Abstract
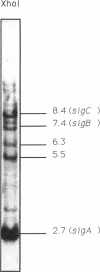
A third putative sigma factor gene, sigC, has been isolated from Myxococcus xanthus by using the sigA gene (formerly rpoD of M. xanthus) as a probe. The nucleotide sequence of sigC has been determined, and an open reading frame of 295 residues (M(r) = 33,430) has been identified. The deduced amino acid sequence of sigC exhibits the features which are characteristic of other bacterial sigma factors. The characterization of a sigC-lacZ strain has demonstrated that sigC expression is induced immediately after cells enter into the developmental cycle and is dramatically reduced at the onset of sporulation. A deletion mutant of sigC grows normally in vegetative culture and is able to develop normally. However, in contrast to the wild-type cells, the sigC deletion mutant cells became capable of forming fruiting bodies and myxospores on semirich agar plates. This suggests that sigC may play a role in expression of genes involved in negatively regulating the initiation of fruiting body formation.
Full text
PDF



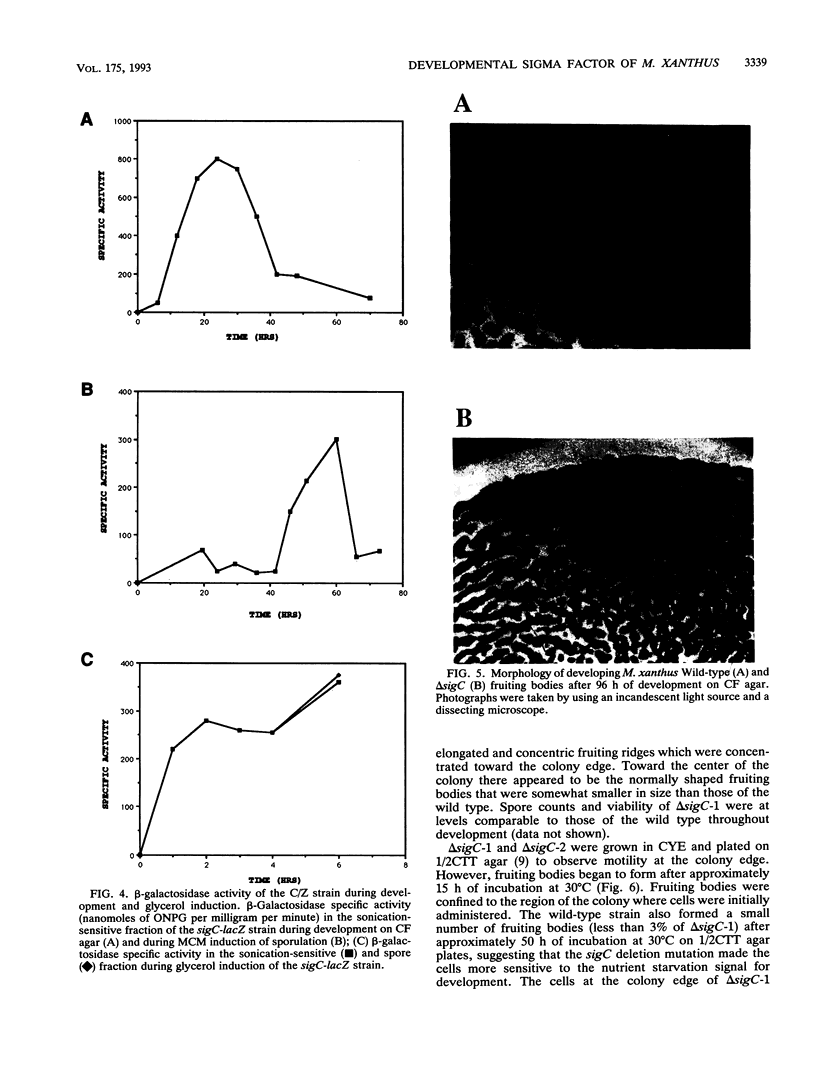



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Apelian D., Inouye S. Development-specific sigma-factor essential for late-stage differentiation of Myxococcus xanthus. Genes Dev. 1990 Aug;4(8):1396–1403. doi: 10.1101/gad.4.8.1396. [DOI] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Campos J. M., Geisselsoder J., Zusman D. R. Isolation of bacteriophage MX4, a generalized transducing phage for Myxococcus xanthus. J Mol Biol. 1978 Feb 25;119(2):167–178. doi: 10.1016/0022-2836(78)90431-x. [DOI] [PubMed] [Google Scholar]
- Clare J. J., Oliver S. G. The regulation of RNA synthesis in yeast IV. Synthesis of double-stranded RNA. Mol Gen Genet. 1979 Mar 20;171(2):161–166. doi: 10.1007/BF00270002. [DOI] [PubMed] [Google Scholar]
- DWORKIN M., GIBSON S. M. A SYSTEM FOR STUDYING MICROBIAL MORPHOGENESIS: RAPID FORMATION OF MICROCYSTS IN MYXOCOCCUS XANTHUS. Science. 1964 Oct 9;146(3641):243–244. doi: 10.1126/science.146.3641.243. [DOI] [PubMed] [Google Scholar]
- Gardella T., Moyle H., Susskind M. M. A mutant Escherichia coli sigma 70 subunit of RNA polymerase with altered promoter specificity. J Mol Biol. 1989 Apr 20;206(4):579–590. doi: 10.1016/0022-2836(89)90567-6. [DOI] [PubMed] [Google Scholar]
- Hagen D. C., Bretscher A. P., Kaiser D. Synergism between morphogenetic mutants of Myxococcus xanthus. Dev Biol. 1978 Jun;64(2):284–296. doi: 10.1016/0012-1606(78)90079-9. [DOI] [PubMed] [Google Scholar]
- Helmann J. D., Chamberlin M. J. Structure and function of bacterial sigma factors. Annu Rev Biochem. 1988;57:839–872. doi: 10.1146/annurev.bi.57.070188.004203. [DOI] [PubMed] [Google Scholar]
