Abstract
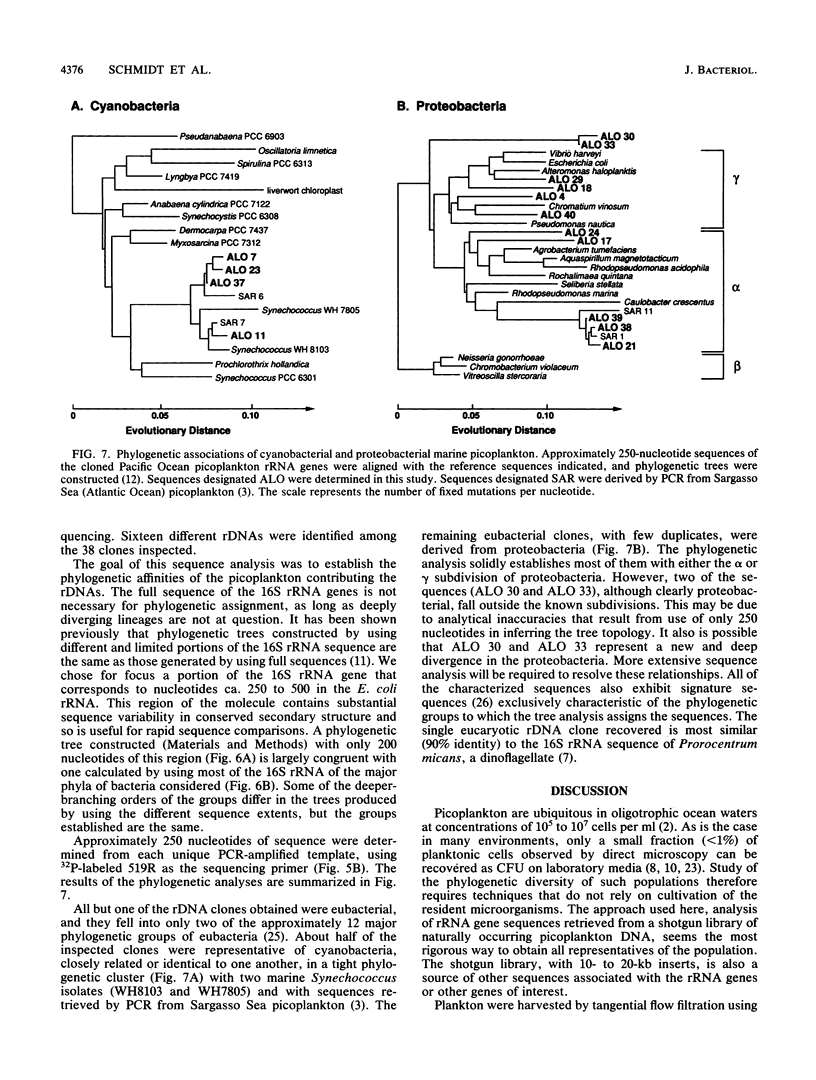
The phylogenetic diversity of an oligotrophic marine picoplankton community was examined by analyzing the sequences of cloned ribosomal genes. This strategy does not rely on cultivation of the resident microorganisms. Bulk genomic DNA was isolated from picoplankton collected in the north central Pacific Ocean by tangential flow filtration. The mixed-population DNA was fragmented, size fractionated, and cloned into bacteriophage lambda. Thirty-eight clones containing 16S rRNA genes were identified in a screen of 3.2 x 10(4) recombinant phage, and portions of the rRNA gene were amplified by polymerase chain reaction and sequenced. The resulting sequences were used to establish the identities of the picoplankton by comparison with an established data base of rRNA sequences. Fifteen unique eubacterial sequences were obtained, including four from cyanobacteria and eleven from proteobacteria. A single eucaryote related to dinoflagellates was identified; no archaebacterial sequences were detected. The cyanobacterial sequences are all closely related to sequences from cultivated marine Synechococcus strains and with cyanobacterial sequences obtained from the Atlantic Ocean (Sargasso Sea). Several sequences were related to common marine isolates of the gamma subdivision of proteobacteria. In addition to sequences closely related to those of described bacteria, sequences were obtained from two phylogenetic groups of organisms that are not closely related to any known rRNA sequences from cultivated organisms. Both of these novel phylogenetic clusters are proteobacteria, one group within the alpha subdivision and the other distinct from known proteobacterial subdivisions. The rRNA sequences of the alpha-related group are nearly identical to those of some Sargasso Sea picoplankton, suggesting a global distribution of these organisms.
