Abstract
Sterol carrier protein-2 (SCP-2) was independently discovered as a soluble protein that binds and transfers cholesterol as well as phospholipids (nonspecific lipid transfer protein, nsLTP) in vitro. Physiological functions of this protein are only now beginning to be resolved. The gene encoding SCP-2 also encodes sterol carrier protein-x (SCP-x) arising from an alternate transcription site. In vitro and in vivo SCP-x serves as a peroxisomal 3-ketoacyl-CoA thiolase in oxidation of branched-chain lipids (cholesterol to form bile acids; branched-chain fatty acid for detoxification). While peroxisomal SCP-2 facilitates branched-chain lipid oxidation, the role(s) of extraperoxisomal (up to 50% of total) are less clear. Studies using transfected fibroblasts overexpressing SCP-2 and hepatocytes from SCP-2/SCP-x gene-ablated mice reveal that SCP-2 selectively remodels the lipid composition, structure, and function of lipid rafts/caveolae. Studies of purified SCP-2 and in cells show that SCP-2 has high affinity for and selectively transfers many lipid species involved in intracellular signaling: fatty acids, fatty acyl CoAs, lysophosphatidic acid, phosphatidylinositols, and sphingolipids (sphingomyelin, ceramide, mono- di- and multi-hexosylceramides, gangliosides). SCP-2 selectively redistributes these signaling lipids between lipid rafts/caveolae and intracellular sites. These findings suggest SCP-2 serves not only in cholesterol and phospholipid transfer, but also in regulating multiple lipid signaling pathways in lipid raft/caveolae microdomains of the plasma membrane.
Keywords: Caveolae, lipid rafts, sterol carrier protein-2, SCP-2, phosphatidylinositol, sphingolipids, cholesterol, fatty acyl CoA, fatty acid
INTRODUCTION
Sterol carrier protein-2 was originally discovered in 1970 by Scallen and associates as one of two proteins in the rat liver soluble fraction which was required for cholesterol biosynthesis (rev. in [1]. Subsequent studies from the Scallen and other labs revealed that the respective purified proteins exhibit very different specific roles in cholesterol biosynthesis: (i) Sterol carrier protein-1 (SCP-1, also called supernatant protein factor or SPF) is a 47 kDa acidic protein originally discovered to enhance the enzymatic conversion of squalene to lanosterol (rev. in [1]. (ii) Sterol carrier protein-2 (SCP-2) is a13 kDa, basic protein with multiple roles in isoprenoid and cholesterol metabolism (rev. in [1,2].
SPF has been cloned and its x-ray crystal structure solved [3]. It is a member of the CRAL_Trio family of proteins. Supernatant protein factor (SPF) promotes the epoxidation of squalene catalyzed by microsomes. Several studies suggest its in vivo role in the cholesterol biosynthetic pathway by a yet unknown mechanism. SPF belongs to a family of lipid binding proteins called CRAL_TRIO, which include yeast phosphatidylinositol transfer protein Sec14 and tocopherol transfer protein TTP. The crystal structure of human SPF at a resolution of 1.9 A reveals a two domain topology. The N-terminal 275 residues form a Sec14-like domain, while the C-terminal 115 residues consist of an eight-stranded jelly-roll barrel similar to that found in many viral protein structures. The ligand binding cavity has a peculiar horseshoe-like shape. Contrary to the Sec14 crystal structure, the lipid-exchange loop is in a closed conformation, suggesting a mechanism for lipid exchange. SPF forms a complex with RRR-alpha-tocopherylquinone, suggesting a link between oxidized Vitamin E and cholesterol biosynthesis [4].
SCP-2 has multiple roles in isoprenoid metabolism including stimulating cis-prenyltransferase to form longer polyisopreoids common to both the dolichol and cholesterol biosynthetic pathways [2] as well as enhancing the final enzymatic steps involved in conversion of lanosterol to cholesterol (rev. in [1]. With a high affinity for cholesterol and phospholipids, sterol carrier protein-2 (SCP-2), also called non-specific lipid transfer protein (nsLTP), most research was initially focused on determining the role(s) of this protein in cholesterol and phospholipid metabolism. In vitro, SCP-2 transfers cholesterol [5] and phospholipids (rev. in [6] between membranes or between membranes and serum lipoproteins [1,7–11]. In keeping with these findings, studies with transfected cells overexpressing SCP-2 demonstrate that this protein increases cholesterol uptake [12], intracellular cholesterol cycling [13,14], cholesterol oxidation (rev. in [1,15], cholesterol esterification [12,16], and phospholipid formation [17,18]. In contrast, SCP-2 overexpression inhibits HDL-mediated cholesterol efflux through lipid rafts/caveolae [19]. In vivo studies with liver from wild-type mice, rats treated with antisense SCP-2 RNA, and mice wherein the SCP-2/SCP-x gene is ablated indicate that SCP-2 stimulates cholesterol esterification in liver [20], cholesterol hypersecretion into bile [21,22], delivery of newly synthesized cholesterol from the endoplasmic reticulum (ER) for secretion into bile [23], and oxidation of branched-side chain lipids not only in vitro [24–26] but also in mice [20,27–30]. Specifically, SCP-x oxidizes the branched-side chain of cholesterol (to form bile acids) as well as the branched-chains of fatty acids (to detoxify) [24,27–29]. In summary, while compelling evidence indicates important direct roles for SCP-2 in mediating cholesterol, phospholipid, and fatty acid metabolism, exciting recent data indicate that this protein may have additional physiological functions, especially in intracellular lipid signaling.
Gene structure
One gene encodes SCP-2 and SCP-x, with the latter arising from an alternate transcription site (rev. in [30,31]. Separate mRNAs are transcribed for full-length 58 kDa SCP-x and a precursor 15 kDa pro-SCP-2 proteins which share the same C-terminus structure. Partial posttranslational modification cleaves the 58 kDa SCP-x into a 46 kDa protein and fully functional 13 kDa SCP-2. Only the 13 kDa SCP-2, but not the 15 kDa pro-SCP-2, is detected in intact cells/tissues indicating that pro-SCP-2 undergoes complete post-translational cleavage to fully functional 13 kDa SCP-2 and a 2 kDa peptide. In vitro and in vivo studies have shown that 58 kDa SCP-x as well as its 46 kDa post-translational cleavage product are peroxisomal 3-ketoacyl-CoA thiolases functioning in oxidation of branched-chain lipids (cholesterol to form bile acids; branched-chain fatty acid detoxification) (rev. in [30,31]. In vitro SCP-2 enhances cholesterol transfer from other intracellular membranes to mitochondria as well as from the outer to the inner mitochondrial membrane for oxidation, albeit less effectively than steroidogenic acute regulatory protein (StAR) [1,32–35]. The function, if any, of the 2 kDa post-translational cleavage product of the 15 kDa pro-SCP-2 is unknown (rev. in [31].
Protein Structure
The structure of SCP-2 has been determined by time-resolved fluorescence spectroscopy, circular dichroism, NMR, and x-ray crystallography as recently reviewed in detail [36]. Therefore, only those aspects relating structure to function will be briefly covered here. SCP-x/SCP-2 is comprised of at least four functionally important domains:
First, while almost nothing is known about the tertiary or secondary structure of SCP-x, its functional significance has become the subject of intense interest. The N-terminal 46 kDa is a 3-ketoacyl-CoA thiolase enzyme, involved not only in peroxisomal oxidation of straight-chain fatty acids, but is the only known 3-ketoacyl-CoA thiolase enzyme specific for oxidizing the branched-chain fatty acids and the branched-side chain of cholesterol. In contrast to mitochondrial fatty acid oxidation (net production of ATP), SCP-x does not mediated oxidations resulting in net energy production, but enhances detoxification and metabolic remodeling of branched-chain lipids. Branched-chain and straight-chain fatty acids differ markedly in their origin (isoprenoid vs classic fatty acid biosynthesis pathway), toxicity (highly toxic branched-chain vs much less toxic straight-chain), and oxidative pathway (peroxisomal vs mitochondrial) (rev. in [20,24,30,37–44]. Peroxisomal SCP-x mediated oxidation of the branched-side chain of cholesterol is essential for bile acid biosynthesis [27,29,45]. Interestingly, SCP-x expression in mice is gender dependent (female ≪ male) and female mice are much more susceptible to the toxic actions of dietary branched-chain fatty acids [40]. The first report of a spontaneous mutation in human SCP-x indicated a lipid metabolic phenotype reflecting loss of 3-ketoacyl-CoA thiolytic activity with branched-chain fatty acids and pathology similar to that noted in SCP-x/SCP-2 gene ablated mice [46].
Second, the N-terminal 20 amino acid presequence of 15 kDa pro-SCP-2 plays a very important role in intracellular targeting of SCP-2 (rev. in [31]. While examination of the amino acid and cDNA sequence of SCP-x/SCP-2 suggests that the 15 kDa pro-SCP-2 is product of an alternate transcription site and/or C-terminal cleavage of SCP-x (rev. in [31], ablation of the SCP-x gene component (but not SCP-2 gene expression) in mice did not reduce expression of SCP-2 in primary cultured hepatocytes, indicating that the SCP-x transcript is not essential for production of SCP-2 [47]. Comparison of the structures (by circular dichroism) and accessibility of the C-terminal AKL peroxisomal targeting sequence (by mass spectrometry) indicated that the C-terminal AKL peroxisomal targeting sequence of the 13 kDa SCP-2 is only slightly exposed to the aqueous buffer [48]. In contrast, presence of the 20 kDa N-terminal presequence in 15 kDa pro-SCP-2 alters the conformation of the protein to increase aqueous exposure of this C-terminal AKL peroxisomal targeting sequence [48]. The following section on intracellular targeting further details the functional consequence of the structural modification elicited by the presence of the N-terminal 20 amino acid presequence.
Third, the N-terminal 32 amino acids of the mature 13 kDa SCP-2 represents an amphipathic α-helical domain required for interaction of the protein with membranes to elicit ligand binding/transfer (rev. in [36]. This amphipathic α-helix directly interacts with the polar head groups of anionic phospholipids [49,50] and, as shown recently, represents the interaction domain for SCP-2 binding to caveolin-1 at the plasma membrane cytofacial leaflet [51,52]. Anionic phospholipids and caveolin-1 are highly enriched in the cytofacial leaflet of caveolae/lipid rafts—plasma membrane microdomains wherein multiple proteins involved in signaling are localized (rev. in [53].
Fourth, the β-strands 4,5 and helix D along with the hydrophobic faces of the N-terminal amphipathic α helix in the mature 13 kDa SCP-2 form a very large ligand binding cavity suitable for accommodating multiple types of ligands—see below (rev. in [36]. Although molecular details comprising the fatty acid binding site in SCP-2 have been established (rev. in [36], nothing is known regarding the specific amino acid residues that interact with other bound ligands in the SCP-2 binding pocket.
Tissue and intracellular distribution
The tissue and intracellular distribution of SCP-x and SCP-2 were previously reviewed (rev. in [7,31,54,55]. Briefly, SCP-x/SCP-2 gene products are: (i) highly expressed in tissues involved in cholesterol trafficking and oxidation: liver, intestine, adrenal, testis, and ovary [7]. (ii) SCP-x is almost exclusively peroxisomal [1,56], (iii) SCP-2 is present at highest concentration in peroxisomes, but half or more is extraperoxisomal in cytoplasm [1,56–58]. The molecular basis for SCP-2 being present in both peroxisomes and cytoplasm was recently resolved by examining structure (see above) and distribution of SCP-2/SCP-x gene products in transfected cells overexpressing the 15 kDa pro-SCP-2 and 13 kDa SCP-2 [31,59]. Overexpression of the 15 kDa pro-SCP-2 resulted in significant localization in peroxisomes, but with nearly 50% localized in extraperoxisomal sites [48]. In contrast, overexpression of the mature 13 kDa SCP-2 resulted in SCP-2 being largely extraperoxisomal [48]. The differential distribution of these proteins was due weak exposure of the C-terminal AKL peroxisomal targeting sequence of the 13 kDa SCP-2 to the aqueous buffer while that of 15 kDa pro-SCP-2 was more exposed (see above). Thus, the 15 kDa pro-SCP-2 (gene product of the second promoter region) may better interact with the peroxisomal membrane AKL receptor and/or peroxisomal membrane proteins involved in 15 kDa pro-SCP-2 translocation into the peroxisomal matrix [48]. These findings suggest that the relative proportion of SCP-2 appearing in peroxisomes vs cytoplasm is likely attributed to the extent of N-terminal 20 amino acid cleavage of the 15 kDa pro-SCP-2 prior to entry into the peroxisomal matrix. However, the specific protease which accomplishes this cleavage and the intracellular localization of this protease are not yet known.
Lipids involved in intracellular signaling
Although lipids were originally construed primarily as components of membrane bilayers and energy storage, more recent data indicate that many lipid molecules also participate in intracellular signaling. Table 1 provides a summary of lipid classes involved in intracellular signaling and some selected examples of physiological responses elicited by these lipids. Because of space limitations in this review, it is not possible to provide a detailed, exhaustive listing of all signaling pathways in which these lipids participate. However, the reader is encouraged to the cited articles and specialized reviews noted in Table 1. Although structures of these lipids are diverse, all include a hydrophobic acyl chain (fatty acids, fatty acyl CoAs, lysophosphatidic acid, phosphatidylinositides, sphingolipids) or a hydrophobic side chain (cholesterol). While there are many other lipidic molecules involved in signaling (isoprenoids, prostaglandins, leukotrienes, thromboxanes, etc), only molecules with evidence of interaction with SCP-2 are listed. For example, by interacting with isoprenoids SCP-2 may influence signaling pathways in both the cytoplasm and nucleus. SCP-2 stimulates cis-prenyltransferase to form longer polyisopreoids [2]. The finding that SCP-2 has high affinity for isoprenyl pyrophosphates [e.g. dimethylallyl pyrophosphate, farnesyl pyrophosphate, geranyl pyrophosphate, geranyl geranyl pyrophosphate [60]] suggests potential roles in regulating the activity of isoprenyl transferase enzymes that covalently modify (e.g. farnesylate, geranylate) proteins—resulting in targeting of these protein to caveolae/lipid rafts, plasma membrane microdomains highly involved in cell signaling. However, there have been no reports further investigating this interesting possibility. Finally, SCP-2 binds [60] and enhances the peroxisomal metabolism of branched-chain fatty acids in cultured cells [61], and mice [20]. Branched-chain fatty acids, isoprenoids derived from cleavage of chlorophyll side chain (rev. in [20,61], are nuclear signaling molecules representing the most potent endogenous inducers of peroxisome proliferator activated receptors known [62–67]. Finally, SCP-2 also facilitates the peroxisomal oxidation of the branched-side chain of cholesterol to form bile acids [1,20,68]. Bile acids are also potent ligand signaling molecules in the nucleus where they interact with and regulate transcriptional activity of nuclear receptors, e.g. FXR (rev. in [31,69,70]. Although these findings suggest that by binding these branched-chain isoprenoids or facilitating their formation SCP-2 may influence their trafficking/availability to the nucleus for nuclear signaling, these possibilities remain to be investigated.
Table 1.
Lipids involved in intracellular signaling.
| Lipid Class | Responses | References |
|---|---|---|
| Straight-chain fatty acids | Regulate diacylglycerol kinase; sodium channel regulator; Tyr phosphorylation; neutrophil function; cell-cell signaling; stimulate transcriptional activity of nuclear receptors such as PPARα | [200–209] |
| Branched-chain fatty acids | Stimulate transcriptional activity of nuclear receptors such as PPARα | [62,62–67] |
| Fatty Acyl CoAs | Activate K+ channel; Stimulate Ca++ release; differentially regulate PKC subtypes; enhance vesicle budding, release and transport; protein transport; stimulate transcriptional activity of nuclear receptors such as PPARα and HNF4α. | [208–223] |
| Lysophosphatidic acid | Membrane fission; vesicle trafficking | [224,225] |
| Phosphatidylinositols | Phospholipase C signaling | [51,165,169,170,172] |
| Sphingomyelin | Substrate for ceramide formation | [149] |
| Ceramide | Enhances cholesterol efflux to apo-A1; Activates eNOS; | [75,226,227] |
| Glucosylceramide | Modulates endocytic trafficking | [228] |
| Galactosylceramide | Ca++ flux; oligodendrocyte development | [229] |
| Lactosylceramide | Activates Ras and MAP kinase phosphorylation of Raf ; simulates endocytosis | [229–231] |
| Globoside | P determinant in the P system of blood groups | [229] |
| Ganglioside GM1 | Tyr phosphorylation; PI turnover; Ca++ flux; Inhibits beta-adrenergic receptor-induced cAMP formation | [149,229] |
| Farnesyl-PP Geranyl-PP | Isoprenylate or geranylate signaling proteins targeted to caveolae/rafts????? | [60] |
| Cholesterol | Required for caveolae/raft; stimulates caveolar endocytosis; Intracellular cholesterol sensor pool regulating release of SREBP and transcriptional activity in nucleus. _ | [75,127,144,230] |
Lipid rafts/caveolae: a conceptual framework forming the basis for lipid signaling at the plasma membrane
Signaling lipid substrates/precursors (e.g. phospholipids containing arachidonic acid, phosphatidylinositols, sphingolipids, etc) and the enzymes that produce/release them (kinases, lipases) both reside in the plasma membrane, yet do not interact to produce signaling cascades until induced by receptor activation at the cell surface. It has been hypothesized that cholesterol-rich lipid rafts/caveolae provide a conceptual organizing framework for these transmembrane signaling pathways (rev. in [53,71–75]. It is thought that at least two types of these cholesterol-rich microdomains exist: (i) lipid rafts, and (ii) caveolae, a subset of lipid rafts (Fig. 1). These plasma membrane microcomains are believed to be enriched not only in signaling proteins, but as indicated in Figure 1 also in signaling lipids.
