Abstract
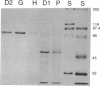
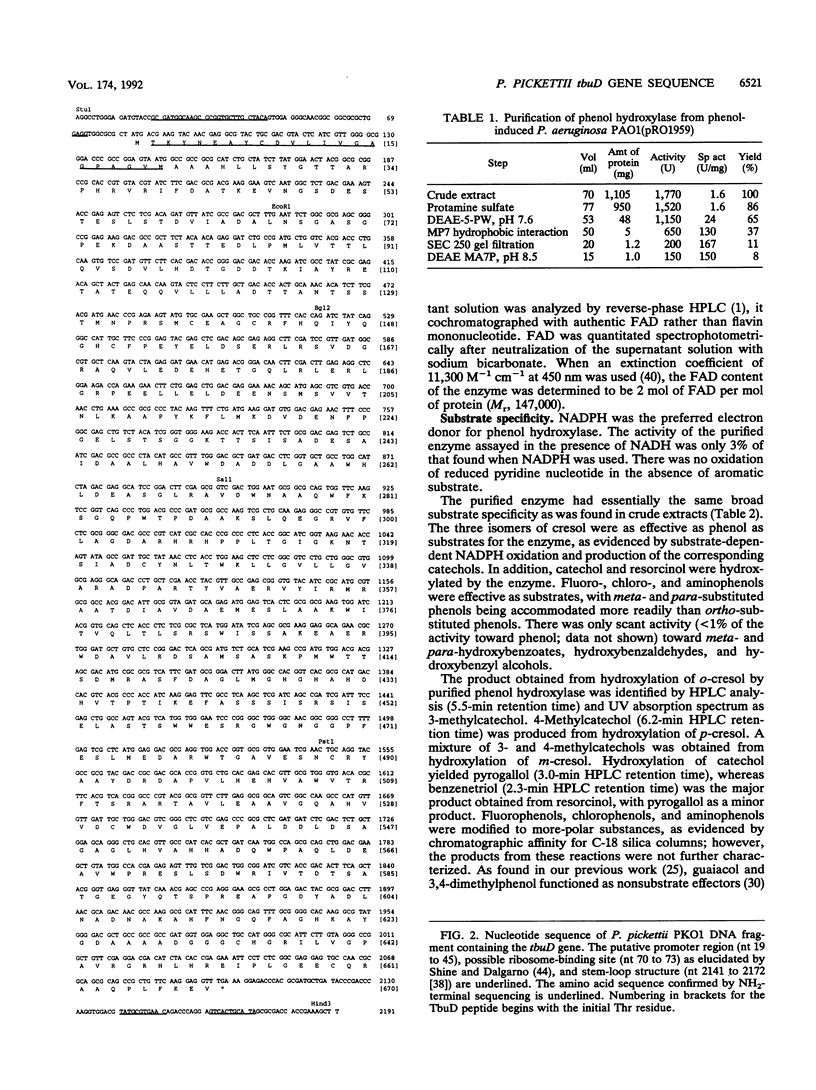
The gene (tbuD) encoding phenol hydroxylase, the enzyme that converts cresols or phenol to the corresponding catechols, has been cloned from Pseudomonas pickettii PKO1 as a 26.5-kbp BamHI-cleaved DNA fragment, designated pRO1957, which allowed the heterogenetic recipient Pseudomonas aeruginosa PAO1c to grow on phenol as the sole source of carbon. Two subclones of pRO1957 carried in trans have shown phenol hydroxylase activity in cell extracts of P. aeruginosa. The nucleotide sequence was determined for one of these subclones, a 3.1-kbp HindIII fragment, and an open reading frame that would encode a peptide of 73 kDa was found. The size of this deduced peptide is consistent with the size of a novel peptide that had been detected in extracts of phenol-induced cells of P. aeruginosa carrying pRO1959, a partial HindIII deletion subclone of pRO1957. Phenol hydroxylase purified from phenol-plus-Casamino Acid-grown cells of P. aeruginosa carrying pRO1959 has an absorbance spectrum characteristic of a simple flavoprotein; moreover, the enzyme exhibits a broad substrate range, accommodating phenol and the three isomers of cresol equally well. Sequence comparisons revealed little overall homology with other flavoprotein hydroxylases, supporting the novelty of this enzyme, although three conserved domains were apparent.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Batie C. J., LaHaie E., Ballou D. P. Purification and characterization of phthalate oxygenase and phthalate oxygenase reductase from Pseudomonas cepacia. J Biol Chem. 1987 Feb 5;262(4):1510–1518. [PubMed] [Google Scholar]
- Beadle C. A., Smith A. R. The purification and properties of 2,4-dichlorophenol hydroxylase from a strain of Acinetobacter species. Eur J Biochem. 1982 Apr 1;123(2):323–332. doi: 10.1111/j.1432-1033.1982.tb19771.x. [DOI] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1006/abio.1976.9999. [DOI] [PubMed] [Google Scholar]
- Detmer K., Massey V. Effect of monovalent anions on the mechanism of phenol hydroxylase. J Biol Chem. 1984 Sep 25;259(18):11265–11272. [PubMed] [Google Scholar]
- Dixon R. The xylABC promoter from the Pseudomonas putida TOL plasmid is activated by nitrogen regulatory genes in Escherichia coli. Mol Gen Genet. 1986 Apr;203(1):129–136. doi: 10.1007/BF00330393. [DOI] [PubMed] [Google Scholar]
- Eggink G., Engel H., Vriend G., Terpstra P., Witholt B. Rubredoxin reductase of Pseudomonas oleovorans. Structural relationship to other flavoprotein oxidoreductases based on one NAD and two FAD fingerprints. J Mol Biol. 1990 Mar 5;212(1):135–142. doi: 10.1016/0022-2836(90)90310-I. [DOI] [PubMed] [Google Scholar]
- Entsch B., Nan Y., Weaich K., Scott K. F. Sequence and organization of pobA, the gene coding for p-hydroxybenzoate hydroxylase, an inducible enzyme from Pseudomonas aeruginosa. Gene. 1988 Nov 30;71(2):279–291. doi: 10.1016/0378-1119(88)90044-3. [DOI] [PubMed] [Google Scholar]
- Garoff H., Ansorge W. Improvements of DNA sequencing gels. Anal Biochem. 1981 Aug;115(2):450–457. doi: 10.1016/0003-2697(81)90031-2. [DOI] [PubMed] [Google Scholar]
