Abstract
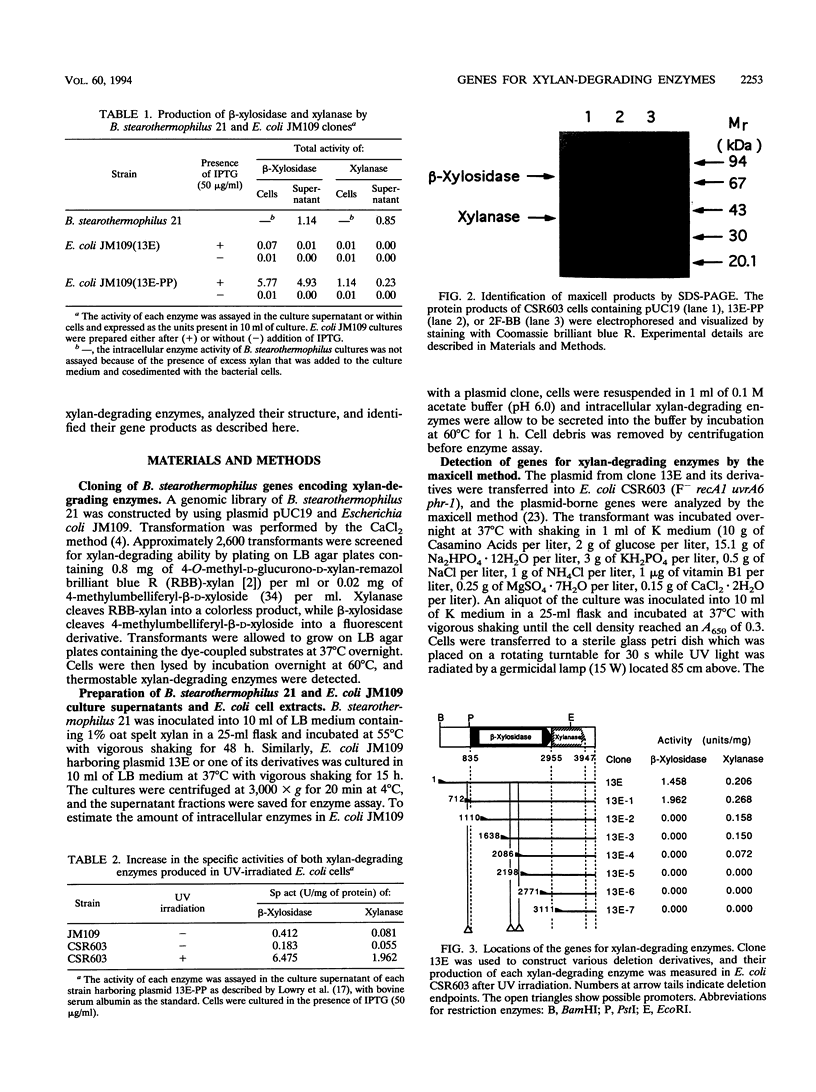
Bacillus stearothermophilus 21 is a gram-positive, facultative thermophilic aerobe that can utilize xylan as a sole source of carbon. We isolated this strain from soil, purified its extracellular xylanase and beta-xylosidase, and analyzed the two-step degradation of xylan by these enzymes (T. Nanmori, T. Watanabe, R. Shinke, A. Kohno, and Y. Kawamura, J. Bacteriol. 172:6669-6672, 1990). An Escherichia coli transformant carrying a 4.2-kbp chromosomal segment of this bacterium as a recombinant plasmid was isolated. It excreted active beta-xylosidase and xylanase into the culture medium. The plasmid was introduced into UV-sensitive E. coli CSR603, and its protein products were analyzed by the maxicell method. Proteins harboring beta-xylosidase and xylanase activities were identified, and their molecular masses were estimated by sodium dodecyl sulfate-polyarylamide gel electrophoresis to be 75 and 40 kDa, respectively. The values were identical to those of proteins prepared from cells of B. stearothermophilus 21. The genes for both enzymes were encoded in a 3.4-kbp PstI fragment derived from the 4.2-kbp chromosomal segment. The nucleotide sequence of the 4.2-kbp segment was accordingly determined. The beta-xylosidase gene (xylA) is located upstream of the xylanase gene (xynA) with a possible promoter and a Shine-Dalgarno sequence. The latter gene is preceded by two possible promoters and a Shine-Dalgarno sequence that are located within the 3'-terminal coding region of the former. The two genes thus appear to be, at least partly, expressed independently, which was experimentally confirmed in E. coli by deletion analysis.(ABSTRACT TRUNCATED AT 250 WORDS)
Full text
PDF



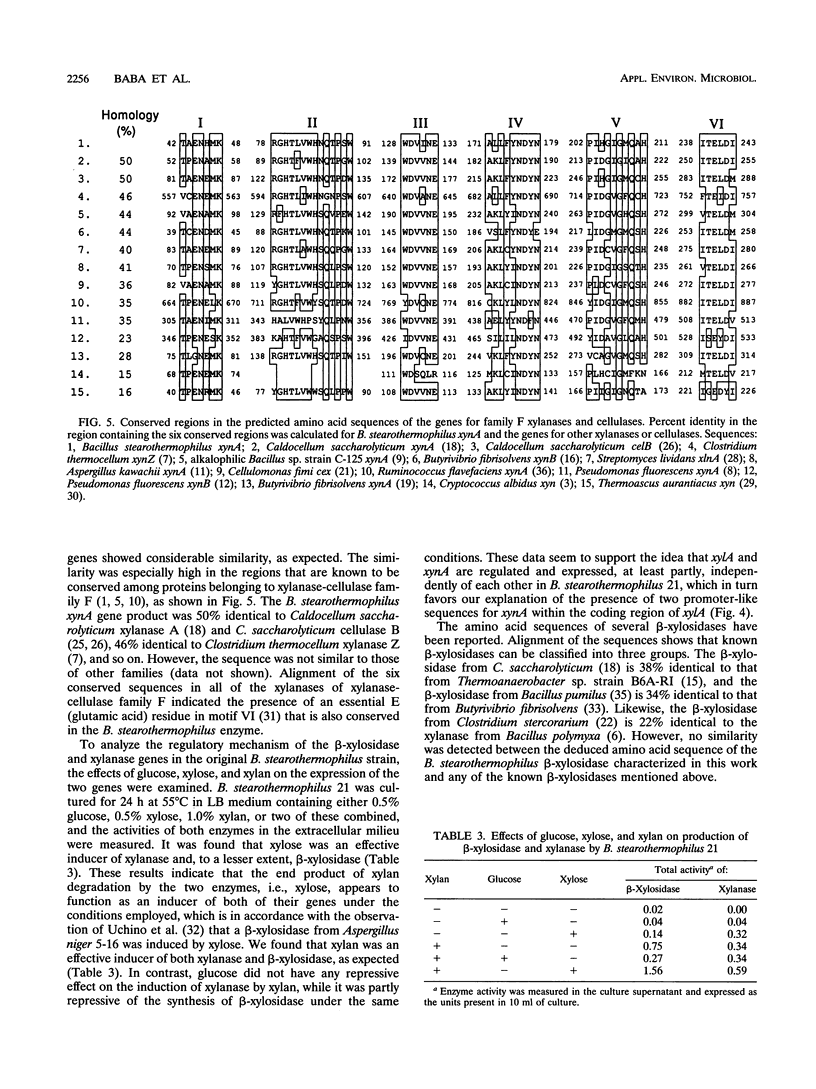


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Biely P., Markovic O., Mislovicová D. Sensitive detection of endo-1,4-beta-glucanases and endo-1,4-beta-xylanases in gels. Anal Biochem. 1985 Jan;144(1):147–151. doi: 10.1016/0003-2697(85)90096-x. [DOI] [PubMed] [Google Scholar]
- Boucher F., Morosoli R., Durand S. Complete nucleotide sequence of the xylanase gene from the yeast Cryptococcus albidus. Nucleic Acids Res. 1988 Oct 25;16(20):9874–9874. doi: 10.1093/nar/16.20.9874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Béguin P. Molecular biology of cellulose degradation. Annu Rev Microbiol. 1990;44:219–248. doi: 10.1146/annurev.mi.44.100190.001251. [DOI] [PubMed] [Google Scholar]