- Hengge-Aronis R., Klein W., Lange R., Rimmele M., Boos W. Trehalose synthesis genes are controlled by the putative sigma factor encoded by rpoS and are involved in stationary-phase thermotolerance in Escherichia coli. J Bacteriol. 1991 Dec;173(24):7918–7924. doi: 10.1128/jb.173.24.7918-7924.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu M. Y., Inouye S., Inouye M. Structural requirements of the RNA precursor for the biosynthesis of the branched RNA-linked multicopy single-stranded DNA of Myxococcus xanthus. J Biol Chem. 1989 Apr 15;264(11):6214–6219. [PubMed] [Google Scholar]
- Inouye M., Inouye S., Zusman D. R. Gene expression during development of Myxococcus xanthus: pattern of protein synthesis. Dev Biol. 1979 Feb;68(2):579–591. doi: 10.1016/0012-1606(79)90228-8. [DOI] [PubMed] [Google Scholar]
- Inouye S. Cloning and DNA sequence of the gene coding for the major sigma factor from Myxococcus xanthus. J Bacteriol. 1990 Jan;172(1):80–85. doi: 10.1128/jb.172.1.80-85.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye S., Hsu M. Y., Eagle S., Inouye M. Reverse transcriptase associated with the biosynthesis of the branched RNA-linked msDNA in Myxococcus xanthus. Cell. 1989 Feb 24;56(4):709–717. doi: 10.1016/0092-8674(89)90593-x. [DOI] [PubMed] [Google Scholar]
- Johnson J. L., Ordal E. J. Deoxyribonucleic acid homology in bacterial taxonomy: effect of incubation temperature on reaction specificity. J Bacteriol. 1968 Mar;95(3):893–900. doi: 10.1128/jb.95.3.893-900.1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaiser D. Control of multicellular development: Dictyostelium and Myxococcus. Annu Rev Genet. 1986;20:539–566. doi: 10.1146/annurev.ge.20.120186.002543. [DOI] [PubMed] [Google Scholar]
- Kroos L., Kuspa A., Kaiser D. A global analysis of developmentally regulated genes in Myxococcus xanthus. Dev Biol. 1986 Sep;117(1):252–266. doi: 10.1016/0012-1606(86)90368-4. [DOI] [PubMed] [Google Scholar]
- Lonetto M., Gribskov M., Gross C. A. The sigma 70 family: sequence conservation and evolutionary relationships. J Bacteriol. 1992 Jun;174(12):3843–3849. doi: 10.1128/jb.174.12.3843-3849.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McBride M. J., Ensign J. C. Effects of intracellular trehalose content on Streptomyces griseus spores. J Bacteriol. 1987 Nov;169(11):4995–5001. doi: 10.1128/jb.169.11.4995-5001.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McBride M. J., Zusman D. R. Trehalose accumulation in vegetative cells and spores of Myxococcus xanthus. J Bacteriol. 1989 Nov;171(11):6383–6386. doi: 10.1128/jb.171.11.6383-6386.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muñoz-Dorado J., Inouye S., Inouye M. A gene encoding a protein serine/threonine kinase is required for normal development of M. xanthus, a gram-negative bacterium. Cell. 1991 Nov 29;67(5):995–1006. doi: 10.1016/0092-8674(91)90372-6. [DOI] [PubMed] [Google Scholar]