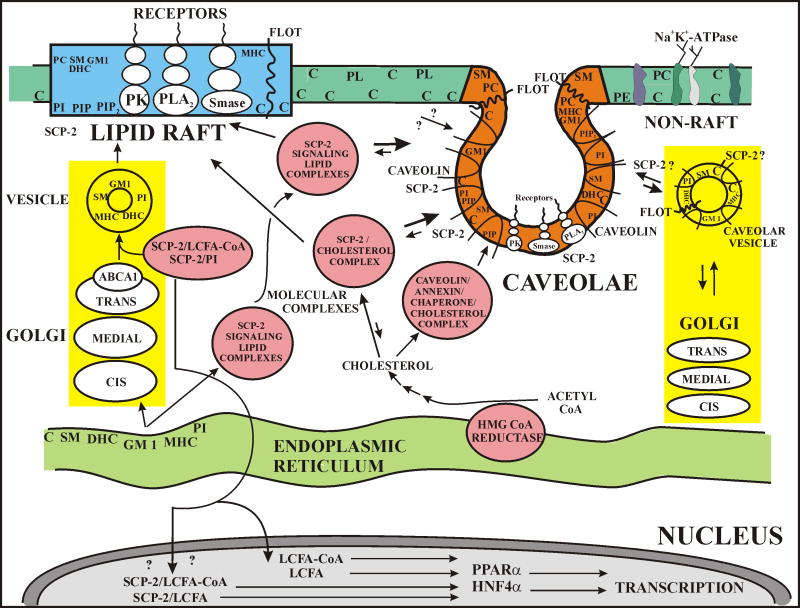
Fig. 1. SCP-2 mediates signaling and lipid trafficking to plasma membrane lipid rafts and caveolae.
At the plasma membrane, blue shading refers to lipid rafts, red shading refers to caveolae (a lipid raft subfraction), and dark green shading refers to non-rafts. Within the cell, yellow shading refers to vesicular pathways, salmon (dark) pink shading refers to molecular complexes of SCP-2 with SM, DHC, GM1, MHC, PI, cholesterol. Light pink shading refers to molecular complexes of SCP-2 with fatty acyl-CoA or PI. Caveolin (∩), flotillin (FLOT), sterol carrier protein-2 (SCP-2), protein kinase (PK), phospholipase A2 (PLA-A2), phospholipase C (PL-C), peroxisome proliferator activated receptor α (PPARα), hepatocyte nuclear factor 4α (HNF4α), cholesterol (C), phosphatidylinositol (PI), phosphatidylinositolphosphate (PIP), phosphatidylinositolbisphosphate (PIP2), sphingomyelin (SM), ceramide (CER), monohexosylceramide (MHC), dihexosylceramide (DHC), globoside (G), ganglioside M1 (GM1), phosphatidylcholine (PC), phosphatidylserine (PS), phosphatidylethanolamine (PE).
While lipid rafts/caveolae provide an attractive conceptual framework for organizing lipid substrates and proteins, the precise definition and even the existence of cholesterol-rich lipid rafts/caveolae is a continuing subject of discussion (rev. in [53,71,75–85]. The idea of lipid domains (rafts?, caveolae?) is based on early model membrane studies which demonstrate the physical immiscibility of different lipid classes into multiple coexisting phases: liquid crystalline (most fluid), liquid ordered (intermediate, detected in biochemically isolated lipid rafts/caveolae), and gel (least fluid) (rev. in [86–88]. Depending on the specific marker used, the size of lipid rafts obtained through morphological studies of model membranes ranges from 260–330 nm for fluorescent tagged GM1 and 230–2000 nm for fluorescent tagged PE (rev. in [53,81,82,89]. Since cholesterol induces formation of phase separated cholesterol-rich and –poor domains in simple two component model membranes, it has been proposed that cholesterol forms the organizing principle of lipid ‘domain’ or ‘microdomain’ formation in membranes (rev. in [90]. The finding that plasma membranes of mammalian cells are highly enriched in cholesterol (70–90% of cell total cholesterol) suggested the potential existence of cholesterol-rich and –poor regions therein. Indeed, early biochemical subfractionation studies resolved cholesterol-rich and –poor plasma membrane subfractions (apical/serosal/sinusoidal, basolateral/contiguous, microvillar/canalicular plasma membrane subfractions) differing in lipid composition, protein distribution, and structure from polarized cells (liver, intestine, kidney) (rev. in [91]. It is important to note that microvillar/brush border/canalicular subfractions represent highly dynamic regions of the plasma membrane where absorption (intestinal microvilli) and secretion (kidney brush border/hepatic canaliculi) of lipids such as cholesterol occur. These microvillar/brush border/canalicular subfractions are (i) highly enriched in cholesterol and sphingomyelin (rev. in [92], (ii) less fluid (rev. in [92], and (iii) rich in proteins involved in reverse cholesterol transport (i.e. SRB1, caveolin-1, ABCA1) which are also enriched in ‘lipid rafts/caveolae’ [85,93–97]. Likewise, spicules/pseudopodia blebbing off the surface of deoxygenated red blood cells are enriched in sterol and exhibit high sterol dynamics [98]. Affinity chromatography, electrophoresis, ionic lysis, and gradient centrifugation also resolve plasma membranes from non-polarized cells into subfractions differing in lipid composition and/or protein distribution [92,99–110]. Concurrent with these studies over three decades ago, morphologically distinct flask-like (60–90 nm diameter) structures were observed in plasma membranes and called ‘caveolae’ or little caves (rev. in [106]. Again, depending on the specific marker used, the size of lipid rafts obtained through morphological studies of biological membranes ranged from 26–330 nm for GPI-anchored proteins and 500–2000 for MHC-1 (rev. in [53,81,82]. This indicates that some lipid rafts may be so small as to constitute an ‘annulus’ surrounding a specific protein, a suggestion supported by studies showing cholesterol forming an annulus around the oxytocin receptor [111–113]. Alternately, other lipid rafts may be sufficiently large (i.e. 230–2000 nm) to be visible by optical microscopy (limit of resolution near 200 nm).
Despite the above findings, morphological studies did not address whether the caveolae were cholesterol-rich or –poor. To begin to resolve these questions, a variety of biochemical techniques were used to isolate cholesterol-rich fractions from cholesterol-containing model membranes and from biological membranes through use of detergents, high-pH carbonate buffer, density gradient centrifugation, and affinity-chromatography (rev. in [53]. These biochemically isolated fractions were selectively lipid rich (especially in cholesterol and sphingomyelin), exhibited a liquid-ordered structure (i.e. less fluid), and were subsequently termed detergent resistant membranes (DRM), lipid rafts, caveolae, etc. Whether caveolin-1 is present in these fractions is cell type dependent [e.g. hepatocytes do not contain caveolin-1 [114]]—indicating that caveolin-1 is not required for formation of these cholesterol- and sphingomyelin-rich fractions (rev. in [53]. Finally, since the existence of cholesterol-rich and –poor domains in plasma membranes of living cells had not been demonstrated, it was unclear whether the biochemically isolated lipid rafts/caveolae, DRM, etc represented artifactually induced enrichments of cholesterol in these fractions. This issue was resolved recently by real-time multiphoton imaging of a naturally-occurring fluorescent sterol (dehdyroergosterol) in the plasma membrane of living cells [53,115–118]. The multiphoton imaging studies showed that sterol was not uniformly distributed in the plasma membrane of L-cell fibroblasts, but instead into sterol-rich and sterol-poor regions in plasma membranes of living cells. The size range of sterol-rich domains was estimated to be from 200 nm (limit of optical microscopy) to 565 nm [116]. This size range was generally smaller than or, at the high range, overlapped the lower range of sizes typically observed for microvilli which were in the range of 500–1000 nm as determined by electron microscopy [110,119–121]. While video imaging of HepG2 cells confirmed the sterol-rich (especially canalicular microvilli) and –poor distribution in the plasma membrane of polarized cells [97]. However, the video imaging studies resolved regions in the size range of 2000–3000 nm and it was concluded that this represented microvilli and that there were no ‘sterol-rich’ vs ‘sterol-poor’ regions [97]. This conclusion is difficult to reconcile with existing data showing that: (i) microvilli (rev. in [92] and pseudopodia [98] isolated by biochemical fractional are sterol-rich; (ii) morphological (electron microscopy, immunofluorescence) studies of fixed cells and biochemical fractionation studies using the non-cytolytic perfringolysin O (highly selective for cholesterol-rich regions) confirmed the existence of cholesterol-rich and –poor domains in microvilli/filopodia in lymphoblastoid cells and platelets [122,123]; (iii) morphological (electron microscopy, immunofluorescence) of fixed cells and biochemical fractionation studies using the non-cytolytic perfringolysin O confirmed the existence of cholesterol-rich and –poor domains in lipid rafts of erythrocytes [123]; (iv) Morphological studies with filipin (disrupts cholesterol-rich lipid rafts) indiscriminately labeled cholesterol (both cholesterol-rich and –poor) regions of the plasma membrane and could not differentiate between them [122,123]. The reasons for the apparent discrepancy with results obtained by video imaging vs multiphoton imaging (as well as morphological studies with the perfringolysin O) is not clear, but may be due in part to video imaging resulting in strong photobleaching of the fluorescent sterol, the use of methyl-beta cyclodextrin to incorporate the fluorescent sterol at high amounts for video imaging, saturation of all the plasma plasma membrane sterol with fluorescent sterol for video imaging, and video imaging’s weak resolution in the Z-axis. In contrast, the multiphoton imaging studies used bioincorporation (small phospholipid/dehydroergosterol vesicles) of small amounts of dehydroergosterol, multiphoton excitation results in much less photobleaching of the fluorescent sterol, and multiphoton imaging has resolution in the Z-axis only slightly less than confocal imaging. Taken together, the bulk of data suggest that cholesterol-rich and poor ‘domains’ exist in plasma membranes of living cells. Microvilli, plasma membrane regions associated with high cholesterol flux, are also enriched in cholesterol, sphingomyelin, and protein markers typically also associated with ‘lipid rafts/caveolae’. The upper range of sizes obtained with multiphoton imaging for some sterol-rich ‘domains’ appears to overlap with the lower range of microvilli/filipodia, suggesting that some cholesterol-rich lipid rafts/caveolae may cluster in microvilli/filipodia. However, the resolution of optical microscopy (200 nm) is insufficient to resolve this issue.
In summary, there is as yet no universal definition of lipid rafts/caveolae is lacking. Thus, for the sake of simplicity and to avoid semantic issues in the current review we will use the term ‘lipid rafts/caveolae’ to refer to cholesterol-rich, specific marker-rich, domains isolated from purified plasma membranes without the use of detergents or high pH carbonate buffers. It is not completely clear whether DRMs are equivalent to lipid rafts/caveolae (rev. in [53].
Lipid rafts/caveolae microdomains involved in lipid signaling: Preparations based on detergents or high pH carbonate buffers
Early biochemical fractionation studies initially suggested that plasma membrane lipid rafts/caveolar microdomains were lipid-rich, especially in cholesterol and sphingolipids, but relatively poor in other structural lipids such as phospholipids (especially phosphatidylcholine) such that the cholesterol/phospholipid molar ratio therein was several-fold higher than the non-raft component (rev. in [124]. The relative proportion of lipid raft/caveolae vs non-raft domains in plasma membranes varied from a few % to nearly all the membrane, depending on the type of detergent used, temperature, pH, and other conditions (rev. in [124]. Thus, it was not clear whether lipid rafts/caveolae represented a minor or a major part of the plasma membrane. Significant to the current review, such detergent- and high pH carbonate- based lipid raft/caveolae preparations were also enriched in high levels of signaling proteins (rev. in [124] and signaling lipids [phosphatidylinositol (PI), phosphatidylinositol-4-phosphate (PIP), phosphatidylinositol-4,5-bisphosphate (PIP2), sphingomyelin, gangliosides, ceramide, and diacylglycerol] (rev. in [72,125–132]. However, detergent or high pH carbonate based lipid raft/caveolae preparations are increasingly being criticized as producing artifactual distributions of proteins and/or lipids into lipid rafts/caveolae, i.e. not reflecting the physiological distribution of these molecules in plasma membranes (rev. in [133–140]. One recent proteomics/mass spectrometry study comparing the purity of detergent- and high pH carbonate-based lipid raft/caveolae preparations showed that almost 33% and 75% of proteins, respectively, were non-specific contaminant copurifying proteins [141].
Lipid rafts/caveolae microdomains involved in lipid signaling: Preparations based on density gradients and affinity chromatography
Density gradients [53,61,114,140,142–149] or density gradients/affinity chromatography methods [53,61,110,114,140,149] do not use detergents or high pH carbonate buffer. Biochemical analysis of marker protein distribution in lipid rafts/caveolae and non-rafts isolated from purified plasma membranes from cultured cells (e..g. L-cell fibroblasts, MDCK cells, endothelial cells, primary hepatocytes) by density gradient centrifugation and affinity chromatography demonstrated that lipid rafts/caveolae were enriched (on the basis/mg protein) in appropriate caveolae/lipid raft protein (caveolin-1, except in hepatocytes; flotillin; SRB1; eNOS in endothelial cells only) and lipid (GM1, cholesterol, sphingomyelin, total lipid) markers while concomitantly being relatively poor in non-raft protein (Na+,K+-ATPase; transferrin in endothelial cells) and poor lipid markers (GM1, cholesterol, sphingomyelin, total lipid) [61,115,140,147]. Furthermore, neither fraction was enriched in intracellular protein markers, e.g. calnexin—an endoplasmic reticulum marker [61]. The presence of caveolin-1 was not required for the existence of lipid rafts, as shown by lipid rafts isolated from cultured mouse primary hepatocytes and Fisher rat thyroid (FRT) cells—which do not contain caveolin-1 [114,150–152]. Lipid markers (cholesterol, phospholipid, sphingomyelin, GM1) were also enriched in lipid rafts/caveolae and did not require the presence of caveolin-1 [114]. Again the latter enrichments did not require the presence of caveolin-1 [114]. A typical biochemical analysis of lipid distribution in lipid rafts/caveolae and non-rafts isolated from plasma membranes purified from cultured fibroblasts is shown in Table 2. In control fibroblasts, lipid rafts/caveolae were enriched 2.5-fold in cholesterol, 3.7-fold in phospholipid, and 3-fold in total lipid as compared to non-rafts (Table 2). Phospholipids, especially phosphatidylcholine, were not depleted in lipid rafts/caveolae isolated from fibroblasts [61] or in lipid rafts isolated from cultured primary hepatocytes [114]. Consequently, the cholesterol/phospholipid molar ratio in lipid rafts/caveolae and lipid rafts did not differ significantly from non-rafts in fibroblasts (Table 2) or in cultured primary hepatocytes [114]—in agreement with other methods not using detergents or high pH carbonate buffers for isolation (rev. in [53]. Taken together, these and earlier findings with non-detergent, non-high pH carbonate buffer based isolation procedures suggest that benchmark markers for the caveolae/lipid raft preparations should include enrichment in caveolin-1, GM1, cholesterol, and sphingomyelin (all based on/mg protein) as compared to non-rafts. As a negative control, the caveolae/lipid rafts should be deficient in non-raft (e.g. Na+,K+-ATPase; clathrin; transferring in some cell types) and intracellular membrane (e.g. calnexin for ER; golgin 97 and giantin for Golgi) markers.
Table 2.
Structural lipids enriched in lipid rafts/caveolae of cultured fibroblastsa.
| Lipid Class | Control | SCP-2 Overexpressor | ||
|---|---|---|---|---|
| Non-raft | Lipid raft/caveolae | Non-raft | Lipid raft/Caveolae | |
| Cholesterol | 380±96 | 963±82* | 442±99 | 857±128*,+ |
| Phospholipid | 330±55 | 1221±150* | 681±116 | 871±136+ |
| C/PL Ratio | 1.1±0.3 | 0.6±0.1 | 0.7±0.1 | 1.0±0.2 |
| Total Lipid | 1016±126 | 3046±196* | 1270±158 | 1910±186*,+ |
Lipid raft/caveolae and non-raft domains were isolated from purified plasma membranes of cultured L-cell fibroblasts using density gradient centrifugation and concanavalin-A affinity chromatography [61]. Values represent nmol/mg protein ± SD (n=3–4). Asterisk = p<0.05 vs control; dagger = p<0.05 vs non-raft.
Lipid rafts/caveolae isolated without detergents/high pH carbonate buffer (see below) were enriched in both signaling proteins and in signaling lipids (rev. in [53]. Taken together, these data alleviated concerns that, in case of select signaling proteins [141] and select signaling lipids such as sphingomyelin and phosphatidylinositol [146], the fundamental qualitative (but not quantitative) conclusions about lipid rafts/caveolae being enriched in signaling proteins and signaling lipids remained valid—regardless of the method of lipid raft/caveolae isolation [61]. This conclusion is underscored by the fact that disruption of cholesterol-rich microdomains inhibits these signaling activities [124].
Signaling lipids are enriched in lipid rafts/caveolae
The enrichment of signaling lipids in caveolae and non-caveolar rafts and their putative intracellular trafficking pathways mediated by SCP-2 are schematically illustrated in Fig. 1. Multiple classes of signaling lipids are enriched in lipid rafts/caveolae isolated without detergents or high pH carbonate buffer (Table 3). For example, fatty acids are enriched 1.6-fold in lipid rafts/caveolae as compared to non-rafts isolated by affinity chromatography from purified plasma membranes derived from cultured fibroblasts (data not shown). Likewise, fatty acids are enriched 4.6-fold in lipid rafts as compared to non-rafts isolated by affinity chromatography from plasma membranes purified from cultured primary hepatocytes (Table 3 and [114]. These findings were consistent with the fact that fatty acid uptake across the plasma membrane into cells occurs through cholesterol-rich, sphingolipid-rich lipid rafts/caveolae [153]. Plasma membranes contain several proteins that bind and transport fatty acids: caveolin-1 [154], FAT/CD36 (rev. in [155–157], and FATP [158]. Within plasma membranes, all these fatty acid transport/translocase proteins are enriched in lipid raft/caveolae microdomains: caveolin-1 [53,61,114,140,142–146,149], FAT/CD36 (rev. in [74,159–161], and FATP (rev. in [160–162].
Table 3.
Signaling lipids are enriched in lipid rafts/caveolae.
| Lipid Class | Fold-Enrichment Relative to Non-Rafts | References |
|---|---|---|
| Fatty acids | 4–12 | [114] |
| PI, PIP, PIP2 | 3–16 | [61,71,124,232,233] |
| Total sphingolipid | 2 to >100 | [149] |
| Sphingomyelin | 1.7 to >100 | [61,131,149] |
| Ceramide | Yes/No | [72,149] |
| Monohexosides | 7 | [149] |
| Dihexosides | Yes/No | [149] |
| Globoside | 5 | [149] |
| Ganglioside GM1 | 2–5 | [53,61,114,149] |
| Total gangliosides | >20 | [131,149] |
| Cholesterol | 2–3 | [53,61,114,131,142,143,146,149,234] |
Lipid rafts/caveolae are highly enriched (3–16 fold, Table 3) in phosphatidylinositol (PI) [61,114], phosphatidylinositol-4-phosphate (PIP) [125], and phosphatidylinositol-4-5-bisphosphate (PIP2) [125]. PI serves as a substrate for kinases that produce phosphorylated forms of PI (PIP, PIP2) while PIP and PIP2 in turn serve as substrates for phospholipase enzymes present in lipid rafts/caveolae resulting in release of phosphoinositide messengers and intracellular signaling (Table 1).
