Abstract
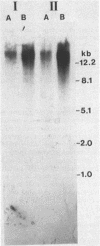
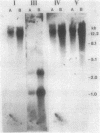
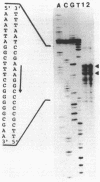
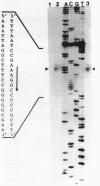
The fdhA and fdhB genes of Methanobacterium formicicum, which code for the alpha and beta subunits of formate dehydrogenase, were cotranscribed as part of a large transcript. By using Northern (RNA) gel blot analysis, the transcription start site was located within a 1.6-kilobase BglII-NcoI fragment 4.3 kilobases upstream from the fdhA gene. The precise transcription start site within the fragment was determined with the aid of primer extension analysis and S1 nuclease protection studies. A putative promoter sequence for structural genes of methanogenic archaebacteria is proposed based on a comparison of DNA sequences of the upstream region of methanogen operons for which transcription initiation sites are known. Comparison of the DNA sequence of the upstream region of the fdh operon of M. formicicum with the sequence upstream of the fdhF gene of Escherichia coli revealed regions of considerable identity.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alam J., Whitaker R. A., Krogmann D. W., Curtis S. E. Isolation and sequence of the gene for ferredoxin I from the cyanobacterium Anabaena sp. strain PCC 7120. J Bacteriol. 1986 Dec;168(3):1265–1271. doi: 10.1128/jb.168.3.1265-1271.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balch W. E., Fox G. E., Magrum L. J., Woese C. R., Wolfe R. S. Methanogens: reevaluation of a unique biological group. Microbiol Rev. 1979 Jun;43(2):260–296. doi: 10.1128/mr.43.2.260-296.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Betlach M., Pfeifer F., Friedman J., Boyer H. W. Bacterio-opsin mutants of Halobacterium halobium. Proc Natl Acad Sci U S A. 1983 Mar;80(5):1416–1420. doi: 10.1073/pnas.80.5.1416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birkmann A., Zinoni F., Sawers G., Böck A. Factors affecting transcriptional regulation of the formate-hydrogen-lyase pathway of Escherichia coli. Arch Microbiol. 1987 Jun;148(1):44–51. doi: 10.1007/BF00429646. [DOI] [PubMed] [Google Scholar]
- Bokranz M., Bäumner G., Allmansberger R., Ankel-Fuchs D., Klein A. Cloning and characterization of the methyl coenzyme M reductase genes from Methanobacterium thermoautotrophicum. J Bacteriol. 1988 Feb;170(2):568–577. doi: 10.1128/jb.170.2.568-577.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown J. W., Thomm M., Beckler G. S., Frey G., Stetter K. O., Reeve J. N. An archaebacterial RNA polymerase binding site and transcription initiation of the hisA gene in Methanococcus vannielii. Nucleic Acids Res. 1988 Jan 11;16(1):135–150. doi: 10.1093/nar/16.1.135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cram D. S., Sherf B. A., Libby R. T., Mattaliano R. J., Ramachandran K. L., Reeve J. N. Structure and expression of the genes, mcrBDCGA, which encode the subunits of component C of methyl coenzyme M reductase in Methanococcus vannielii. Proc Natl Acad Sci U S A. 1987 Jun;84(12):3992–3996. doi: 10.1073/pnas.84.12.3992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cue D., Beckler G. S., Reeve J. N., Konisky J. Structure and sequence divergence of two archaebacterial genes. Proc Natl Acad Sci U S A. 1985 Jun;82(12):4207–4211. doi: 10.1073/pnas.82.12.4207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox G. E., Magrum L. J., Balch W. E., Wolfe R. S., Woese C. R. Classification of methanogenic bacteria by 16S ribosomal RNA characterization. Proc Natl Acad Sci U S A. 1977 Oct;74(10):4537–4541. doi: 10.1073/pnas.74.10.4537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox G. E., Stackebrandt E., Hespell R. B., Gibson J., Maniloff J., Dyer T. A., Wolfe R. S., Balch W. E., Tanner R. S., Magrum L. J. The phylogeny of prokaryotes. Science. 1980 Jul 25;209(4455):457–463. doi: 10.1126/science.6771870. [DOI] [PubMed] [Google Scholar]
