Abstract
We report the amplification of bacterial genes from uninoculated surface and subsurface sediments by the polymerase chain reaction (PCR). PCR amplification of indigenous bacterial 16S ribosomal DNA genes was unsuccessful when subsurface sediment containing approximately 10(7) cells.g-1 was added directly to a PCR mixture. However, when 10 mg of sediment was inoculated with approximately 10(5) cells of Pseudomonas putida G7, the nahAc naphthalene dioxygenase gene characteristic of the P. putida G7 NAH7 plasmid was detected by PCR amplification. Southern blotting of the PCR amplification product improved sensitivity to 10(3) to 10(4) cells from samples inoculated with P. putida G7, but controls with no sediment added showed that the PCR was partially inhibited by the sediments. Lysozyme-sodium dodecyl sulfate-freeze-thaw DNA extraction was combined with gel electrophoretic partial purification in the presence of polyvinylpyrrolidone to render DNA from indigenous bacteria in surface or subsurface sediment samples amplifiable by PCR using eubacterial 16S ribosomal DNA primers. The nahAc gene could also be amplified from indigenous bacteria by using nahAc-specific primers when PCR conditions were modified by increasing Taq and primer concentrations. Restriction digests of the nahAc amplification products from surface and subsurface sediments revealed polymorphism relative to P. putida G7. The procedures for DNA extraction, purification, and PCR amplification described here demonstrate that the PCR is a potentially useful tool in studies of function- and taxon-specific DNA from indigenous microbial communities in sediment and groundwater environments.
Full text
PDF







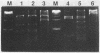
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Amann R., Springer N., Ludwig W., Görtz H. D., Schleifer K. H. Identification in situ and phylogeny of uncultured bacterial endosymbionts. Nature. 1991 May 9;351(6322):161–164. doi: 10.1038/351161a0. [DOI] [PubMed] [Google Scholar]
- Balkwill D. L., Ghiorse W. C. Characterization of subsurface bacteria associated with two shallow aquifers in oklahoma. Appl Environ Microbiol. 1985 Sep;50(3):580–588. doi: 10.1128/aem.50.3.580-588.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barkay T., Gillman M., Liebert C. Genes encoding mercuric reductases from selected gram-negative aquatic bacteria have a low degree of homology with merA of transposon Tn501. Appl Environ Microbiol. 1990 Jun;56(6):1695–1701. doi: 10.1128/aem.56.6.1695-1701.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giovannoni S. J., Britschgi T. B., Moyer C. L., Field K. G. Genetic diversity in Sargasso Sea bacterioplankton. Nature. 1990 May 3;345(6270):60–63. doi: 10.1038/345060a0. [DOI] [PubMed] [Google Scholar]
- Harshman L., Riley M. Conservation and variation of nucleotide sequences in Escherichia coli strains isolated from nature. J Bacteriol. 1980 Nov;144(2):560–568. doi: 10.1128/jb.144.2.560-568.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holben W. E., Schroeter B. M., Calabrese V. G., Olsen R. H., Kukor J. K., Biederbeck V. O., Smith A. E., Tiedje J. M. Gene probe analysis of soil microbial populations selected by amendment with 2,4-dichlorophenoxyacetic acid. Appl Environ Microbiol. 1992 Dec;58(12):3941–3948. doi: 10.1128/aem.58.12.3941-3948.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurkela S., Lehväslaiho H., Palva E. T., Teeri T. H. Cloning, nucleotide sequence and characterization of genes encoding naphthalene dioxygenase of Pseudomonas putida strain NCIB9816. Gene. 1988 Dec 20;73(2):355–362. doi: 10.1016/0378-1119(88)90500-8. [DOI] [PubMed] [Google Scholar]
- Lanzillo J. J. Preparation of digoxigenin-labeled probes by the polymerase chain reaction. Biotechniques. 1990 Jun;8(6):620–622. [PubMed] [Google Scholar]