Sphingolipids represent a complex group of subclasses involved in lipid signaling and many are highly enriched in lipid rafts/caveolae, 1.7 to >100-fold (Table 3). Sphingomyelin, ceramide, neutral glycosphingolipids (monohexoside, dihexoside, globoside), and acidic glycosphingolipids (gangliosides) are enriched 2–20+ fold (Table 3) in lipid rafts/caveolae as compared to non-raft domains of fibroblasts, primary hepatocytes, and/or MDCK cells [53,61,114,140,149]. Interestingly, incorporation of a photoactivatable ganglioside GM1 into lipid rafts/caveolae, followed by photocross-linking demonstrated that ganglioside cross-links to caveolin-1 in plasma membrane caveolae [163]. Thus, caveolin-1 may represent a close neighbor of gangliosides, a ganglioside binding protein within lipid rafts/caveolae, and/or may possibly transport/target gangliosides to caveolae through vesicular or chaperone-complexed caveolin-1 docking/interaction with plasma membrane caveolae [163]. Since caveolin-1 is not present in hepatocytes, caveolin-1 may be important for regulating the distribution and/or trafficking of signaling lipids to caveolae in tissues/cells (e.g. fibroblasts, endothelial cells) highly enriched in caveolin [78,79,164]. Although SCP-2 is not normally present in significant amounts in fibroblasts and endothelial cells, in fibroblasts overexpressing SCP-2 this protein is detected in caveolae in very close proximity to caveolin-1 [51,165]. These data suggest that both of caveolin-1 and SCP-2 may in part regulate the distribution of signaling lipids within caveolae (Fig. 1). Although caveolin-1 is not present in plasma membranes or plasma membrane lipid rafts isolated from cultured primary hepatocytes, hepatocytes contain high level of SCP-2 and signaling lipids (PI, GM1) are enriched in plasma membrane lipid rafts [114]. Thus, depending on the cell type SCP-2 and/or caveolin-1 may contribute to retention, binding, or trafficking of signaling lipids to lipid rafts (Fig. 1).
SCP-2 exhibits high affinity for lipids participating in intracellular signaling
Lipid signaling molecules are in general highly membrane-bound, since these lipophilic molecules are relatively insoluble in the aqueous environment of cytoplasm. Cells have evolved multiple mechanisms (Fig. 1) to transfer lipid signaling molecules between intracellular sites of synthesis (ER) and utilization (lipid rafts/caveolae) (rev. in [53,71,79,124,166,167]. Newly synthesized cholesterol and signaling lipids (or their substrates) move between Golgi and plasma membrane by one or more mechanisms: (i) relatively slowly (10–20 min) as vesicles from golgi to lipid rafts (Fig. 1, Vesicle) or caveolae (Fig. 1, Caveolar Vesicle), or (ii) more rapidly (1–2 min) by protein-mediated molecular transport (Fig. 1, SCP-2/signaling lipid complexes; Fig. 2A) (rev. in [19,168]. Lipid binding proteins may also regulate vesicular trafficking [169–173], by binding signaling molecules involved in vesicle budding such as fatty acyl CoA and phosphatidylinositides (Fig. 1, SCP-2/LCFA-CoA and SCP-2/PI mediating vesicle budding from golgi). Consistent with these possibilities, overexpression of SCP-2 in cultured fibroblasts enhanced molecular transfer of a fluorescent cholesterol (i.e. SCP-2/sterol complex), while concomitantly decreasing vesicular transfer of cholesterol to lipid rafts/caveolae for efflux to HDL—likely by binding fatty acyl CoA and/or phosphatidylinositides necessary for vesicle budding [19].
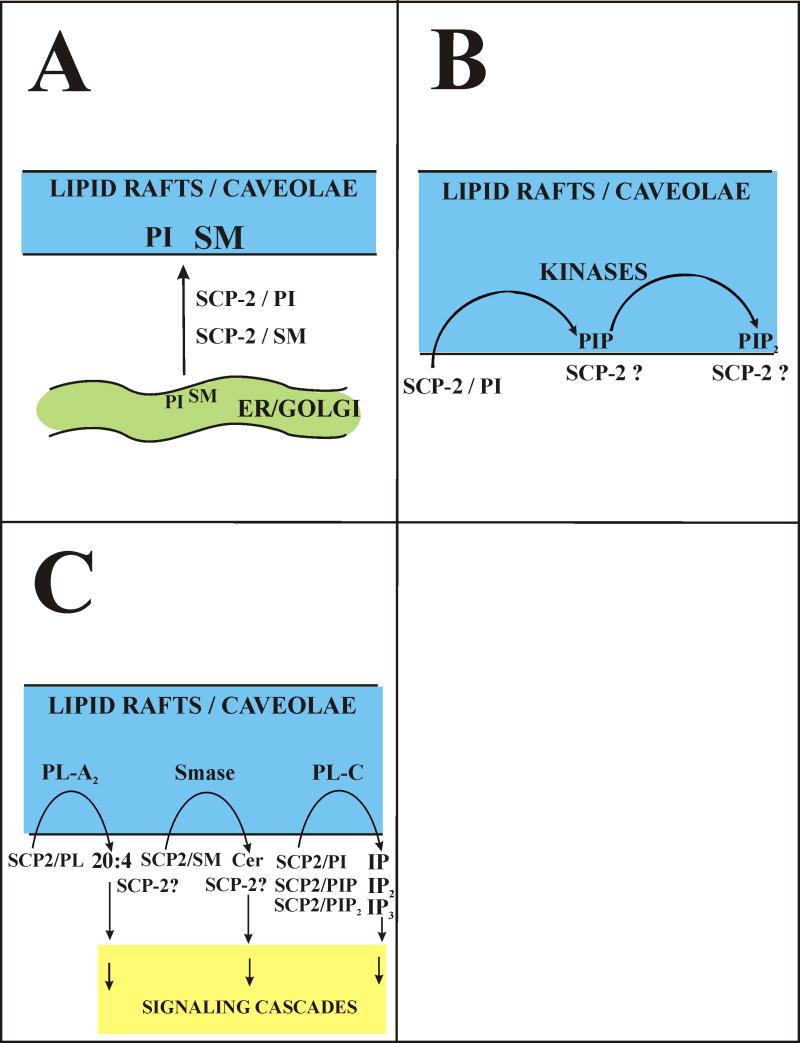
Fig. 2. Putative mechanisms whereby SCP-2 mediates signaling and lipid trafficking to plasma membrane lipid rafts/caveolae.
Panel A: Complexes of SCP-2/PI or SCP-2/SM transport signaling lipids and/or signaling lipid precursors (i.e. PI, SM) from intracellular sites of synthesis (ER, Golgi) to plasma membrane lipid rafts/caveolae. Panel B: Complexes of SCP-2/PI or SCP-2/PIP are better substrates than the unbound PI or PIP for kinases that produces PIP and PIP2, respectively. Panel C: Complexes of SCP-2/PL, SCP-2/SM, SCP-2/PI, SCP-2/PIP, and SCP-2/PIP2 are better substrates than the unbound lipids for lipases that release signaling lipids (arachidonic acid, ceramide, IP, IP2, IP3). Phospholipase A2 (PLA-A2), sphingomyelinase (SMase), phospholipase C (PLA-C), phospholipid (PL), phosphatidylinositol (PI), phosphatidylinositolphosphate (PIP), phosphatidylinositolbisphosphate (PIP2), inositol phosphate (IP), inositol bisphosphate (IP2), inositol triphosphate (IP3), sphingomyelin (SM), ceramide (CER).
A fundamental requirement for protein-mediated lipid transport is that the protein binds the signaling lipid with high affinity. Mammalian cells express not only specific proteins that directly interact with select, signaling lipids in select tissues/cells (fatty acid binding proteins, phosphatidylinositol binding protein, glycolipid binding protein, etc), but also the more promiscuous SCP-2 which binds multiple signaling lipids, is ubiquitously distributed in all tissues, and is expressed at highest level in liver, steroidogenic tissues (adrenal, testis, ovary), and intestine [31]. Sterol carrier protein-2 (SCP-2) exhibits high affinity for many lipid classes involved in intracellular signaling (Table 4). Its broad range of affinities for several of these ligands reflects the influence of acyl chain composition as well as from use of fundamentally different assays. Classical radioligand binding and separation assays typically yield much weaker affinities (i.e. higher Kds) than those obtained with fluorescence and isothermal titration calorimetry techniques that do not require separation of free from bound ligand (rev. in [174–179]. Some of these signaling lipid binding proteins are highlighted below:
Table 4.
SCP-2 exhibits high affinity for lipids participating in intracellular signaling.
| Signaling Lipid Ligand | Kd or Ki (nM)a | References |
|---|---|---|
| Fatty acids | 234–650 | [174,235–238] |
| Fatty acyl CoAs | 3–250 | [20,149,175,237] |
| Lysophosphatidic acid | 114±16 | [149] |
| Phosphatidylinositols | PIP2>PIP>PI | [51,181] |
| Sphingomyelin | 101±16 | [149] |
| Ceramide | 5±1 | [149] |
| Glucosylceramide | 101±16 | [149] |
| Galactosylceramide | 110±10 | [149] |
| Lactosylceramide | 116±9 | [149] |
| Globoside | 121±2 | [149] |
| Ganglioside | 43±6 | [149] |
| Cholesterol | 4–2600 | [15,20,149,187,236–238] |
Affinities depend on specific acyl chain esterified to the lipid and on type of binding assay used.
SCP-2 exhibits high affinity for fatty acids, with Kds as low as 234 nM (Table 4). While SCP-2 has not classically been thought of as a fatty acid binding protein, its affinity for these ligands is in the same range as exhibited by members of the large family of fatty acid binding proteins (rev. in [177]. SCP-2 also has high affinity for fatty acyl CoAs, as low as 3 nM Kds (Table 4). These affinities are in the same range as acyl CoA binding protein (ACBP) (rev. in [176,180] and PITP [181]. SCP-2 shows high affinity for phosphatidylinositol (rev. in [165]. The affinities of SCP-2 for phosphorylated PI species are in the order PIP2>PIP>PI (Table 4), where SCP-2 has highest affinity for the most negatively-charged PI species. These affinities/functions of SCP-2 in PI binding are in the same range as for phosphatidylinositol transfer proteins such as PITPα and PITPβ (rev. in [165]. Although PITPα and less so PITPβ are both cytoplasmic, significant amounts are detected in perinuclear Golgi (rev. in [165]. PITP is hypothesized to directly present bound substrate to phospholipase C and interact with phosphoinositide kinase signaling complex in plasma membrane caveolae (rev. in [165]. As indicated above, nearly half of total SCP-2 is extraperoxisomal in cytoplasm, ER, and/or mitochondria (rev. in [15,48,57]. The finding that some SCP-2, but not PITP, is present at the plasma membrane and directly interacts with caveolin-1 in caveolae [51,165], suggests that in tissues rich in SCP-2 this protein may contribute significantly to protein-mediated PI transfer. SCP-2 is one of the few proteins known to exhibit high affinity for sphingolipids including sphingomyelin, ceramide, neutral glycosphingolipids (monohexosides, dihexosides, globosides), and acidic glycosphingolipids (gangliosides) (Table 4). For example, human recombinant SCP-2 has high affinity (Kds of 5–215 nM) for all sphingolipid classes tested and this affinity was selective, decreasing in the order ceramide > acidic glycosphingolipid (ganglioside) > neutral glycosphingolipid (monohexoside, globoside) > sphingomyelin (Table 4) [149].
In summary, as shown in Table 3, SCP-2 binds a variety of lipids that could serve as second messengers in several intracellular signaling pathways. To date, very little is known regarding specific signaling pathways wherein SCP-2 may function. However, at least two examples indicate that SCP-2 regulates signaling in both cytoplasm and nucleus. For example, SCP-2 overexpression stimulates insulin-mediated inositol-triphosphate production [165]. Likewise, SCP-2/SCP-x gene ablation enhances transcriptional activity of PPARα to induce formation of fatty acid oxidative enzymes [20,47]. However, it remains to be determined what is the specific (or predominant) intracellular signaling pathway(s) that SCP-2 regulates.
SCP-2 transfers intracellular signaling lipids in vitro and intact cells
SCP-2 mediates intracellular transport/targeting (Fig. 1, Fig. 2A) of many lipid classes involved in intracellular signaling (Table 5). For example, SCP-2 enhances not only cellular uptake of signaling lipids such as fatty acids [37,38,177,182], but also enhances fatty acid intracellular diffusion/transport [177,182] and fatty acid targeting to ER for incorporation into select phospholipid classes [18] or peroxisomes for oxidation [38]. Whether SCP-2 also regulates transport/presentation of fatty acids released by phospholipase A2 activity in the lipid rafts caveolae (Fig. 2C, arachidonic acid) to intracellular sites where such fatty acids (e.g. arachidonic, docosahexaenoic, eicosapentaenoic) are converted to signaling lipids (e.g. prostaglandins, leucotrienes, and/or thromboxanes) is not known. Precedent for possible involvement of SCP-2 in this process is based on studies in vitro demonstrating that a fatty acid binding protein with high affinity for linoleic acid and arachidonic acid inhibits 15-lipoxygenase induced oxygenation of these fatty acids to signaling lipids [183].
Table 5.
SCP-2 stimulates transfer of intracellular signaling lipids.
SCP-2 also binds fatty acyl CoAs with high affinity (Table 4) and enhances their transport and/or presentation to microsomal enzymes involved in fatty acyl CoA transacylation. For example, SCP-2 stimulates microsomal fatty acyl CoA incorporation into phosphatidic acid in vitro [17] and in intact cells [18]. Since fatty acyl CoAs regulate vesicle budding at the Golgi (see above) by binding acyl CoAs the SCP-2 may potentially be involved in regulating vesicular trafficking of signaling lipids as well as in their molecular transfer to the plasma membrane lipid rafts/caveolae (Fig. 1, Vesicle and Caveolar Vesicle budding from Golgi). Overexpression of SCP-2 in cultured fibroblasts not only enhances molecular sterol transfer to plasma membrane lipid rafts/caveolae for HDL-mediated efflux but simultaneously decreases vesicular sterol transfer to plasma membrane lipid rafts/caveolae to inhibit efflux to HDL [19]. Since vesicular sterol transfer predominates over molecular sterol transfer in L-cell fibroblasts, the net effect of these opposing influences was that SCP-2 overexpression inhibited HDL-mediated sterol efflux [19].
SCP-2 may also mediates phosphatidylinositol molecular transport from its site of synthesis in ER to plasma membrane lipid rafts/caveolae (rev. in [53]. In vitro studies [165] show that SCP-2 stimulates PI transport from microsomal donor membranes to acceptor membranes by 13-fold (Fig. 2A). Direct comparisons of SCP-2 mediated enhancement of PI transfer with that of another phosphatidylinositol transfer protein, PITP, revealed that the fold-stimulation of these proteins was not significantly different—suggesting that both proteins are highly effective in transferring PI from microsomes [165]. Both PITP isoforms (PITPα and PITPβ) are well known to enhance PI transfer (rev. in [165]. Further, SCP-2 overexpression significantly redistributes PI from intracellular membranes (mitochondria, ER) to plasma membranes and concomitantly downregulates expression of the other major PI binding protein (PITP) nearly 6-fold [165]. The finding that SCP-2 stimulates insulin-mediated inositol-triphosphate production [165] suggests that SCP-2 either: (i) binds PI and PIP to present these molecules as substrates for kinase enzymes to produce more phosphorylated PI in the lipid rafts/caveolae (Fig. 2B), or (ii) binds PI, PIP, and PIP2 to present these bound substrates for phospholipase C in the lipid rafts/caveolae (Fig. 2C). Finally, since phosphoinositides regulate vesicle budding at the golgi (see above), by binding PI, PIP, and PIP2 the SCP-2 may potentially be involved in regulating vesicular trafficking of signaling lipid substrates as well as in molecular transfer of these lipids (Fig. 1).
Early studies with soluble fractions from liver homogenates suggested that SCP-2 enhances molecular transfer of sphingomyelin (Fig. 1, 2A), neutral (globoside) glycosphingolipids (Fig. 1), and acidic (ganglioside) glycosphingolipids (Fig. 1) in vitro [184,185]. However, these supernatants contain not only SCP-2 but also several other lipid transfer proteins (PITP, L-FABP, etc) [186]. PITP also binds/transfers sphingomyelin (rev. in [169,170]. Likewise, it is not known if SCP-2/sphingomyelin complexes are better substrates for sphingomyelinase to release ceramide in lipid rafts/caveolae (Fig. 2C). Thus, it remains to be shown if highly purified recombinant SCP-2 enhances sphingolipid transfer and/or alters sphingomyelinase activity (see below).
SCP-2 not only enhances the cellular uptake of cholesterol [12,19], but also increases cholesterol transport/trafficking to ER for esterification in vitro [1,187] and in intact cells [16]. Cholesterol delivery to ER regulates release of sterol regulatory element binding protein (SREBP), which in turn trafficks to the nucleus and regulates transcription of the gene encoding HMG-CoA reductase—the rate limiting enzyme in cholesterol synthesis [188–193]. The finding that SCP-2 overexpression in hepatoma cells enhances uptake of exogenous cholesterol hydroperoxide and elicits hypersensitivity of mitochondria to its peroxidative damage, suggests that SCP-2 binds and targets this cholesterol oxidation product to mitochondria [194]. These data, plus the fact that many oxidized forms of cholesterol are potent inhibitors of cholesterol synthesis [195], suggests that SCP-2 may also regulate intracellular cholesterol synthesis by transporting these potent regulatory lipids to sites of inhibition.
