Abstract
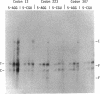
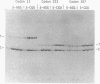
A system for testing the effects of specific codons on gene expression is described. Tandem test and control genes are contained in a transcription unit for bacteriophage T7 RNA polymerase in a multicopy plasmid, and nearly identical test and control mRNAs are generated from the primary transcript by RNase III cleavages. Their coding sequences, derived from T7 gene 9, are translated efficiently and have few low-usage codons of Escherichia coli. The upstream test gene contains a site for insertion of test codons, and the downstream control gene has a 45-codon deletion that allows test and control mRNAs and proteins to be separated by gel electrophoresis. Codons can be inserted among identical flanking codons after codon 13, 223, or 307 in codon test vectors pCT1, pCT2, and pCT3, respectively, the third site being six codons from the termination codon. The insertion of two to five consecutive AGG (low-usage) arginine codons selectively reduced the production of full-length test protein to extents that depended on the number of AGG codons, the site of insertion, and the amount of test mRNA. Production of aberrant proteins was also stimulated at high levels of mRNA. The effects occurred primarily at the translational level and were not produced by CGU (high-usage) arginine codons. Our results are consistent with the idea that sufficiently high levels of the AGG mRNA can cause essentially all of the tRNA(AGG) in the cell to become sequestered in translating peptidyl-tRNA(AGG) -mRNA-ribosome complexes stalled at the first of two consecutive AGG codons and that the approach of an upstream translating ribosome stimulates a stalled ribosome of frameshift, hop, or terminate translation.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andersson S. G., Kurland C. G. Codon preferences in free-living microorganisms. Microbiol Rev. 1990 Jun;54(2):198–210. doi: 10.1128/mr.54.2.198-210.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Atkins J. F., Weiss R. B., Gesteland R. F. Ribosome gymnastics--degree of difficulty 9.5, style 10.0. Cell. 1990 Aug 10;62(3):413–423. doi: 10.1016/0092-8674(90)90007-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonekamp F., Andersen H. D., Christensen T., Jensen K. F. Codon-defined ribosomal pausing in Escherichia coli detected by using the pyrE attenuator to probe the coupling between transcription and translation. Nucleic Acids Res. 1985 Jun 11;13(11):4113–4123. doi: 10.1093/nar/13.11.4113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brinkmann U., Mattes R. E., Buckel P. High-level expression of recombinant genes in Escherichia coli is dependent on the availability of the dnaY gene product. Gene. 1989 Dec 21;85(1):109–114. doi: 10.1016/0378-1119(89)90470-8. [DOI] [PubMed] [Google Scholar]
- Chen G. F., Inouye M. Suppression of the negative effect of minor arginine codons on gene expression; preferential usage of minor codons within the first 25 codons of the Escherichia coli genes. Nucleic Acids Res. 1990 Mar 25;18(6):1465–1473. doi: 10.1093/nar/18.6.1465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunn J. J., Studier F. W. Complete nucleotide sequence of bacteriophage T7 DNA and the locations of T7 genetic elements. J Mol Biol. 1983 Jun 5;166(4):477–535. doi: 10.1016/s0022-2836(83)80282-4. [DOI] [PubMed] [Google Scholar]
- Ikemura T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes. J Mol Biol. 1981 Feb 15;146(1):1–21. doi: 10.1016/0022-2836(81)90363-6. [DOI] [PubMed] [Google Scholar]
- Komine Y., Adachi T., Inokuchi H., Ozeki H. Genomic organization and physical mapping of the transfer RNA genes in Escherichia coli K12. J Mol Biol. 1990 Apr 20;212(4):579–598. doi: 10.1016/0022-2836(90)90224-A. [DOI] [PubMed] [Google Scholar]
- Menninger J. R. Computer simulation of ribosome editing. J Mol Biol. 1983 Dec 25;171(4):383–399. doi: 10.1016/0022-2836(83)90036-0. [DOI] [PubMed] [Google Scholar]
