Abstract
The induction of cat-86 by chloramphenicol has been proposed to follow the translational attenuation model. In the absence of inducer, the cat-86 gene is transcribed but remains phenotypically unexpressed because the transcripts sequester the ribosome binding site for the cat coding sequence in a stable stem-loop structure, preventing translation initiation. The translational attenuation model proposes that the natural inducer, chloramphenicol, stalls a ribosome in the leader region of cat transcripts, which causes localized melting of the downstream stem-loop structure, allowing initiation of translation of the cat-86 coding sequence. Although it is established that ribosome stalling in the cat-86 leader can induce translation of the coding sequence, several subsequent steps predicted by the model remain to be experimentally confirmed. As a consequence, the present evidence for cat-86 regulation can also be explained by two other potential control devices, ribosome hopping and translational frameshifting. Here we describe experiments designed to determine whether the alternatives to translational attenuation regulate cat-86. The results obtained are inconsistent with both competing models and are consistent with predictions made by the translational attenuation model.
Full text
PDF
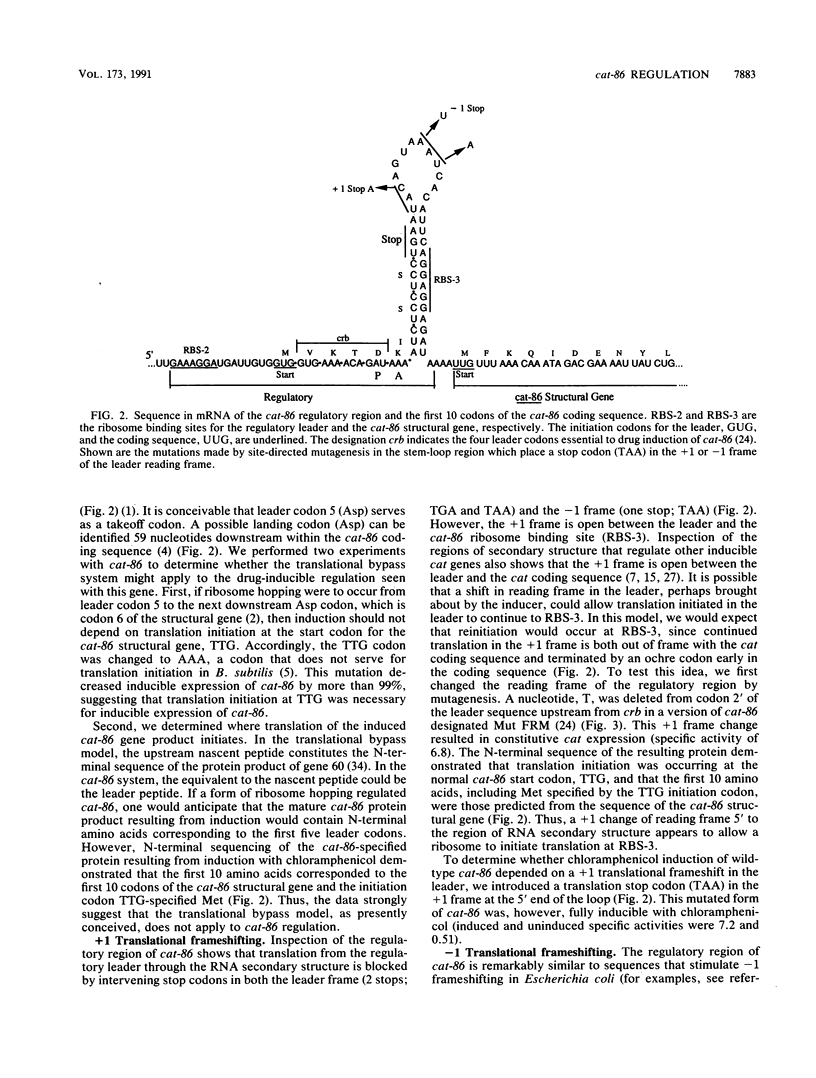





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alexieva Z., Duvall E. J., Ambulos N. P., Jr, Kim U. J., Lovett P. S. Chloramphenicol induction of cat-86 requires ribosome stalling at a specific site in the leader. Proc Natl Acad Sci U S A. 1988 May;85(9):3057–3061. doi: 10.1073/pnas.85.9.3057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ambulos N. P., Jr, Duvall E. J., Lovett P. S. Analysis of the regulatory sequences needed for induction of the chloramphenicol acetyltransferase gene cat-86 by chloramphenicol and amicetin. J Bacteriol. 1986 Sep;167(3):842–849. doi: 10.1128/jb.167.3.842-849.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ambulos N. P., Jr, Kim U. J., Rogers E. J., Lovett P. S. Constitutive expression of cat-86 associated with a change in the transcription start point. Gene. 1991 Aug 30;105(1):113–117. doi: 10.1016/0378-1119(91)90521-c. [DOI] [PubMed] [Google Scholar]
- Ambulos N. P., Jr, Mongkolsuk S., Kaufman J. D., Lovett P. S. Chloramphenicol-induced translation of cat-86 mRNA requires two cis-acting regulatory regions. J Bacteriol. 1985 Nov;164(2):696–703. doi: 10.1128/jb.164.2.696-703.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ambulos N. P., Jr, Smith T., Mulbry W., Lovett P. S. CUG as a mutant start codon for cat-86 and xylE in Bacillus subtilis. Gene. 1990 Sep 28;94(1):125–128. doi: 10.1016/0378-1119(90)90478-a. [DOI] [PubMed] [Google Scholar]
- Atkins J. F., Weiss R. B., Gesteland R. F. Ribosome gymnastics--degree of difficulty 9.5, style 10.0. Cell. 1990 Aug 10;62(3):413–423. doi: 10.1016/0092-8674(90)90007-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brückner R., Matzura H. Regulation of the inducible chloramphenicol acetyltransferase gene of the Staphylococcus aureus plasmid pUB112. EMBO J. 1985 Sep;4(9):2295–2300. doi: 10.1002/j.1460-2075.1985.tb03929.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubnau D. Translational attenuation: the regulation of bacterial resistance to the macrolide-lincosamide-streptogramin B antibiotics. CRC Crit Rev Biochem. 1984;16(2):103–132. doi: 10.3109/10409238409102300. [DOI] [PubMed] [Google Scholar]