SCP-2 expression regulates the distribution of signaling lipids to caveolae/lipid rafts
Significant amounts of SCP-2 are associated with lipid rafts/caveolae in plasma membranes. Cell fractionation of fibroblasts overexpressing SCP-2 and western blotting detected SCP-2 in purified plasma membranes, especially in caveolae/lipid rafts as compared to the non-raft fraction [51]. Dual immunolabeling immunofluorescence microscopy showed SCP-2 at the plasma membrane was significantly colocalized (37–72%) with caveolin-1 in multiple cell types [165]. Fluorescence resonance energy transfer (FRET) demonstrated that at the plasma membrane SCP-2, but not PITP, was located in close proximity (48±4 Å) to caveolin-1 [51]. Similar findings were confirmed by double immunogold labeling and electron microscopy [51]. Interaction of SCP-2 with caveolin-1 was specific since FRET showed PITP further away from SCP-2, not close to caveolin-1 and not coimmunoprecipitated with caveolin-1 [51]. Yeast two hybrid analysis showed SCP-2 directly interacting with caveolin-1 in cell culture. Thus, data has now shown for the first time that SCP-2 (but not PITP) selectively interacts with caveolin-1, both within cytoplasm and at the plasma membrane [51]. SCP-2 overexpression in transfected fibroblasts significantly altered intracellular distribution of signaling lipids to plasma membrane lipid rafts/caveolae (Table 6). These data are consistent with SCP-2 mediating trafficking of bound lipids between lipid rafts/caveolae and intracellular sites.
Table 6.
Effect of SCP-2 expression on distribution of signaling lipids to caveolae/lipid rafts.
| Cell type | SCP-2 Level | Lipid Class | Response | References |
|---|---|---|---|---|
| Fibroblasts | Overexpression | Fatty acids | Decrease | Current data |
| Hepatocyte | Ablation | Fatty acids | No effect | [114] |
| Fibroblasts | Overexpression | PI | Decrease | [61,149] |
| Hepatocyte | Ablation | PI | Increase | [114] |
| Fibroblasts | Overexpression | Total sphingolipid | Increase | [149] |
| Fibroblasts | Overexpression | Sphingomyelin | Decrease | [61,149] |
| Hepatocyte | Ablation | Sphingomyelin | Small Increase | [114] |
| Fibroblasts | Overexpression | Ceramide | Increase | [149] |
| Fibroblasts | Overexpression | Glucosylceramide | Increase | [149] |
| Fibroblasts | Overexpression | Galactosylceramide | No change | [149] |
| Fibroblasts | Overexpression | Lactosylceramide | No change | [149] |
| Fibroblasts | Overexpression | Globoside | Increase | [149] |
| Fibroblasts | Overexpression | Ganglioside GM1 | Decrease | [61] |
| Fibroblasts | Overexpression | Total gangliosides | No effect | [149] |
| Hepatocyte | Ablation | Ganglioside GM1 | Small Decrease | [114] |
| Fibroblasts | Overexpression | Cholesterol | No change | [61,149] |
| Hepatocyte | Ablation | Cholesterol | No change | [114] |
Lipid raft/caveolae content of unesterified fatty acids is cell-type and SCP-2 expression dependent (Table 6). In cultured primary hepatocytes from SCP-2/SCP-x gene-ablated mice (complete loss of SCP-2) there was no effect on fatty acid content of lipid rafts [114]. This observation may be due hepatocytes having a high content of plasma membrane lipid raft proteins (FAT/CD36, FATP) that bind/transport fatty acids into the hepatocyte. In contrast, increased level of SCP-2 as in transfected L-cell fibroblasts resulted in a 2.9-fold decreased fatty acid level in lipid rafts/caveolae (data not shown). The latter finding may be due to fibroblast plasma membrane lipid rafts/caveolae expressing lower levels of fatty acid binding/transport proteins. Once fatty acids are taken up into cells, SCP-2 binds and stimulates their intracellular diffusion/transport (rev. in [177,182] and enhances their transacylation to complex lipids (phosphatidic acid, cholesteryl ester) (rev. in [1].
SCP-2 altered intracellular distribution of phosphatidylinositol to plasma membranes and lipid rafts/caveolae therein (Table 6). SCP-2 overexpression in cultured fibroblasts significantly redistributed phosphatidylinositol away from intracellular sites (ER, mitochondria, lysosomes) to plasma membranes such that plasma membrane mass of PI was increased by 32% [165]. However, this increase was not reflected in the fibroblast plasma membrane lipid rafts/caveolae [61]. This observation may be explained in part by the fact that SCP-2 overexpression also enhanced insulin-stimulated inositol triphosphate production—an activity localized in lipid rafts/caveolae [165]. Conversely, ablation of the SCP-2/SCP-x gene completely abolished the synthesis of SCP-2 and increased the content of PI by 2.5-fold in lipid rafts isolated from plasma membranes of cultured primary hepatocytes [114]. These data suggest that SCP-2 enhances distribution of PI to the plasma membrane and lipid rafts/caveolae therein, while concomitantly enhancing PI utilization by enzymes involved in PI signaling in lipid rafts/caveolae. An analogous function has been proposed for another PI transfer protein, PITP (rev. in [170,172].
SCP-2 overexpression in cultured L-cells further enriched the content several sphingolipid classes in lipid rafts/caveolae (Table 6). Neutral sphingolipids such as monohexosides and globosides were increased 2- and 2.4-fold, respectively (Table 6) [61]. Although SCP-2 overexpression decreased GM1 content, the mass of total gangliosides increased slightly [61]. SCP-2 overexpression decreased the lipid raft/caveolae content of sphingomyelin by nearly 50% [61,149] but increased ceramide content nearly 2-fold [149]. The reduced sphingomyelin concomitant with increased ceramide in lipid rafts/caveolae of SCP-2 overexpressing cells may reflect increased signaling activity, i.e. SCP-2/sphingomyelin facilitates sphingomyelinase to release ceramide (Fig. 2C). Since ceramide is both an intermediate in the synthesis of multiple sphingolipids and the product of sphingomyelinase-mediated sphingomyelin hydrolysis, SCP-2 may influence production/utilization of this sphingolipid intermediate during signaling reactions in lipid rafts/caveolae. Thus the reduced sphingomyelin in lipid rafts/caveolae of SCP-2 overexpressing cells may arise from: (i) SCP-2 stimulating sphingomyelin hydrolysis to ceramide, (ii) SCP-2 facilitating removal of ceramide from lipid rafts/caveolae to intracellular sites, or (iii) SCP-2 stimulating utilization of ceramide by enzymes that produce more complex sphingolipids. Consistent with the latter interpretation, SCP-2 overexpression significantly increased the cell content of neutral glycosphingolipids (1.7-fold higher lactosylceramide and neutral glycosyl ceramide) and acidic total gangliosides [196]. This effect was specific since the levels of one of the gangliosides (GM1) as well as glucosylceramide were decreased [196]. These findings suggest that SCP-2 may enhance the utilization of sphingomyelin for signaling in lipid rafts, analogous to SCP-2-enhanced insulin stimulation of inositol triphosphate production from phosphorylated PI [165], a process which depletes lipid rafts/caveolae of PI [61].
Since SCP-2 transfers both cholesterol and bulk phospholipids between intracellular sites and plasma membrane lipid rafts/caveolae, it may change the absolute level of these lipids at the plasma membrane. However, SCP-2 affects these lipids similarly in plasma membrane lipid rafts/caveolae such that neither SCP-2 overexpression in cultured fibroblasts nor SCP-2/SCP-x gene ablation in cultured primary hepatocytes significantly altered the cholesterol/phospholipid ratio (Table 2) [61,114].
Studies with SCP-2 overexpression fibroblasts and SCP-2/SCP-x gene ablated mice
SCP-2 plays a role in modulating cellular content, distribution, and metabolism of several classes of signaling lipids including cholesterol, fatty acids, phosphatidylinositol, and sphingolipids: (i) SCP-2 regulated cholesterol uptake, intracellular trafficking and esterification in SCP-2 overexpressing cultured cells [1,12,13,16], transgenic mice overexpressing SCP-2 [14], and rats treated with SCP-2 antisense RNA [23]; (ii) SCP-2 enhanced the uptake and metabolism of fatty acids, especially branched-chain fatty acids in cultured cells [38] and primary hepatocytes from SCP-2/SCP-x gene ablated mice [41]; (iii) SCP-2 overexpression significantly altered the distribution and content of phosphatidylinositol and sphingolipids in lipid rafts/caveolae as well as altered the structure and cholesterol dynamics in lipid rafts/caveolae [61,118,140,147,149]; (iv) SCP-2 may have important roles in providing SCP-2-bound signaling lipids as substrates to enzymes that metabolize these lipids during signaling reactions in lipid rafts/caveolae. Precedent for the latter possibilities is based in part on the fact that SCP-2-bound fatty acyl CoA is a better substrate than free fatty acyl CoA for transacylase reactions [1,16,17,197]. Furthermore, SCP-2 overexpression in cultured fibroblasts stimulated insulin-mediated inositol-triphosphate production [165]. Thus, cultured cell data as well as studies with SCP-2 overexpression and siRNA SCP-2 knockdown in animals suggest that SCP-2 may have important roles in intracellular trafficking of signaling lipids, remodeling lipid raft/caveolae lipid composition, and providing SCP-2-bound signaling lipids as substrates to enzymes that metabolize these lipids during signaling reactions in lipid rafts/caveolae. Nevertheless, the full physiological significance of SCP-2 in intracellular signaling is yet to be resolved in vivo. Part of the challenge is that gene-targeted mice may not necessarily exhibit a strong phenotype under control-chow and otherwise unchallenged conditions. For example, despite the apparent promiscuity as a lipid binding and regulatory protein, SCP-2/SCP-x gene ablated mice generated by disruption of Exon 14 have yielded phenotypic changes that are mostly related to defects in branched chain lipid (i.e. fatty acid, cholesterol) metabolism [20,43]. Unfortunately, there have been no studies to date examining any of the known signaling pathways in these mice. Moreover, it must be noted that in this specific SCP-2/SCP-x ablation of Exon 14 concomitantly resulted in 5–6-fold upregulation of another promiscuous lipid binding protein (L-FABP) which also binds many signaling lipids (rev. in [177,178,183,198]. Whether additional proteins that bind/transfer signaling molecules (e.g. phosphatidylinositol binding protein, glycolipid transfer protein) or other component proteins of the signaling cascades were also upregulated to compensate for loss of SCP-2 in these mice is not known. In contrast, another study generated SCP-2/SCP-x gene ablated mice through targeted disruption of exon 16 [114]. The resultant knockout mice exhibited very little change in L-FABP expression [114]. Furthermore, cultured primary hepatocytes from these SCP-2/SCP-x gene ablated mice (i.e. exon 16 knockout) confirmed that ablating SCP-2 regulates lipid raft properties as evidenced by: (i) Increased proportion of lipid rafts 1.5-fold; (ii) Increased raft total phospholipid; (iii) Decreased fluorescence polarization indicating a decreased liquid disordered phase and a lesser transbilayer fluidity gradient in hepatocyte lipid rafts. Thus, lipid rafts present in primary cultured mouse hepatocyte plasma membranes displayed unique protein, lipid, signaling lipid, structure, and biophysical properties which were differentially regulated by SCP-2 expression. The availability of mouse models overexpressing SCP-2 [23,199], ablated in SCP-x [47], or ablated in SCP-2/SCP-x [20,114] now provides the means to examine these possibilities in more physiologically relevant systems. However, dietary or other stress (age, exercise, etc) challenge may have to be performed to overcome compensatory upregulation of other proteins that may prevent observation of specific signaling phenotypes under control conditions.
SUMMARY AND CONCLUSIONS
Although SCP-2 was established as a protein that transfers cholesterol and phospholipids decades ago, recent findings with lipid rafts/caveolae and SCP-2 suggest that they may provide a conceptual link to metabolic processes that might be regulated through the respective signaling pathways. The molecular basis for this possibility resides in three major factors:
First, SCP-2’s lipid binding pocket exhibits high affinity for a large variety of lipid molecules that are directly or indirectly involved in signaling: (i) fatty acids which are precursors of prostaglandins, leukotrienes, and thromboxanes (i.e. arachidonic, docosahexaenoic, and eicosapentaenoic acids), (ii) fatty acids which when transported into nuclei regulate transcriptional activity of receptors (e.g. peroxisome proliferator activated receptors); (iii) fatty acyl CoAs which regulate vesicular budding from the Golgi, (iv) fatty acyl CoAs when transported into nuclei regulate transcriptional activity of receptors (e.g. peroxisome proliferator activated receptors, hepatocyte nuclear factor 4α); (v) phosphoinositides (PI, PIP, PIP2) which regulate vesicular budding or as a result of phospholipase C action yield second messengers (IP, IP2, IP3); (vi) sphingolipids (sphingomyelin, ceramides, globosides, gangliosides) which are involved in multiple signaling pathways, and (vii) cholesterol which regulates production of SREBP at the endoplasmic reticulum and in turn trafficks to the nuclei to regulate transcription.
Second, cholesterol-rich lipid rafts/caveolae exist in part to compartmentalize signaling lipids with select proteins that use them as substrates. Although early studies suggested that lipid rafts/caveolae isolated with detergents or high pH carbonate buffers are highly enriched in proteins and lipid involved in signaling, increasing concern that these fractionation methods may induce artifacts has introduced uncertainty. This issue was resolved by development of new methods that do not use detergents or high pH carbonate buffers to isolate lipid rafts/caveolae. Biochemical analysis of the proteins and lipids isolated by the latter methods confirmed that many of the basic conclusions obtained by the earlier detergent and high pH carbonate buffer based methods were often qualitatively, if not quantitatively, correct. Furthermore, newly developed fluorescence imaging technologies for the first time provided real-time data indicating that plasma membranes of living cells contain sterol-rich and –poor domains.
Third, SCP-2 provides a conceptual link between physico-chemical data and metabolic processes that might be regulated through multiple signaling pathways. SCP-2 may exhibit new roles in lipid signaling mediated through plasma membrane lipid rafts/caveolae by mechanisms that involve molecular transfer of signaling lipids (Fig. 1, 2A), vesicular transfer of signaling lipids (Fig. 1, Vesicle and Caveolar Vesicle), presentation of lipid substrates to enzymes that produce more complex signaling molecules/substrates (Fig. 2B), and presentation of signaling lipid substrates to enzymes for hydrolysis and release of product signaling molecules (Fig. 2C).
In summary, despite the fact that SCP-2 was known for nearly thirty years for its ability to bind and transfer cholesterol and phospholipids, SCP-2’s apparent promiscuity in binding other classes of lipids (especially lipids involved in intracellular signaling) suggest more complex physiological function(s). The challenge of future studies will be to begin to resolve the physiological function of SCP-2 in mediating these lipid signaling pathways. Because of the likelihood of compensatory upregulation of other proteins (e.g. PITP, etc) with overlapping functions in the signaling cascades, studies with SCP-2/SCP-x gene-targeted mice will require additional challenges (nutritional, endocrine, exercise, etc. stress). To further elucidate mechanistic details, it is highly likely that these findings will need to be supplemented/complemented by studies with cultured cells and in vitro systems.
Acknowledgments
This work was supported in part by the USPHS, NIH grants GM31651 and DK41402 (FS and ABK) and GM63236 (JMB).
