Abstract
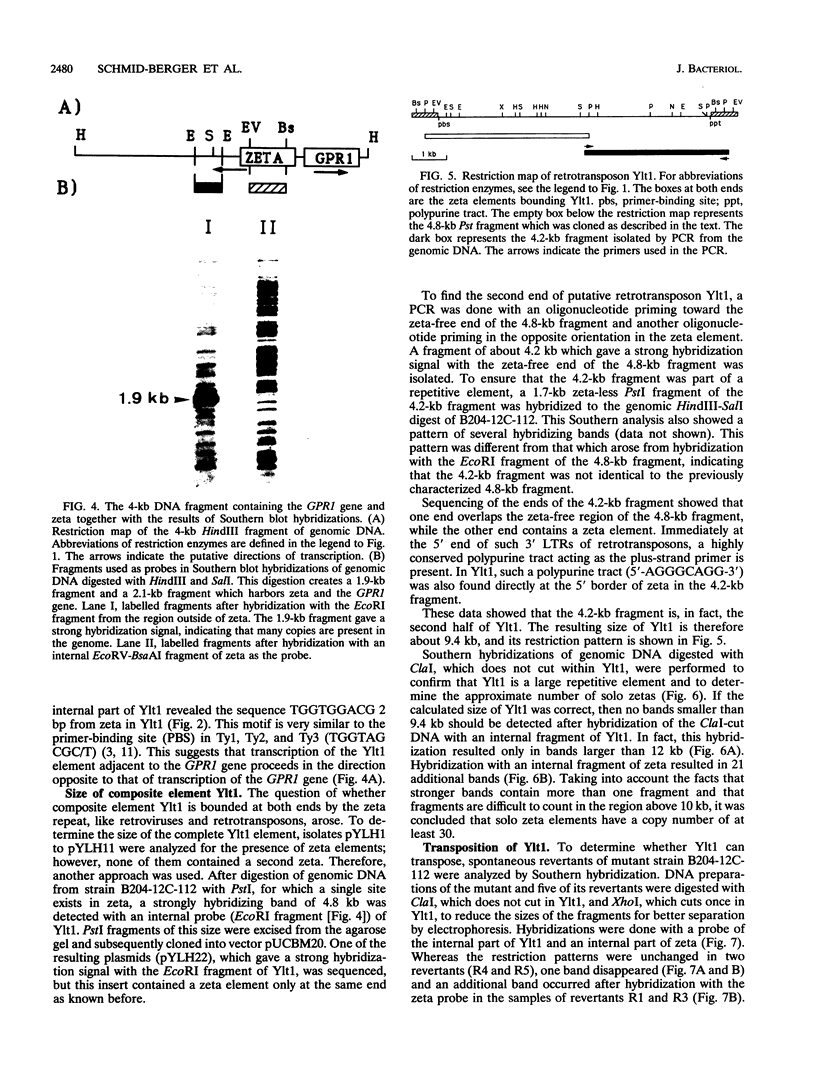
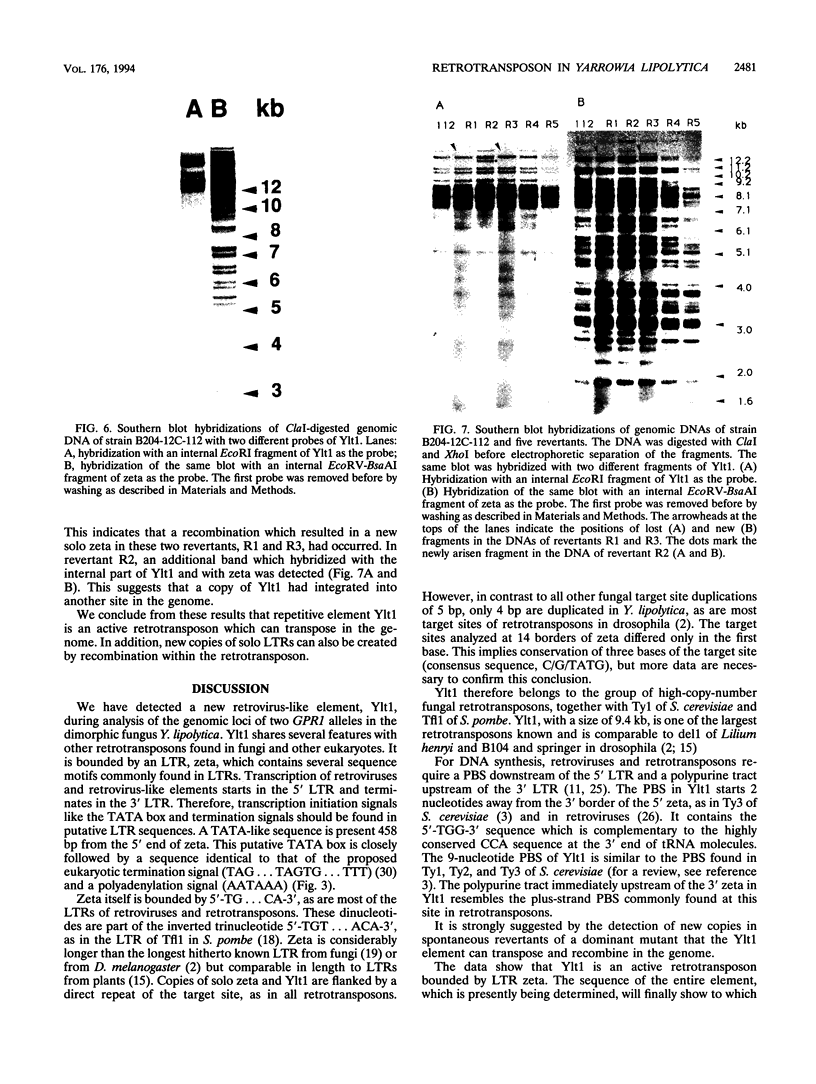
A highly repetitive composite element, Ylt1, was detected in the genome of the dimorphic fungus Yarrowia lipolytica. Ylt1 resembles retrotransposons found in other eukaryotes. It is about 9.4 kb long and can transpose in the genome. The Ylt1 element is bounded by a long terminal repeat (LTR), the zeta element. Several copies of zeta were isolated and sequenced. The sequence of this element is well conserved. It is 714 bp long and is bounded by nucleotides 5'-TG...CA-3', which are part of a short inverted repeat, a feature conserved in the LTRs of retroviruses and retrotransposons. Sequence analysis revealed motifs commonly found in LTR elements, like signals for the start and termination of transcription. The zeta element exists as part of retrotransposon Ylt1, as well as a solo element in the genome. Ylt1 and solo zeta elements are flanked by a 4-bp directly repeated genomic sequence. The copy numbers of Ylt1 and solo zeta are dependent on the strain examined, but at least 35 copies of the composite Ylt1 element and more than 30 copies of the solo zeta element per haploid genome have been observed.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barth G., Weber H. Genetic studies on the yeast Saccharomycopsis lipolytica. Inactivation and mutagenesis. Z Allg Mikrobiol. 1983;23(3):147–157. doi: 10.1002/jobm.3630230302. [DOI] [PubMed] [Google Scholar]
- Brodeur G. M., Sandmeyer S. B., Olson M. V. Consistent association between sigma elements and tRNA genes in yeast. Proc Natl Acad Sci U S A. 1983 Jun;80(11):3292–3296. doi: 10.1073/pnas.80.11.3292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cameron J. R., Loh E. Y., Davis R. W. Evidence for transposition of dispersed repetitive DNA families in yeast. Cell. 1979 Apr;16(4):739–751. doi: 10.1016/0092-8674(79)90090-4. [DOI] [PubMed] [Google Scholar]
- Chaleff D. T., Fink G. R. Genetic events associated with an insertion mutation in yeast. Cell. 1980 Aug;21(1):227–237. doi: 10.1016/0092-8674(80)90130-0. [DOI] [PubMed] [Google Scholar]