- Nakamura K., Masui Y., Inouye M. Use of a lac promoter-operator fragment as a transcriptional control switch for expression of the constitutive lpp gene in Escherichia coli. J Mol Appl Genet. 1982;1(4):289–299. [PubMed] [Google Scholar]
- Peden K. W. Revised sequence of the tetracycline-resistance gene of pBR322. Gene. 1983 May-Jun;22(2-3):277–280. doi: 10.1016/0378-1119(83)90112-9. [DOI] [PubMed] [Google Scholar]
- Rosenbluh A., Rosenberg E. Sporulation of Myxococcus xanthus in liquid shake flask cultures. J Bacteriol. 1989 Aug;171(8):4521–4524. doi: 10.1128/jb.171.8.4521-4524.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosner J. L. Formation, induction, and curing of bacteriophage P1 lysogens. Virology. 1972 Jun;48(3):679–689. doi: 10.1016/0042-6822(72)90152-3. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimkets L. J., Gill R. E., Kaiser D. Developmental cell interactions in Myxococcus xanthus and the spoC locus. Proc Natl Acad Sci U S A. 1983 Mar;80(5):1406–1410. doi: 10.1073/pnas.80.5.1406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimkets L. J. Social and developmental biology of the myxobacteria. Microbiol Rev. 1990 Dec;54(4):473–501. doi: 10.1128/mr.54.4.473-501.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siegele D. A., Hu J. C., Walter W. A., Gross C. A. Altered promoter recognition by mutant forms of the sigma 70 subunit of Escherichia coli RNA polymerase. J Mol Biol. 1989 Apr 20;206(4):591–603. doi: 10.1016/0022-2836(89)90568-8. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Stragier P., Losick R. Cascades of sigma factors revisited. Mol Microbiol. 1990 Nov;4(11):1801–1806. doi: 10.1111/j.1365-2958.1990.tb02028.x. [DOI] [PubMed] [Google Scholar]
- Teintze M., Thomas R., Furuichi T., Inouye M., Inouye S. Two homologous genes coding for spore-specific proteins are expressed at different times during development of Myxococcus xanthus. J Bacteriol. 1985 Jul;163(1):121–125. doi: 10.1128/jb.163.1.121-125.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- Yee T., Inouye M. Reexamination of the genome size of myxobacteria, including the use of a new method for genome size analysis. J Bacteriol. 1981 Mar;145(3):1257–1265. doi: 10.1128/jb.145.3.1257-1265.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang W., Munoz-Dorado J., Inouye M., Inouye S. Identification of a putative eukaryotic-like protein kinase family in the developmental bacterium Myxococcus xanthus. J Bacteriol. 1992 Aug;174(16):5450–5453. doi: 10.1128/jb.174.16.5450-5453.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuber P., Healy J., Carter H. L., 3rd, Cutting S., Moran C. P., Jr, Losick R. Mutation changing the specificity of an RNA polymerase sigma factor. J Mol Biol. 1989 Apr 20;206(4):605–614. doi: 10.1016/0022-2836(89)90569-x. [DOI] [PubMed] [Google Scholar]