Abstract
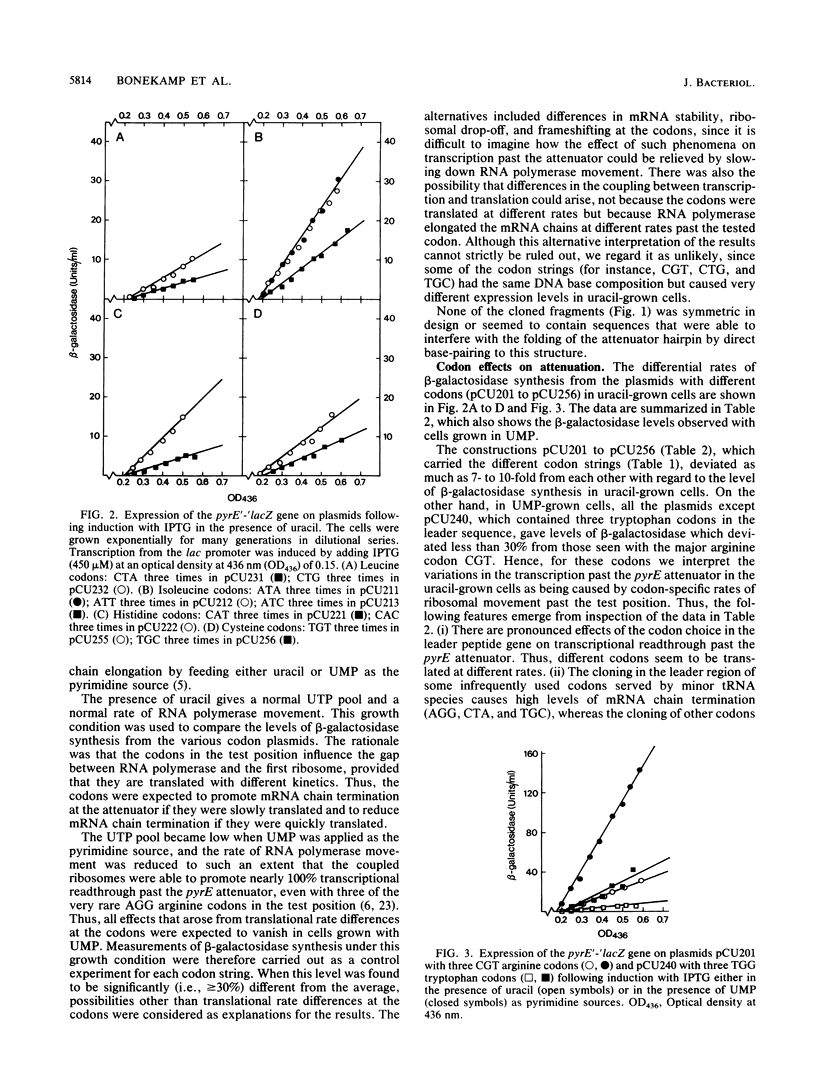
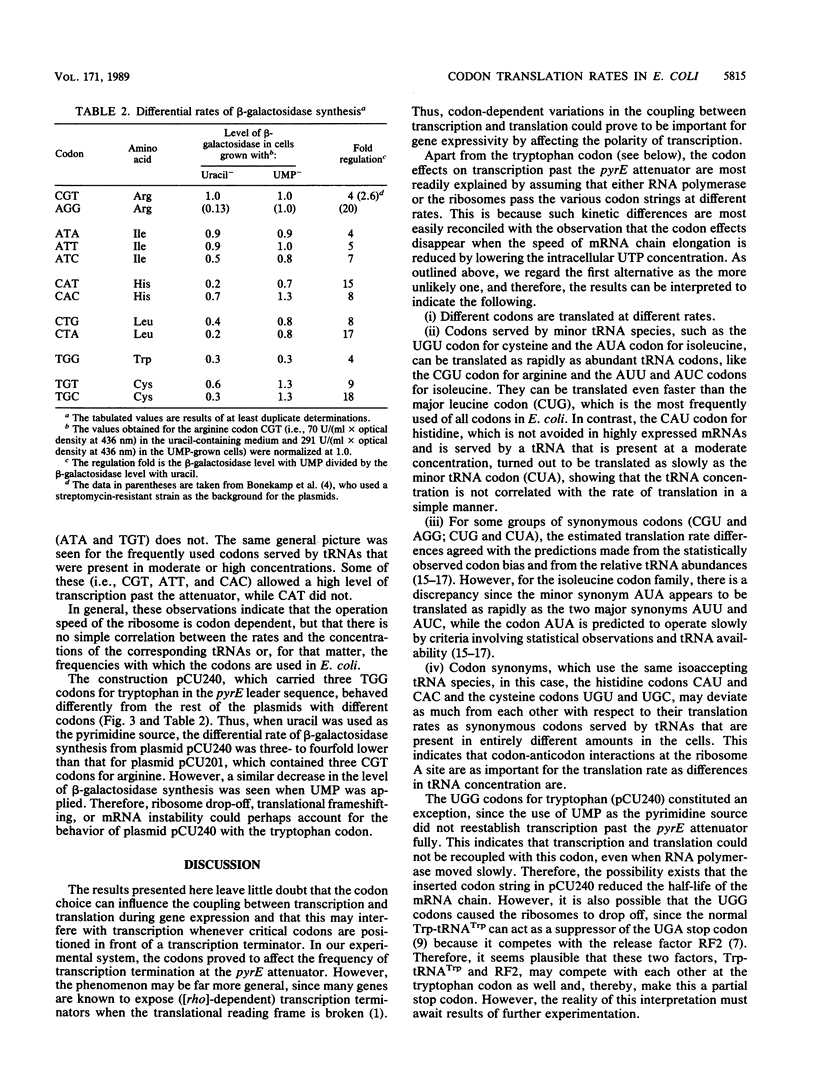
We analyzed 12 individual codons, which differed widely with respect to the frequency of use in Escherichia coli and the abundance of the corresponding tRNAs, for their influence on the coupling between transcription and translation. This was probed by determining the effects of codon substitutions in the leader peptide gene on transcription past the pyrE attenuator, as described previously by Bonekamp et al. (F. Bonekamp, H. D. Andersen, T. Christensen, and K. F. Jensen, Nucleic Acids Res. 13:4113-4123, 1985). In principle, the results revealed that either RNA polymerase or the (leading) ribosomes pass the different codon strings at different rates. However, under the assumption that the rate of transcription elongation is unaffected by the sequence changes, the results may be interpreted as indicating that different codons are translated at different rates and that these rates do not generally reflect the concentrations of the corresponding tRNAs or the frequencies with which the codons are used in E. coli. Moreover, it seems that codon synonyms that are served by the same isoaccepting tRNA species can deviate as much from each other in translational behavior as synonymous codons that are served by isoacceptors present in the cell in widely different amounts can.
Full text
PDF


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adhya S., Gottesman M. Control of transcription termination. Annu Rev Biochem. 1978;47:967–996. doi: 10.1146/annurev.bi.47.070178.004535. [DOI] [PubMed] [Google Scholar]
- Alff-Steinberger C. Evidence for a coding pattern on the non-coding strand of the E. coli genome. Nucleic Acids Res. 1984 Mar 12;12(5):2235–2241. doi: 10.1093/nar/12.5.2235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bennetzen J. L., Hall B. D. Codon selection in yeast. J Biol Chem. 1982 Mar 25;257(6):3026–3031. [PubMed] [Google Scholar]
- Bonekamp F., Andersen H. D., Christensen T., Jensen K. F. Codon-defined ribosomal pausing in Escherichia coli detected by using the pyrE attenuator to probe the coupling between transcription and translation. Nucleic Acids Res. 1985 Jun 11;13(11):4113–4123. doi: 10.1093/nar/13.11.4113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonekamp F., Clemmesen K., Karlström O., Jensen K. F. Mechanism of UTP-modulated attenuation at the pyrE gene of Escherichia coli: an example of operon polarity control through the coupling of translation to transcription. EMBO J. 1984 Dec 1;3(12):2857–2861. doi: 10.1002/j.1460-2075.1984.tb02220.