Abstract
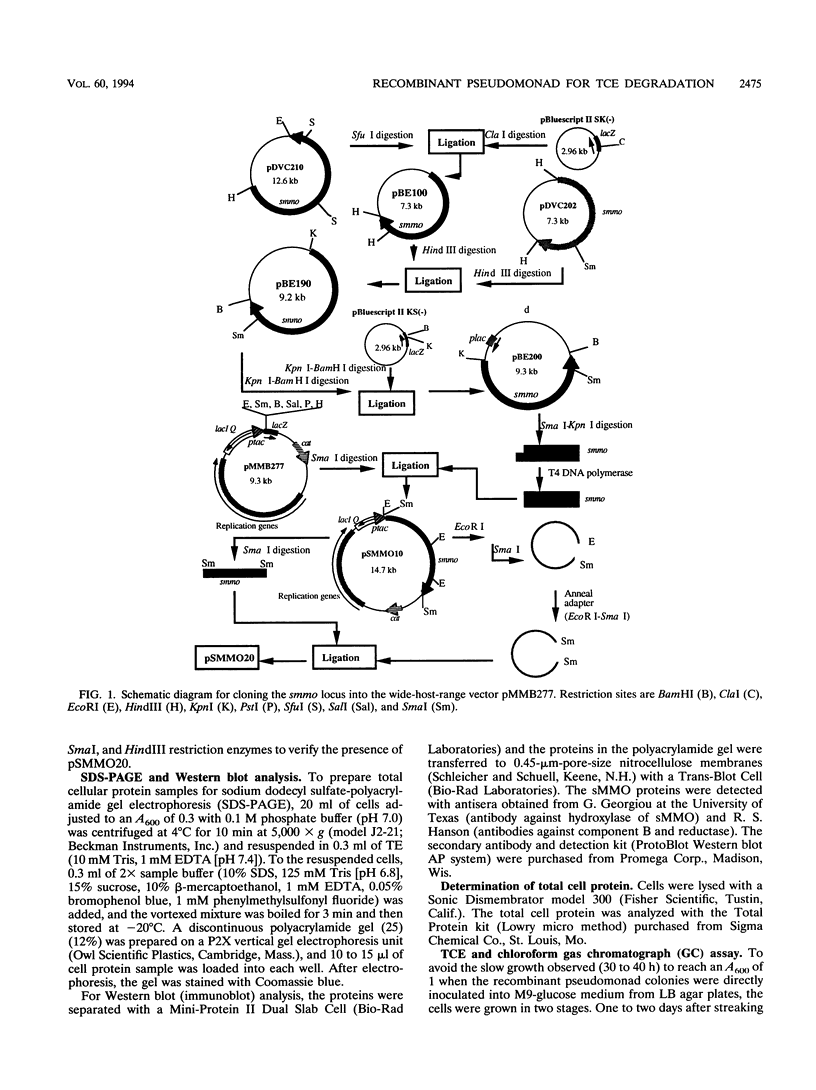
Soluble methane monooxygenase (sMMO) from Methylosinus trichosporium OB3b can degrade many halogenated aliphatic compounds that are found in contaminated soil and groundwater. This enzyme oxidizes the most frequently detected pollutant, trichloroethylene (TCE), at least 50 times faster than other enzymes. However, slow growth of the strain, strong competition between TCE and methane for sMMO, and repression of the smmo locus by low concentrations of copper ions limit the use of this bacterium. To overcome these obstacles, the 5.5-kb smmo locus of M. trichosporium OB3b was cloned into a wide-host-range vector (to form pSMMO20), and this plasmid was electroporated into five Pseudomonas strains. The best TCE degradation results were obtained with Pseudomonas putida F1/pSMMO20. The plasmid was maintained stably, and all five of the sMMO proteins (alpha, beta, and gamma hydroxylase proteins, reductase, and component B) were observed clearly by both sodium dodecyl sulfate-polyacrylamide gel electrophoresis and Western immunoblotting. TCE degradation rates were quantified for P. putida F1/pSMMO20 with a gas chromatograph (Vmax = 5 nmol per min per mg of protein), and the recombinant strain mineralized 55% of the TCE (10 microM) as indicated by measuring chloride ion concentrations with a chloride ion-specific electrode. The maximum TCE degradation rate obtained with the recombinant strain was lower than that of M. trichosporium OB3b but greater than other TCE-degrading recombinants and most well-studied pseudomonads. In addition, this recombinant strain mineralizes chloroform (a specific substrate for sMMO), grows much faster than M. trichosporium OB3b, and degrades TCE without competitive inhibition from the growth substrate.
