Abstract
The xth gene of Escherichia coli K-12, which encodes exonuclease III, has been sequenced. Exonuclease III from a cloned copy of the E. coli K-12 gene has been purified and characterized. The molecular weight (30,921), the amino-terminal amino acid sequence, and the amino acid composition of the polypeptide predicted from the nucleotide sequence are in excellent agreement with those properties determined for the purified enzyme. The xth promoter was mapped by primer extension of in vivo transcripts. Inspection of the nucleotide sequence reveals that a region of dyad symmetry which could form a hairpin stem-loop structure in RNA characteristic of a rho-dependent terminator lies immediately downstream from the xth gene.
Full text
PDF
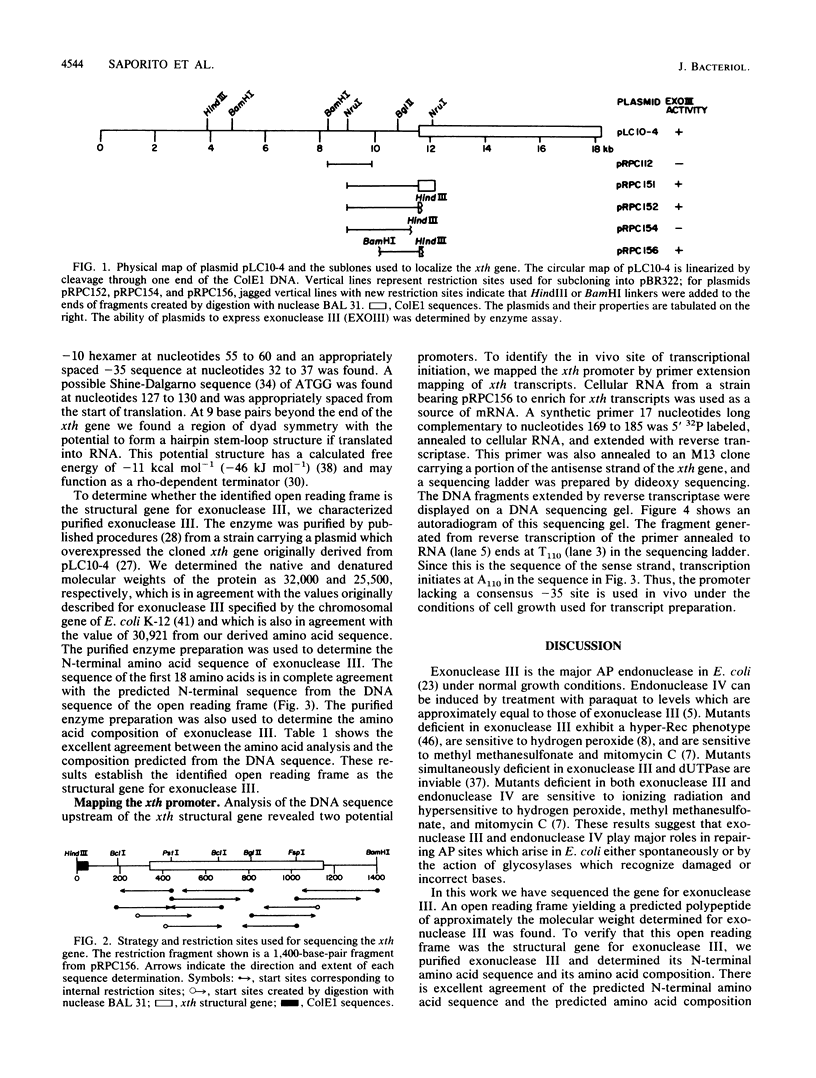




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bailly V., Verly W. G. Escherichia coli endonuclease III is not an endonuclease but a beta-elimination catalyst. Biochem J. 1987 Mar 1;242(2):565–572. doi: 10.1042/bj2420565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belfort M., Pedersen-Lane J., West D., Ehrenman K., Maley G., Chu F., Maley F. Processing of the intron-containing thymidylate synthase (td) gene of phage T4 is at the RNA level. Cell. 1985 Jun;41(2):375–382. doi: 10.1016/s0092-8674(85)80010-6. [DOI] [PubMed] [Google Scholar]
- Breimer L. H., Lindahl T. DNA glycosylase activities for thymine residues damaged by ring saturation, fragmentation, or ring contraction are functions of endonuclease III in Escherichia coli. J Biol Chem. 1984 May 10;259(9):5543–5548. [PubMed] [Google Scholar]
- Breimer L., Lindahl T. A DNA glycosylase from Escherichia coli that releases free urea from a polydeoxyribonucleotide containing fragments of base residues. Nucleic Acids Res. 1980 Dec 20;8(24):6199–6211. doi: 10.1093/nar/8.24.6199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan E., Weiss B. Endonuclease IV of Escherichia coli is induced by paraquat. Proc Natl Acad Sci U S A. 1987 May;84(10):3189–3193. doi: 10.1073/pnas.84.10.3189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clarke L., Carbon J. A colony bank containing synthetic Col El hybrid plasmids representative of the entire E. coli genome. Cell. 1976 Sep;9(1):91–99. doi: 10.1016/0092-8674(76)90055-6. [DOI] [PubMed] [Google Scholar]
- Cunningham R. P., Saporito S. M., Spitzer S. G., Weiss B. Endonuclease IV (nfo) mutant of Escherichia coli. J Bacteriol. 1986 Dec;168(3):1120–1127. doi: 10.1128/jb.168.3.1120-1127.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demple B., Halbrook J., Linn S. Escherichia coli xth mutants are hypersensitive to hydrogen peroxide. J Bacteriol. 1983 Feb;153(2):1079–1082. doi: 10.1128/jb.153.2.1079-1082.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demple B., Johnson A., Fung D. Exonuclease III and endonuclease IV remove 3' blocks from DNA synthesis primers in H2O2-damaged Escherichia coli. Proc Natl Acad Sci U S A. 1986 Oct;83(20):7731–7735. doi: 10.1073/pnas.83.20.7731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demple B., Linn S. DNA N-glycosylases and UV repair. Nature. 1980 Sep 18;287(5779):203–208. doi: 10.1038/287203a0. [DOI] [PubMed] [Google Scholar]
- Doetsch P. W., Helland D. E., Haseltine W. A. Mechanism of action of a mammalian DNA repair endonuclease. Biochemistry. 1986 Apr 22;25(8):2212–2220. doi: 10.1021/bi00356a054. [DOI] [PubMed] [Google Scholar]