- Gurujeyalakshmi G., Oriel P. Isolation of phenol-degrading Bacillus stearothermophilus and partial characterization of the phenol hydroxylase. Appl Environ Microbiol. 1989 Feb;55(2):500–502. doi: 10.1128/aem.55.2.500-502.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Holloway B. W., Krishnapillai V., Morgan A. F. Chromosomal genetics of Pseudomonas. Microbiol Rev. 1979 Mar;43(1):73–102. doi: 10.1128/mr.43.1.73-102.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Howell L. G., Spector T., Massey V. Purification and properties of p-hydroxybenzoate hydroxylase from Pseudomonas fluorescens. J Biol Chem. 1972 Jul 10;247(13):4340–4350. [PubMed] [Google Scholar]
- Inouye S., Ebina Y., Nakazawa A., Nakazawa T. Nucleotide sequence surrounding transcription initiation site of xylABC operon on TOL plasmid of Pseudomonas putida. Proc Natl Acad Sci U S A. 1984 Mar;81(6):1688–1691. doi: 10.1073/pnas.81.6.1688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye S., Nakazawa A., Nakazawa T. Determination of the transcription initiation site and identification of the protein product of the regulatory gene xylR for xyl operons on the TOL plasmid. J Bacteriol. 1985 Sep;163(3):863–869. doi: 10.1128/jb.163.3.863-869.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson K., Parker M. L., Lory S. Nucleotide sequence and transcriptional initiation site of two Pseudomonas aeruginosa pilin genes. J Biol Chem. 1986 Nov 25;261(33):15703–15708. [PubMed] [Google Scholar]
- Kivisaar M. A., Habicht J. K., Heinaru A. L. Degradation of phenol and m-toluate in Pseudomonas sp. strain EST1001 and its Pseudomonas putida transconjugants is determined by a multiplasmid system. J Bacteriol. 1989 Sep;171(9):5111–5116. doi: 10.1128/jb.171.9.5111-5116.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kukor J. J., Olsen R. H. Genetic organization and regulation of a meta cleavage pathway for catechols produced from catabolism of toluene, benzene, phenol, and cresols by Pseudomonas pickettii PKO1. J Bacteriol. 1991 Aug;173(15):4587–4594. doi: 10.1128/jb.173.15.4587-4594.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kukor J. J., Olsen R. H. Molecular cloning, characterization, and regulation of a Pseudomonas pickettii PKO1 gene encoding phenol hydroxylase and expression of the gene in Pseudomonas aeruginosa PAO1c. J Bacteriol. 1990 Aug;172(8):4624–4630. doi: 10.1128/jb.172.8.4624-4630.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu T., Chapman P. J. Purification and properties of a plasmid-encoded 2,4-dichlorophenol hydroxylase. FEBS Lett. 1984 Aug 6;173(2):314–318. doi: 10.1016/0014-5793(84)80797-8. [DOI] [PubMed] [Google Scholar]
- Mandel M. Deoxyribonucleic acid base composition in the genus Pseudomonas. J Gen Microbiol. 1966 May;43(2):273–292. doi: 10.1099/00221287-43-2-273. [DOI] [PubMed] [Google Scholar]
- Mercer A. A., Loutit J. S. Transformation and transfection of Pseudomonas aeruginosa: effects of metal ions. J Bacteriol. 1979 Oct;140(1):37–42. doi: 10.1128/jb.140.1.37-42.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neujahr H. Y., Gaal A. Phenol hydroxylase from yeast. Purification and properties of the enzyme from Trichosporon cutaneum. Eur J Biochem. 1973 Jun;35(2):386–400. doi: 10.1111/j.1432-1033.1973.tb02851.x. [DOI] [PubMed] [Google Scholar]
- Nordlund I., Powlowski J., Shingler V. Complete nucleotide sequence and polypeptide analysis of multicomponent phenol hydroxylase from Pseudomonas sp. strain CF600. J Bacteriol. 1990 Dec;172(12):6826–6833. doi: 10.1128/jb.172.12.6826-6833.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nurk A., Kasak L., Kivisaar M. Sequence of the gene (pheA) encoding phenol monooxygenase from Pseudomonas sp. EST1001: expression in Escherichia coli and Pseudomonas putida. Gene. 1991 Jun 15;102(1):13–18. doi: 10.1016/0378-1119(91)90531-f. [DOI] [PubMed] [Google Scholar]