Full text
PDF




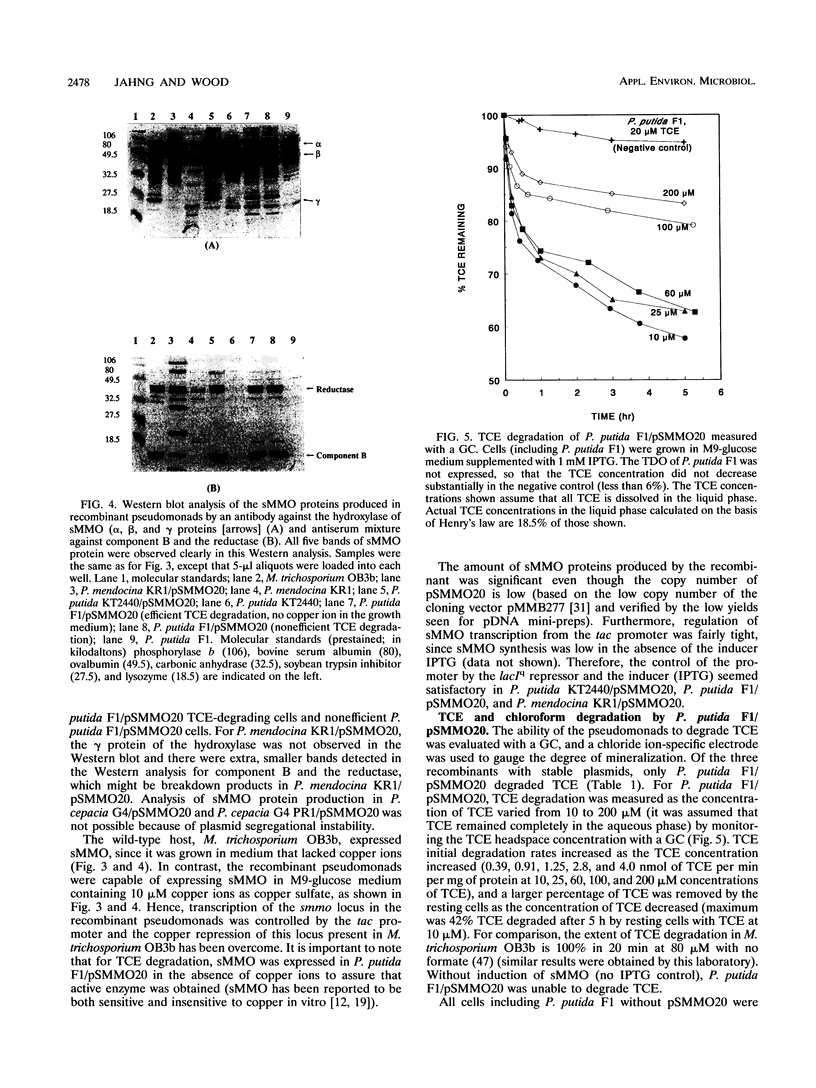
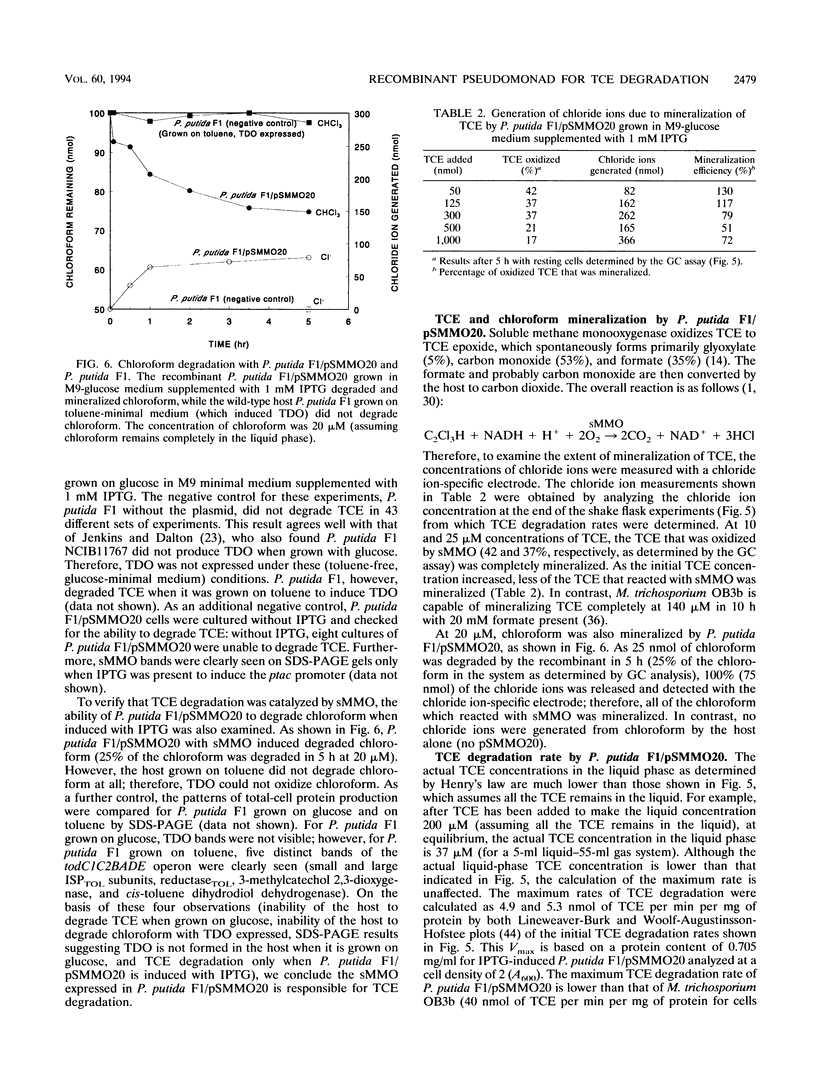



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alvarez-Cohen L., McCarty P. L., Boulygina E., Hanson R. S., Brusseau G. A., Tsien H. C. Characterization of a methane-utilizing bacterium from a bacterial consortium that rapidly degrades trichloroethylene and chloroform. Appl Environ Microbiol. 1992 Jun;58(6):1886–1893. doi: 10.1128/aem.58.6.1886-1893.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alvarez-Cohen L., McCarty P. L. Product toxicity and cometabolic competitive inhibition modeling of chloroform and trichloroethylene transformation by methanotrophic resting cells. Appl Environ Microbiol. 1991 Apr;57(4):1031–1037. doi: 10.1128/aem.57.4.1031-1037.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arciero D., Vannelli T., Logan M., Hooper A. B. Degradation of trichloroethylene by the ammonia-oxidizing bacterium Nitrosomonas europaea. Biochem Biophys Res Commun. 1989 Mar 15;159(2):640–643. doi: 10.1016/0006-291x(89)90042-9. [DOI] [PubMed] [Google Scholar]
- Brusseau G. A., Tsien H. C., Hanson R. S., Wackett L. P. Optimization of trichloroethylene oxidation by methanotrophs and the use of a colorimetric assay to detect soluble methane monooxygenase activity. Biodegradation. 1990;1(1):19–29. doi: 10.1007/BF00117048. [DOI] [PubMed] [Google Scholar]
- Cardy D. L., Laidler V., Salmond G. P., Murrell J. C. Molecular analysis of the methane monooxygenase (MMO) gene cluster of Methylosinus trichosporium OB3b. Mol Microbiol. 1991 Feb;5(2):335–342. doi: 10.1111/j.1365-2958.1991.tb02114.x. [DOI] [PubMed] [Google Scholar]
- Cardy D. L., Laidler V., Salmond G. P., Murrell J. C. The methane monooxygenase gene cluster of Methylosinus trichosporium: cloning and sequencing of the mmoC gene. Arch Microbiol. 1991;156(6):477–483. doi: 10.1007/BF00245395. [DOI] [PubMed] [Google Scholar]
- Chen L. Y., Leu W. M., Wang K. T., Lee Y. H. Copper transfer and activation of the Streptomyces apotyrosinase are mediated through a complex formation between apotyrosinase and its trans-activator MelC1. J Biol Chem. 1992 Oct 5;267(28):20100–20107. [PubMed] [Google Scholar]
- Ensley B. D. Biochemical diversity of trichloroethylene metabolism. Annu Rev Microbiol. 1991;45:283–299. doi: 10.1146/annurev.mi.45.100191.001435. [DOI] [PubMed] [Google Scholar]