- Doetsch P. W., Henner W. D., Cunningham R. P., Toney J. H., Helland D. E. A highly conserved endonuclease activity present in Escherichia coli, bovine, and human cells recognizes oxidative DNA damage at sites of pyrimidines. Mol Cell Biol. 1987 Jan;7(1):26–32. doi: 10.1128/mcb.7.1.26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gossard F., Verly W. G. Properties of the main endonuclease specific for apurinic sites of Escherichia coli (endonuclease VI). Mechanism of apurinic site excision from DNA. Eur J Biochem. 1978 Jan 16;82(2):321–332. doi: 10.1111/j.1432-1033.1978.tb12026.x. [DOI] [PubMed] [Google Scholar]
- Helland D. E., Doetsch P. W., Haseltine W. A. Substrate specificity of a mammalian DNA repair endonuclease that recognizes oxidative base damage. Mol Cell Biol. 1986 Jun;6(6):1983–1990. doi: 10.1128/mcb.6.6.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henner W. D., Grunberg S. M., Haseltine W. A. Enzyme action at 3' termini of ionizing radiation-induced DNA strand breaks. J Biol Chem. 1983 Dec 25;258(24):15198–15205. [PubMed] [Google Scholar]
- Katcher H. L., Wallace S. S. Characterization of the Escherichia coli X-ray endonuclease, endonuclease III. Biochemistry. 1983 Aug 16;22(17):4071–4081. doi: 10.1021/bi00286a013. [DOI] [PubMed] [Google Scholar]
- Keilty S., Rosenberg M. Constitutive function of a positively regulated promoter reveals new sequences essential for activity. J Biol Chem. 1987 May 5;262(13):6389–6395. [PubMed] [Google Scholar]
- Kow Y. W., Wallace S. S. Exonuclease III recognizes urea residues in oxidized DNA. Proc Natl Acad Sci U S A. 1985 Dec;82(24):8354–8358. doi: 10.1073/pnas.82.24.8354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. Comparison of initiation of protein synthesis in procaryotes, eucaryotes, and organelles. Microbiol Rev. 1983 Mar;47(1):1–45. doi: 10.1128/mr.47.1.1-45.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kyte J., Doolittle R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982 May 5;157(1):105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lipman D. J., Pearson W. R. Rapid and sensitive protein similarity searches. Science. 1985 Mar 22;227(4693):1435–1441. doi: 10.1126/science.2983426. [DOI] [PubMed] [Google Scholar]
- Ljungquist S., Lindahl T., Howard-Flanders P. Methyl methane sulfonate-sensitive mutant of Escherichia coli deficient in an endonuclease specific for apurinic sites in deoxyribonucleic acid. J Bacteriol. 1976 May;126(2):646–653. doi: 10.1128/jb.126.2.646-653.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKenney K., Shimatake H., Court D., Schmeissner U., Brady C., Rosenberg M. A system to study promoter and terminator signals recognized by Escherichia coli RNA polymerase. Gene Amplif Anal. 1981;2:383–415. [PubMed] [Google Scholar]
- Povirk L. F., Goldberg I. H. Endonuclease-resistant apyrimidinic sites formed by neocarzinostatin at cytosine residues in DNA: evidence for a possible role in mutagenesis. Proc Natl Acad Sci U S A. 1985 May;82(10):3182–3186. doi: 10.1073/pnas.82.10.3182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raibaud O., Schwartz M. Positive control of transcription initiation in bacteria. Annu Rev Genet. 1984;18:173–206. doi: 10.1146/annurev.ge.18.120184.001133. [DOI] [PubMed] [Google Scholar]
- Rao R. N., Rogers S. G. A thermoinducible lambda phage-ColE1 plasmid chimera for the overproduction of gene products from cloned DNA segments. Gene. 1978 May;3(3):247–263. doi: 10.1016/0378-1119(78)90035-5. [DOI] [PubMed] [Google Scholar]
- Rogers S. G., Weiss B. Cloning of the exonuclease III gene of Escherichia coli. Gene. 1980 Nov;11(3-4):187–195. doi: 10.1016/0378-1119(80)90059-1. [DOI] [PubMed] [Google Scholar]