- Chen J. Y., Fonzi W. A. A temperature-regulated, retrotransposon-like element from Candida albicans. J Bacteriol. 1992 Sep;174(17):5624–5632. doi: 10.1128/jb.174.17.5624-5632.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clarke L., Carbon J. A colony bank containing synthetic Col El hybrid plasmids representative of the entire E. coli genome. Cell. 1976 Sep;9(1):91–99. doi: 10.1016/0092-8674(76)90055-6. [DOI] [PubMed] [Google Scholar]
- Cryer D. R., Eccleshall R., Marmur J. Isolation of yeast DNA. Methods Cell Biol. 1975;12:39–44. doi: 10.1016/s0091-679x(08)60950-4. [DOI] [PubMed] [Google Scholar]
- Eibel H., Gafner J., Stotz A., Philippsen P. Characterization of the yeast mobile element Ty1. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 2):609–617. doi: 10.1101/sqb.1981.045.01.079. [DOI] [PubMed] [Google Scholar]
- Errede B., Cardillo T. S., Wever G., Sherman F., Stiles J. I., Friedman L. R., Sherman F. Studies on transposable elements in yeast. I. ROAM mutations causing increased expression of yeast genes: their activation by signals directed toward conjugation functions and their formation by insertion of Ty1 repetitive elements. II. deletions, duplications, and transpositions of the COR segment that encompasses the structural gene of yeast iso-1-cytochrome c. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 2):593–607. [PubMed] [Google Scholar]
- Farabaugh P. J., Fink G. R. Insertion of the eukaryotic transposable element Ty1 creates a 5-base pair duplication. Nature. 1980 Jul 24;286(5771):352–356. doi: 10.1038/286352a0. [DOI] [PubMed] [Google Scholar]
- Gafner J., Philippsen P. The yeast transposon Ty1 generates duplications of target DNA on insertion. Nature. 1980 Jul 24;286(5771):414–418. doi: 10.1038/286414a0. [DOI] [PubMed] [Google Scholar]
- Genbauffe F. S., Chisholm G. E., Cooper T. G. Tau, sigma, and delta. A family of repeated elements in yeast. J Biol Chem. 1984 Aug 25;259(16):10518–10525. [PubMed] [Google Scholar]
- Grandbastien M. A. Retroelements in higher plants. Trends Genet. 1992 Mar;8(3):103–108. doi: 10.1016/0168-9525(92)90198-d. [DOI] [PubMed] [Google Scholar]