Abbreviations
- SCP-2
sterol carrier protein-2
- PITP
phosphatidylinositol transfer protein
- L-FABP
liver fatty acid binding protein
- HDL
high density lipoprotein
- SRB1
scavenger receptor B1
- P-gp
P- glycoprotein
- ABCA1
ABCA1 transporter
- conA
concanavalin A
- SL
Sphingolipid
- SM
sphingomyelin
- Chol
cholesterol (C)
- Cer
ceramide
- Glo
globosides
- GlcCer
glucosylceramide
- Gal
galactose
- GalCer
galactosylceramide
- CGT
ceramide glucosyltransferase
- LacCer
lactosylceramide
- GSL
glycosphingolipid
- LBPA
lysobisphosphatidic acid
- NANA
N- acetylneuraminic acid
- LCFA
long chain fatty acid
- LCFA-CoA
long chain fatty acyl CoA
- OlLyso
oleoyl lysophosphatidic acid
- PI
phosphatidylinositol
- PIP
phosphatidylinositolphosphate
- PIP2
phosphatidylinositolbisphosphate
- MHC
monohexosylceramide
- DHC
dihexosylceramide
- G
globoside
- GM1
ganglioside M1
- PC
phosphatidylcholine
- PS
phosphatidylserine
- PE
phosphatidylethanolamine
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Moncecchi DM, Nemecz G, Schroeder F, Scallen TJ. The participation of sterol carrier protein-2 (SCP-2) in cholesterol metabolism. In: Patterson GW, Nes WD, editors. Physiology and Biochemistry of Sterols. Champaign, IL: 1991. pp. 1–27. [Google Scholar]
- 2.Ericsson J, Appelkvist EL, Runquist M, Dallner G. Biosynthesis of dolichol and cholesterol in rat liver peroxisomes. . Biochimie. 1993;75:167–173. doi: 10.1016/0300-9084(93)90074-3. [Review] [DOI] [PubMed] [Google Scholar]
- 3.Stocker A, Tomizaki T, Schulze-Briese C, Baumann U. Crystal structure of the human supernatant protein factor. Structure. 2002;10:1533–1540. doi: 10.1016/s0969-2126(02)00884-5. [DOI] [PubMed] [Google Scholar]
- 4.Stocker A, Baumann U. Supernatant protein factor in complex with RRR-alpha-tocopherylquinone: a link between oxidized vitamin E and cholesterol biosynthesis. J Mol Biol. 2003;332:759–765. doi: 10.1016/s0022-2836(03)00924-0. [DOI] [PubMed] [Google Scholar]
- 5.Chanderbhan R, Noland BJ, Scallen TJ, Vahouny GV. Sterol carrier protein2. Delivery of cholesterol from adrenal lipid droplets to mitochondria for pregnenolone synthesis. J Biol Chem. 1982;257:8928–8934. [PubMed] [Google Scholar]
- 6.Helmkamp GM. Phospholipid transfer proteins: mechanism of action. J Bioenergetic Biomembranes. 1986;18:71–91. doi: 10.1007/BF00743477. [Review] [97 refs] [DOI] [PubMed] [Google Scholar]
- 7.Vahouny GV, Chanderbhan R, Kharoubi A, Noland BJ, Pastuszyn A, Scallen TJ. Sterol carrier and lipid transfer proteins. Adv Lipid Res. 1987;22:83–113. doi: 10.1016/b978-0-12-024922-0.50007-2. [DOI] [PubMed] [Google Scholar]
- 8.Wirtz KW. Phospholipid transfer proteins. Annu Rev Biochem J. 1991;60:73–99. doi: 10.1146/annurev.bi.60.070191.000445. [Review] [DOI] [PubMed] [Google Scholar]
- 9.Ossendorp BC, Snoek GT, Wirtz KWA. Intracellular Phospholipid Transfer Proteins, Current Topics in Membranes. Vol. 40. New York: 1994. pp. 217–259. [Google Scholar]
- 10.Wirtz KWA. Phospholipid transfer proteins revisited. Biochem J. 1997;324:353–360. doi: 10.1042/bj3240353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bloj B, Zilversmit DB. Rat liver proteins capable of transferring phosphatidylethanolamine. J Biol Chem. 1977;252:1613–1619. [PubMed] [Google Scholar]
- 12.Moncecchi DM, Murphy EJ, Prows DR, Schroeder F. Sterol carrier protein-2 expression in mouse L-cell fibroblasts alters cholesterol uptake. Biochim Biophys Acta. 1996;1302:110–116. doi: 10.1016/0005-2760(96)00044-6. [DOI] [PubMed] [Google Scholar]
- 13.Baum CL, Reschly EJ, Gayen AK, Groh ME, Schadick K. Sterol carrier protein-2 overexpression enhances sterol cycling and inhibits cholesterol ester synthesis and high density lipoprotein cholesterol secretion. J Biol Chem. 1997;272:6490–6498. doi: 10.1074/jbc.272.10.6490. [DOI] [PubMed] [Google Scholar]
- 14.Puglielli L, Rigotti A, Greco AV, Santos MJ, Nervi F. Sterol carrier protein-2 is involved in cholesterol transfer from the endoplasmic reticulum to the plasma membrane in human fibroblasts. J Biol Chem. 1995;270:18723–18726. doi: 10.1074/jbc.270.32.18723. [DOI] [PubMed] [Google Scholar]
- 15.Schroeder F, Frolov A, Schoer J, Gallegos A, Atshaves BP, Stolowich NJ, Scott AI, Kier AB. Intracellular sterol binding proteins, cholesterol transport and membrane domains. In: Chang TY, Freeman DA, editors. Intracellular Cholesterol Trafficking. Boston: 1998. pp. 213–234. [Google Scholar]
- 16.Murphy EJ, Schroeder F. Sterol carrier protein-2 mediated cholesterol esterification in transfected L-cell fibroblasts. Biochim Biophys Acta. 1997;1345:283–292. doi: 10.1016/s0005-2760(97)00003-9. [DOI] [PubMed] [Google Scholar]
- 17.Starodub O, Jolly CA, Atshaves BP, Roths JB, Murphy EJ, Kier AB, Schroeder F. Sterol carrier protein-2 immunolocalization in endoplasmic reticulum and stimulation of phospholipid formation. Am J Physiol. 2000;279:C1259–C1269. doi: 10.1152/ajpcell.2000.279.4.C1259. [DOI] [PubMed] [Google Scholar]
- 18.Murphy EJ, Stiles T, Schroeder F. Sterol carrier protein-2 expression alters phospholipid content and fatty acid composition in L-cell fibroblasts. J Lipid Res. 2000;41:788–796. [PubMed] [Google Scholar]
- 19.Atshaves BP, Starodub O, McIntosh AL, Roths JB, Kier AB, Schroeder F. Sterol carrier protein-2 alters HDL-mediated cholesterol efflux. J Biol Chem. 2000;275:36852–36861. doi: 10.1074/jbc.M003434200. [DOI] [PubMed] [Google Scholar]
- 20.Seedorf U, Raabe M, Ellinghaus P, Kannenberg F, Fobker M, Engel T, Denis S, Wouters F, Wirtz KWA, Wanders RJA, Maeda N, Assmann G. Defective peroxisomal catabolism of branched fatty acyl coenzyme A in mice lacking the sterol carrier protein-2/sterol carrier protein-x gene function. Genes and Development. 1998;12:1189–1201. doi: 10.1101/gad.12.8.1189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Fuchs M, Lammert F, Wang DQH, Paigen B, Carey MC, Cohen DE. Sterol carrier protein-2 participates in hypersecretion of biliary cholesterol during cholesterol gallstone formation in genetically gallstone susceptible mice. Biochem J. 1998;336:33–37. doi: 10.1042/bj3360033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Fuchs M, Lammert F, Wang DQH, Paigen B, Carey MC, Cohen DE. Lith genes induce overexpression of sterol carrier proteing 2 during cholesterol gallstone formation. FASEB J. 1997;11:A1060. [Google Scholar]
- 23.Puglielli L, Rigotti A, Amigo L, Nunez L, Greco AV, Santos MJ, Nervi F. Modulation on intrahepatic cholesterol trafficking: Evidence by in vivo antisense treatment for the involvement of sterol carrier protein-2 in newly synthesized cholesterol transfer into bile. Biochem J. 1996;317:681–687. doi: 10.1042/bj3170681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Seedorf U, Brysch P, Engel T, Schrage K, Assmann G. Sterol carrier protein X is peroxisomal 3-oxoacyl coenzyme A thiolase with intrinsic sterol carrier and lipid transfer activity. J Biol Chem. 1994;269:21277–21283. [PubMed] [Google Scholar]
- 25.Antonenkov VD, Van Veldhoven PP, Waelkens E, Mannaerts GP. Substrate specificities of 3-oxoacyl-CoA thiolase A and sterol carrier protein 2/3-oxoacyl-CoA thiolase purified from normal rat liver peroxisomes. J Biol Chem. 1997;272:26023–26031. doi: 10.1074/jbc.272.41.26023. [DOI] [PubMed] [Google Scholar]
- 26.Antonenkov VD, Van Veldhoven PP, Mannaerts GP. Isolation and subunit composition of native sterol carrier protein-2/3-oxoacyl-coenzyme A thiolase from normal rat liver peroxisomes. Protein Exp Purif. 2000;18:249–256. doi: 10.1006/prep.2000.1192. [DOI] [PubMed] [Google Scholar]
- 27.Kannenberg F, Ellinghaus P, Assmann G, Seedorf U. Aberrant oxidation of the cholesterol side chain in bile acid synthesis of sterol carrier protein-2/sterol carrier protein-x knockout mice. J Biol Chem. 1999;274:35455–35460. doi: 10.1074/jbc.274.50.35455. [DOI] [PubMed] [Google Scholar]
- 28.Muench C, Hafer A, Katzberg N, Scheibner J, Stange EF, Seedorf U, Fuchs M. Relevance of the sterol carrier protein-2 gene for bile acid synthesis and gallstone formation in genetically susceptible mice. Gastroenterology. 2000;118(4 Part 1 Supplement 2):1167. Ref Type: Abstract. [Google Scholar]
- 29.Fuchs M, Hafer A, Muench C, Kannenberg F, Teichmann S, Scheibner J, Stange EF, Seedorf U. Disruption of the sterol carrier protein 2 gene in mice impairs biliary lipid and hepatic cholesterol metabolism. J Biol Chem. 2001;276:48058–48065. doi: 10.1074/jbc.M106732200. [DOI] [PubMed] [Google Scholar]
- 30.Seedorf U, Ellinghaus P, Nofer JR. Sterol carrier protein-2. Biochim Biophys Acta. 2000;1486:45–54. doi: 10.1016/s1388-1981(00)00047-0. [DOI] [PubMed] [Google Scholar]
- 31.Gallegos AM, Atshaves BP, Storey SM, Starodub O, Petrescu AD, Huang H, McIntosh A, Martin G, Chao H, Kier AB, Schroeder F. Gene structure, intracellular localization, and functional roles of sterol carrier protein-2. Prog Lipid Res. 2001;40:498–563. doi: 10.1016/s0163-7827(01)00015-7. [DOI] [PubMed] [Google Scholar]
- 32.Gallegos AM, Schoer J, Starodub O, Kier AB, Billheimer JT, Schroeder F. A potential role for sterol carrier protein-2 in cholesterol transfer to mitochondria. Chem Phys Lipids. 2000;105:9–29. doi: 10.1016/s0009-3084(99)00128-0. [DOI] [PubMed] [Google Scholar]
- 33.Frolov AA, Woodford JK, Murphy EJ, Billheimer JT, Schroeder F. Fibroblast membrane sterol kinetic domains: modulation by sterol carrier protein 2 and liver fatty acid binding protein. J Lipid Res. 1996;37:1862–1874. [PubMed] [Google Scholar]
- 34.Frolov A, Woodford JK, Murphy EJ, Billheimer JT, Schroeder F. Spontaneous and protein-mediated sterol transfer between intracellular membranes. J Biol Chem. 1996;271:16075–16083. doi: 10.1074/jbc.271.27.16075. [DOI] [PubMed] [Google Scholar]
- 35.Petrescu AD, Gallegos AM, Okamura Y, Strauss IJF, Schroeder F. Steroidogenic acute regulatory protein binds cholesterol and modulaes mitochondrial membrane sterol domain dynamics. J Biol Chem. 2001;276:36970–36982. doi: 10.1074/jbc.M101939200. [DOI] [PubMed] [Google Scholar]
- 36.Stolowich NJ, Petrescu AD, Huang H, Martin G, Scott AI, Schroeder F. Sterol carrier protein-2: structure reveals function. Cell Mol Life Sci. 2002;59:193–212. doi: 10.1007/s00018-002-8416-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Atshaves BP, Storey SM, Petrescu AD, Greenberg CC, Lyuksyutova OI, Smith R, Schroeder F. Expression of fatty acid binding proteins inhibits lipid accumulation and alters toxicity in L-cell fibroblasts. Am J Physiol. 2002;283:C688–C703. doi: 10.1152/ajpcell.00586.2001. [DOI] [PubMed] [Google Scholar]
- 38.Atshaves BP, Storey SM, Schroeder F. Sterol carrier protein-2/sterol carrier protein-x expression differentially alters fatty acid metabolism in L-cell fibroblasts. J Lipid Res. 2003;44:1751–1762. doi: 10.1194/jlr.M300141-JLR200. [DOI] [PubMed] [Google Scholar]
- 39.Atshaves BP, Storey S, Huang H, Schroeder F. Liver fatty acid binding protein expression enhances branched-chain fatty acid metabolism. Mol Cell Biochem. 2004;259:115–129. doi: 10.1023/b:mcbi.0000021357.97765.f2. [DOI] [PubMed] [Google Scholar]
- 40.Atshaves BP, Payne HR, McIntosh AL, Tichy SE, Russell D, Kier AB, Schroeder F. Sexually dimorphic metabolism of branched chain lipids in C57BL/6J mice. J Lipid Res. 2004;45:812–830. doi: 10.1194/jlr.M300408-JLR200. [DOI] [PubMed] [Google Scholar]
- 41.Atshaves BP, McIntosh AL, Lyuksyutova OI, Zipfel WR, Webb WW, Schroeder F. Liver fatty acid binding protein gene ablation inhibits branched-chain fatty acid metabolism in cultured primary hepatocytes. J Biol Chem. 2004;279:30954–30965. doi: 10.1074/jbc.M313571200. [DOI] [PubMed] [Google Scholar]
- 42.Atshaves BP, McIntosh AL, Payne HR, Mackie J, Kier AB, Schroeder F. Effect of branched-chain fatty acid on lipid dynamics in mice lacking liver fatty acid binding protein gene. Am J Physiol. 2005;288:C543–C558. doi: 10.1152/ajpcell.00359.2004. [DOI] [PubMed] [Google Scholar]
- 43.Monnig G, Wiekowski J, Kirchhof P, Stypmann J, Plenz G, Fabritz L, Bruns HJ, Eckardt L, Assmann G, Haverkamp W, Breithardt G, Seedorf U. Phytanic acid accumulation is associated with conduction delay and sudden cardiac death in sterol carrier protein-2/sterol carrier protein-x deficient mice. J Cardiovasc Electrophysiol. 2004;15:1310–1316. doi: 10.1046/j.1540-8167.2004.03679.x. [DOI] [PubMed] [Google Scholar]
- 44.Idel S, Ellinghaus P, Wolfrum C, Nofer JR, Gloerich J, Assmann G, Spener F, Seedorf U. Branched chain fatty acids induce nitric oxide-dependent apoptosis in vascular smooth muscle cells. J Biol Chem. 2002;277:49319–49325. doi: 10.1074/jbc.M204639200. [DOI] [PubMed] [Google Scholar]
- 45.Wanders RJ, Denis S, van Berkel E, Wouters F, Wirtz KWA, Seedorf U. Identification of the newly discovered 58 kDa peroxisomal thiolase SCP-x as the main thiolase involved in both pristanic acid and trihydroyxycholestanoic acid oxidation: Implications for peroxisomal beta-oxidation disorders. J Inher Metab Dis. 1998;21:302–305. doi: 10.1023/a:1005349028853. [DOI] [PubMed] [Google Scholar]
- 46.Ferdinandusse S, Kostopoulos P, Denis S, Rusch R, Overmars H, Dillman U, Reith W, Haas D, Wanders RJA, Duran M, Marziniak M. Mutations in the gene encoding peroxisomal sterol carrier protein-x (SCP-x) cause leukencephalopathy with dystonia and motor neuropathy. Am J Hum Genet. 2006;78:1046–1052. doi: 10.1086/503921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Atshaves BP, McIntosh AL, Landrock D, Payne HR, Mackie J, Maeda N, Ball JM, Kier AB, Schroeder F. Effect of SCP-x gene ablation on branched-chain fatty acid metabolism. Am J Physiol. 2007;292:939–951. doi: 10.1152/ajpgi.00308.2006. [DOI] [PubMed] [Google Scholar]
- 48.Schroeder F, Frolov A, Starodub O, Russell W, Atshaves BP, Petrescu AD, Huang H, Gallegos A, McIntosh A, Tahotna D, Russell D, Billheimer JT, Baum CL, Kier AB. Pro-sterol carrier protein-2: role of the N-terminal presequence in structure, function, and peroxisomal targeting. J Biol Chem. 2000;275:25547–25555. doi: 10.1074/jbc.M000431200. [DOI] [PubMed] [Google Scholar]
- 49.Huang H, Ball JA, Billheimer JT, Schroeder F. The sterol carrier protein-2 amino terminus: a membrane interaction domain. Biochemistry. 1999;38:13231–13243. doi: 10.1021/bi990870x. [DOI] [PubMed] [Google Scholar]
- 50.Huang H, Ball JA, Billheimer JT, Schroeder F. Interaction of the N-terminus of sterol carrier protein-2 with membranes: role of membrane curvature. Biochem J. 1999;344:593–603. [PMC free article] [PubMed] [Google Scholar]
- 51.Zhou M, Parr RD, Petrescu AD, Payne HR, Atshaves BP, Kier AB, Ball JA, Schroeder F. Sterol carrier protein-2 directly interacts with caveolin-1 in vitro and in vivo. Biochem. 2004;43:7288–7306. doi: 10.1021/bi035914n. [DOI] [PubMed] [Google Scholar]
- 52.Parr RD, Martin GG, Schroeder ME, Mir KD, Kier AB, Schroeder F, Ball JM. A new N-terminal recognition domain in caveolin-1 interacts with sterol carrier protein-2 (SCP-2) Biochemistry. 2007 doi: 10.1021/bi7002636. submitted. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Schroeder F, Atshaves BP, Gallegos AM, McIntosh AL, Liu JC, Kier AB, Huang H, Ball JM. Lipid rafts and caveolae organization. In: Frank PG, Lisanti MP, editors. Advances in Molecular and Cell Biology. Vol. 36. Advances in Molecular and Cell Biology; Amsterdam: 2005. pp. 3–36. [Google Scholar]
- 54.Pastuszyn A, Noland BJ, Bazan F, Fletterick RJ, Scallen TJ. Primary sequence and structural analysis of sterol protein-2 from rat liver: homology with immunoglobins. J Biol Chem. 1987;262:13219–13227. [PubMed] [Google Scholar]
- 55.Moncecchi DM, Pastuszyn A, Scallen TJ. cDNA sequence and bacterial expression of mouse liver sterol carrier protein-2. J Biol Chem. 1991;266:9885–9892. [PubMed] [Google Scholar]
- 56.Mendis-Handagama SM, Watkins PA, Gelber SJ, Scallen TJ. Leydig cell peroxisomes and sterol carrier protein-2 in luteinizing hormone-deprived rats. Endocrinology. 1992;131:2839–2845. doi: 10.1210/endo.131.6.1446622. [DOI] [PubMed] [Google Scholar]
