Abstract
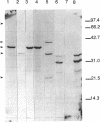
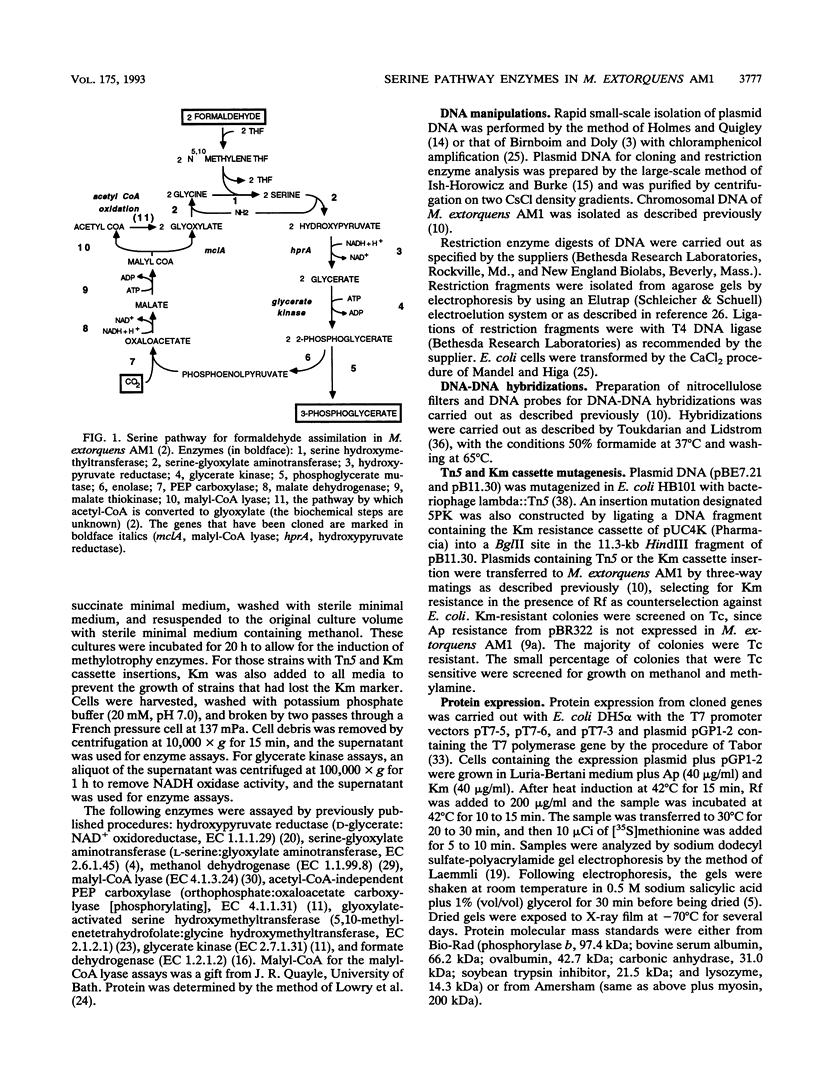
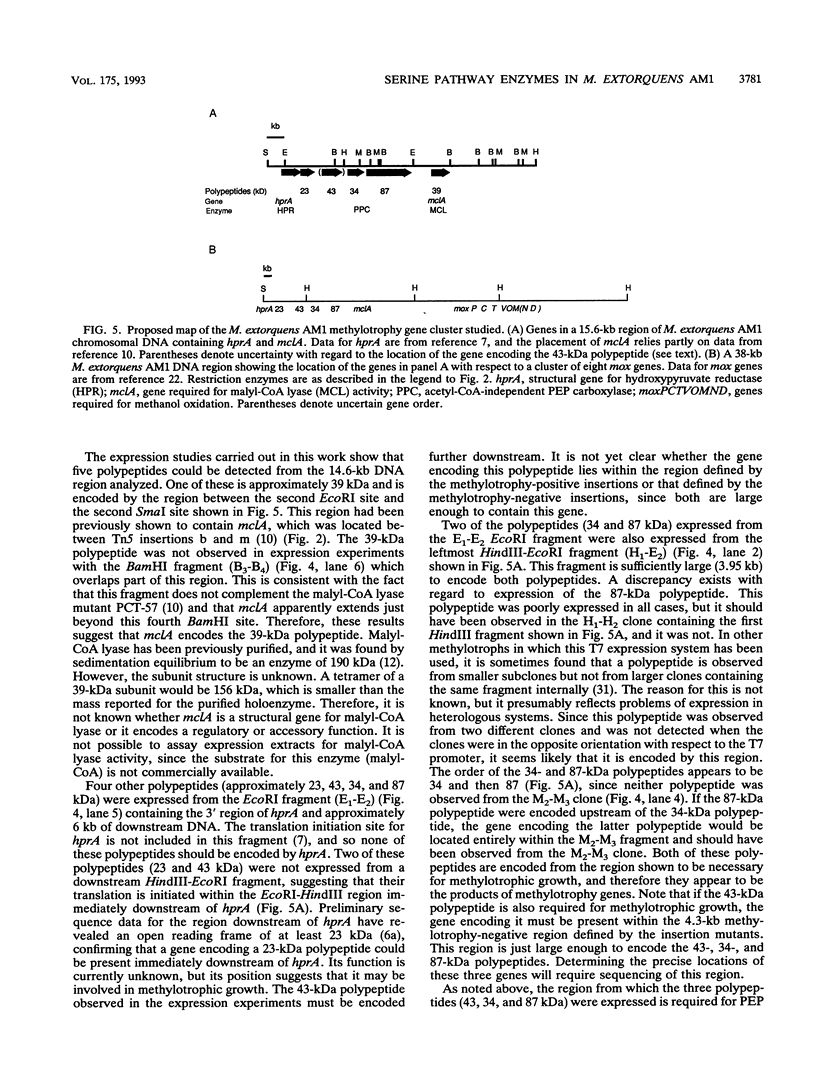
Methylobacterium extorquens AM1 is a facultative methylotrophic bacterium that uses the serine pathway for formaldehyde incorporation as its assimilation pathway during growth on one-carbon compounds. A DNA region from M. extorquens AM1 previously shown to contain genes for the serine pathway enzymes malyl coenzyme A (CoA) lyase and hydroxypyruvate reductase has been characterized in more detail. Insertion mutagenesis revealed an additional region required for growth on one-carbon compounds, and all of the insertion mutants in this region lacked activity for another serine pathway enzyme, the acetyl-CoA-independent phosphoenolpyruvate (PEP) carboxylase. Expression analysis with Escherichia coli of DNA fragments that included the malyl-CoA lyase and PEP carboxylase regions identified five polypeptides, all transcribed in the same direction. Three of these polypeptides were expressed from the region necessary for the acetyl-CoA-independent PEP carboxylase, one was expressed from the region containing the malyl-CoA lyase gene, and the fifth was expressed from a region immediately downstream from the gene encoding hydroxypyruvate reductase. All six genes are transcribed in the same direction, but the transposon insertion data suggest that they are not all cotranscribed.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anderson D. J., Lidstrom M. E. The moxFG region encodes four polypeptides in the methanol-oxidizing bacterium Methylobacterium sp. strain AM1. J Bacteriol. 1988 May;170(5):2254–2262. doi: 10.1128/jb.170.5.2254-2262.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blackmore M. A., Quayle J. R. Microbial growth on oxalate by a route not involving glyoxylate carboligase. Biochem J. 1970 Jun;118(1):53–59. doi: 10.1042/bj1180053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chamberlain J. P. Fluorographic detection of radioactivity in polyacrylamide gels with the water-soluble fluor, sodium salicylate. Anal Biochem. 1979 Sep 15;98(1):132–135. doi: 10.1016/0003-2697(79)90716-4. [DOI] [PubMed] [Google Scholar]
- Chistoserdov A. Y., Tsygankov Y. D., Lidstrom M. E. Genetic organization of methylamine utilization genes from Methylobacterium extorquens AM1. J Bacteriol. 1991 Sep;173(18):5901–5908. doi: 10.1128/jb.173.18.5901-5908.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chistoserdova L. V., Lidstrom M. E. Cloning, mutagenesis, and physiological effect of a hydroxypyruvate reductase gene from Methylobacterium extorquens AM1. J Bacteriol. 