Full text
PDF


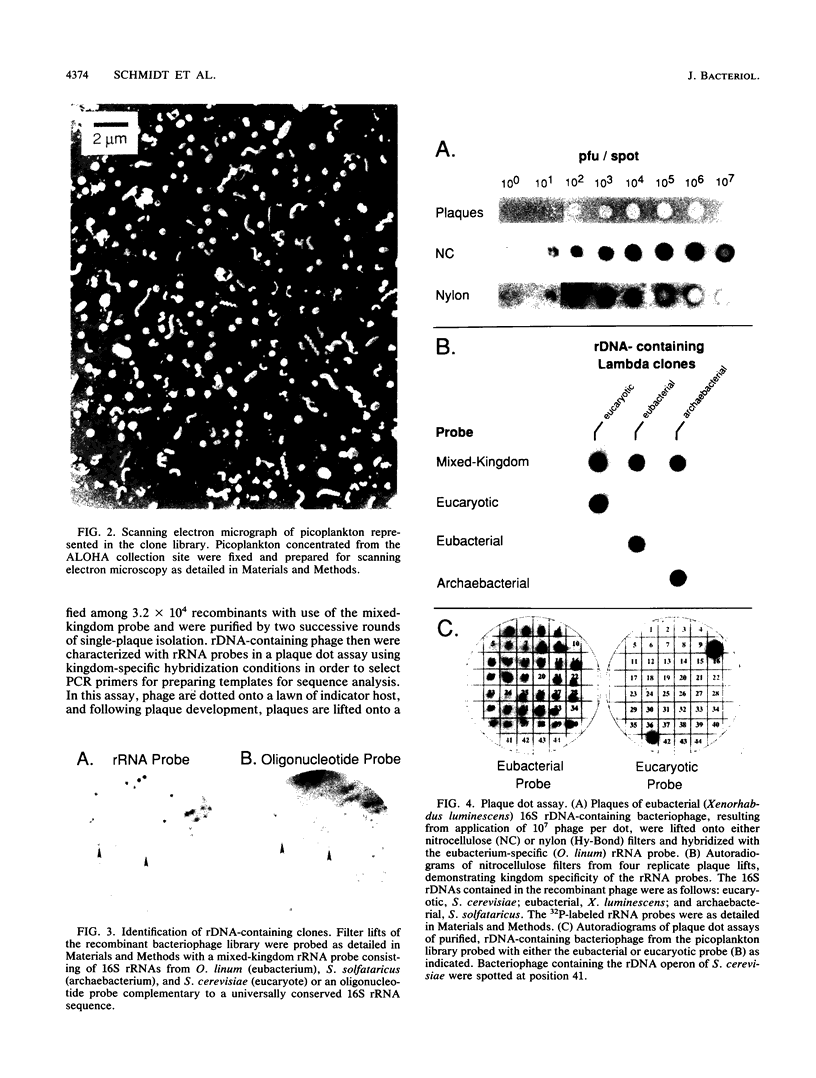




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Giovannoni S. J., Britschgi T. B., Moyer C. L., Field K. G. Genetic diversity in Sargasso Sea bacterioplankton. Nature. 1990 May 3;345(6270):60–63. doi: 10.1038/345060a0. [DOI] [PubMed] [Google Scholar]
- Giovannoni S. J., DeLong E. F., Olsen G. J., Pace N. R. Phylogenetic group-specific oligodeoxynucleotide probes for identification of single microbial cells. J Bacteriol. 1988 Feb;170(2):720–726. doi: 10.1128/jb.170.2.720-726.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giovannoni S. J., DeLong E. F., Schmidt T. M., Pace N. R. Tangential flow filtration and preliminary phylogenetic analysis of marine picoplankton. Appl Environ Microbiol. 1990 Aug;56(8):2572–2575. doi: 10.1128/aem.56.8.2572-2575.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helms C., Graham M. Y., Dutchik J. E., Olson M. V. A new method for purifying lambda DNA from phage lysates. DNA. 1985 Feb;4(1):39–49. doi: 10.1089/dna.1985.4.39. [DOI] [PubMed] [Google Scholar]
- Herzog M., Maroteaux L. Dinoflagellate 17S rRNA sequence inferred from the gene sequence: Evolutionary implications. Proc Natl Acad Sci U S A. 1986 Nov;83(22):8644–8648. doi: 10.1073/pnas.83.22.8644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kogure K., Simidu U., Taga N. A tentative direct microscopic method for counting living marine bacteria. Can J Microbiol. 1979 Mar;25(3):415–420. doi: 10.1139/m79-063. [DOI] [PubMed] [Google Scholar]
- Lane D. J., Pace B., Olsen G. J., Stahl D. A., Sogin M. L., Pace N. R. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci U S A. 1985 Oct;82(20):6955–6959. doi: 10.1073/pnas.82.20.6955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olsen G. J. Phylogenetic analysis using ribosomal RNA. Methods Enzymol. 1988;164:793–812. doi: 10.1016/s0076-6879(88)64084-5. [DOI] [PubMed] [Google Scholar]
- Pace N. R. Structure and synthesis of the ribosomal ribonucleic acid of prokaryotes. Bacteriol Rev. 1973 Dec;37(4):562–603. doi: 10.1128/br.37.4.562-603.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Powell R., Neilan J., Gannon F. Plaque dot assay. Nucleic Acids Res. 1986 Feb 11;14(3):1541–1541. doi: 10.1093/nar/14.3.1541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saiki R. K., Gelfand D. H., Stoffel S., Scharf S. J., Higuchi R., Horn G. T., Mullis K. B., Erlich H. A. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988 Jan 29;239(4839):487–491. doi: 10.1126/science.2448875. [DOI] [PubMed] [Google Scholar]