- Cohen S. N., Chang A. C., Hsu L. Nonchromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-factor DNA. Proc Natl Acad Sci U S A. 1972 Aug;69(8):2110–2114. doi: 10.1073/pnas.69.8.2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilkes N. R., Henrissat B., Kilburn D. G., Miller R. C., Jr, Warren R. A. Domains in microbial beta-1, 4-glycanases: sequence conservation, function, and enzyme families. Microbiol Rev. 1991 Jun;55(2):303–315. doi: 10.1128/mr.55.2.303-315.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gosalbes M. J., Pérez-González J. A., González R., Navarro A. Two beta-glycanase genes are clustered in Bacillus polymyxa: molecular cloning, expression, and sequence analysis of genes encoding a xylanase and an endo-beta-(1,3)-(1,4)-glucanase. J Bacteriol. 1991 Dec;173(23):7705–7710. doi: 10.1128/jb.173.23.7705-7710.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grépinet O., Chebrou M. C., Béguin P. Nucleotide sequence and deletion analysis of the xylanase gene (xynZ) of Clostridium thermocellum. J Bacteriol. 1988 Oct;170(10):4582–4588. doi: 10.1128/jb.170.10.4582-4588.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall J., Hazlewood G. P., Huskisson N. S., Durrant A. J., Gilbert H. J. Conserved serine-rich sequences in xylanase and cellulase from Pseudomonas fluorescens subspecies cellulosa: internal signal sequence and unusual protein processing. Mol Microbiol. 1989 Sep;3(9):1211–1219. doi: 10.1111/j.1365-2958.1989.tb00271.x. [DOI] [PubMed] [Google Scholar]
- Henrissat B., Claeyssens M., Tomme P., Lemesle L., Mornon J. P. Cellulase families revealed by hydrophobic cluster analysis. Gene. 1989 Sep 1;81(1):83–95. doi: 10.1016/0378-1119(89)90339-9. [DOI] [PubMed] [Google Scholar]
- Ito K., Ikemasu T., Ishikawa T. Cloning and sequencing of the xynA gene encoding xylanase A of Aspergillus kawachii. Biosci Biotechnol Biochem. 1992 Jun;56(6):906–912. doi: 10.1271/bbb.56.906. [DOI] [PubMed] [Google Scholar]
- Kellett L. E., Poole D. M., Ferreira L. M., Durrant A. J., Hazlewood G. P., Gilbert H. J. Xylanase B and an arabinofuranosidase from Pseudomonas fluorescens subsp. cellulosa contain identical cellulose-binding domains and are encoded by adjacent genes. Biochem J. 1990 Dec 1;272(2):369–376. doi: 10.1042/bj2720369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khasin A., Alchanati I., Shoham Y. Purification and characterization of a thermostable xylanase from Bacillus stearothermophilus T-6. Appl Environ Microbiol. 