Abstract
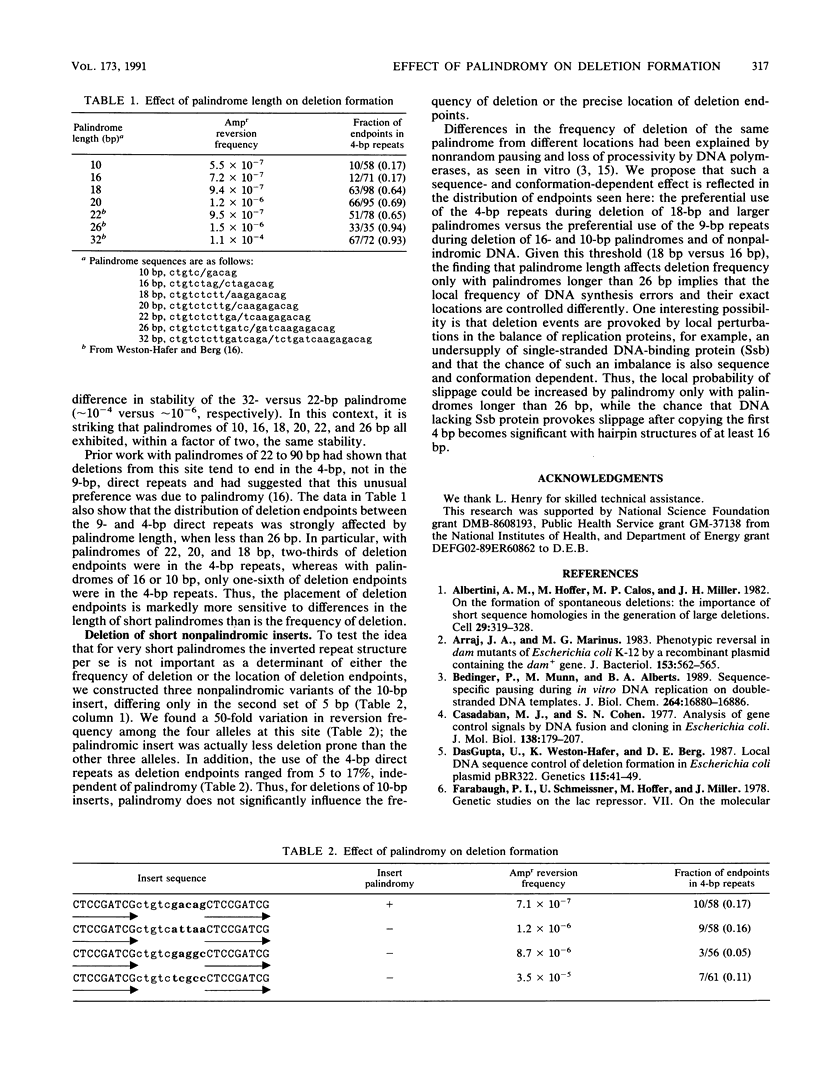
We tested the effect of palindromy on deletion formation. This involved a study of reversion of insertion mutations in the pBR322 amp gene at a site where deletions end either in 9-bp direct repeats or in adjoining 4-bp direct repeats. Inserts of palindromic DNAs ranging from 10 to more than 26 bp and related nonpalindromic DNAs were compared. The frequency of deletions (selected as Ampr revertants) was stimulated by palindromy only at lengths greater than 26 bp. The 4-bp direct repeats, one component of which is located in the palindromic insert, were used preferentially as deletion endpoints with palindromes of at least 18 bp but not of 16 or 10 bp. We interpret these results with a model of slippage during DNA replication. Because deletion frequency and deletion endpoint location depend differently on palindrome length, we propose that different factors commit a molecule to undergo deletion and determine exactly where deletion endpoints will be.
Full text
PDF


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Albertini A. M., Hofer M., Calos M. P., Miller J. H. On the formation of spontaneous deletions: the importance of short sequence homologies in the generation of large deletions. Cell. 1982 Jun;29(2):319–328. doi: 10.1016/0092-8674(82)90148-9. [DOI] [PubMed] [Google Scholar]
- Arraj J. A., Marinus M. G. Phenotypic reversal in dam mutants of Escherichia coli K-12 by a recombinant plasmid containing the dam+ gene. J Bacteriol. 1983 Jan;153(1):562–565. doi: 10.1128/jb.153.1.562-565.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bedinger P., Munn M., Alberts B. M. Sequence-specific pausing during in vitro DNA replication on double-stranded DNA templates. J Biol Chem. 1989 Oct 5;264(28):16880–16886. [PubMed] [Google Scholar]
- Casadaban M. J., Cohen S. N. Analysis of gene control signals by DNA fusion and cloning in Escherichia coli. J Mol Biol. 1980 Apr;138(2):179–207. doi: 10.1016/0022-2836(80)90283-1. [DOI] [PubMed] [Google Scholar]
- DasGupta U., Weston-Hafer K., Berg D. E. Local DNA sequence control of deletion formation in Escherichia coli plasmid pBR322. Genetics. 1987 Jan;115(1):41–49. doi: 10.1093/genetics/115.1.41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Glickman B. W., Ripley L. S. Structural intermediates of deletion mutagenesis: a role for palindromic DNA. Proc Natl Acad Sci U S A. 1984 Jan;81(2):512–516. doi: 10.1073/pnas.81.2.512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nag D. K., DasGupta U., Adelt G., Berg D. E. IS50-mediated inverse transposition: specificity and precision. Gene. 1985;34(1):17–26. doi: 10.1016/0378-1119(85)90290-2. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaaper R. M., Danforth B. N., Glickman B. W. Mechanisms of spontaneous mutagenesis: an analysis of the spectrum of spontaneous mutation in the Escherichia coli lacI gene. J Mol Biol. 1986 May 20;189(2):273–284. doi: 10.1016/0022-2836(86)90509-7. [DOI] [PubMed] [Google Scholar]
- Weisman-Shomer P., Dube D. K., Perrino F. W., Stokes K., Loeb L. A., Fry M. Sequence specificity of pausing by DNA polymerases. Biochem Biophys Res Commun. 1989 Nov 15;164(3):1149–1156. doi: 10.1016/0006-291x(89)91789-0. [DOI] [PubMed] [Google Scholar]
- Weston-Hafer K., Berg D. E. Palindromy and the location of deletion endpoints in Escherichia coli. Genetics. 1989 Apr;121(4):651–658. doi: 10.1093/genetics/121.4.651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weston-Hafer K., Berg D. E. Specificity of deletion events in pBR322. Plasmid. 1989 May;21(3):251–253. doi: 10.1016/0147-619x(89)90050-4. [DOI] [PubMed] [Google Scholar]


