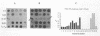Abstract
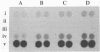
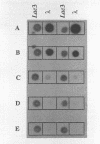
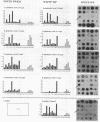
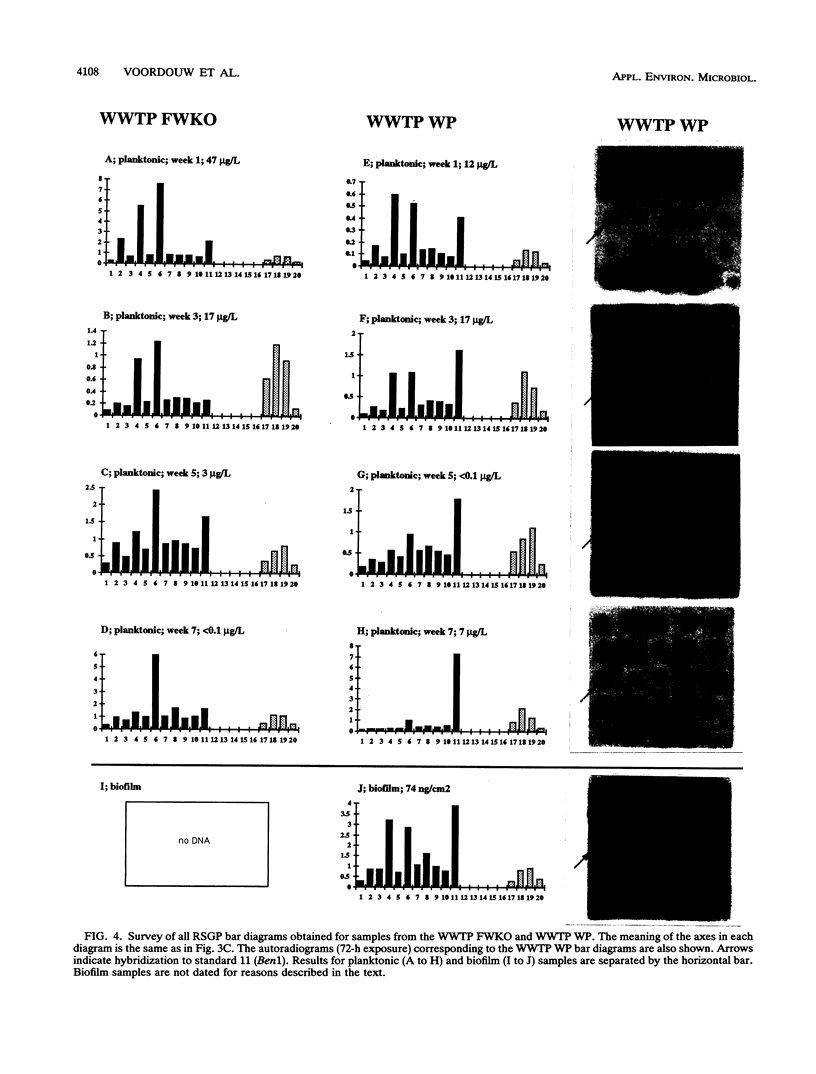
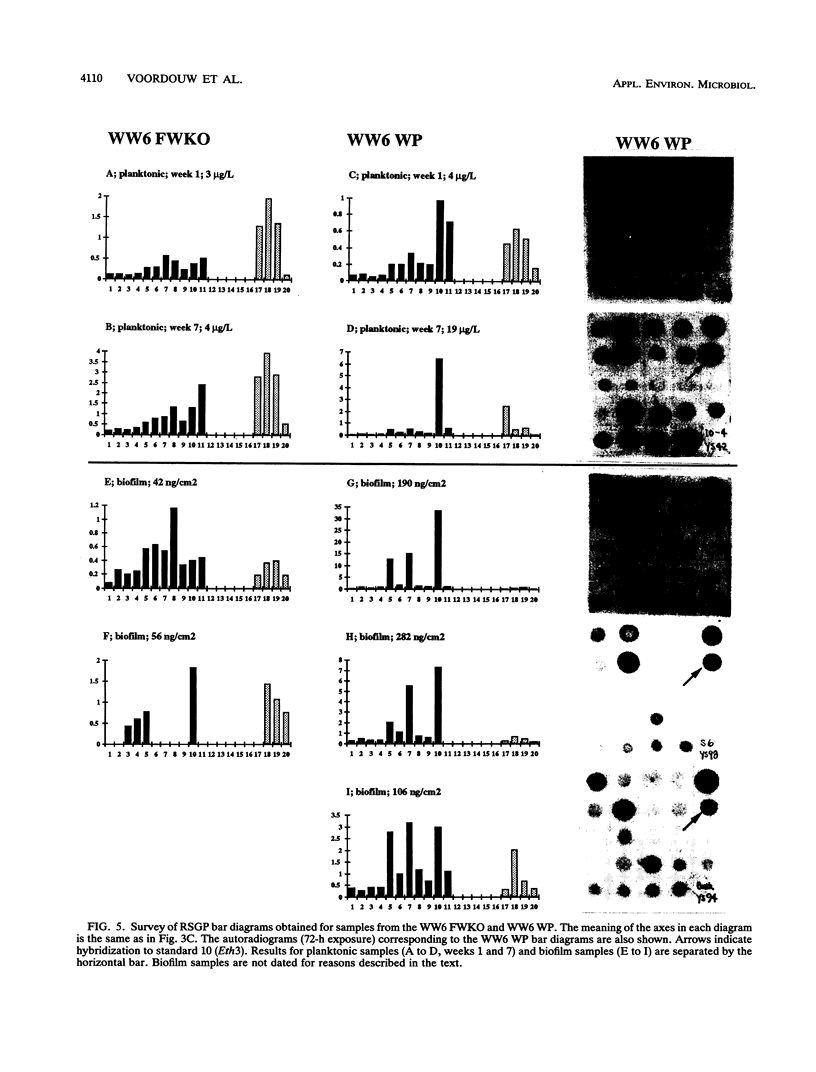
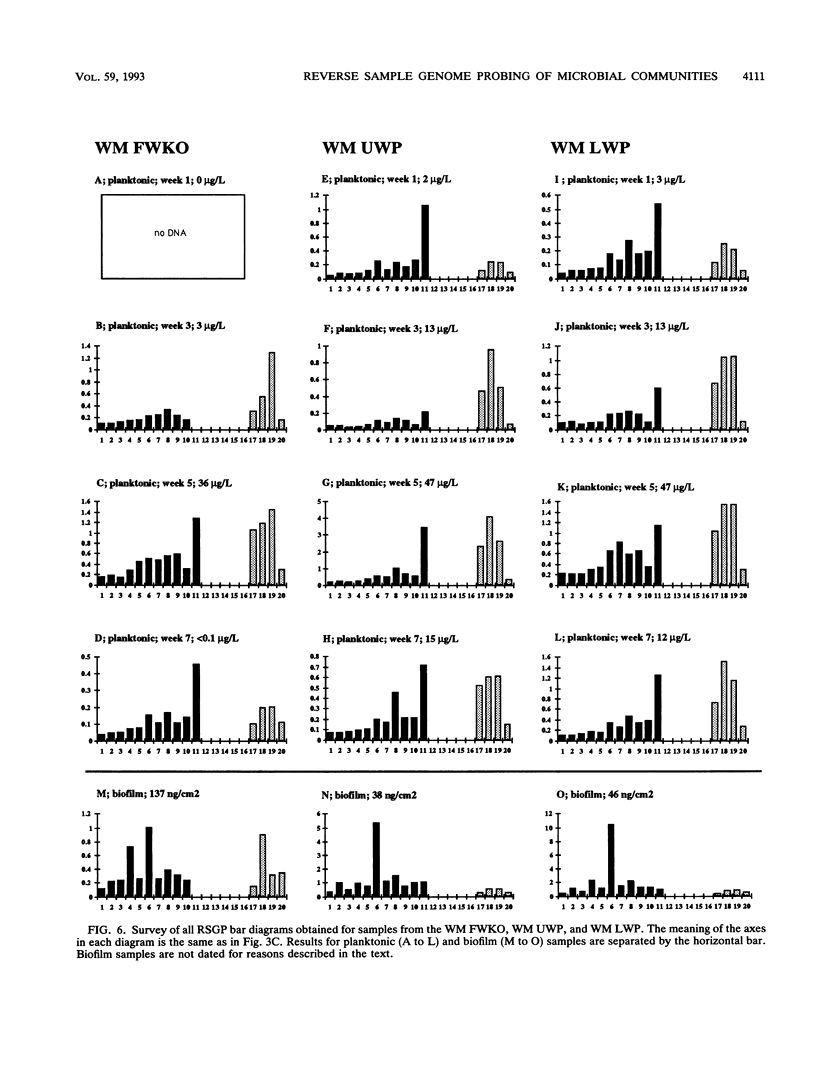
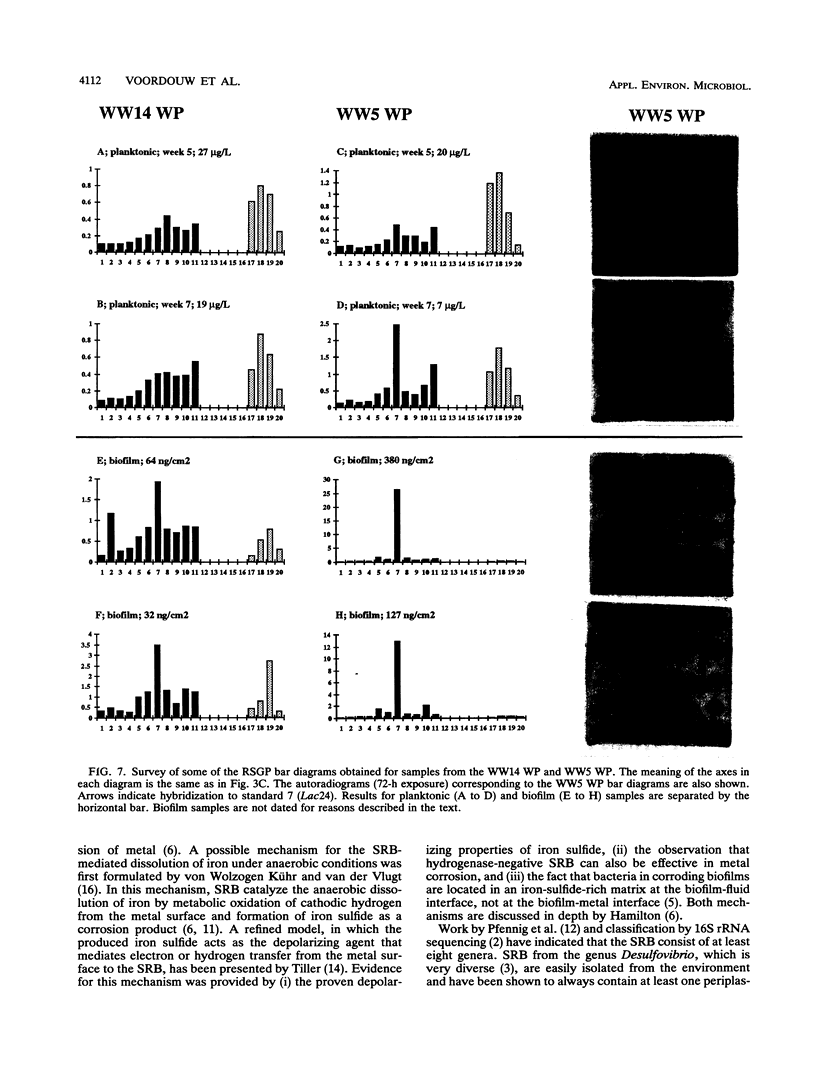
This paper presents a protocol for quantitative analysis of microbial communities by reverse sample genome probing is presented in which (i) whole community DNA is isolated and labeled in the presence of a known amount of an added internal standard and (ii) the resulting spiked reverse genome probe is hybridized with a master filter on which denatured genomic DNAs from bacterial standards isolated from the target environment were spotted in large amounts (up to 1,500 ng) in order to improve detection sensitivity. This protocol allowed reproducible fingerprinting of the microbial community in oil field production waters at 19 sites from which water and biofilm samples were collected. It appeared that selected sulfate-reducing bacteria were significantly enhanced in biofilms covering the metal surfaces in contact with the production waters.
