Abstract
Uracil phosphoribosyltransferase catalyzes the key reaction in the salvage of uracil in many microorganisms. The gene encoding uracil phosphoribosyltransferase (upp) was cloned from Lactococcus lactis subsp. cremoris MG1363 by complementation of an Escherichia coli mutant. The gene was sequenced, and the putative amino acid sequence was deduced. The promoter was mapped by both primer extension and analysis of beta-galactosidase expressed from strains carrying fusion between upp promoter fragments and the lacLM gene. The results showed that the upp gene was expressed from its own promoter. After in vitro construction of an internal deletion, a upp mutant was constructed by a double-crossover event. This implicated the utilization of a plasmid with a thermosensitive origin of replication and a new and easy way to screen for double crossover events in both gram-positive and gram-negative bacterial strains. The phenotype of the uracil phosphoribosyltransferase-deficient strain was established. Surprisingly, the upp strain is resistant only to very low concentrations of 5-fluorouracil. Secondary mutants in thymidine phosphorylase and thymidine kinase were isolated by selection for resistance to high concentrations of 5-fluorouracil.
Full text
PDF
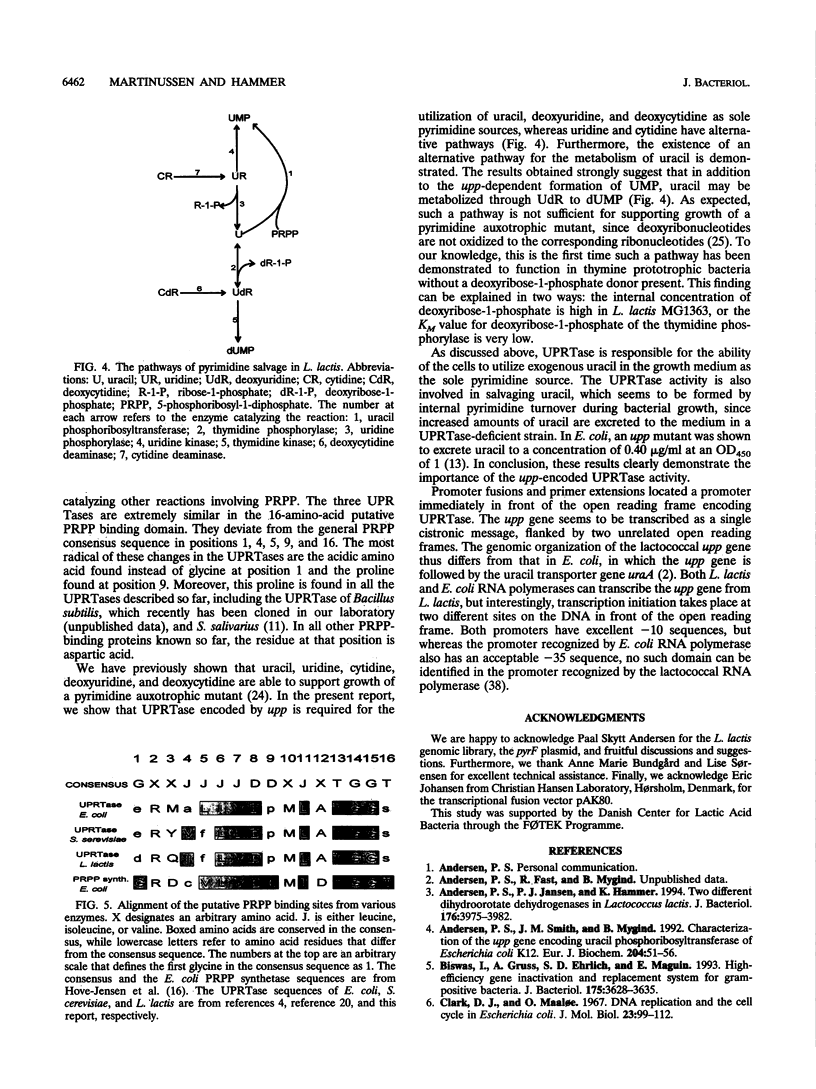





Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andersen P. S., Jansen P. J., Hammer K. Two different dihydroorotate dehydrogenases in Lactococcus lactis. J Bacteriol. 1994 Jul;176(13):3975–3982. doi: 10.1128/jb.176.13.3975-3982.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biswas I., Gruss A., Ehrlich S. D., Maguin E. High-efficiency gene inactivation and replacement system for gram-positive bacteria. J Bacteriol. 1993 Jun;175(11):3628–3635. doi: 10.1128/jb.175.11.3628-3635.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gasson M. J. Plasmid complements of Streptococcus lactis NCDO 712 and other lactic streptococci after protoplast-induced curing. J Bacteriol. 1983 Apr;154(1):1–9. doi: 10.1128/jb.154.1.1-9.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerdes K., Thisted T., Martinussen J. Mechanism of post-segregational killing by the hok/sok system of plasmid R1: sok antisense RNA regulates formation of a hok mRNA species correlated with killing of plasmid-free cells. Mol Microbiol. 1990 Nov;4(11):1807–1818. doi: 10.1111/j.1365-2958.1990.tb02029.x. [DOI] [PubMed] [Google Scholar]
- Giffard P. M., Rathsam C., Kwan E., Kwan D. W., Bunny K. L., Koo S. P., Jacques N. A. The ftf gene encoding the cell-bound fructosyltransferase of Streptococcus salivarius ATCC 25975 is preceded by an insertion sequence and followed by FUR1 and clpP homologues. J Gen Microbiol. 1993 May;139(5):913–920. doi: 10.1099/00221287-139-5-913. [DOI] [PubMed] [Google Scholar]