- Duvall E. J., Ambulos N. P., Jr, Lovett P. S. Drug-free induction of a chloramphenicol acetyltransferase gene in Bacillus subtilis by stalling ribosomes in a regulatory leader. J Bacteriol. 1987 Sep;169(9):4235–4241. doi: 10.1128/jb.169.9.4235-4241.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duvall E. J., Lovett P. S. Chloramphenicol induces translation of the mRNA for a chloramphenicol-resistance gene in Bacillus subtilis. Proc Natl Acad Sci U S A. 1986 Jun;83(11):3939–3943. doi: 10.1073/pnas.83.11.3939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duvall E. J., Williams D. M., Lovett P. S., Rudolph C., Vasantha N., Guyer M. Chloramphenicol-inducible gene expression in Bacillus subtilis. Gene. 1983 Oct;24(2-3):171–177. doi: 10.1016/0378-1119(83)90077-x. [DOI] [PubMed] [Google Scholar]
- Gryczan T. J., Grandi G., Hahn J., Grandi R., Dubnau D. Conformational alteration of mRNA structure and the posttranscriptional regulation of erythromycin-induced drug resistance. Nucleic Acids Res. 1980 Dec 20;8(24):6081–6097. doi: 10.1093/nar/8.24.6081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harwood C. R., Bell D. E., Winston A. K. The effects of deletions in the leader sequence of cat-86, a chloramphenicol-resistance gene isolated from Bacillus pumilus. Gene. 1987;54(2-3):267–273. doi: 10.1016/0378-1119(87)90496-3. [DOI] [PubMed] [Google Scholar]
- Horinouchi S., Weisblum B. Nucleotide sequence and functional map of pC194, a plasmid that specifies inducible chloramphenicol resistance. J Bacteriol. 1982 May;150(2):815–825. doi: 10.1128/jb.150.2.815-825.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horinouchi S., Weisblum B. Posttranscriptional modification of mRNA conformation: mechanism that regulates erythromycin-induced resistance. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7079–7083. doi: 10.1073/pnas.77.12.7079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang W. M., Ao S. Z., Casjens S., Orlandi R., Zeikus R., Weiss R., Winge D., Fang M. A persistent untranslated sequence within bacteriophage T4 DNA topoisomerase gene 60. Science. 1988 Feb 26;239(4843):1005–1012. doi: 10.1126/science.2830666. [DOI] [PubMed] [Google Scholar]
- Laredo J., Wolff V. L., Lovett P. S. Chloramphenicol acetyltransferase specified by cat-86: relationship between the gene and the protein. Gene. 1988 Dec 15;73(1):209–214. doi: 10.1016/0378-1119(88)90327-7. [DOI] [PubMed] [Google Scholar]
- Matsudaira P. Sequence from picomole quantities of proteins electroblotted onto polyvinylidene difluoride membranes. J Biol Chem. 1987 Jul 25;262(21):10035–10038. [PubMed] [Google Scholar]
- Moos M., Jr, Nguyen N. Y., Liu T. Y. Reproducible high yield sequencing of proteins electrophoretically separated and transferred to an inert support. J Biol Chem. 1988 May 5;263(13):6005–6008. [PubMed] [Google Scholar]