- Liesack W., Stackebrandt E. Occurrence of novel groups of the domain Bacteria as revealed by analysis of genetic material isolated from an Australian terrestrial environment. J Bacteriol. 1992 Aug;174(15):5072–5078. doi: 10.1128/jb.174.15.5072-5078.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madsen E. L., Sinclair J. L., Ghiorse W. C. In situ biodegradation: microbiological patterns in a contaminated aquifer. Science. 1991 May 10;252(5007):830–833. doi: 10.1126/science.2028258. [DOI] [PubMed] [Google Scholar]
- Nelson K., Whittam T. S., Selander R. K. Nucleotide polymorphism and evolution in the glyceraldehyde-3-phosphate dehydrogenase gene (gapA) in natural populations of Salmonella and Escherichia coli. Proc Natl Acad Sci U S A. 1991 Aug 1;88(15):6667–6671. doi: 10.1073/pnas.88.15.6667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sayler G. S., Layton A. C. Environmental application of nucleic acid hybridization. Annu Rev Microbiol. 1990;44:625–648. doi: 10.1146/annurev.mi.44.100190.003205. [DOI] [PubMed] [Google Scholar]
- Schmidt T. M., DeLong E. F., Pace N. R. Analysis of a marine picoplankton community by 16S rRNA gene cloning and sequencing. J Bacteriol. 1991 Jul;173(14):4371–4378. doi: 10.1128/jb.173.14.4371-4378.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stahl D. A., Lane D. J., Olsen G. J., Pace N. R. Characterization of a Yellowstone hot spring microbial community by 5S rRNA sequences. Appl Environ Microbiol. 1985 Jun;49(6):1379–1384. doi: 10.1128/aem.49.6.1379-1384.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steffan R. J., Atlas R. M. DNA amplification to enhance detection of genetically engineered bacteria in environmental samples. Appl Environ Microbiol. 1988 Sep;54(9):2185–2191. doi: 10.1128/aem.54.9.2185-2191.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steffan R. J., Goksøyr J., Bej A. K., Atlas R. M. Recovery of DNA from soils and sediments. Appl Environ Microbiol. 1988 Dec;54(12):2908–2915. doi: 10.1128/aem.54.12.2908-2915.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsai Y. L., Olson B. H. Rapid method for direct extraction of DNA from soil and sediments. Appl Environ Microbiol. 1991 Apr;57(4):1070–1074. doi: 10.1128/aem.57.4.1070-1074.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsai Y. L., Olson B. H. Rapid method for separation of bacterial DNA from humic substances in sediments for polymerase chain reaction. Appl Environ Microbiol. 1992 Jul;58(7):2292–2295. doi: 10.1128/aem.58.7.2292-2295.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward D. M., Weller R., Bateson M. M. 16S rRNA sequences reveal numerous uncultured microorganisms in a natural community. Nature. 1990 May 3;345(6270):63–65. doi: 10.1038/345063a0. [DOI] [PubMed] [Google Scholar]
- Weisburg W. G., Barns S. M., Pelletier D. A., Lane D. J. 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol. 1991 Jan;173(2):697–703. doi: 10.1128/jb.173.2.697-703.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weller R., Weller J. W., Ward D. M. 16S rRNA sequences of uncultivated hot spring cyanobacterial mat inhabitants retrieved as randomly primed cDNA. Appl Environ Microbiol. 1991 Apr;57(4):1146–1151. doi: 10.1128/aem.57.4.1146-1151.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson K. H., Blitchington R. B., Greene R. C. Amplification of bacterial 16S ribosomal DNA with polymerase chain reaction. J Clin Microbiol. 1990 Sep;28(9):1942–1946. doi: 10.1128/jcm.28.9.1942-1946.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R. Bacterial evolution. Microbiol Rev. 1987 Jun;51(2):221–271. doi: 10.1128/mr.51.2.221-271.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yen K. M., Gunsalus I. C. Plasmid gene organization: naphthalene/salicylate oxidation. Proc Natl Acad Sci U S A. 1982 Feb;79(3):874–878. doi: 10.1073/pnas.79.3.874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zylstra G. J., Gibson D. T. Toluene degradation by Pseudomonas putida F1. Nucleotide sequence of the todC1C2BADE genes and their expression in Escherichia coli. J Biol Chem. 1989 Sep 5;264(25):14940–14946. [PubMed] [Google Scholar]