Abstract
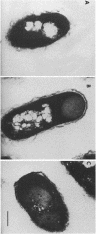
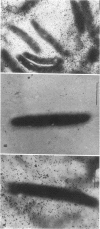
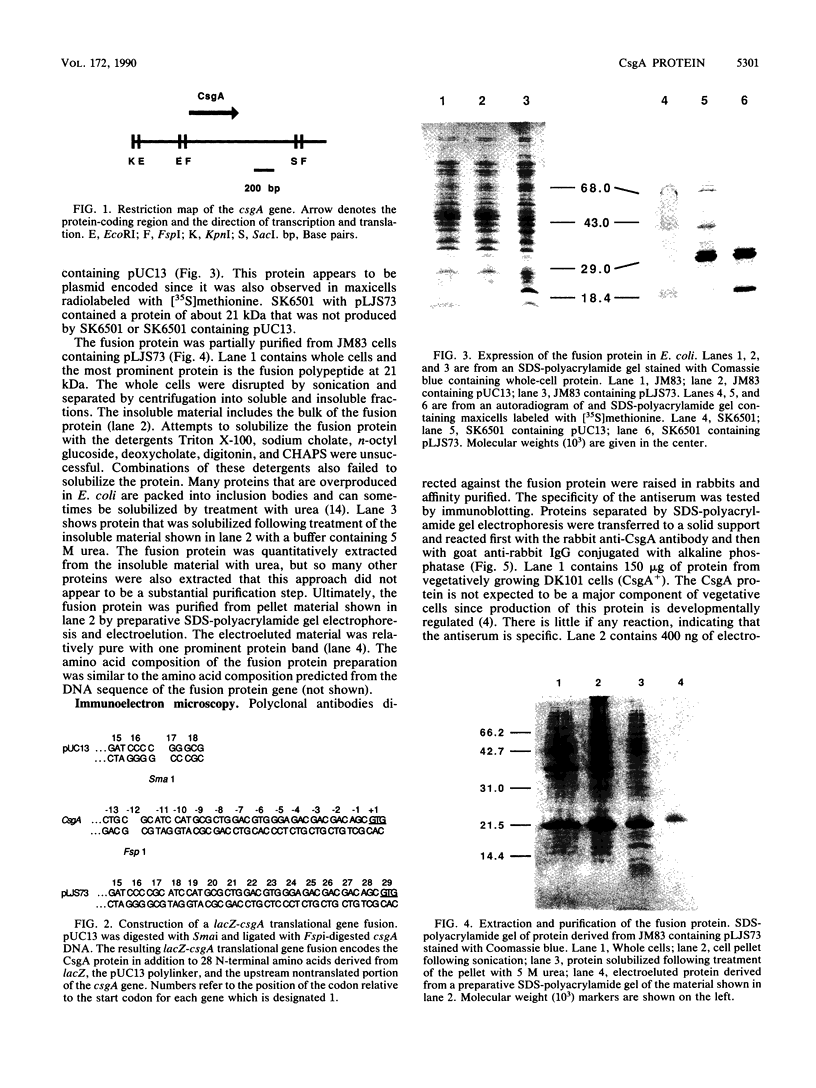
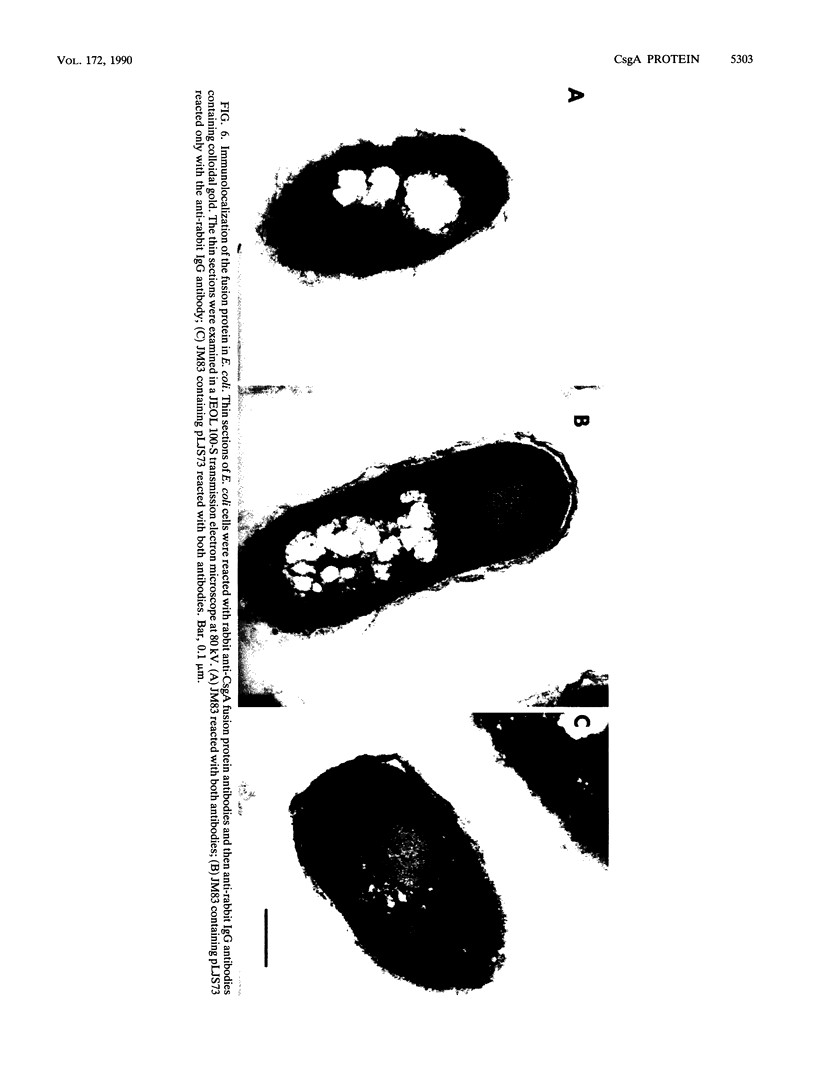
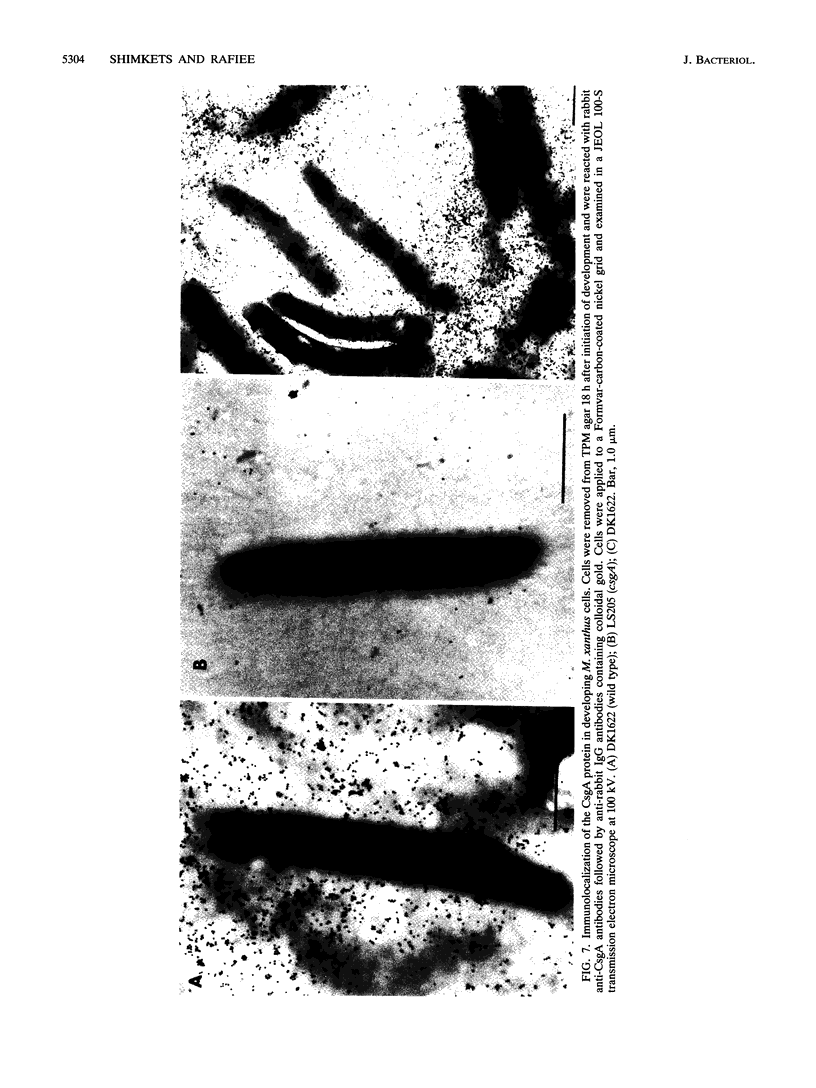
CsgA mutants of Myxococcus xanthus appear to be defective in producing an extracellular molecule essential for the developmental behaviors of this bacterium. The csgA gene encodes a 17.7-kilodalton polypeptide whose function and cellular location were investigated with immunological probes. Large quantities of the CsgA gene product were obtained from a lacZ-csgA translational gene fusion expressed in Escherichia coli. The chimeric 21-kilodalton protein was purified by preparative sodium dodecyl sulfate-polyacrylamide gel electrophoresis. Affinity-purified polyclonal antibodies raised against the fusion protein were used to determine the cellular location of the native CsgA protein by colloidal gold labeling and transmission electron microscopy. Between 1,100 and 2,200 extracellular molecules of CsgA per developing M. xanthus cell were detected, most of which were associated with the extracellular matrix. The anti-CsgA antibodies inhibited wild-type development unless they were first neutralized with the fusion protein. Together these results suggest that the CsgA gene product has an essential, extracellular function during development, possibly as a pheromone.