- Glatron M. F., Rapoport G. Biosynthesis of the parasporal inclusion of Bacillus thuringiensis: half-life of its corresponding messenger RNA. Biochimie. 1972;54(10):1291–1301. doi: 10.1016/s0300-9084(72)80070-1. [DOI] [PubMed] [Google Scholar]
- Hammer-Jespersen K., Munch-Petersen A. Mutants of Escherichia coli unable to metabolize cytidine: isolation and characterization. Mol Gen Genet. 1973 Nov 2;126(2):177–186. doi: 10.1007/BF00330992. [DOI] [PubMed] [Google Scholar]
- Hawley D. K., McClure W. R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983 Apr 25;11(8):2237–2255. doi: 10.1093/nar/11.8.2237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holo H., Nes I. F. High-Frequency Transformation, by Electroporation, of Lactococcus lactis subsp. cremoris Grown with Glycine in Osmotically Stabilized Media. Appl Environ Microbiol. 1989 Dec;55(12):3119–3123. doi: 10.1128/aem.55.12.3119-3123.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hove-Jensen B., Harlow K. W., King C. J., Switzer R. L. Phosphoribosylpyrophosphate synthetase of Escherichia coli. Properties of the purified enzyme and primary structure of the prs gene. J Biol Chem. 1986 May 25;261(15):6765–6771. [PubMed] [Google Scholar]
- Jensen P. R., Hammer K. Minimal Requirements for Exponential Growth of Lactococcus lactis. Appl Environ Microbiol. 1993 Dec;59(12):4363–4366. doi: 10.1128/aem.59.12.4363-4366.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kern L., de Montigny J., Jund R., Lacroute F. The FUR1 gene of Saccharomyces cerevisiae: cloning, structure and expression of wild-type and mutant alleles. Gene. 1990 Apr 16;88(2):149–157. doi: 10.1016/0378-1119(90)90026-n. [DOI] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Ludwig W., Seewaldt E., Kilpper-Bälz R., Schleifer K. H., Magrum L., Woese C. R., Fox G. E., Stackebrandt E. The phylogenetic position of Streptococcus and Enterococcus. J Gen Microbiol. 1985 Mar;131(3):543–551. doi: 10.1099/00221287-131-3-543. [DOI] [PubMed] [Google Scholar]
- Maguin E., Duwat P., Hege T., Ehrlich D., Gruss A. New thermosensitive plasmid for gram-positive bacteria. J Bacteriol. 1992 Sep;174(17):5633–5638. doi: 10.1128/jb.174.17.5633-5638.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martinussen J., Andersen P. S., Hammer K. Nucleotide metabolism in Lactococcus lactis: salvage pathways of exogenous pyrimidines. J Bacteriol. 1994 Mar;176(5):1514–1516. doi: 10.1128/jb.176.5.1514-1516.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nilsson B., Uhlén M., Josephson S., Gatenbeck S., Philipson L. An improved positive selection plasmid vector constructed by oligonucleotide mediated mutagenesis. Nucleic Acids Res. 1983 Nov 25;11(22):8019–8030. doi: 10.1093/nar/11.22.8019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pedersen M. L., Arnved K. R., Johansen E. Genetic analysis of the minimal replicon of the Lactococcus lactis subsp. lactis biovar diacetylactis citrate plasmid. Mol Gen Genet. 1994 Aug 15;244(4):374–382. doi: 10.1007/BF00286689. [DOI] [PubMed] [Google Scholar]