1992 Jan;174(1):71–77. doi: 10.1128/jb.174.1.71-77.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunstan P. M., Anthony C., Drabble W. T. Microbial metabolism of C 1 and C 2 compounds. The role of glyoxylate, glycollate and acetate in the growth of Pseudomonas AM1 on ethanol and on C 1 compounds. Biochem J. 1972 Jun;128(1):107–115. doi: 10.1042/bj1280107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Figurski D. H., Helinski D. R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1648–1652. doi: 10.1073/pnas.76.4.1648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fulton G. L., Nunn D. N., Lidstrom M. E. Molecular cloning of a malyl coenzyme A lyase gene from Pseudomonas sp. strain AM1, a facultative methylotroph. J Bacteriol. 1984 Nov;160(2):718–723. doi: 10.1128/jb.160.2.718-723.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hacking A. J., Quayle J. R. Purification and properties of malyl-coenzyme A lyase from Pseudomonas AM1. Biochem J. 1974 May;139(2):399–405. doi: 10.1042/bj1390399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Ish-Horowicz D., Burke J. F. Rapid and efficient cosmid cloning. Nucleic Acids Res. 1981 Jul 10;9(13):2989–2998. doi: 10.1093/nar/9.13.2989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson P. A., Quayle J. R. Microbial growth on C-1 compounds. 6. Oxidation of methanol, formaldehyde and formate by methanol-grown Pseudomonas AM-1. Biochem J. 1964 Nov;93(2):281–290. doi: 10.1042/bj0930281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jorgensen R. A., Rothstein S. J., Reznikoff W. S. A restriction enzyme cleavage map of Tn5 and location of a region encoding neomycin resistance. Mol Gen Genet. 1979;177(1):65–72. doi: 10.1007/BF00267254. [DOI] [PubMed] [Google Scholar]
- Knauf V. C., Nester E. W. Wide host range cloning vectors: a cosmid clone bank of an Agrobacterium Ti plasmid. Plasmid. 1982 Jul;8(1):45–54. doi: 10.1016/0147-619x(82)90040-3. [DOI] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Large P. J., Peel D., Quayle J. R. Microbial growth on C(1) compounds. 4. Carboxylation of phosphoenolpyruvate in methanol-grown Pseudomonas AM1. Biochem J. 1962 Oct;85(1):243–250. doi: 10.1042/bj0850243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Large P. J., Quayle J. R. Microbial growth on C(1) compounds. 5. Enzyme activities in extracts of Pseudomonas AM1. Biochem J. 1963 May;87(2):386–396. doi: 10.1042/bj0870386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mandel M., Higa A. Calcium-dependent bacteriophage DNA infection. J Mol Biol. 1970 Oct 14;53(1):159–162. doi: 10.1016/0022-2836(70)90051-3. [DOI] [PubMed] [Google Scholar]