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonekamp F., Jensen K. F. The AGG codon is translated slowly in E. coli even at very low expression levels. Nucleic Acids Res. 1988 Apr 11;16(7):3013–3024. doi: 10.1093/nar/16.7.3013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chavancy G., Chevallier A., Fournier A., Garel J. P. Adaptation of iso-tRNA concentration to mRNA codon frequency in the eukaryote cell. Biochimie. 1979;61(1):71–78. doi: 10.1016/s0300-9084(79)80314-4. [DOI] [PubMed] [Google Scholar]
- Engelberg-Kulka H. UGA suppression by normal tRNA Trp in Escherichia coli: codon context effects. Nucleic Acids Res. 1981 Feb 25;9(4):983–991. doi: 10.1093/nar/9.4.983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fickett J. W. Recognition of protein coding regions in DNA sequences. Nucleic Acids Res. 1982 Sep 11;10(17):5303–5318. doi: 10.1093/nar/10.17.5303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gouy M., Gautier C. Codon usage in bacteria: correlation with gene expressivity. Nucleic Acids Res. 1982 Nov 25;10(22):7055–7074. doi: 10.1093/nar/10.22.7055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grantham R., Gautier C., Gouy M. Codon frequencies in 119 individual genes confirm consistent choices of degenerate bases according to genome type. Nucleic Acids Res. 1980 May 10;8(9):1893–1912. doi: 10.1093/nar/8.9.1893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grantham R., Gautier C., Gouy M., Jacobzone M., Mercier R. Codon catalog usage is a genome strategy modulated for gene expressivity. Nucleic Acids Res. 1981 Jan 10;9(1):r43–r74. doi: 10.1093/nar/9.1.213-b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gribskov M., Devereux J., Burgess R. R. The codon preference plot: graphic analysis of protein coding sequences and prediction of gene expression. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 2):539–549. doi: 10.1093/nar/12.1part2.539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ikemura T. Codon usage and tRNA content in unicellular and multicellular organisms. Mol Biol Evol. 1985 Jan;2(1):13–34. doi: 10.1093/oxfordjournals.molbev.a040335. [DOI] [PubMed] [Google Scholar]