- Rogers S. G., Weiss B. Exonuclease III of Escherichia coli K-12, an AP endonuclease. Methods Enzymol. 1980;65(1):201–211. doi: 10.1016/s0076-6879(80)65028-9. [DOI] [PubMed] [Google Scholar]
- Rosenberg M., Court D. Regulatory sequences involved in the promotion and termination of RNA transcription. Annu Rev Genet. 1979;13:319–353. doi: 10.1146/annurev.ge.13.120179.001535. [DOI] [PubMed] [Google Scholar]
- Sancar A., Hack A. M., Rupp W. D. Simple method for identification of plasmid-coded proteins. J Bacteriol. 1979 Jan;137(1):692–693. doi: 10.1128/jb.137.1.692-693.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp P. M., Li W. H. Codon usage in regulatory genes in Escherichia coli does not reflect selection for 'rare' codons. Nucleic Acids Res. 1986 Oct 10;14(19):7737–7749. doi: 10.1093/nar/14.19.7737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. The 3'-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci U S A. 1974 Apr;71(4):1342–1346. doi: 10.1073/pnas.71.4.1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stormo G. D., Schneider T. D., Gold L. M. Characterization of translational initiation sites in E. coli. Nucleic Acids Res. 1982 May 11;10(9):2971–2996. doi: 10.1093/nar/10.9.2971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. DNA sequence analysis with a modified bacteriophage T7 DNA polymerase. Proc Natl Acad Sci U S A. 1987 Jul;84(14):4767–4771. doi: 10.1073/pnas.84.14.4767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor A. F., Weiss B. Role of exonuclease III in the base excision repair of uracil-containing DNA. J Bacteriol. 1982 Jul;151(1):351–357. doi: 10.1128/jb.151.1.351-357.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tinoco I., Jr, Borer P. N., Dengler B., Levin M. D., Uhlenbeck O. C., Crothers D. M., Bralla J. Improved estimation of secondary structure in ribonucleic acids. Nat New Biol. 1973 Nov 14;246(150):40–41. doi: 10.1038/newbio246040a0. [DOI] [PubMed] [Google Scholar]
- Verly W. G., Rassart E. Purification of Escherichia coli endonuclease specific for apurinic sites in DNA. J Biol Chem. 1975 Oct 25;250(20):8214–8219. [PubMed] [Google Scholar]
- Warner H. R., Demple B. F., Deutsch W. A., Kane C. M., Linn S. Apurinic/apyrimidinic endonucleases in repair of pyrimidine dimers and other lesions in DNA. Proc Natl Acad Sci U S A. 1980 Aug;77(8):4602–4606. doi: 10.1073/pnas.77.8.4602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss B. Endonuclease II of Escherichia coli is exonuclease III. J Biol Chem. 1976 Apr 10;251(7):1896–1901. [PubMed] [Google Scholar]
- Weiss R. B., Duker N. J. Endonucleolytic incision of UVB-irradiated DNA. Photochem Photobiol. 1987 Jun;45(6):763–768. doi: 10.1111/j.1751-1097.1987.tb07879.x. [DOI] [PubMed] [Google Scholar]
- Weiss R. B., Duker N. J. Photoalkylated DNA and ultraviolet-irradiated DNA are incised at cytosines by endonuclease III. Nucleic Acids Res. 1986 Aug 26;14(16):6621–6631. doi: 10.1093/nar/14.16.6621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yajko D. M., Weiss B. Mutations simultaneously affecting endonuclease II and exonuclease III in Escherichia coli. Proc Natl Acad Sci U S A. 1975 Feb;72(2):688–692. doi: 10.1073/pnas.72.2.688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zieg J., Maples V. F., Kushner S. R. Recombinant levels of Escherichia coli K-12 mutants deficient in various replication, recombination, or repair genes. J Bacteriol. 1978 Jun;134(3):958–966. doi: 10.1128/jb.134.3.958-966.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]