- Nunn D. N., Lidstrom M. E. Isolation and complementation analysis of 10 methanol oxidation mutant classes and identification of the methanol dehydrogenase structural gene of Methylobacterium sp. strain AM1. J Bacteriol. 1986 May;166(2):581–590. doi: 10.1128/jb.166.2.581-590.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salem A. R., Hacking A. J., Quayle J. R. Cleavage of malyl-Coenzyme A into acetyl-Coenzyme A and glyoxylate by Pseudomonas AM1 and other C1-unit-utilizing bacteria. Biochem J. 1973 Sep;136(1):89–96. doi: 10.1042/bj1360089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stephens R. L., Haygood M. G., Lidstrom M. E. Identification of putative methanol dehydrogenase (moxF) structural genes in methylotrophs and cloning of moxF genes from Methylococcus capsulatus bath and Methylomonas albus BG8. J Bacteriol. 1988 May;170(5):2063–2069. doi: 10.1128/jb.170.5.2063-2069.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. A bacteriophage T7 RNA polymerase/promoter system for controlled exclusive expression of specific genes. Proc Natl Acad Sci U S A. 1985 Feb;82(4):1074–1078. doi: 10.1073/pnas.82.4.1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terada K., Murata T., Izui K. Site-directed mutagenesis of phosphoenolpyruvate carboxylase from E. coli: the role of His579 in the catalytic and regulatory functions. J Biochem. 1991 Jan;109(1):49–54. [PubMed] [Google Scholar]
- Toukdarian A. E., Lidstrom M. E. DNA hybridization analysis of the nif region of two methylotrophs and molecular cloning of nif-specific DNA. J Bacteriol. 1984 Mar;157(3):925–930. doi: 10.1128/jb.157.3.925-930.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuji K., Tsien H. C., Hanson R. S., DePalma S. R., Scholtz R., LaRoche S. 16S ribosomal RNA sequence analysis for determination of phylogenetic relationship among methylotrophs. J Gen Microbiol. 1990 Jan;136(1):1–10. doi: 10.1099/00221287-136-1-1. [DOI] [PubMed] [Google Scholar]
- Weaver C. A., Redborg A. H., Konisky J. Plasmid-determined immunity of Escherichia coli K-12 to colicin Ia Is mediated by a plasmid-encoded membrane protein. J Bacteriol. 1981 Dec;148(3):817–828. doi: 10.1128/jb.148.3.817-828.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto T., Yokota T. Construction of a physical map of a kanamycin (Km) transposon, Tn5, and a comparison to another Km transposon, Tn903. Mol Gen Genet. 1980 Apr;178(1):77–83. doi: 10.1007/BF00267215. [DOI] [PubMed] [Google Scholar]