- Panayotatos N., Truong K. Cleavage within an RNase III site can control mRNA stability and protein synthesis in vivo. Nucleic Acids Res. 1985 Apr 11;13(7):2227–2240. doi: 10.1093/nar/13.7.2227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson M., Lilley R., Little S., Emtage J. S., Yarranton G., Stephens P., Millican A., Eaton M., Humphreys G. Codon usage can affect efficiency of translation of genes in Escherichia coli. Nucleic Acids Res. 1984 Sep 11;12(17):6663–6671. doi: 10.1093/nar/12.17.6663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosenberg A. H., Lade B. N., Chui D. S., Lin S. W., Dunn J. J., Studier F. W. Vectors for selective expression of cloned DNAs by T7 RNA polymerase. Gene. 1987;56(1):125–135. doi: 10.1016/0378-1119(87)90165-x. [DOI] [PubMed] [Google Scholar]
- Rosenberg A. H., Studier F. W. T7 RNA polymerase can direct expression of influenza virus cap-binding protein (PB2) in Escherichia coli. Gene. 1987;59(2-3):191–200. doi: 10.1016/0378-1119(87)90327-1. [DOI] [PubMed] [Google Scholar]
- Spanjaard R. A., Chen K., Walker J. R., van Duin J. Frameshift suppression at tandem AGA and AGG codons by cloned tRNA genes: assigning a codon to argU tRNA and T4 tRNA(Arg). Nucleic Acids Res. 1990 Sep 11;18(17):5031–5036. doi: 10.1093/nar/18.17.5031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spanjaard R. A., van Duin J. Translation of the sequence AGG-AGG yields 50% ribosomal frameshift. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7967–7971. doi: 10.1073/pnas.85.21.7967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steitz J. A. Polypeptide chain initiation: nucleotide sequences of the three ribosomal binding sites in bacteriophage R17 RNA. Nature. 1969 Dec 6;224(5223):957–964. doi: 10.1038/224957a0. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Maizel J. V., Jr T7-directed protein synthesis. Virology. 1969 Nov;39(3):575–586. doi: 10.1016/0042-6822(69)90105-6. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Moffatt B. A. Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol. 1986 May 5;189(1):113–130. doi: 10.1016/0022-2836(86)90385-2. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Rosenberg A. H., Dunn J. J., Dubendorff J. W. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 1990;185:60–89. doi: 10.1016/0076-6879(90)85008-c. [DOI] [PubMed] [Google Scholar]
- Studier F. W. Use of bacteriophage T7 lysozyme to improve an inducible T7 expression system. J Mol Biol. 1991 May 5;219(1):37–44. doi: 10.1016/0022-2836(91)90855-z. [DOI] [PubMed] [Google Scholar]
- Sørensen M. A., Kurland C. G., Pedersen S. Codon usage determines translation rate in Escherichia coli. J Mol Biol. 1989 May 20;207(2):365–377. doi: 10.1016/0022-2836(89)90260-x. [DOI] [PubMed] [Google Scholar]
- Sørensen M. A., Pedersen S. Absolute in vivo translation rates of individual codons in Escherichia coli. The two glutamic acid codons GAA and GAG are translated with a threefold difference in rate. J Mol Biol. 1991 Nov 20;222(2):265–280. doi: 10.1016/0022-2836(91)90211-n. [DOI] [PubMed] [Google Scholar]
- TAKANAMI M., ZUBAY G. AN ESTIMATE OF THE SIZE OF THE RIBOSOMAL SITE FOR MESSENGER RNA BINDING. Proc Natl Acad Sci U S A. 1964 May;51:834–839. doi: 10.1073/pnas.51.5.834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varenne S., Baty D., Verheij H., Shire D., Lazdunski C. The maximum rate of gene expression is dependent on the downstream context of unfavourable codons. Biochimie. 1989 Nov-Dec;71(11-12):1221–1229. doi: 10.1016/0300-9084(89)90027-8. [DOI] [PubMed] [Google Scholar]
- Varenne S., Lazdunski C. Effect of distribution of unfavourable codons on the maximum rate of gene expression by an heterologous organism. J Theor Biol. 1986 May 7;120(1):99–110. doi: 10.1016/s0022-5193(86)80020-0. [DOI] [PubMed] [Google Scholar]
- Wolin S. L., Walter P. Ribosome pausing and stacking during translation of a eukaryotic mRNA. EMBO J. 1988 Nov;7(11):3559–3569. doi: 10.1002/j.1460-2075.1988.tb03233.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang S. P., Zubay G., Goldman E. Low-usage codons in Escherichia coli, yeast, fruit fly and primates. Gene. 1991 Aug 30;105(1):61–72. doi: 10.1016/0378-1119(91)90514-c. [DOI] [PubMed] [Google Scholar]