1993 Jun;59(6):1725–1730. doi: 10.1128/aem.59.6.1725-1730.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Lee S. F., Forsberg C. W., Gibbins L. N. Xylanolytic Activity of Clostridium acetobutylicum. Appl Environ Microbiol. 1985 Oct;50(4):1068–1076. doi: 10.1128/aem.50.4.1068-1076.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin L. L., Thomson J. A. Cloning, sequencing and expression of a gene encoding a 73 kDa xylanase enzyme from the rumen anaerobe Butyrivibrio fibrisolvens H17c. Mol Gen Genet. 1991 Aug;228(1-2):55–61. doi: 10.1007/BF00282447. [DOI] [PubMed] [Google Scholar]
- Lüthi E., Love D. R., McAnulty J., Wallace C., Caughey P. A., Saul D., Bergquist P. L. Cloning, sequence analysis, and expression of genes encoding xylan-degrading enzymes from the thermophile "Caldocellum saccharolyticum". Appl Environ Microbiol. 1990 Apr;56(4):1017–1024. doi: 10.1128/aem.56.4.1017-1024.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mannarelli B. M., Evans S., Lee D. Cloning, sequencing, and expression of a xylanase gene from the anaerobic ruminal bacterium Butyrivibrio fibrisolvens. J Bacteriol. 1990 Aug;172(8):4247–4254. doi: 10.1128/jb.172.8.4247-4254.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Neill G., Goh S. H., Warren R. A., Kilburn D. G., Miller R. C., Jr Structure of the gene encoding the exoglucanase of Cellulomonas fimi. Gene. 1986;44(2-3):325–330. doi: 10.1016/0378-1119(86)90197-6. [DOI] [PubMed] [Google Scholar]
- Sakka K., Yoshikawa K., Kojima Y., Karita S., Ohmiya K., Shimada K. Nucleotide sequence of the Clostridium stercorarium xylA gene encoding a bifunctional protein with beta-D-xylosidase and alpha-L-arabinofuranosidase activities, and properties of the translated product. Biosci Biotechnol Biochem. 1993 Feb;57(2):268–272. [PubMed] [Google Scholar]
- Sancar A., Hack A. M., Rupp W. D. Simple method for identification of plasmid-coded proteins. J Bacteriol. 1979 Jan;137(1):692–693. doi: 10.1128/jb.137.1.692-693.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saul D. J., Williams L. C., Grayling R. A., Chamley L. W., Love D. R., Bergquist P. L. celB, a gene coding for a bifunctional cellulase from the extreme thermophile "Caldocellum saccharolyticum". Appl Environ Microbiol. 1990 Oct;56(10):3117–3124. doi: 10.1128/aem.56.10.3117-3124.