- 57.Keller GA, Scallen TJ, Clarke D, Maher PA, Krisans SK, Singer SJ. Subcellular localization of sterol carrier protein-2 in rat hepatocytes: its primary localization to peroxisomes. J Cell Biol. 1989;108:1353–1361. doi: 10.1083/jcb.108.4.1353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Mendis-Handagama SM, Aten RF, Watkins PA, Scallen TJ, Berhman HR. Peroxisomes and sterol carrier protein-2 in luteal cell steroidogenesis: a possible role in cholesterol transport from lipid droplets to mitochondria. Tissue Cell. 1995;27:483–490. doi: 10.1016/s0040-8166(05)80056-4. [DOI] [PubMed] [Google Scholar]
- 59.Atshaves BP, Petrescu A, Starodub O, Roths J, Kier AB, Schroeder F. Expression and Intracellular Processing of the 58 kDa Sterol Carrier Protein 2/3-Oxoacyl-CoA Thiolase in Transfected Mouse L-cell Fibroblasts. J Lipid Res. 1999;40:610–622. [PubMed] [Google Scholar]
- 60.Frolov A, Miller K, Billheimer JT, Cho TC, Schroeder F. Lipid specificity and location of the sterol carrier protein-2 fatty acid binding site: A fluorescence displacement and energy transfer study. Lipids. 1997;32:1201–1209. doi: 10.1007/s11745-997-0154-5. [DOI] [PubMed] [Google Scholar]
- 61.Atshaves BP, Gallegos A, McIntosh AL, Kier AB, Schroeder F. Sterol carrier protein-2 selectively alters lipid composition and cholesterol dynamics of caveolae/lipid raft vs non-raft domains in L-cell fibroblast plasma membranes. Biochemistry. 2003;42:14583–14598. doi: 10.1021/bi034966+. [DOI] [PubMed] [Google Scholar]
- 62.Lassen D, Lücke C, Kromminga A, Lezius A, Spener F, Rüterjans H. Solution structure of bovine heart FABP. Mol Cell Biochem. 1993;123:15–22. doi: 10.1007/BF01076470. [DOI] [PubMed] [Google Scholar]
- 63.Hanhoff T, Wolfrum C, Ellinghaus P, Seedorf U, Spener F. Pristanic acid is activator of PPARalpha. Eur J Lipid Sci. 2001;103:75–80. [Google Scholar]
- 64.Wolfrum C, Spener F. Fatty acids as regulators in lipid metabolism. Eur J Lip Sci Technol. 2000;102:746–762. [Google Scholar]
- 65.Ellinghaus P, Wolfrum C, Assmann G, Spener F, Seedorf U. Phytanic acid activates the peroxisome proliferator-activated receptor alpha (PPARalpha) in sterol carrier protein-2-/sterol carrier protein x-deficient mice. J Biol Chem. 1999;274:2766–2772. doi: 10.1074/jbc.274.5.2766. [DOI] [PubMed] [Google Scholar]
- 66.Wolfrum C, Ellinghaus P, Fobker M, Seedorf U, Assmann G, Borchers T, Spener F. Phytanic acid is ligand and transcriptional activator of murine liver fatty acid binding protein. J Lipid Res. 1999;40:708–714. [PubMed] [Google Scholar]
- 67.Wolfrum C, Borchers T, Sacchettini JC, Spener F. Binding of fatty acids and perooxisome proliferators to orthologous fatty acid binding proteins from human, murine, and bovine liver. Biochemistry. 2000;39:1469–1474. doi: 10.1021/bi991638u. [DOI] [PubMed] [Google Scholar]
- 68.Wouters F, Bastiaens PI, Wirtz KW, Jovin TM. FRET microscopy demonstrates molecular association of non-specific lipid transfer protein (nsL-TP) with fatty acid oxidation enzymes. EMBO J. 1998;17:7179–7189. doi: 10.1093/emboj/17.24.7179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Martin GG, Atshaves BP, McIntosh AL, Mackie JT, Kier AB, Schroeder F. Liver fatty acid binding protein (L-FABP) gene ablation alters liver bile acid metabolism in male mice. Biochem J. 2005;391:549–560. doi: 10.1042/BJ20050296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Martin GG, Atshaves BP, McIntosh AL, Mackie JT, Kier AB, Schroeder F. Liver fatty acid binding protein (L-FABP) gene ablation potentiates hepatic cholesterol accumulation in cholesterol-fed female mice. Am J Physiol. 2006;290:G36–G48. doi: 10.1152/ajpgi.00510.2004. [DOI] [PubMed] [Google Scholar]
- 71.Anderson RGW. Caveolae: Where incoming and outgoing messengers meet. Proc Natl Acad Sci. 1993;90:10909–10913. doi: 10.1073/pnas.90.23.10909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Liu P, Anderson RGW. Compartmentalized production of ceramide at the cell surface. J Biol Chem. 1995;270:27179–27185. doi: 10.1074/jbc.270.45.27179. [DOI] [PubMed] [Google Scholar]
- 73.Lisanti MP, Tang Z, Scherer PE, Kubler E, Koleske AJ, Sargiacomo M. Caveolae, transmembrane signalling and cellular transformation. Mol Memb Biol. 1995;12:121–124. doi: 10.3109/09687689509038506. [DOI] [PubMed] [Google Scholar]
- 74.Stralfors P. Insulin Signaling and Caveolae. In: Lisanti MP, Frank PG, editors. Caveolae and Lipid Rafts: Roles in Signal Transduction and Human Disease. Vol. 36. Advances in Molecular and Cell Biology; San Diego: 2005. pp. 141–169. [Google Scholar]
- 75.Smart EJ, Graf GA, McNiven MA, Sessa WC, Engelman JA, Scherer PE, Okamoto T, Lisanti MP. Caveolins, Liquid-ordered domains, and signal transduction. Mol Cell Biol. 1999;19:7289–7304. doi: 10.1128/mcb.19.11.7289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Lisanti MP, Tang Z, Scherer PE, Kubler E, Koleske AJ, Sargiacomo M. Caveolae, transmembrane signalling and cellular transformation. Molecular Membrane Biology. 1995;12:121–124. doi: 10.3109/09687689509038506. [Review] [25 refs] [DOI] [PubMed] [Google Scholar]
- 77.Schlegel A, Volonte D, Engelman JA, Galbiati F, Mehta PP, Zhang XL, Scherer PE, Lisanti MP. Crowded little caves: structure and function of caveolae. Cell Signal. 1998;10:457–463. doi: 10.1016/s0898-6568(98)00007-2. [DOI] [PubMed] [Google Scholar]
- 78.Smart EJ. Caveolae and the regulation of cellular cholesterol homeostasis. In: Lisanti MP, Frank PG, editors. Advances in Molecular and Cell Biology. Vol. 36. Advances in Molecular and Cell Biology; Amsterdam: 2005. p. 35. [Google Scholar]
- 79.Everson WV, Smart EJ. Caveolae and the regulation of cellular cholesterol homeostasis. In: Lisanti MP, Frank PG, editors. Caveolae and Lipid Rafts: Roles in Signal Transduction and the Pathogenesis of Human Disease. Vol. 36. Advances in Molecular and Cell Biology; San Diego: 2005. pp. 37–55. [Google Scholar]
- 80.Jacobson K, Vaz WL. Domains in biological membranes. Comments Mol Cell Biophys. 1992;8:1–15. [Google Scholar]
- 81.Jacobson K, Dietrich C. Looking at rafts? Trends Cell Biol. 1999;9:87–91. doi: 10.1016/s0962-8924(98)01495-0. [DOI] [PubMed] [Google Scholar]
- 82.Anderson RGW, Jacobson K. A role for lipid shells in targeting proteins to caveolae, rafts, and other lipid domains. Science. 2002;296:1821–1825. doi: 10.1126/science.1068886. [DOI] [PubMed] [Google Scholar]
- 83.Dietrich C, Bagatolli LA, Volovyk ZN, Thompson NL, Levi M, Jacobson K, Gratton E. Lipid rafts reconstituted in model membranes. Biophysical Journal. 2001;80:1417–1428. doi: 10.1016/S0006-3495(01)76114-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Munro S. Lipid rafts: Elusive or Illusive? Cell. 2003;115:377–388. doi: 10.1016/s0092-8674(03)00882-1. [DOI] [PubMed] [Google Scholar]
- 85.Wustner D. Plasma membrane sterol distribution resembles the surface topolography of living cells. Mol Biol Cell. 2007;18:211–228. doi: 10.1091/mbc.E06-05-0445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Kaiser RD, London E. Location of Diphenylhexatriene (DPH) and Its Derivatives within Membranes: Comparison of Different Fluorescence Quenching Analyses of Membrane Depth. Biochemistry. 1998;37:8180–8190. doi: 10.1021/bi980064a. [DOI] [PubMed] [Google Scholar]
- 87.Brown DA, London E. Structure and function of sphingolipid- and cholesterol-rich membrane rafts. J Biol Chem. 2000;275:17221–17224. doi: 10.1074/jbc.R000005200. [DOI] [PubMed] [Google Scholar]
- 88.Brown DA, London E. Structure and origin of ordered lipid domains in biological membranes. J Membr Biol. 1998;164:103–114. doi: 10.1007/s002329900397. [DOI] [PubMed] [Google Scholar]
- 89.Schutz GJ, Kada G, Pastushenko VPh, Schindler H. Properties of lipid microdomains in a muscle cell membrane visualized by single molecule microscopy. EMBO J. 2000;19:892–901. doi: 10.1093/emboj/19.5.892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Bretscher MS, Munro S. Cholesterol and the Golgi apparatus. Science. 1993;261:1280–1281. doi: 10.1126/science.8362242. [DOI] [PubMed] [Google Scholar]
- 91.Wisher MH, Evans WH. Functional polarity of the rat hepatocyte surface membrane. Biochem J. 1975;146:375–388. doi: 10.1042/bj1460375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Sweet WD, Schroeder F. Lipid domains and enzyme activity. In: Aloia RC, Cirtain CC, Gordon LM, editors. Advances in Membrane Fluidity: Lipid Domains and the Relationship to Membrane Function. New York, NY: 1988. pp. 17–42. [Google Scholar]
- 93.Sehayek E, Wang R, Ono JG, Zinchuck VS, Duncan EM, Shefer S, Vance DE, Ananthanarayanan M, Chait BT, Breslow JL. Localization of the PE methylation pathway and SR-B1 to the canalicular membrane: evidence for apical PC biosynthesis that may promote biliary excretion of phospholipid and cholesterol. J Lipid Res. 2003;44:1605–1613. doi: 10.1194/jlr.M200488-JLR200. [DOI] [PubMed] [Google Scholar]
- 94.Peng Y, Akmentin W, Conneely MA, Lund-Katz S, Phillips MC, Williams DL. Scavenger receptor B1 (SR-B1) clustered on microvillar extensions suggests that this plasma membrane domain is a way station for cholesterol trafficking between cells and high density lipoprotein. Mol Biol Cell. 2003;15:384–396. doi: 10.1091/mbc.E03-06-0445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Reaven E, Leers-Sucheta S, Nomoto A, Azhar S. Expression of scavenger receptor class B type 1 (SR-B1) promotes microvillar channel formation and selective cholesteryl ester transport in a heterologous reconstituted system. Proc Natl Acad Sci. 2001;98:1613–1618. doi: 10.1073/pnas.98.4.1613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Ikemoto M, Arai H, Feng D, Tanaka K, Aoki J, Dohmae N, Takio K, Adachi H, Tsujimoto M, Inoue K. Identification of a PDZ-domain-containing protein that interacts with the scavenger receptor class B type 1. Proc Natl Acad Sci. 2000;97:6538–6543. doi: 10.1073/pnas.100114397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Wustner D. Improved visualization and quantitative analysis of fluorescent membrane sterol in polarized hepatic cells. J Microscopy. 2005;220:47–64. doi: 10.1111/j.1365-2818.2005.01516.x. [DOI] [PubMed] [Google Scholar]
- 98.Kavecansky J, Schroeder F, Joiner CH. Deoxygenation-induced alterations in sickle cell membrane cholesterol exchange. Am J Physiol. 1995;269:C1105–C1111. doi: 10.1152/ajpcell.1995.269.5.C1105. [DOI] [PubMed] [Google Scholar]
- 99.Brunner G, Ferber E, Resch K. Fractionation of membrane vesicles I. Differentiation. 1976;5:161–164. doi: 10.1111/j.1432-0436.1976.tb00908.x. [DOI] [PubMed] [Google Scholar]
- 100.Brunner G, Ferber E, Resch K. Fractionation of membrane vesicles I. A separation method for different populations of membrane vesicles of thymocytes by affinity chromatography on Con A-Sepharose. Anal Biochem. 1977;80:420–429. doi: 10.1016/0003-2697(77)90664-9. [DOI] [PubMed] [Google Scholar]
- 101.Brunner G, Heidrich HG, Golecki JR, Bauer HC, Suter D, Pluckhahn P, Ferber E. Fractionation of membrane vesicles. II. A method for separation of membrane vesicles bearing different enzymes by free-flow electrophoresis. Biochim Biophys Acta. 1977;471:195–212. doi: 10.1016/0005-2736(77)90250-4. [DOI] [PubMed] [Google Scholar]
- 102.Soderman DD, Germerschausen J, Katzen HJ. Affinity binding of intact fat cells and their ghosts to immobilized insulin. Proc Natl Acad Sci USA. 1973;70:792–796. doi: 10.1073/pnas.70.3.792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Johansson G, Gysin R, Flanagan SD. Affinity partitioning of membranes: Evidence for discrete membrane domains containing cholinergic receptor. J Bio Chem. 1981;256:9126–9135. [PubMed] [Google Scholar]
- 104.Ozer J, Wallach DFH. H-2 components and cellular membranes: Distinctions between plasma membrane and endoplasmic reticulum governed by the H-2 region in the mouse. Transplantation. 1967;5:652–667. [PubMed] [Google Scholar]
- 105.Steck TL. Preparation of impermeable inside out and right side out vesicles from eryhtorcyte membranes. In: Korn EP, editor. Methods in Membrane Biology. New York: 1974. pp. 245–281. [Google Scholar]
- 106.Kremmer T, Wisher MH, Evans WH. The lipid composition of plasma membrane subfractions originating from the three major functional domains of the rat hepatocyte cell surface. Biochim Biophys Acta. 1976;455:655–664. doi: 10.1016/0005-2736(76)90039-0. [DOI] [PubMed] [Google Scholar]
- 107.Evans WH. A biochemical dissection of the functional polarity of the plasma membrane of the hepatocyte. Biochim Biophys Acta. 1980;604:27–64. doi: 10.1016/0005-2736(80)90584-2. [DOI] [PubMed] [Google Scholar]
- 108.Brasitus TA, Schachter D. Lipid composition and fluidity of rat enterocyte basolateral membranes. Regional differences. Biochim Biophys Acta. 1984;774:138–146. doi: 10.1016/0005-2736(84)90284-0. [DOI] [PubMed] [Google Scholar]
- 109.Schroeder F, Sweet WD. The role of membrane lipid and structure asymmetry on transport systems. In: Jorgensen PL, Verna R, editors. Advances in Biotechnology of Membrane Ion Transport. Vol. 51. New York, NY: 1988. pp. 183–195. [Google Scholar]
- 110.Schroeder F, Fontaine RN, Kinden DA. LM fibroblast plasma membrane subfractionation by affinity chromatography on ConA-sepharose. Biochim Biophys Acta. 1982;690:231–242. doi: 10.1016/0005-2736(82)90327-3. [DOI] [PubMed] [Google Scholar]
- 111.Gimpl G, Burger K, Fahrenholz F. Cholesterol as modulator of receptor function. Biochemistry. 1997;36:10959–10974. doi: 10.1021/bi963138w. [DOI] [PubMed] [Google Scholar]
- 112.Gimpl G, Fahrenholz F. Human oxytocin receptors in cholesterol-rich vs cholesterol-poor microdomains of the plasma membrane. Eur J Biochem. 2000;267:2483–2497. doi: 10.1046/j.1432-1327.2000.01280.x. [DOI] [PubMed] [Google Scholar]
- 113.Burger K, Gimpl G, Fahrenholz F. Regulation of receptor function by cholesterol. Cell Mol Life Sci. 2000;57:1577–1592. doi: 10.1007/PL00000643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Atshaves BP, McIntosh AL, Gallegos AM, Payne HR, Landrock K, Kier AB, Schroeder F. Sterol carrier protein-2/sterol carrier protein-x gene ablation alters primary cultured hepatocyte plasma membrane lipid raft domains. J Lipid Res. 2007 doi: 10.1194/jlr.M700102-JLR200. submitted. [DOI] [PubMed] [Google Scholar]
- 115.McIntosh A, Gallegos A, Atshaves BP, Storey S, Kannoju D, Schroeder F. Fluorescence and multiphoton imaging resolve unique structural forms of sterol in membranes of living cells. J Biol Chem. 2003;278:6384–6403. doi: 10.1074/jbc.M205472200. [DOI] [PubMed] [Google Scholar]
- 116.Zhang W, McIntosh A, Xu H, Wu D, Gruninger T, Atshaves BP, Liu JCS, Schroeder F. Structural analysis of sterol distribution in the plasma membrane of living cells. Biochemistry. 2005;44:2864–2984. doi: 10.1021/bi048172m. [DOI] [PubMed] [Google Scholar]
- 117.McIntosh A, Atshaves BP, Huang H, Gallegos AM, Kier AB, Schroeder F, Xu H, Zhang W, Liu S. Multiphoton laser scanning microscopy and spatial analysis of dehydroergosterol distributions on plasma membranes of living cells. In: McIntosh T, editor. Lipid Rafts, Methods in Molecular Biology. Totowa, N.J.: 2006. [DOI] [PubMed] [Google Scholar]
- 118.Gallegos AM, McIntosh AL, Kier AB, Schroeder F. Membrane domain distributions: Analysis of fluorescent sterol exchange kinetics. Currents in Analytical Chemistry. 2006 in press. [Google Scholar]
- 119.Schroeder F, Kinden DA. Measurement of phagocytosis using fluorescent latex beads. J Biochem Biophys Meth. 1983;8:15–27. doi: 10.1016/0165-022x(83)90017-9. [DOI] [PubMed] [Google Scholar]