- Guide to molecular cloning techniques. Methods Enzymol. 1987;152:1–812. [PubMed] [Google Scholar]
- Huet J., Schnabel R., Sentenac A., Zillig W. Archaebacteria and eukaryotes possess DNA-dependent RNA polymerases of a common type. EMBO J. 1983;2(8):1291–1294. doi: 10.1002/j.1460-2075.1983.tb01583.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krol A. J., Hontelez J. G., Roozendaal B., van Kammen A. On the operon structure of the nitrogenase genes of Rhizobium leguminosarum and Azotobacter vinelandii. Nucleic Acids Res. 1982 Jul 24;10(14):4147–4157. doi: 10.1093/nar/10.14.4147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- May H. D., Patel P. S., Ferry J. G. Effect of molybdenum and tungsten on synthesis and composition of formate dehydrogenase in Methanobacterium formicicum. J Bacteriol. 1988 Aug;170(8):3384–3389. doi: 10.1128/jb.170.8.3384-3389.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McMaster G. K., Carmichael G. G. Analysis of single- and double-stranded nucleic acids on polyacrylamide and agarose gels by using glyoxal and acridine orange. Proc Natl Acad Sci U S A. 1977 Nov;74(11):4835–4838. doi: 10.1073/pnas.74.11.4835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mierendorf R. C., Pfeffer D. Direct sequencing of denatured plasmid DNA. Methods Enzymol. 1987;152:556–562. doi: 10.1016/0076-6879(87)52061-4. [DOI] [PubMed] [Google Scholar]
- Schauer N. L., Ferry J. G. Composition of the coenzyme F420-dependent formate dehydrogenase from Methanobacterium formicicum. J Bacteriol. 1986 Feb;165(2):405–411. doi: 10.1128/jb.165.2.405-411.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schauer N. L., Ferry J. G., Honek J. F., Orme-Johnson W. H., Walsh C. Mechanistic studies of the coenzyme F420 reducing formate dehydrogenase from Methanobacterium formicicum. Biochemistry. 1986 Nov 4;25(22):7163–7168. doi: 10.1021/bi00370a059. [DOI] [PubMed] [Google Scholar]
- Schauer N. L., Ferry J. G. Metabolism of formate in Methanobacterium formicicum. J Bacteriol. 1980 Jun;142(3):800–807. doi: 10.1128/jb.142.3.800-807.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shuber A. P., Orr E. C., Recny M. A., Schendel P. F., May H. D., Schauer N. L., Ferry J. G. Cloning, expression, and nucleotide sequence of the formate dehydrogenase genes from Methanobacterium formicicum. J Biol Chem. 1986 Oct 5;261(28):12942–12947. [PubMed] [Google Scholar]
- Thomm M., Madon J., Stetter K. O. DNA-dependent RNA polymerases of the three orders of methanogens. Biol Chem Hoppe Seyler. 1986 Jun;367(6):473–481. doi: 10.1515/bchm3.1986.367.1.473. [DOI] [PubMed] [Google Scholar]
- Thomm M., Sherf B. A., Reeve J. N. RNA polymerase-binding and transcription initiation sites upstream of the methyl reductase operon of Methanococcus vannielii. J Bacteriol. 1988 Apr;170(4):1958–1961. doi: 10.1128/jb.170.4.1958-1961.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomm M., Wich G. An archaebacterial promoter element for stable RNA genes with homology to the TATA box of higher eukaryotes. Nucleic Acids Res. 1988 Jan 11;16(1):151–163. doi: 10.1093/nar/16.1.151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Wang S. Z., Chen J. S., Johnson J. L. The presence of five nifH-like sequences in Clostridium pasteurianum: sequence divergence and transcription properties. Nucleic Acids Res. 1988 Jan 25;16(2):439–454. doi: 10.1093/nar/16.2.439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wich G., Hummel H., Jarsch M., Bär U., Böck A. Transcription signals for stable RNA genes in Methanococcus. Nucleic Acids Res. 1986 Mar 25;14(6):2459–2479. doi: 10.1093/nar/14.6.2459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wich G., Jarsch M., Böck A. Apparent operon for a 5S ribosomal RNA gene and for tRNA genes in the archaebacterium Methanococcus vannielii. Mol Gen Genet. 1984;196(1):146–151. doi: 10.1007/BF00334107. [DOI] [PubMed] [Google Scholar]