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saul D. J., Williams L. C., Love D. R., Chamley L. W., Bergquist P. L. Nucleotide sequence of a gene from Caldocellum saccharolyticum encoding for exocellulase and endocellulase activity. Nucleic Acids Res. 1989 Jan 11;17(1):439–439. doi: 10.1093/nar/17.1.439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shao W., Wiegel J. Purification and characterization of a thermostable beta-xylosidase from Thermoanaerobacter ethanolicus. J Bacteriol. 1992 Sep;174(18):5848–5853. doi: 10.1128/jb.174.18.5848-5853.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shareck F., Roy C., Yaguchi M., Morosoli R., Kluepfel D. Sequences of three genes specifying xylanases in Streptomyces lividans. Gene. 1991 Oct 30;107(1):75–82. doi: 10.1016/0378-1119(91)90299-q. [DOI] [PubMed] [Google Scholar]
- Srinivasa B. R., Swaminathan K. R., Ganapathy C., Roy R. P., Murthy S. K., Vithayathil P. J. The primary structure of xylanase from Thermoascus aurantiacus. Protein Seq Data Anal. 1991 Jul;4(1):15–20. [PubMed] [Google Scholar]
- Tull D., Withers S. G., Gilkes N. R., Kilburn D. G., Warren R. A., Aebersold R. Glutamic acid 274 is the nucleophile in the active site of a "retaining" exoglucanase from Cellulomonas fimi. J Biol Chem. 1991 Aug 25;266(24):15621–15625. [PubMed] [Google Scholar]
- Utt E. A., Eddy C. K., Keshav K. F., Ingram L. O. Sequencing and expression of the Butyrivibrio fibrisolvens xylB gene encoding a novel bifunctional protein with beta-D-xylosidase and alpha-L-arabinofuranosidase activities. Appl Environ Microbiol. 1991 Apr;57(4):1227–1234. doi: 10.1128/aem.57.4.1227-1234.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitehead T. R., Hespell R. B. The genes for three xylan-degrading activities from Bacteroides ovatus are clustered in a 3.8-kilobase region. J Bacteriol. 1990 May;172(5):2408–2412. doi: 10.1128/jb.172.5.2408-2412.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu W. Z., Shima Y., Negoro S., Urabe I. Sequence and properties of beta-xylosidase from Bacillus pumilus IPO. Contradiction of the previous nucleotide sequence. Eur J Biochem. 1991 Dec 18;202(3):1197–1203. doi: 10.1111/j.1432-1033.1991.tb16490.x. [DOI] [PubMed] [Google Scholar]
- Zhang J. X., Flint H. J. A bifunctional xylanase encoded by the xynA gene of the rumen cellulolytic bacterium Ruminococcus flavefaciens 17 comprises two dissimilar domains linked by an asparagine/glutamine-rich sequence. Mol Microbiol. 1992 Apr;6(8):1013–1023. doi: 10.1111/j.1365-2958.1992.tb02167.x. [DOI] [PubMed] [Google Scholar]