- Rogers E. J., Ambulos N. P., Jr, Lovett P. S. Complementarity of Bacillus subtilis 16S rRNA with sites of antibiotic-dependent ribosome stalling in cat and erm leaders. J Bacteriol. 1990 Nov;172(11):6282–6290. doi: 10.1128/jb.172.11.6282-6290.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogers E. J., Kim U. J., Ambulos N. P., Jr, Lovett P. S. Four codons in the cat-86 leader define a chloramphenicol-sensitive ribosome stall sequence. J Bacteriol. 1990 Jan;172(1):110–115. doi: 10.1128/jb.172.1.110-115.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogers E. J., Kim U. J., Ambulos N. P., Jr, Lovett P. S. Four codons in the cat-86 leader define a chloramphenicol-sensitive ribosome stall sequence. J Bacteriol. 1990 Jan;172(1):110–115. doi: 10.1128/jb.172.1.110-115.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharrock W. J., Rabinowitz J. C. Protein synthesis in Bacillus subtilis. I. Hydrodynamics and in vitro functional properties of ribosomes from B. subtilis W168. J Mol Biol. 1979 Dec 15;135(3):611–626. doi: 10.1016/0022-2836(79)90167-0. [DOI] [PubMed] [Google Scholar]
- Shaw W. V., Brenner D. G., LeGrice S. F., Skinner S. E., Hawkins A. R. Chloramphenicol acetyltransferase gene of staphylococcal plasmid pC221. Nucleotide sequence analysis and expression studies. FEBS Lett. 1985 Jan 1;179(1):101–106. doi: 10.1016/0014-5793(85)80200-3. [DOI] [PubMed] [Google Scholar]
- Taylor J. W., Ott J., Eckstein F. The rapid generation of oligonucleotide-directed mutations at high frequency using phosphorothioate-modified DNA. Nucleic Acids Res. 1985 Dec 20;13(24):8765–8785. doi: 10.1093/nar/13.24.8765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Towbin H., Staehelin T., Gordon J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci U S A. 1979 Sep;76(9):4350–4354. doi: 10.1073/pnas.76.9.4350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuchihashi Z., Kornberg A. Translational frameshifting generates the gamma subunit of DNA polymerase III holoenzyme. Proc Natl Acad Sci U S A. 1990 Apr;87(7):2516–2520. doi: 10.1073/pnas.87.7.2516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuchihashi Z. Translational frameshifting in the Escherichia coli dnaX gene in vitro. Nucleic Acids Res. 1991 May 11;19(9):2457–2462. doi: 10.1093/nar/19.9.2457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss R. B., Huang W. M., Dunn D. M. A nascent peptide is required for ribosomal bypass of the coding gap in bacteriophage T4 gene 60. Cell. 1990 Jul 13;62(1):117–126. doi: 10.1016/0092-8674(90)90245-A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams D. M., Duvall E. J., Lovett P. S. Cloning restriction fragments that promote expression of a gene in Bacillus subtilis. J Bacteriol. 1981 Jun;146(3):1162–1165. doi: 10.1128/jb.146.3.1162-1165.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wulczyn F. G., Bölker M., Kahmann R. Translation of the bacteriophage Mu mom gene is positively regulated by the phage com gene product. Cell. 1989 Jun 30;57(7):1201–1210. doi: 10.1016/0092-8674(89)90057-3. [DOI] [PubMed] [Google Scholar]
- Wulczyn F. G., Kahmann R. Translational stimulation: RNA sequence and structure requirements for binding of Com protein. Cell. 1991 Apr 19;65(2):259–269. doi: 10.1016/0092-8674(91)90160-z. [DOI] [PubMed] [Google Scholar]
- Yang M. Y., Ferrari E., Henner D. J. Cloning of the neutral protease gene of Bacillus subtilis and the use of the cloned gene to create an in vitro-derived deletion mutation. J Bacteriol. 1984 Oct;160(1):15–21. doi: 10.1128/jb.160.1.15-21.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zoller M. J., Smith M. Oligonucleotide-directed mutagenesis of DNA fragments cloned into M13 vectors. Methods Enzymol. 1983;100:468–500. doi: 10.1016/0076-6879(83)00074-9. [DOI] [PubMed] [Google Scholar]