Full text
PDF



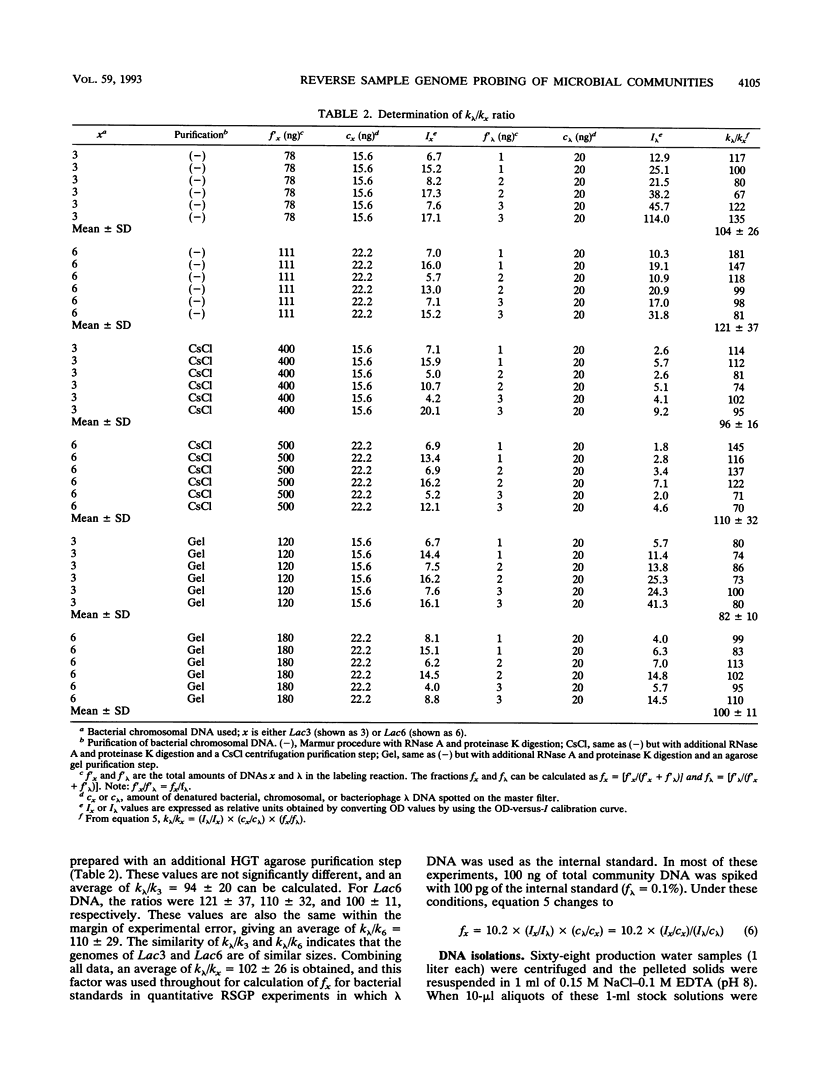

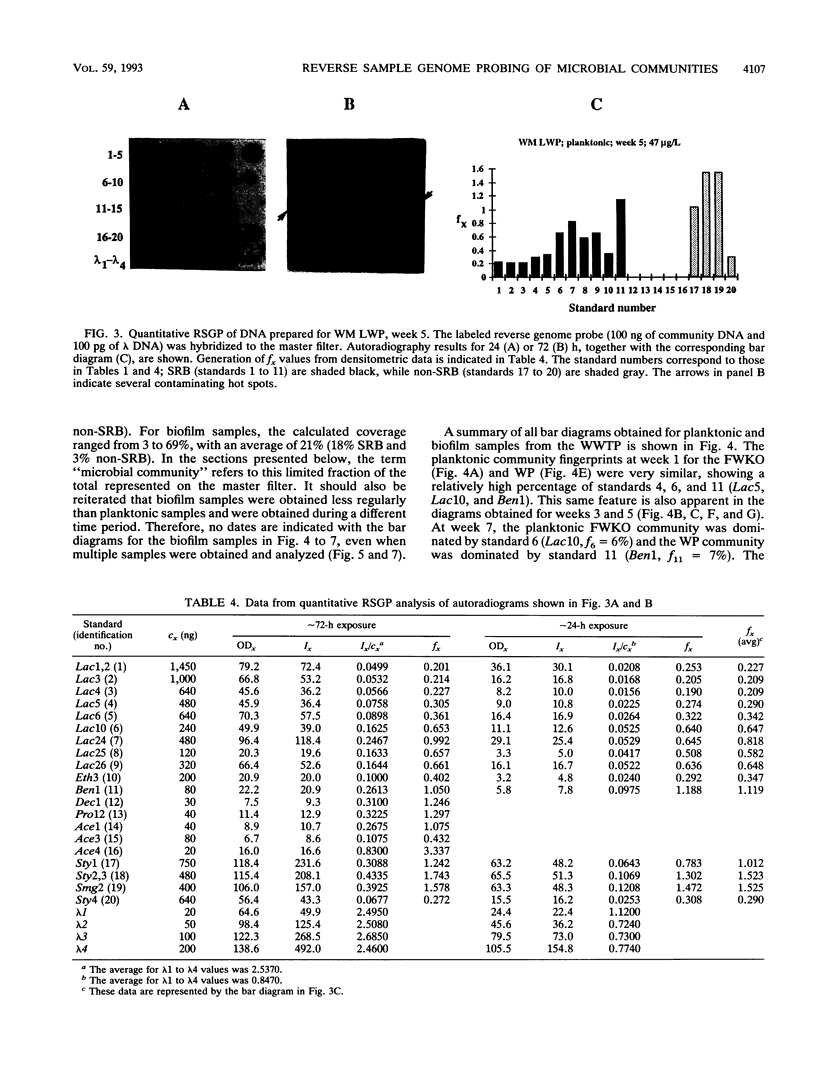



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Amann R. I., Stromley J., Devereux R., Key R., Stahl D. A. Molecular and microscopic identification of sulfate-reducing bacteria in multispecies biofilms. Appl Environ Microbiol. 1992 Feb;58(2):614–623. doi: 10.1128/aem.58.2.614-623.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux R., Delaney M., Widdel F., Stahl D. A. Natural relationships among sulfate-reducing eubacteria. J Bacteriol. 1989 Dec;171(12):6689–6695. doi: 10.1128/jb.171.12.6689-6695.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux R., He S. H., Doyle C. L., Orkland S., Stahl D. A., LeGall J., Whitman W. B. Diversity and origin of Desulfovibrio species: phylogenetic definition of a family. J Bacteriol. 1990 Jul;172(7):3609–3619. doi: 10.1128/jb.172.7.3609-3619.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamilton W. A. Sulphate-reducing bacteria and anaerobic corrosion. Annu Rev Microbiol. 1985;39:195–217. doi: 10.1146/annurev.mi.39.100185.001211. [DOI] [PubMed] [Google Scholar]
- Kafatos F. C., Jones C. W., Efstratiadis A. Determination of nucleic acid sequence homologies and relative concentrations by a dot hybridization procedure. Nucleic Acids Res. 1979 Nov 24;7(6):1541–1552. doi: 10.1093/nar/7.6.1541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Postgate J. R., Kent H. M., Robson R. L., Chesshyre J. A. The genomes of Desulfovibrio gigas and D. vulgaris. J Gen Microbiol. 1984 Jul;130(7):1597–1601. doi: 10.1099/00221287-130-7-1597. [DOI] [PubMed] [Google Scholar]