- Ikemura T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes: a proposal for a synonymous codon choice that is optimal for the E. coli translational system. J Mol Biol. 1981 Sep 25;151(3):389–409. doi: 10.1016/0022-2836(81)90003-6. [DOI] [PubMed] [Google Scholar]
- Ikemura T., Ozeki H. Codon usage and transfer RNA contents: organism-specific codon-choice patterns in reference to the isoacceptor contents. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):1087–1097. doi: 10.1101/sqb.1983.047.01.123. [DOI] [PubMed] [Google Scholar]
- Jensen K. F. Hyper-regulation of pyr gene expression in Escherichia coli cells with slow ribosomes. Evidence for RNA polymerase pausing in vivo? Eur J Biochem. 1988 Aug 15;175(3):587–593. doi: 10.1111/j.1432-1033.1988.tb14232.x. [DOI] [PubMed] [Google Scholar]
- Jensen K. F., Larsen J. N., Schack L., Sivertsen A. Studies on the structure and expression of Escherichia coli pyrC, pyrD, and pyrF using the cloned genes. Eur J Biochem. 1984 Apr 16;140(2):343–352. doi: 10.1111/j.1432-1033.1984.tb08107.x. [DOI] [PubMed] [Google Scholar]
- Post L. E., Nomura M. DNA sequences from the str operon of Escherichia coli. J Biol Chem. 1980 May 25;255(10):4660–4666. [PubMed] [Google Scholar]
- Post L. E., Strycharz G. D., Nomura M., Lewis H., Dennis P. P. Nucleotide sequence of the ribosomal protein gene cluster adjacent to the gene for RNA polymerase subunit beta in Escherichia coli. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1697–1701. doi: 10.1073/pnas.76.4.1697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poulsen P., Jensen K. F. Effect of UTP and GTP pools on attenuation at the pyrE gene of Escherichia coli. Mol Gen Genet. 1987 Jun;208(1-2):152–158. doi: 10.1007/BF00330436. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp P. M., Li W. H. The codon Adaptation Index--a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 1987 Feb 11;15(3):1281–1295. doi: 10.1093/nar/15.3.1281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp P. M., Tuohy T. M., Mosurski K. R. Codon usage in yeast: cluster analysis clearly differentiates highly and lowly expressed genes. Nucleic Acids Res. 1986 Jul 11;14(13):5125–5143. doi: 10.1093/nar/14.13.5125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staden R., McLachlan A. D. Codon preference and its use in identifying protein coding regions in long DNA sequences. Nucleic Acids Res. 1982 Jan 11;10(1):141–156. doi: 10.1093/nar/10.1.141. [DOI] [PMC free article] [PubMed] [Google Scholar]