- Fitch M. W., Graham D. W., Arnold R. G., Agarwal S. K., Phelps P., Speitel G. E., Jr, Georgiou G. Phenotypic characterization of copper-resistant mutants of Methylosinus trichosporium OB3b. Appl Environ Microbiol. 1993 Sep;59(9):2771–2776. doi: 10.1128/aem.59.9.2771-2776.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Folsom B. R., Chapman P. J., Pritchard P. H. Phenol and trichloroethylene degradation by Pseudomonas cepacia G4: kinetics and interactions between substrates. Appl Environ Microbiol. 1990 May;56(5):1279–1285. doi: 10.1128/aem.56.5.1279-1285.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox B. G., Borneman J. G., Wackett L. P., Lipscomb J. D. Haloalkene oxidation by the soluble methane monooxygenase from Methylosinus trichosporium OB3b: mechanistic and environmental implications. Biochemistry. 1990 Jul 10;29(27):6419–6427. doi: 10.1021/bi00479a013. [DOI] [PubMed] [Google Scholar]
- Fox B. G., Froland W. A., Dege J. E., Lipscomb J. D. Methane monooxygenase from Methylosinus trichosporium OB3b. Purification and properties of a three-component system with high specific activity from a type II methanotroph. J Biol Chem. 1989 Jun 15;264(17):10023–10033. [PubMed] [Google Scholar]
- Fox B. G., Liu Y., Dege J. E., Lipscomb J. D. Complex formation between the protein components of methane monooxygenase from Methylosinus trichosporium OB3b. Identification of sites of component interaction. J Biol Chem. 1991 Jan 5;266(1):540–550. [PubMed] [Google Scholar]
- Fox B. G., Surerus K. K., Münck E., Lipscomb J. D. Evidence for a mu-oxo-bridged binuclear iron cluster in the hydroxylase component of methane monooxygenase. Mössbauer and EPR studies. J Biol Chem. 1988 Aug 5;263(22):10553–10556. [PubMed] [Google Scholar]
- Green J., Dalton H. Substrate specificity of soluble methane monooxygenase. Mechanistic implications. J Biol Chem. 1989 Oct 25;264(30):17698–17703. [PubMed] [Google Scholar]
- Green J., Prior S. D., Dalton H. Copper ions as inhibitors of protein C of soluble methane monooxygenase of Methylococcus capsulatus (Bath). Eur J Biochem. 1985 Nov 15;153(1):137–144. doi: 10.1111/j.1432-1033.1985.tb09279.x. [DOI] [PubMed] [Google Scholar]
- Hanson R. S., Tsien H. C., Tsuji K., Brusseau G. A., Wackett L. P. Biodegradation of low-molecular-weight halogenated hydrocarbons by methanotrophic bacteria. FEMS Microbiol Rev. 1990 Dec;7(3-4):273–278. doi: 10.1111/j.1574-6968.1990.tb04924.x. [DOI] [PubMed] [Google Scholar]
- Koh S. C., Bowman J. P., Sayler G. S. Soluble Methane Monooxygenase Production and Trichloroethylene Degradation by a Type I Methanotroph, Methylomonas methanica 68-1. Appl Environ Microbiol. 1993 Apr;59(4):960–967. doi: 10.1128/aem.59.4.960-967.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lee S. K., Nesheim J. C., Lipscomb J. D. Transient intermediates of the methane monooxygenase catalytic cycle. J Biol Chem. 1993 Oct 15;268(29):21569–21577. [PubMed] [Google Scholar]