- Sgaramella V., Khorana H. G. CXII. Total synthesis of the structural gene for an alanine transfer RNA from yeast. Enzymic joining of the chemically synthesized polydeoxynucleotides to form the DNA duplex representing nucleotide sequence 1 to 20. J Mol Biol. 1972 Dec 28;72(2):427–444. doi: 10.1016/0022-2836(72)90155-6. [DOI] [PubMed] [Google Scholar]
- Stahl D. A., Lane D. J., Olsen G. J., Pace N. R. Analysis of hydrothermal vent-associated symbionts by ribosomal RNA sequences. Science. 1984 Apr 27;224(4647):409–411. doi: 10.1126/science.224.4647.409. [DOI] [PubMed] [Google Scholar]
- Stahl D. A., Lane D. J., Olsen G. J., Pace N. R. Characterization of a Yellowstone hot spring microbial community by 5S rRNA sequences. Appl Environ Microbiol. 1985 Jun;49(6):1379–1384. doi: 10.1128/aem.49.6.1379-1384.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward D. M., Weller R., Bateson M. M. 16S rRNA sequences reveal numerous uncultured microorganisms in a natural community. Nature. 1990 May 3;345(6270):63–65. doi: 10.1038/345063a0. [DOI] [PubMed] [Google Scholar]
- Watson S. W., Novitsky T. J., Quinby H. L., Valois F. W. Determination of bacterial number and biomass in the marine environment. Appl Environ Microbiol. 1977 Apr;33(4):940–946. doi: 10.1128/aem.33.4.940-946.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weller R., Ward D. M. Selective Recovery of 16S rRNA Sequences from Natural Microbial Communities in the Form of cDNA. Appl Environ Microbiol. 1989 Jul;55(7):1818–1822. doi: 10.1128/aem.55.7.1818-1822.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R. Bacterial evolution. Microbiol Rev. 1987 Jun;51(2):221–271. doi: 10.1128/mr.51.2.221-271.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R., Stackebrandt E., Macke T. J., Fox G. E. A phylogenetic definition of the major eubacterial taxa. Syst Appl Microbiol. 1985;6:143–151. doi: 10.1016/s0723-2020(85)80047-3. [DOI] [PubMed] [Google Scholar]