- Olsen R. H., DeBusscher G., McCombie W. R. Development of broad-host-range vectors and gene banks: self-cloning of the Pseudomonas aeruginosa PAO chromosome. J Bacteriol. 1982 Apr;150(1):60–69. doi: 10.1128/jb.150.1.60-69.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olsen R. H., Hansen J. Evolution and utility of a Pseudomonas aeruginosa drug resistance factor. J Bacteriol. 1976 Mar;125(3):837–844. doi: 10.1128/jb.125.3.837-844.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perkins E. J., Gordon M. P., Caceres O., Lurquin P. F. Organization and sequence analysis of the 2,4-dichlorophenol hydroxylase and dichlorocatechol oxidative operons of plasmid pJP4. J Bacteriol. 1990 May;172(5):2351–2359. doi: 10.1128/jb.172.5.2351-2359.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Platt T. Transcription termination and the regulation of gene expression. Annu Rev Biochem. 1986;55:339–372. doi: 10.1146/annurev.bi.55.070186.002011. [DOI] [PubMed] [Google Scholar]
- Porter T. D., Kasper C. B. Coding nucleotide sequence of rat NADPH-cytochrome P-450 oxidoreductase cDNA and identification of flavin-binding domains. Proc Natl Acad Sci U S A. 1985 Feb;82(4):973–977. doi: 10.1073/pnas.82.4.973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Powlowski J., Shingler V. In vitro analysis of polypeptide requirements of multicomponent phenol hydroxylase from Pseudomonas sp. strain CF600. J Bacteriol. 1990 Dec;172(12):6834–6840. doi: 10.1128/jb.172.12.6834-6840.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russel M., Model P. Sequence of thioredoxin reductase from Escherichia coli. Relationship to other flavoprotein disulfide oxidoreductases. J Biol Chem. 1988 Jun 25;263(18):9015–9019. [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. Determinant of cistron specificity in bacterial ribosomes. Nature. 1975 Mar 6;254(5495):34–38. doi: 10.1038/254034a0. [DOI] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. DNA sequence analysis with a modified bacteriophage T7 DNA polymerase. Proc Natl Acad Sci U S A. 1987 Jul;84(14):4767–4771. doi: 10.1073/pnas.84.14.4767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wierenga R. K., Terpstra P., Hol W. G. Prediction of the occurrence of the ADP-binding beta alpha beta-fold in proteins, using an amino acid sequence fingerprint. J Mol Biol. 1986 Jan 5;187(1):101–107. doi: 10.1016/0022-2836(86)90409-2. [DOI] [PubMed] [Google Scholar]
- You I. S., Ghosal D., Gunsalus I. C. Nucleotide sequence analysis of the Pseudomonas putida PpG7 salicylate hydroxylase gene (nahG) and its 3'-flanking region. Biochemistry. 1991 Feb 12;30(6):1635–1641. doi: 10.1021/bi00220a028. [DOI] [PubMed] [Google Scholar]
- Zylstra G. J., Gibson D. T. Toluene degradation by Pseudomonas putida F1. Nucleotide sequence of the todC1C2BADE genes and their expression in Escherichia coli. J Biol Chem. 1989 Sep 5;264(25):14940–14946. [PubMed] [Google Scholar]
- Zylstra G. J., Olsen R. H., Ballou D. P. Genetic organization and sequence of the Pseudomonas cepacia genes for the alpha and beta subunits of protocatechuate 3,4-dioxygenase. J Bacteriol. 1989 Nov;171(11):5915–5921. doi: 10.1128/jb.171.11.5915-5921.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]