- Lee Y. H., Chen B. F., Wu S. Y., Leu W. M., Lin J. J., Chen C. W., Lo S. C. A trans-acting gene is required for the phenotypic expression of a tyrosinase gene in Streptomyces. Gene. 1988 May 15;65(1):71–81. doi: 10.1016/0378-1119(88)90418-0. [DOI] [PubMed] [Google Scholar]
- Little C. D., Palumbo A. V., Herbes S. E., Lidstrom M. E., Tyndall R. L., Gilmer P. J. Trichloroethylene biodegradation by a methane-oxidizing bacterium. Appl Environ Microbiol. 1988 Apr;54(4):951–956. doi: 10.1128/aem.54.4.951-956.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morales V. M., Bäckman A., Bagdasarian M. A series of wide-host-range low-copy-number vectors that allow direct screening for recombinants. Gene. 1991 Jan 2;97(1):39–47. doi: 10.1016/0378-1119(91)90007-x. [DOI] [PubMed] [Google Scholar]
- Nelson M. J., Montgomery S. O., Mahaffey W. R., Pritchard P. H. Biodegradation of trichloroethylene and involvement of an aromatic biodegradative pathway. Appl Environ Microbiol. 1987 May;53(5):949–954. doi: 10.1128/aem.53.5.949-954.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson M. J., Montgomery S. O., Pritchard P. H. Trichloroethylene metabolism by microorganisms that degrade aromatic compounds. Appl Environ Microbiol. 1988 Feb;54(2):604–606. doi: 10.1128/aem.54.2.604-606.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newman L. M., Wackett L. P. Fate of 2,2,2-trichloroacetaldehyde (chloral hydrate) produced during trichloroethylene oxidation by methanotrophs. Appl Environ Microbiol. 1991 Aug;57(8):2399–2402. doi: 10.1128/aem.57.8.2399-2402.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oldenhuis R., Oedzes J. Y., van der Waarde J. J., Janssen D. B. Kinetics of chlorinated hydrocarbon degradation by Methylosinus trichosporium OB3b and toxicity of trichloroethylene. Appl Environ Microbiol. 1991 Jan;57(1):7–14. doi: 10.1128/aem.57.1.7-14.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oldenhuis R., Vink R. L., Janssen D. B., Witholt B. Degradation of chlorinated aliphatic hydrocarbons by Methylosinus trichosporium OB3b expressing soluble methane monooxygenase. Appl Environ Microbiol. 1989 Nov;55(11):2819–2826. doi: 10.1128/aem.55.11.2819-2826.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phelps P. A., Agarwal S. K., Speitel G. E., Georgiou G. Methylosinus trichosporium OB3b Mutants Having Constitutive Expression of Soluble Methane Monooxygenase in the Presence of High Levels of Copper. Appl Environ Microbiol. 1992 Nov;58(11):3701–3708. doi: 10.1128/aem.58.11.3701-3708.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothmel R. K., Chakrabarty A. M., Berry A., Darzins A. Genetic systems in Pseudomonas. Methods Enzymol. 1991;204:485–514. doi: 10.1016/0076-6879(91)04025-j. [DOI] [PubMed] [Google Scholar]
