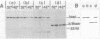Abstract
The nusA134 mutation was isolated from a sup0 strain as a temperature-sensitive mutant which grew at 32 degrees C but not at 42 degrees C. Immunoblot analysis showed that this mutant produced a 31,000-dalton nusA-encoded protein instead of the full-size 54,500-dalton product. Sequence and genetic analyses of the mutant nusA gene revealed a substitution of T for C at the PstI site (i.e., CTGCAG to CTGTAG), thereby creating a nonsense UAG codon. These results indicate that nusA134 is an amber mutation and that the 31,000-dalton amber fragment is active for Escherichia coli growth at 32 degrees C but not at 42 degrees C. Most lambda bacteriophage variants tested grew normally on the nusA134 mutant both at permissive and at nonpermissive temperatures. However, lambda r32, which carries an IS2 insertion beyond the tR1 terminator, was restricted at 42 degrees C. Defects in the transcriptional antitermination process, but not in transcription termination, were observed. A comparative study of nusA134 protein and a PstI-truncated protein suggests that truncation of the peptide chain at the PstI site by the amber mutation, rather than the loss of the glutamine residue, is primarily responsible for the defect in antitermination. The mode of the involvement of mutant nusA proteins in the N-mediated antitermination reaction is discussed.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adhya S., Miller W. Modulation of the two promoters of the galactose operon of Escherichia coli. Nature. 1979 Jun 7;279(5713):492–494. doi: 10.1038/279492a0. [DOI] [PubMed] [Google Scholar]
- Appleyard R K. Segregation of Lambda Lysogenicity during Bacterial Recombination in Escherichia Coli K12. Genetics. 1954 Jul;39(4):429–439. doi: 10.1093/genetics/39.4.429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bachmann B. J. Linkage map of Escherichia coli K-12, edition 7. Microbiol Rev. 1983 Jun;47(2):180–230. doi: 10.1128/mr.47.2.180-230.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- CAMPBELL A. Sensitive mutants of bacteriophage lambda. Virology. 1961 May;14:22–32. doi: 10.1016/0042-6822(61)90128-3. [DOI] [PubMed] [Google Scholar]
- Court D., Brady C., Rosenberg M., Wulff D. L., Behr M., Mahoney M., Izumi S. U. Control of transcription termination: a rho-dependent termination site in bacteriophage lambda. J Mol Biol. 1980 Apr;138(2):231–254. doi: 10.1016/0022-2836(80)90285-5. [DOI] [PubMed] [Google Scholar]
- Farnham P. J., Greenblatt J., Platt T. Effects of NusA protein on transcription termination in the tryptophan operon of Escherichia coli. Cell. 1982 Jul;29(3):945–951. doi: 10.1016/0092-8674(82)90457-3. [DOI] [PubMed] [Google Scholar]
- Friedman D. I., Baron L. S. Genetic characterization of a bacterial locus involved in the activity of the N function of phage lambda. Virology. 1974 Mar;58(1):141–148. doi: 10.1016/0042-6822(74)90149-4. [DOI] [PubMed] [Google Scholar]
- Friedman D. I., Olson E. R., Georgopoulos C., Tilly K., Herskowitz I., Banuett F. Interactions of bacteriophage and host macromolecules in the growth of bacteriophage lambda. Microbiol Rev. 1984 Dec;48(4):299–325. doi: 10.1128/mr.48.4.299-325.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greenblatt J., McLimont M., Hanly S. Termination of transcription by nusA gene protein of Escherichia coli. Nature. 1981 Jul 16;292(5820):215–220. doi: 10.1038/292215a0. [DOI] [PubMed] [Google Scholar]
- Howe J. G., Hershey J. W. A sensitive immunoblotting method for measuring protein synthesis initiation factor levels in lysates of Escherichia coli. J Biol Chem. 1981 Dec 25;256(24):12836–12839. [PubMed] [Google Scholar]
- Kassavetis G. A., Chamberlin M. J. Pausing and termination of transcription within the early region of bacteriophage T7 DNA in vitro. J Biol Chem. 1981 Mar 25;256(6):2777–2786. [PubMed] [Google Scholar]
- Kingston R. E., Chamberlin M. J. Pausing and attenuation of in vitro transcription in the rrnB operon of E. coli. Cell. 1981 Dec;27(3 Pt 2):523–531. doi: 10.1016/0092-8674(81)90394-9. [DOI] [PubMed] [Google Scholar]
