Whitmore, and B. A. Wallace. 2002. DICHROWEB: an interactive website for the analysis of protein secondary structure from circular dichroism spectra. Bioinformatics. 18:211–212. [DOI] [PubMed] [Google Scholar]
- 36.Bates, I. R., J. M. Boggs, J. B. Feix, and G. Harauz. 2003. Membrane-anchoring and charge effects in the interaction of myelin basic protein with lipid bilayers studied by site-directed spin labeling. J. Biol. Chem. 278:29041–29047. [DOI] [PubMed] [Google Scholar]
- 37.Altenbach, C., S. L. Flitsch, H. G. Khorana, and W. L. Hubbell. 1989. Structural studies on transmembrane proteins. 2. Spin labeling of bacteriorhodopsin mutants at unique cysteines. Biochemistry. 28:7806–7812. [DOI] [PubMed] [Google Scholar]
- 38.Feix, J. B., and C. S. Klug. 1998. Site-directed spin labeling of membrane proteins and peptide-membrane interactions. In Biological Magnetic Resonance, Vol. 14: Spin Labeling: The Next Millennium. L. J. Berliner, editor. Plenum Press, New York. 252–281.
- 39.Altenbach, C., D. A. Greenhalgh, H. G. Khorana, and W. L. Hubbell. 1994. A collision gradient method to determine the immersion depth of nitroxides in lipid bilayers: application to spin-labeled mutants of bacteriorhodopsin. Proc. Natl. Acad. Sci. USA. 91:1667–1671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yu, Y. G., T. Thorgeirsson, and Y.-K. Shin. 1994. Topology of an amphiphilic mitochondrial signal sequence in the membrane-inserted state: a spin labeling study. Biochemistry. 33:14221–14226. [DOI] [PubMed] [Google Scholar]
- 41.Langen, R., K. J. Oh, D. Cascio, and W. L. Hubbell. 2000. Crystal structures of spin labeled T4 lysozyme mutants: implications for the interpretation of EPR spectra in terms of structure. Biochemistry. 39:8396–8405. [DOI] [PubMed] [Google Scholar]
- 42.Columbus, L., T. Kalai, J. Jeko, K. Hideg, and W. L. Hubbell. 2001. Molecular motion of spin labeled side chains in alpha-helices: analysis by variation of side chain structure. Biochemistry. 40:3828–3846. [DOI] [PubMed] [Google Scholar]
- 43.Altenbach, C., K. J. Oh, R. J. Trabanino, K. Hideg, and W. L. Hubbell. 2001. Estimation of inter-residue distances in spin labeled proteins at physiological temperatures: experimental strategies and practical limitations. Biochemistry. 40:15471–15482. [DOI] [PubMed] [Google Scholar]
- 44.Subczynski, W. K., and J. S. Hyde. 1981. The diffusion-concentration product of oxygen in lipid bilayers using the spin-label T1 method. Biochim. Biophys. Acta. 643:283–291. [PubMed] [Google Scholar]
- 45.Heller, W. T., A. J. Waring, R. I. Lehrer, T. A. Harroun, T. M. Weiss, L. Yang, and H. W. Huang. 2000. Membrane thinning effect of the beta-sheet antimicrobial protegrin. Biochemistry. 39:139–145. [DOI] [PubMed] [Google Scholar]
- 46.Chen, F. Y., M. T. Lee, and H. W. Huang. 2003. Evidence for membrane thinning effect as the mechanism for peptide-induced pore formation. Biophys. J. 84:3751–3758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mecke, A., D. K. Lee, A. Ramamoorthy, B. G. Orr, and M. M. Banaszak Holl. 2005. Membrane thinning due to antimicrobial peptide binding—an atomic force microscopy study of MSI-78 in lipid bilayers. Biophys. J. 89:4043–4050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hung, S. C., W. Wang, S. I. Chan, and H. M. Chen. 1999. Membrane lysis by the antibacterial peptides cecropins B1 and B3: a spin-label electron spin resonance study on phospholipid bilayers. Biophys. J. 77:3120–3133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Ludtke, S. J., K. He, W. T. Heller, T. A. Harroun, L. Yang, and H. W. Huang. 1996. Membrane pores induced by magainin. Biochemistry. 35:13723–13728. [DOI] [PubMed] [Google Scholar]
- 50.Hallock, K. J., D. K. Lee, and A. Ramamoorthy. 2003. MSI-78, an analogue of the magainin antimicrobial peptides, disrupts lipid bilayer structure via positive curvature strain. Biophys. J. 84:3052–3060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Henzler-Wildman, K. A., G. V. Martinez, M. F. Brown, and A. Ramamoorthy. 2004. Perturbation of the hydrophobic core of lipid bilayers by the human antimicrobial peptide LL-37. Biochemistry. 43:8459–8469. [DOI] [PubMed] [Google Scholar]
- 52.Rabenstein, M. D., and Y. K. Shin. 1995. Determination of the distance between two spin labels attached to a macromolecule. Proc. Natl. Acad. Sci. USA. 92:8239–8243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Pannier, M., S. Veit, A. Godt, G. Jeschke, and H. W. Spiess. 2000. Dead-time free measurement of dipole-dipole interactions between electron spins. J. Magn. Reson. 142:331–340. [DOI] [PubMed] [Google Scholar]
- 54.Zhou, Z., S. C. DeSensi, R. A. Stein, S. Brandon, M. Dixit, E. J. McArdle, E. M. Warren, H. K. Kroh, L. Song, C. E. Cobb, E. J. Hustedt, and A. H. Beth. 2005. Solution structure of the cytoplasmic domain of erythrocyte membrane band 3 determined by site-directed spin labeling. Biochemistry. 44:15115–15128. [DOI] [PubMed] [Google Scholar]