- 120.Yonemura S, Tsukita S, Tsukita S. Direct involvement of ezrin/radixin/moesin (ERM)-binding membrane proteins in the organization of microvilli in collaboration with activated ERM proteins. J Cell Biol. 1999;145:1497–1509. doi: 10.1083/jcb.145.7.1497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Corbeil D, Roper K, Hannah MJ, Hellwig A, Huttner WB. Selective localization of the polytopic membrane protein prominin in microvilli of ephithelial cells--a combinationof apical sorting and retention in plasma membrane protrusions. J Cell Sci. 1999;112:1023–1033. doi: 10.1242/jcs.112.7.1023. [DOI] [PubMed] [Google Scholar]
- 122.Mobius W, Ohno-Iwashita Y, van Donselaar E, Oorschot VMJ, Shimada Y, Fujimoto T, Heijnen HFG, Geuze HJ, Slot JW. Immunoelectron microscopic localization of cholesterol using biotinylated and non-cytolytic perfringolysin O. J Histochem Cytochem. 2002;50:43–55. doi: 10.1177/002215540205000105. [DOI] [PubMed] [Google Scholar]
- 123.Waheed AA, Shimada Y, Heijnen HFG, Nakamura M, Inomata M, Hayashi M, Iwashita S, Slot JW, Ohno-Iwashita Y. Selective binding of perfringolysin O derivative to cholesterol-rich membrane microdomains (rafts) Proc Natl Acad Sci USA. 2001;98:4926–4931. doi: 10.1073/pnas.091090798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Anderson R. The caveolae membrane system. Ann Rev Biochem. 1998;67:199–225. doi: 10.1146/annurev.biochem.67.1.199. [DOI] [PubMed] [Google Scholar]
- 125.Mescher MF, Apgar JR. The plasma membrane ‘skeleton’ of tumor and lymphoid cells: a role in cell lysis. Adv Exper Med Biol. 1985;184:387–400. doi: 10.1007/978-1-4684-8326-0_26. [DOI] [PubMed] [Google Scholar]
- 126.Brown DA, Rose JK. Sorting of GPI anchored proteins to glycolipid-enriched membrane subdomains during transport to the apical cell surface. Cell. 1992;68:533–544. doi: 10.1016/0092-8674(92)90189-j. [DOI] [PubMed] [Google Scholar]
- 127.Smart EJ, Ying Y, Conrad PA, Anderson RGW. Caveolin moves from caveolae to the golgi apparatus in response to cholesterol oxidation. J Cell Biol. 1994;127:1185–1197. doi: 10.1083/jcb.127.5.1185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Pike LJ, Casey L. Localization and turnover of phosphatidylinositol 4,5-bisphsophate in caveolin-enriched membrane domains. J Bio Chem. 1996:26453–26456. doi: 10.1074/jbc.271.43.26453. [DOI] [PubMed] [Google Scholar]
- 129.Fridricksson EK, Shipkova PA, Sheets ED, Holowka D, Baird B, McLafferty FW. Quantitative analysis fo phospholipids in functionally important membrane domains from RBL-2H3 mast cells using tandem high-resolution mass spectrometry. Biochemistry. 1999;38:8056–8063. doi: 10.1021/bi9828324. [DOI] [PubMed] [Google Scholar]
- 130.Wang XQ, Sun P, Paller AS. Ganglioside induces caveolin-1 redistribution and interaction with the epidermal growth factor receptor. J Biol Chem. 2002;277:47028–47034. doi: 10.1074/jbc.M208257200. [DOI] [PubMed] [Google Scholar]
- 131.Ortegren U, Karlsson M, Blazic N, Blomqvist M, Nysrom FH, Gustavsson J, Fredman P, Stralfors P. Lipids and glycosphingolipids in caveolae and surrounding plasma membrane of primary rat adipocytes. Eur J Biochem. 2004;271:2028–2036. doi: 10.1111/j.1432-1033.2004.04117.x. [DOI] [PubMed] [Google Scholar]
- 132.Tietz P, Jefferson JR, Pagano R, LaRusso NF. Membrane microdomains in hepatocytes: potential target areas for proteins involved in canalicular bile secretion. J Lipid Res. 2005;46:1426–1432. doi: 10.1194/jlr.M400412-JLR200. [DOI] [PubMed] [Google Scholar]
- 133.Everson WV, Smart EJ. Influence of caveolin, cholesterol, and lipoproteins on nitric oxide synthase. TCM. 2001;11:246–250. doi: 10.1016/s1050-1738(01)00119-0. [DOI] [PubMed] [Google Scholar]
- 134.Shaul PW, Chambliss KL. Caveolae and Estrogen Receptor Signaling. In: Lisanti MP, Frank PG, editors. Caveolae and Lipid Rafts: Roles in Signal Transduction and the Pathogenesis of Human Disease. Vol. 36. Advances in Molecular and Cell Biology; San Diego: 2005. pp. 109–123. [Google Scholar]
- 135.Feron O. The Caveolin Interaction with Endothelial Nitric Oxide Synthase (eNOS) In: Lisanti MP, Frank PG, editors. Caveolae and Lipid Rafts: Roles in Signal Transduction and the Pathogenesis of Human Disease. Vol. 36. Advances in Molecular and Cell Biology; San Diego: 2005. pp. 91–108. [Google Scholar]
- 136.Skwarek M. Recent controversy surrounding lipid rafts. Arch Immunol Ther Exp. 2004;52:427–431. [PubMed] [Google Scholar]
- 137.McIntosh TJ, Vidal A, Simon SA. Sorting of lipids and trasnmembrane peptides between detergent-soluble bilayers and detergent resistant rafts. Biophys J. 2003;85:1656–1666. doi: 10.1016/S0006-3495(03)74595-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Heerklotz H. Triton promotes domain formation in lipid raft mixtures. Biophys J. 2002;83:2693–2701. doi: 10.1016/S0006-3495(02)75278-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Radeva G, Sharom FJ. Isolation and characterization of lipid rafts with different properteis from RBL-2H3 (rat basophilic leukaemia) cells. Biochem J. 2004;380:219–230. doi: 10.1042/BJ20031348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Gallegos AM, Storey SM, Kier AB, Schroeder F, Ball JM. Stucture and cholesterol dynamics of caveolae/raft and nonraft plasma membrane domains. Biochem. 2006;45:12100–12116. doi: 10.1021/bi0602720. [DOI] [PubMed] [Google Scholar]
- 141.Foster LJ, de Hoog CL, Mann M. Unbiased quantitative proteomics of lipid rafts reveals high specificity for signaling factors. Proc Natl Acad Sci USA. 2003;100:5813–5818. doi: 10.1073/pnas.0631608100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Smart EJ, Ying Y, Mineo C, Anderson RGW. A detergent-free method for purifying caveolae membrane from tissue culture cells. Proc Natl Acad Sci. 1995;92:10404–10408. doi: 10.1073/pnas.92.22.10104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Eckert GP, Igbavboa U, Muller W, Wood WG. Lipid rafts of purified mouse brain synaptosomes prepared with or without detergent reveal different lipid and protein domains. Brain Res. 2003;962:144–150. doi: 10.1016/s0006-8993(02)03986-0. [DOI] [PubMed] [Google Scholar]
- 144.Kirsch C, Eckert GP, Mueller WE. Statins affect cholesterol micro-domains in brain plasma membranes. Biochem Pharmacol. 2002;65:843–856. doi: 10.1016/s0006-2952(02)01654-4. [DOI] [PubMed] [Google Scholar]
- 145.Gallegos AM, Atshaves BP, Storey S, McIntosh A, Petrescu AD, Schroeder F. Sterol carrier protein-2 expression alters plasma membrane lipid distribution and cholesterol dynamics. Biochemistry. 2001;40:6493–6506. doi: 10.1021/bi010217l. [DOI] [PubMed] [Google Scholar]
- 146.Pike LJ, Han X, Chung KN, Gross RW. Lipid rafts are enriched in arachidonic acid and plasmenylethanolamine and their composition is independent of caveolin-1 expression: a quantitative electrospray ionization/mass spectrometric analysis. Biochemistry. 2002;41:2075–2088. doi: 10.1021/bi0156557. [DOI] [PubMed] [Google Scholar]
- 147.Gallegos AM, McIntosh AL, Atshaves BP, Schroeder F. Structure and cholesterol domain dynamics of an enriched caveolae/raft isolate. Biochem J. 2004;382:451–461. doi: 10.1042/BJ20031562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.McIntosh AL, Atshaves BP, Huang H, Gallegos AM, Kier AB, Schroeder F, Xu H, Zhang W, Liu S. Multiphoton laser scanning microscopy and spatial analysis of dehydroergosterol distributions on plasma membrane of living cells. In: McIntosh TJ, editor. Lipid Rafts, Methods in Molecular Biology. Totowa, NJ: 2006. in press. [DOI] [PubMed] [Google Scholar]
- 149.Atshaves BP, Jefferson JR, Kier AB, Schroeder F. Sterol carrier protein-2, a new sphingolipid binding protein, alters sphingolipid distribution in plasma membrane caveolae/lipid raft domains. Am J Physiol. 2007 doi: 10.1007/s11745-007-3091-z. submitted. [DOI] [PubMed] [Google Scholar]
- 150.Lipardi C, Mora R, Colomer V, Paladino S, Nitsch L, Rodriguez-Boulan E, Zurzolo C. Caveolin transfection results in caveolae formation but not apical sorting of glycosylphosphatidylinositol (GPI)-anchored proteins in epithelial cells. Journal of Cell Biology. 1998;140:617–626. doi: 10.1083/jcb.140.3.617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Wang L, Connelly MA, Ostermeyer AG, Chen H, Williams DL, Brown DA. Caveolin-1 does not affect SR-B1 mediated cholesterol efflux or selective uptake of cholesteryl ester in two cell lines. J Lipid Res. 2003;44:807–815. doi: 10.1194/jlr.M200449-JLR200. [DOI] [PubMed] [Google Scholar]
- 152.Mora R, Bonilha VL, Marmorstein A, Scherer PE, Brown D, Lisanti MP, Rodriguez-Boulan E. Caveolin-2 localizes to the Golgi complex but redistributes to the plasma membrane, caveolae, and rafts when co-expressed with caveolin-1. J Biol Chem. 1999;274:25708–25717. doi: 10.1074/jbc.274.36.25708. [DOI] [PubMed] [Google Scholar]
- 153.Pohl J, Ring A, Stremmel W. Uptake of long chain fatty acids in HepG2 cells involves caveolae: analysis of a novel pathway. J Lipid Res. 2002;43:1390–1399. doi: 10.1194/jlr.m100404-jlr200. [DOI] [PubMed] [Google Scholar]
- 154.Trigatti B, Anderson RGW, Gerber GE. Identification of caveolin-1 as a fatty acid binding protein. Biochem Biophys Res Commun. 1999;255:34–39. doi: 10.1006/bbrc.1998.0123. [DOI] [PubMed] [Google Scholar]
- 155.Febbraio M, Abumrad N, Hajjar DP, Sharma K, Cheng W, Pearce SFA, Silverstein RL. A null mutation in murine CD36 reveals an important role in fatty acid and lipoprotein metabolism. J Biol Chem. 1999;274:19055–19062. doi: 10.1074/jbc.274.27.19055. [DOI] [PubMed] [Google Scholar]
- 156.Harmon CM, Luce P, Abumrad NA. Labelling of an 88 kDa adipocyte membrane protein by sulpho-N- succinimidyl long-chain fatty acids: inhibition of fatty acid transport. Biochem Soc Trans. 1992;20:811–813. doi: 10.1042/bst0200811. [DOI] [PubMed] [Google Scholar]
- 157.Ibrahimi A, Sfeir Z, Magharaie H, Amri EZ, Grimaldi P, Abumrad NA. Expression of the CD36 homolog (FAT) in fibroblast cells: Effects on fatty acid transport. Proc Natl Acad Sci. 1996;93:2646–2651. doi: 10.1073/pnas.93.7.2646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Schaffer JE, Lodish HF. Molecular mechanism of long-chain fatty acid uptake. Trends Cardiovasc Med. 1995;5:218–224. doi: 10.1016/1050-1738(95)00102-6. [DOI] [PubMed] [Google Scholar]
- 159.Kolleck I, Guthmann F, Ladhoff AM, Tandon NN, Schlame M, Rustow B. Cellular cholesterol stimulates acute uptake of palmitate by redistribution of fatty acid translocase in type II pneumocytes. Biochem. 2006;41:6369–6375. doi: 10.1021/bi015980u. [DOI] [PubMed] [Google Scholar]
- 160.Pohl J, Ring A, Schulze-Bergkamen H, Schad A, Verkade P, Stremmel W. Long chain fatty acid uptake into adipocytes depends on lipid raft function. Biochem. 2004;43:4179–4187. doi: 10.1021/bi035743m. [DOI] [PubMed] [Google Scholar]
- 161.Pohl J, Ring A, Korkmaz U, Ehehalt R, Stremmel W. FAT/CD36-mediated long chain fatty acid uptake in adipocytes requires plasma membrane rafts. Mol Biol Cell. 2005;16:24–31. doi: 10.1091/mbc.E04-07-0616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Pohl J, Ring A, Ehehalt R, Hermann A, Stremmel W. New concepts of cellular fatty acid uptake: role of fatty acid transport proteins and of caveolae. Proc Nutrition Society. 2004;63:259–262. doi: 10.1079/PNS2004341. [DOI] [PubMed] [Google Scholar]
- 163.Pitto M, Brunner J, Ferraretto A, Ravasi D, Palestini P, Masserini M. Use of a photoactivatable GM1 ganglioside analogue to assess lipid distribution in caveolae bilayer. Glycoconjugate Journal. 2000;17:215–222. doi: 10.1023/a:1026593307882. [DOI] [PubMed] [Google Scholar]
- 164.Smart EJ, van der Westhuyzen DR. Scavenger receptors, caveolae, caveolin, and cholesterol trafficking. In: Chang TY, Freeman DA, editors. Intracellular Cholesterol Trafficking. Boston: 1998. pp. 253–272. [Google Scholar]
- 165.Schroeder F, Zhou M, Swaggerty CL, Atshaves BP, Petrescu AD, Storey S, Martin GG, Huang H, Helmkamp GM, Ball JM. Sterol carrier protein-2 functions in phosphatidylinositol transfer and signaling. Biochemistry. 2003;42:3189–3202. doi: 10.1021/bi026904+. [DOI] [PubMed] [Google Scholar]
- 166.Conrad PA, Smart EJ, Ying Y, Anderson RGW, Bloom GS. Caveolin cycles between plasma membrane caveolae and the golgi complex by microtubule-dependent and microtubule-independent steps. J Cell Biol. 1995;131:1421–1433. doi: 10.1083/jcb.131.6.1421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Shaul PW, Anderson RGW. Role of plasmalemmal caveolae in signal transduction. Am J Physiol. 1998;19:L843–L851. doi: 10.1152/ajplung.1998.275.5.L843. [DOI] [PubMed] [Google Scholar]
- 168.Wustner D, Herrmann A, Hao M, Maxfield FR. Rapid nonvesicular transport of sterol between the plasma membrane domains of polarized hepatic cells. J Biol Chem. 2002;277:30325–30336. doi: 10.1074/jbc.M202626200. [DOI] [PubMed] [Google Scholar]
- 169.Cockcroft S. Phosphatidylinositol transfer proteins: a requirement in signal transduction and vesicle traffick. BioEssays. 1998;20:423–432. doi: 10.1002/(SICI)1521-1878(199805)20:5<423::AID-BIES9>3.0.CO;2-O. [DOI] [PubMed] [Google Scholar]
- 170.Cockcroft S. Mammalian phosphatidylinositol transfer protein: emerging roles in signal transduction and vesicular traffick. Chem Phys Lipids. 1999;98:23–33. doi: 10.1016/s0009-3084(99)00015-8. [DOI] [PubMed] [Google Scholar]
- 171.Bankaitis VA, Aitken JR, Cleves AE, Dowhan W. An essential role for a phospholipid transfer protein in yeast Golgi function. Nature. 1990;347:561–562. doi: 10.1038/347561a0. see comments. [DOI] [PubMed] [Google Scholar]
- 172.Jones SM, Jr, Alb JG, Phillips SE, Bankaitis VA, Howell KE. A Phosphatidylinositol 3-Kinase and Phosphatidylinositol Transfer Protein Act Synergistically in Formation of Constitutive Transport Vesicles from the Trans-Golgi Network. The Journal of Biological Chemistry. 1998;273:10349–10354. doi: 10.1074/jbc.273.17.10349. [DOI] [PubMed] [Google Scholar]
- 173.Fang M, Rivas MP, Bankaitis VA. The contribution of lipids and lipid metabolism to cellular functions of the Golgi complex. Biochim Biophys Acta. 1998;1404:85–100. doi: 10.1016/s0167-4889(98)00049-4. [DOI] [PubMed] [Google Scholar]
- 174.Schroeder F, Myers-Payne SC, Billheimer JT, Wood WG. Probing the ligand binding sites of fatty acid and sterol carrier proteins: effects of ethanol. Biochemistry. 1995;34:11919–11927. doi: 10.1021/bi00037a033. [DOI] [PubMed] [Google Scholar]
- 175.Frolov A, Cho TH, Billheimer JT, Schroeder F. Sterol carrier protein-2, a new fatty acyl coenzyme A-binding protein. J Biol Chem. 1996;271:31878–31884. doi: 10.1074/jbc.271.50.31878. [DOI] [PubMed] [Google Scholar]
- 176.Gossett RE, Frolov AA, Roths JB, Behnke WD, Kier AB, Schroeder F. Acyl Co A binding proteins: multiplicity and function. Lipids. 1996;31:895–918. doi: 10.1007/BF02522684. [DOI] [PubMed] [Google Scholar]
- 177.McArthur MJ, Atshaves BP, Frolov A, Foxworth WD, Kier AB, Schroeder F. Cellular uptake and intracellular trafficking of long chain fatty acids. J Lipid Res. 1999;40:1371–1383. [PubMed] [Google Scholar]
- 178.Frolov A, Cho TH, Murphy EJ, Schroeder F. Isoforms of rat liver fatty acid binding protein differ in structure and affinity for fatty acids and fatty acyl CoAs. Biochemistry. 1997;36:6545–6555. doi: 10.1021/bi970205t. [DOI] [PubMed] [Google Scholar]
- 179.Frolov AA, Schroeder F. Acyl coenzyme A binding protein: conformational sensitivity to long chain fatty acyl-CoA. J Biol Chem. 1998;273:11049–11055. doi: 10.1074/jbc.273.18.11049. [DOI] [PubMed] [Google Scholar]
- 180.Knudsen J, Jensen MV, Hansen JK, Faergeman NJ, Neergard T, Gaigg B. Role of acyl CoA binding protein in acyl CoA transport, metabolism, and cell signaling. Mol Cell Biochem. 1999;192:95–103. doi: 10.1007/978-1-4615-4929-1_11. [DOI] [PubMed] [Google Scholar]
- 181.Gadella TW, Jr, Wirtz KW. The low-affinity lipid binding site of the non-specific lipid transfer protein. Implications for its mode of action. Biochim Biophys Acta. 1991;1070:237–245. doi: 10.1016/0005-2736(91)90170-d. [DOI] [PubMed] [Google Scholar]
- 182.Murphy EJ. Sterol carrier protein-2 expression increases fatty acid uptake and cytoplasmic diffusion in L-cell fibroblasts. Am J Physiol. 1998;275:G237–G243. doi: 10.1152/ajpgi.1998.275.2.G237. [DOI] [PubMed] [Google Scholar]