- Kinsey J. A., Helber J. Isolation of a transposable element from Neurospora crassa. Proc Natl Acad Sci U S A. 1989 Mar;86(6):1929–1933. doi: 10.1073/pnas.86.6.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kujau M., Weber H., Barth G. Characterization of mutants of the yeast Yarrowia lipolytica defective in acetyl-coenzyme A synthetase. Yeast. 1992 Mar;8(3):193–203. doi: 10.1002/yea.320080305. [DOI] [PubMed] [Google Scholar]
- Levin H. L., Weaver D. C., Boeke J. D. Two related families of retrotransposons from Schizosaccharomyces pombe. Mol Cell Biol. 1990 Dec;10(12):6791–6798. doi: 10.1128/mcb.10.12.6791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McHale M. T., Roberts I. N., Noble S. M., Beaumont C., Whitehead M. P., Seth D., Oliver R. P. CfT-I: an LTR-retrotransposon in Cladosporium fulvum, a fungal pathogen of tomato. Mol Gen Genet. 1992 Jun;233(3):337–347. doi: 10.1007/BF00265429. [DOI] [PubMed] [Google Scholar]
- Roeder G. S., Fink G. R. DNA rearrangements associated with a transposable element in yeast. Cell. 1980 Aug;21(1):239–249. doi: 10.1016/0092-8674(80)90131-2. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Temin H. M. Structure, variation and synthesis of retrovirus long terminal repeat. Cell. 1981 Nov;27(1 Pt 2):1–3. doi: 10.1016/0092-8674(81)90353-6. [DOI] [PubMed] [Google Scholar]
- Voytas D. F., Boeke J. D. Yeast retrotransposon revealed. Nature. 1992 Aug 27;358(6389):717–717. doi: 10.1038/358717a0. [DOI] [PubMed] [Google Scholar]
- Weaver D. C., Shpakovski G. V., Caputo E., Levin H. L., Boeke J. D. Sequence analysis of closely related retrotransposon families from fission yeast. Gene. 1993 Sep 6;131(1):135–139. doi: 10.1016/0378-1119(93)90682-s. [DOI] [PubMed] [Google Scholar]
- Xiong Y., Eickbush T. H. Similarity of reverse transcriptase-like sequences of viruses, transposable elements, and mitochondrial introns. Mol Biol Evol. 1988 Nov;5(6):675–690. doi: 10.1093/oxfordjournals.molbev.a040521. [DOI] [PubMed] [Google Scholar]
- Zaret K. S., Sherman F. DNA sequence required for efficient transcription termination in yeast. Cell. 1982 Mar;28(3):563–573. doi: 10.1016/0092-8674(82)90211-2. [DOI] [PubMed] [Google Scholar]