- Kung H., Spears C., Weissbach H. Purification and properties of a soluble factor required for the deoxyribonucleic acid-directed in vitro synthesis of beta-galactosidase. J Biol Chem. 1975 Feb 25;250(4):1556–1562. [PubMed] [Google Scholar]
- Kurihara T., Nakamura Y. Cloning of the nusA gene of Escherichia coli. Mol Gen Genet. 1983;190(2):189–195. doi: 10.1007/BF00330639. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lau L. F., Roberts J. W., Wu R. Transcription terminates at lambda tR1 in three clusters. Proc Natl Acad Sci U S A. 1982 Oct;79(20):6171–6175. doi: 10.1073/pnas.79.20.6171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S. C., Squires C. L., Squires C. Antitermination of E. coli rRNA transcription is caused by a control region segment containing lambda nut-like sequences. Cell. 1984 Oct;38(3):851–860. doi: 10.1016/0092-8674(84)90280-0. [DOI] [PubMed] [Google Scholar]
- McKenney K., Shimatake H., Court D., Schmeissner U., Brady C., Rosenberg M. A system to study promoter and terminator signals recognized by Escherichia coli RNA polymerase. Gene Amplif Anal. 1981;2:383–415. [PubMed] [Google Scholar]
- Morgan E. A. Insertions of Tn 10 into an E. coli ribosomal RNA operon are incompletely polar. Cell. 1980 Aug;21(1):257–265. doi: 10.1016/0092-8674(80)90133-6. [DOI] [PubMed] [Google Scholar]
- Nakamura Y., Mizusawa S., Court D. L., Tsugawa A. Regulatory defects of a conditionally lethal nusAts mutant of Escherichia coli. Positive and negative modulator roles of NusA protein in vivo. J Mol Biol. 1986 May 5;189(1):103–111. doi: 10.1016/0022-2836(86)90384-0. [DOI] [PubMed] [Google Scholar]
- Nakamura Y., Mizusawa S. In vivo evidence that the nusA and infB genes of E. coli are part of the same multi-gene operon which encodes at least four proteins. EMBO J. 1985 Feb;4(2):527–532. doi: 10.1002/j.1460-2075.1985.tb03660.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakamura Y., Mizusawa S., Tsugawa A., Imai M. Conditionally lethal nusAts mutation of Escherichia coli reduces transcription termination but does not affect antitermination of bacteriophage lambda. Mol Gen Genet. 1986 Jul;204(1):24–28. doi: 10.1007/BF00330182. [DOI] [PubMed] [Google Scholar]
- Nakamura Y., Plumbridge J., Dondon J., Grunberg-Manago M. Evidence for autoregulation of the nusA-infB operon of Escherichia coli. Gene. 1985;36(1-2):189–193. doi: 10.1016/0378-1119(85)90085-x. [DOI] [PubMed] [Google Scholar]
- Nakamura Y., Uchida H. Isolation of conditionally lethal amber mutations affecting synthesis of the nusA protein of Escherichia coli. Mol Gen Genet. 1983;190(2):196–203. doi: 10.1007/BF00330640. [DOI] [PubMed] [Google Scholar]
- Nakamura Y., Yura T. Induction of sigma factor synthesis in Escherichia coli by the N gene product of bacteriophage lambda. Proc Natl Acad Sci U S A. 1976 Dec;73(12):4405–4409. doi: 10.1073/pnas.73.12.4405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosenberg M., Court D., Shimatake H., Brady C., Wulff D. L. The relationship between function and DNA sequence in an intercistronic regulatory region in phage lambda. Nature. 1978 Mar 30;272(5652):414–423. doi: 10.1038/272414a0. [DOI] [PubMed] [Google Scholar]
- Saito M., Tsugawa A., Egawa K., Nakamura Y. Revised sequence of the nusA gene of Escherichia coli and identification of nusA11 (ts) and nusA1 mutations which cause changes in a hydrophobic amino acid cluster. Mol Gen Genet. 1986 Nov;205(2):380–382. doi: 10.1007/BF00430455. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shiba K., Ito K., Nakamura Y., Dondon J., Grunberg-Manago M. Altered translation initiation factor 2 in the cold-sensitive ssyG mutant affects protein export in Escherichia coli. EMBO J. 1986 Nov;5(11):3001–3006. doi: 10.1002/j.1460-2075.1986.tb04598.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stewart V., Yanofsky C. Evidence for transcription antitermination control of tryptophanase operon expression in Escherichia coli K-12. J Bacteriol. 1985 Nov;164(2):731–740. doi: 10.1128/jb.164.2.731-740.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]