- 183.Ek BA, Cistola DP, Hamilton JA, Kaduce TL, Spector AA. Fatty acid binding proteins reduced 15-lipoxygenase-induced oxygenation of linoleic acid and arachidonic acid. Biochim Biophys Acta. 1997;1346:75–85. doi: 10.1016/s0005-2760(97)00021-0. [DOI] [PubMed] [Google Scholar]
- 184.Bloj B, Zilversmit DB. Accelerated transfer of neutral glycosphingolipids and ganglioside GM1 by a purified lipid transfer protein. J Biol Chem. 1981;256:5988–5991. [PubMed] [Google Scholar]
- 185.Fong TH, Wang SM, Lin HS. Immunocytochemical demonstration of a lipid droplet specific capsule in cultured Leydig cells of golden hamsters. J Cell Biochem. 1996;63:366–373. doi: 10.1002/(sici)1097-4644(19961201)63:3<366::aid-jcb12>3.0.co;2-f. [DOI] [PubMed] [Google Scholar]
- 186.DiCorleto PE, Warach JB, Zilversmit DB. Purification and characterization of two phospholipid exchange proteins from bovine heart. J Biol Chem. 1979;254:7795–7802. [PubMed] [Google Scholar]
- 187.Colles SM, Woodford JK, Moncecchi D, Myers-Payne SC, McLean LR, Billheimer JT, Schroeder F. Cholesterol interactions with recombinant human sterol carrier protein-2. Lipids. 1995;30:795–804. doi: 10.1007/BF02533954. [DOI] [PubMed] [Google Scholar]
- 188.Brown MS, Goldstein JL. Sterol regulatory element binding proteins (SREBPs): controllers of lipid synthesis and cellular uptake. Nutrition Reviews. 1998;56:S54–S75. doi: 10.1111/j.1753-4887.1998.tb01680.x. [DOI] [PubMed] [Google Scholar]
- 189.Sakai J, Duncan EA, Rawson RB, Hua X, Brown MS, Goldstein JL. Sterol-regulated release of SREBP-2 from cell membranes requires two sequential cleavages, one within a transmembrane segment. Cell. 1996;85:1037–1046. doi: 10.1016/s0092-8674(00)81304-5. [DOI] [PubMed] [Google Scholar]
- 190.Shimomura I, Bashmakov Y, Shimano H, Horton JD, Goldstein JL, Brown MS. Cholesterol feeding reduces nuclear forms of sterol regulatory element binding proteins in hamster liver. Proceedings of the National Academy of Sciences of the United States of America. 1997;94:12354–12359. doi: 10.1073/pnas.94.23.12354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Wang X, Zelenski NG, Yang J, Sakai J, Brown MS, Goldstein JL. Cleavage of sterol regulatory element binding proteins (SREBPs) by CPP32 during apoptosis. EMBO Journal. 1996;15:1012–1020. [PMC free article] [PubMed] [Google Scholar]
- 192.Brown MS, Goldstein JL. Sterol regulatory element binding proteins (SREBPs): controllers of lipid synthesis and cellular uptake. Nutrition Reviews. 1998;56:S1–3. doi: 10.1111/j.1753-4887.1998.tb01680.x. [Review] [10 refs] discussion. [DOI] [PubMed] [Google Scholar]
- 193.Wang X, Sato R, Brown MS, Hua X, Goldstein JL. SREBP-1, a membrane-bound transcription factor released by sterol-regulated proteolysis. Cell. 1994;77:53–62. doi: 10.1016/0092-8674(94)90234-8. [DOI] [PubMed] [Google Scholar]
- 194.Kriska T, Levchenko VV, Korytowski W, Atshaves BP, Schroeder F, Girotti AW. Hypersensitivity of sterol carrier protein-2 overexpressing hepatoma cells to lethal peroxidative damage induced by an exogenous cholesterol hydroperoxide. J Biol Chem. 2006;281:23643–23651. doi: 10.1074/jbc.M600744200. [DOI] [PubMed] [Google Scholar]
- 195.Brown MS, Goldstein JL. Suppression of 3-hydroxy-3-methylglutaryl coenzyme A reductase activity and inhibition of growth of human fibroblasts by 7- ketocholesterol. J Biol Chem. 1974;249:7306–7314. [PubMed] [Google Scholar]
- 196.Milis DG, Moore MK, Atshaves BP, Schroeder F, Jefferson JR. Sterol carrier protein-2 expression alters sphingolipid metabolism in transfected mouse L-cell fibroblasts. Mol Cell Biochem. 2006;283:57–66. doi: 10.1007/s11010-006-2270-1. [DOI] [PubMed] [Google Scholar]
- 197.Gavey KL, Noland BJ, Scallen TJ. The participation of sterol carrier protein2 in the conversion of cholesterol to cholesterol ester by rat liver microsomes. J Biol Chem. 1981;256:2993–2999. [PubMed] [Google Scholar]
- 198.Paulussen RJA, Veerkamp JH. Intracellular fatty acid-binding proteins characteristics and function. In: Hilderson HJ, editor. Subcellular Biochemistry. Vol. 16. New York: 1990. pp. 175–226. [DOI] [PubMed] [Google Scholar]
- 199.Atshaves BP, McIntosh AL, Landrock D, Payne HR, Schroeder F, Kier AB. Transgenic expressio of sterol carrier protein-2 potentiates diet-induced hepatic cholesterol accumulation in mice. Lipids. 2007 submitted. [Google Scholar]
- 200.Kelleher JA, Sun GY. Effects of free fatty acids and acyl coenzyme A on diacylglycerol kinase in rat brain. J Neurosci Res. 1989;23:87–94. doi: 10.1002/jnr.490230112. [DOI] [PubMed] [Google Scholar]
- 201.Wieland SJ, Gong Q, Poblete H, Fletcher JE, Chen L, Kallen RG. Modulation of human muscle sodium channels by intracellular fatty acids is dependent of the channel isoform. J Biol Chem. 1996;271:19037–19041. doi: 10.1074/jbc.271.32.19037. [DOI] [PubMed] [Google Scholar]
- 202.Anel A, Richieri GV, Kleinfeld AM. A tyrosine phosphorylation requirement for cytotoxic T lymphocyte degranulation. J Biol Chem. 1994;269:9506–9513. [PubMed] [Google Scholar]
- 203.Hawley HP, Gordon GB. The effects of long chain free fatty acids on human neutrophil function and structure. Lab Invest. 1976;34:216–221. [PubMed] [Google Scholar]
- 204.Downard J, Toal D. Branched-chain fatty acids: the case for a novel form of cell-cell signalling during M. xanthus development. Molec Microbiol. 1995;16:171–175. doi: 10.1111/j.1365-2958.1995.tb02290.x. [DOI] [PubMed] [Google Scholar]
- 205.Ailhaud GP, Abumrad N, Amri EZ, Grimaldi PA. A new look at fatty acids as signal-transduction molecules. In: Galli C, Simopoulos AP, Tremoli E, editors. Fatty Acids and Lipids: Biological Aspects. World Rev. Nutr. Diet.; Basel: 1995. pp. 35–45. [Google Scholar]
- 206.Huang H, Starodub O, McIntosh A, Kier AB, Schroeder F. Liver fatty acid binding protein targets fatty acids to the nucleus: real-time confocal and multiphoton fluorescence imaging in living cells. J Biol Chem. 2002;277:29139–29151. doi: 10.1074/jbc.M202923200. [DOI] [PubMed] [Google Scholar]
- 207.Huang H, Starodub O, McIntosh A, Atshaves BP, Woldegiorgis G, Kier AB, Schroeder F. Liver fatty acid binding protein colocalizes with peroxisome proliferator receptor alpha and enhances ligand distribution to nuclei of living cells. Biochemistry. 2004;43:2484–2500. doi: 10.1021/bi0352318. [DOI] [PubMed] [Google Scholar]
- 208.Hostetler HA, Petrescu AD, Kier AB, Schroeder F. Peroxisome proliferator activated receptor alpha (PPARalpha) interacts with high affinity and is conformationally responsive to endogenous ligands. J Biol Chem. 2005;280:18667–18682. doi: 10.1074/jbc.M412062200. [DOI] [PubMed] [Google Scholar]
- 209.Hostetler HA, Kier AB, Schroeder F. Very-long-chain and branched-chain fatty acyl CoAs are high affinity ligands for the peroxisome proliferator-activated receptor alpha (PPARalpha) Biochemistry. 2006;45:7669–7681. doi: 10.1021/bi060198l. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 210.Branstrom R, Leibiger IB, Leibiger B, Corkey BE, Berggren PO. Long chain coenzyme A esters activate the pore forming subunit (Kir6.2) of the ATP-regulated potassium channel. J Biol Chem. 1998;273:31395–31400. doi: 10.1074/jbc.273.47.31395. [DOI] [PubMed] [Google Scholar]
- 211.Holian O, Kumar R, Nyhus LM. Fatty acyl esters potentiate fatty acid induced activation of protein kinase C. Biochem Biophys Res Commun. 1989;160:1110–1116. doi: 10.1016/s0006-291x(89)80118-4. [DOI] [PubMed] [Google Scholar]
- 212.Faergeman NJ, Knudsen J. Role of long-chain fatty acyl-CoA esters in the regulation of metabolism and in cell signalling. Biochem J. 1997;323:1–12. doi: 10.1042/bj3230001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Corkey BE, Deeney JT. Acyl CoA regulation of metabolism and signal transduction. Prog Clin Biol Res. 1990;321:217–232. [PubMed] [Google Scholar]
- 214.Deeney JT, Tornheim K, Korchak HM, Prentki M, Corkey BE. Acyl-CoA esters modulate intracellular Ca2+ handling by permeabilized clonal pancreatic b-cells. J Biol Chem. 1992;267:19840–19845. [PubMed] [Google Scholar]
- 215.Majumdar S, Rossi MW, Fujiki T, Phillips WA, Disa S, Queen CF, Johnston RB, Jr, Rosen OM, Corkey BE, Korchak HM. Protein kinase C isotypes and signaling in neutrophils. Differential substrate specificities of a translocatable calcium- and phospholipid-dependent b-protein kinase C and a novel calcium independent, phospholipid-dependent protein kinase which is inhibited by long chain fatty acyl coenzyme A. J Biol Chem. 1991;266:9285–9294. [PubMed] [Google Scholar]
- 216.Pfanner N, Glick BS, Arden SR, Rothman JE. Fatty acylation promotes fusion of transport vesicles with Golgi cisternae. J Cell Biol. 1990;110:955–961. doi: 10.1083/jcb.110.4.955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Pfanner N, Orci L, Glick BS, Amherdt M, Arden SR, Malhotra V, Rothman JE. Fatty acyl CoA is required for budding of transport vesicles from Golgi cisternae. Cell. 1989;59:95–102. doi: 10.1016/0092-8674(89)90872-6. [DOI] [PubMed] [Google Scholar]
- 218.Glick BS, Rothman JE. Possible role for fatty acyl-coenzyme A in intracellular protein transport. Nature. 1987;326:309–312. doi: 10.1038/326309a0. [DOI] [PubMed] [Google Scholar]
- 219.Hertz R, Magenheim J, Berman I, Bar-Tana J. Fatty acyl-CoA thioesters are ligands of hepatic nuclear factor-4a. Nature. 1998;392:512–516. doi: 10.1038/33185. [DOI] [PubMed] [Google Scholar]
- 220.Petrescu AD, Hertz R, Bar-Tana J, Schroeder F, Kier AB. Ligand specificity and conformational dependence of the hepatic nuclear factor-4alpha (HNF-4a) J Biol Chem. 2002;277:23988–23999. doi: 10.1074/jbc.M201241200. [DOI] [PubMed] [Google Scholar]
- 221.Hertz R, Ben-Haim M, Petrescu A, Kalderon B, Berman I, Eldad N, Schroeder F, Bar-Tana J. Rescue of MODY-1 by agonist ligands of HNF4alpha. J Biol Chem. 2003;278:22578–22585. doi: 10.1074/jbc.M212138200. [DOI] [PubMed] [Google Scholar]
- 222.Petrescu AD, Payne HR, Boedeker AL, Chao H, Hertz R, Bar-Tana J, Schroeder F, Kier AB. Physical and functional interaction of acyl CoA binding protein (ACBP) with hepatocyte nuclear factor-4alpha (HNF4alpha) J Biol Chem. 2003;278:51813–51824. doi: 10.1074/jbc.M303858200. [DOI] [PubMed] [Google Scholar]
- 223.Hertz R, Kalderon B, Byk T, Berman I, Za’tara G, Mayer R, Bar-Tana J. Thioesterase activity and acyl-CoA/fatty acid cross talk of hepatocyte nuclear factor-4a. J Biol Chem. 2005;280:24451–24461. doi: 10.1074/jbc.M500732200. [DOI] [PubMed] [Google Scholar]
- 224.Weigert R, Silletta MG, Spano S, Turacchio G, Cericola C, Colanzi A, Senatore S, Mancini R, Polishchuk EV, Salmona M, Facchiano F, Burger KNJ, Mironov A, Luini A, Corda D. CtBP/BARS induces fission of Golgi membranes by acylating lysophosphatidic acid. Nature. 1999;402:429–433. doi: 10.1038/46587. [DOI] [PubMed] [Google Scholar]
- 225.Schmidt A, Wolde M, Thiele C, Fest W, Kratzin H, Podtelejnikov AV, Witke W, Huttner WB, Soling HD. Endophilin mediates vesicle formation by transfer of arachidonate to lysophosphatidic acid. Nature. 1999;401:133–141. doi: 10.1038/43613. [DOI] [PubMed] [Google Scholar]
- 226.Witting SR, Maiorano JN, Davidson WS. Ceramide enhances cholesteorl efflux to apolipoprotein A-I by increasing the cell surface presence of ATP-binding casette transporter A1. J Biol Chem. 2003;278:40121–40127. doi: 10.1074/jbc.M305193200. [DOI] [PubMed] [Google Scholar]
- 227.Igarashi J. eNOS regulation by sphingosine 1 phosphate and caveolin. In: Lisanti MP, Frank PG, editors. Caveolae and Lipid Rafts: Roles in Signal Transduction and the Pathogenesis of Human Disease. Advances in Molecular and Cell Biology; New York: 2005. pp. 125–140. [Google Scholar]
- 228.Sillence DJ, Puri V, Marks DL, Butters TD, Dwek RA, Pagano RE, Platt FM. Glucosylceramide modulates membrane traffic along the endocytic pathway. J Lipid Res. 2002;43:1837–1845. doi: 10.1194/jlr.m200232-jlr200. [DOI] [PubMed] [Google Scholar]
- 229.Kasahara K, Sanai Y. Functional roles of glycosphingolipids in signal transduction via lipid rafts. Glycoconjugate Journal. 2006;17:153–162. doi: 10.1023/a:1026576804247. [DOI] [PubMed] [Google Scholar]
- 230.Sharma DK, Brown JC, Choudhury A, Peterson TE, Holicky E, Marks DL, Simari R, Parton RG, Pagano RE. Selective stimulation of caveolar endocytosis by glycosphingolipids and cholesterol. Mol Biol Cell. 2004;15:3114–3122. doi: 10.1091/mbc.E04-03-0189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 231.Singh RD, Puri V, Valiyaveettil JT, Marks DL, Bittman R, Pagano RE. Selective caveolin-1 dependent endocytosis of glycosphingolipids. Mol Biol Cell. 2003;14:3254–3265. doi: 10.1091/mbc.E02-12-0809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 232.Fong TH, Wang SM. Dissection of the signaling mechanism for capsule detachment of lipid droplets in rat adrenocortical cells. J Cell Biochem. 1997;65:67–74. [PubMed] [Google Scholar]
- 233.Waugh MG, Lawson D, Tan SK, Hsuan JJ. Phosphatidyl 4-phosphate synthesis in immunoisolated caveolae-like vesicles and low buoyant non-caveolar membranes. J Biol Chem. 1998;273:17115–17121. doi: 10.1074/jbc.273.27.17115. [DOI] [PubMed] [Google Scholar]
- 234.Fujimoto T, Hayashi M, Iwamoto M, Ohno-Iwashita Y. Crosslinked plasmalemmal cholesterol is sequestered to caveolae: analysis with a new cytochemical probe. J Histochem Cytochem. 1997;45:1197–1205. doi: 10.1177/002215549704500903. [DOI] [PubMed] [Google Scholar]
- 235.Stolowich NJ, Frolov A, Atshaves BP, Murphy E, Jolly CA, Billheimer JT, Scott AI, Schroeder F. The sterol carrier protein-2 fatty acid binding site: an NMR, Circular Dichroic, and Fluorescence spectroscopic determination. Biochemistry. 1997;36:1719–1729. doi: 10.1021/bi962317a. [DOI] [PubMed] [Google Scholar]
- 236.Avdulov NA, Chochina SV, Myers-Payne S, Hubbell T, Igbavboa U, Schroeder F, Wood WG. Expression and lipid binding of sterol carrier protein-2 and liver fatty acid binding proteins: differential effects of ethanol in vivo and in vitro. In: Riemersma RAARKRWaWR., editor. Essential Fatty Acids and Eicosanoids: Invited Papers from the Fourth International Congress. Champaign, IL: 1998. pp. 324–327. [Google Scholar]
- 237.Stolowich NJ, Frolov A, Petrescu AD, Scott AI, Billheimer JT, Schroeder F. Holo-sterol carrier protein-2: 13C-NMR investigation of cholesterol and fatty acid binding sites. J Biol Chem. 1999;274:35425–35433. doi: 10.1074/jbc.274.50.35425. [DOI] [PubMed] [Google Scholar]
- 238.Avdulov NA, Chochina SV, Igbavboa U, Warden CH, Schroeder F, Wood WG. Lipid binding to sterol carrier protein-2 is inhibited by ethanol. Biochim Biophys Acta. 1999;1437:37–45. doi: 10.1016/s0005-2760(98)00178-7. [DOI] [PubMed] [Google Scholar]
- 239.Bordewick U, Heese M, Borchers T, Robenek H, Spener F. Compartmentation of hepatic fatty-acid-binding protein in liver cells and its effect on microsomal phosphatidic acid biosynthesis. Biol Chem Hoppe-Seyler. 1989;370:229–238. doi: 10.1515/bchm3.1989.370.1.229. [DOI] [PubMed] [Google Scholar]
- 240.Schroeder F, Jefferson JR, Kier AB, Knittell J, Scallen TJ, Wood WG, Hapala I. Membrane cholesterol dynamics: cholesterol domains and kinetic pools. Proc Soc Exp Biol Med. 1991;196:235–252. doi: 10.3181/00379727-196-43185. [DOI] [PubMed] [Google Scholar]
- 241.Schroeder F, Frolov AA, Murphy EJ, Atshaves BP, Jefferson JR, Pu L, Wood WG, Foxworth WB, Kier AB. Recent advances in membrane cholesterol domain dynamics and intracellular cholesterol trafficking. Proc Soc Exp Biol Med. 1996;213:150–177. doi: 10.3181/00379727-213-44047. [DOI] [PubMed] [Google Scholar]
- 242.Schroeder F, Gallegos AM, Atshaves BP, McIntosh A, Petrescu AD, Huang H, Chao H, Yang H, Frolov A, Kier AB. Recent advances in membrane microdomains: rafts, caveolae and intracellular cholesterol trafficking. Exp Biol Med. 2001;226:873–890. doi: 10.1177/153537020122601002. [DOI] [PubMed] [Google Scholar]