- Schwartz R. D., McCoy C. J. Pseudomonas oleovorans hydroxylation-epoxidation system: additional strain improvements. Appl Microbiol. 1973 Aug;26(2):217–218. doi: 10.1128/am.26.2.217-218.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shields M. S., Reagin M. J. Selection of a Pseudomonas cepacia strain constitutive for the degradation of trichloroethylene. Appl Environ Microbiol. 1992 Dec;58(12):3977–3983. doi: 10.1128/aem.58.12.3977-3983.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith A. W., Iglewski B. H. Transformation of Pseudomonas aeruginosa by electroporation. Nucleic Acids Res. 1989 Dec 25;17(24):10509–10509. doi: 10.1093/nar/17.24.10509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsien H. C., Brusseau G. A., Hanson R. S., Waclett L. P. Biodegradation of trichloroethylene by Methylosinus trichosporium OB3b. Appl Environ Microbiol. 1989 Dec;55(12):3155–3161. doi: 10.1128/aem.55.12.3155-3161.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsien H. C., Hanson R. S. Soluble methane monooxygenase component B gene probe for identification of methanotrophs that rapidly degrade trichloroethylene. Appl Environ Microbiol. 1992 Mar;58(3):953–960. doi: 10.1128/aem.58.3.953-960.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wackett L. P., Brusseau G. A., Householder S. R., Hanson R. S. Survey of microbial oxygenases: trichloroethylene degradation by propane-oxidizing bacteria. Appl Environ Microbiol. 1989 Nov;55(11):2960–2964. doi: 10.1128/aem.55.11.2960-2964.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wackett L. P., Gibson D. T. Degradation of trichloroethylene by toluene dioxygenase in whole-cell studies with Pseudomonas putida F1. Appl Environ Microbiol. 1988 Jul;54(7):1703–1708. doi: 10.1128/aem.54.7.1703-1708.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West C. A., Salmond G. P., Dalton H., Murrell J. C. Functional expression in Escherichia coli of proteins B and C from soluble methane monooxygenase of Methylococcus capsulatus (Bath). J Gen Microbiol. 1992 Jul;138(7):1301–1307. doi: 10.1099/00221287-138-7-1301. [DOI] [PubMed] [Google Scholar]
- Yen K. M., Karl M. R., Blatt L. M., Simon M. J., Winter R. B., Fausset P. R., Lu H. S., Harcourt A. A., Chen K. K. Cloning and characterization of a Pseudomonas mendocina KR1 gene cluster encoding toluene-4-monooxygenase. J Bacteriol. 1991 Sep;173(17):5315–5327. doi: 10.1128/jb.173.17.5315-5327.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zylstra G. J., Wackett L. P., Gibson D. T. Trichloroethylene degradation by Escherichia coli containing the cloned Pseudomonas putida F1 toluene dioxygenase genes. Appl Environ Microbiol. 1989 Dec;55(12):3162–3166. doi: 10.1128/aem.55.12.3162-3166.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]