- Voordouw G., Hagen W. R., Krüse-Wolters K. M., van Berkel-Arts A., Veeger C. Purification and characterization of Desulfovibrio vulgaris (Hildenborough) hydrogenase expressed in Escherichia coli. Eur J Biochem. 1987 Jan 2;162(1):31–36. doi: 10.1111/j.1432-1033.1987.tb10537.x. [DOI] [PubMed] [Google Scholar]
- Voordouw G., Niviere V., Ferris F. G., Fedorak P. M., Westlake D. W. Distribution of Hydrogenase Genes in Desulfovibrio spp. and Their Use in Identification of Species from the Oil Field Environment. Appl Environ Microbiol. 1990 Dec;56(12):3748–3754. doi: 10.1128/aem.56.12.3748-3754.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voordouw G., Strang J. D., Wilson F. R. Organization of the genes encoding [Fe] hydrogenase in Desulfovibrio vulgaris subsp. oxamicus Monticello. J Bacteriol. 1989 Jul;171(7):3881–3889. doi: 10.1128/jb.171.7.3881-3889.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voordouw G., Voordouw J. K., Jack T. R., Foght J., Fedorak P. M., Westlake D. W. Identification of distinct communities of sulfate-reducing bacteria in oil fields by reverse sample genome probing. Appl Environ Microbiol. 1992 Nov;58(11):3542–3552. doi: 10.1128/aem.58.11.3542-3552.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voordouw G., Voordouw J. K., Karkhoff-Schweizer R. R., Fedorak P. M., Westlake D. W. Reverse sample genome probing, a new technique for identification of bacteria in environmental samples by DNA hybridization, and its application to the identification of sulfate-reducing bacteria in oil field samples. Appl Environ Microbiol. 1991 Nov;57(11):3070–3078. doi: 10.1128/aem.57.11.3070-3078.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]