- Perez-Martinez G., Kok J., Venema G., van Dijl J. M., Smith H., Bron S. Protein export elements from Lactococcus lactis. Mol Gen Genet. 1992 Sep;234(3):401–411. doi: 10.1007/BF00538699. [DOI] [PubMed] [Google Scholar]
- Piérard A., Glansdorff N., Yashphe J. Mutations affecting uridine monophosphate pyrophosphorylase or the argR gene in Escherichia coli. Effects on carbamoyl phosphate and pyrimidine biosynthesis and on uracil uptake. Mol Gen Genet. 1972;118(3):235–245. doi: 10.1007/BF00333460. [DOI] [PubMed] [Google Scholar]
- Rasmussen U. B., Mygind B., Nygaard P. Purification and some properties of uracil phosphoribosyltransferase from Escherichia coli K12. Biochim Biophys Acta. 1986 Apr 11;881(2):268–275. doi: 10.1016/0304-4165(86)90013-9. [DOI] [PubMed] [Google Scholar]
- Rima B. K., Takahashi I. Metabolism of pyrimidine bases and nucleosides in Bacillus subtilis. J Bacteriol. 1977 Feb;129(2):574–579. doi: 10.1128/jb.129.2.574-579.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salser W. Globin mRNA sequences: analysis of base pairing and evolutionary implications. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 2):985–1002. doi: 10.1101/sqb.1978.042.01.099. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwartz E., Kröger M., Rak B. IS150: distribution, nucleotide sequence and phylogenetic relationships of a new E. coli insertion element. Nucleic Acids Res. 1988 Jul 25;16(14B):6789–6802. doi: 10.1093/nar/16.14.6789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terzaghi B. E., Sandine W. E. Improved medium for lactic streptococci and their bacteriophages. Appl Microbiol. 1975 Jun;29(6):807–813. doi: 10.1128/am.29.6.807-813.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waser M., Hess-Bienz D., Davies K., Solioz M. Cloning and disruption of a putative NaH-antiporter gene of Enterococcus hirae. J Biol Chem. 1992 Mar 15;267(8):5396–5400. [PubMed] [Google Scholar]
- d'Aubenton Carafa Y., Brody E., Thermes C. Prediction of rho-independent Escherichia coli transcription terminators. A statistical analysis of their RNA stem-loop structures. J Mol Biol. 1990 Dec 20;216(4):835–858. doi: 10.1016/s0022-2836(99)80005-9. [DOI] [PubMed] [Google Scholar]
- van de Guchte M., Kok J., Venema G. Gene expression in Lactococcus lactis. FEMS Microbiol Rev. 1992 Feb;8(2):73–92. doi: 10.1111/j.1574-6968.1992.tb04958.x. [DOI] [PubMed] [Google Scholar]