Full text
PDF
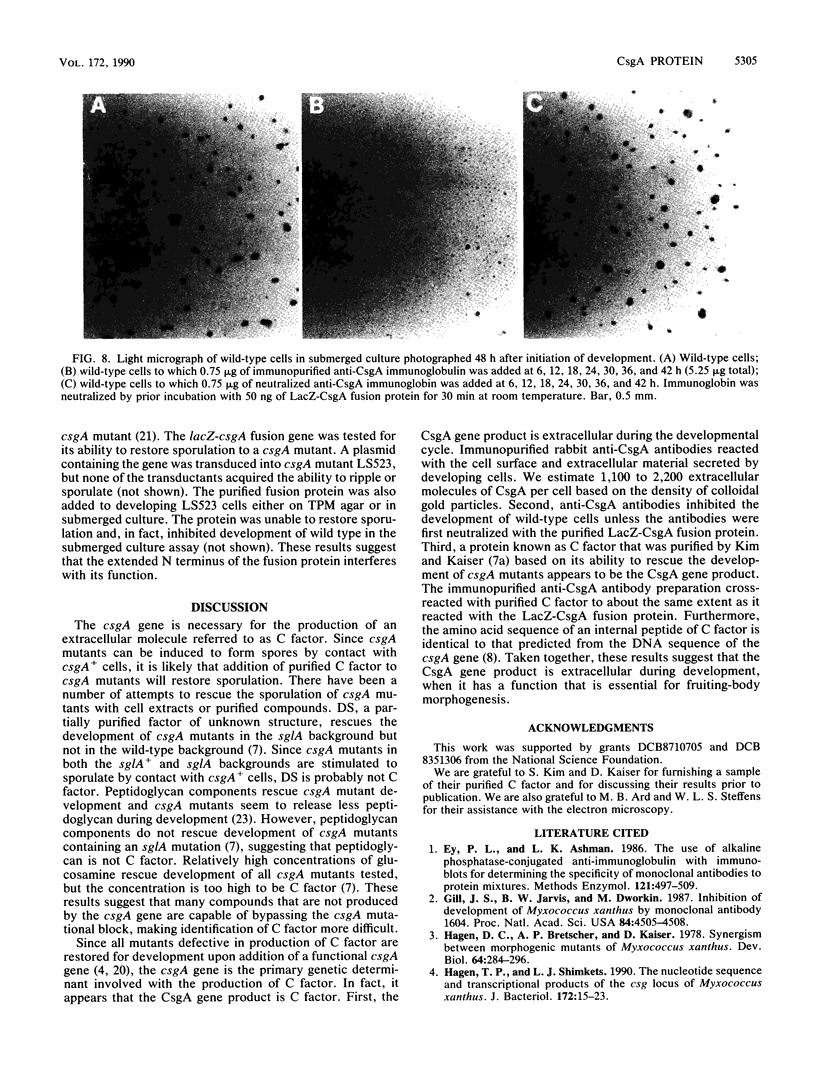




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ey P. L., Ashman L. K. The use of alkaline phosphatase-conjugated anti-immunoglobulin with immunoblots for determining the specificity of monoclonal antibodies to protein mixtures. Methods Enzymol. 1986;121:497–509. doi: 10.1016/0076-6879(86)21050-2. [DOI] [PubMed] [Google Scholar]
- Gill J. S., Jarvis B. W., Dworkin M. Inhibition of development in Myxococcus xanthus by monoclonal antibody 1604. Proc Natl Acad Sci U S A. 1987 Jul;84(13):4505–4508. doi: 10.1073/pnas.84.13.4505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hagen D. C., Bretscher A. P., Kaiser D. Synergism between morphogenetic mutants of Myxococcus xanthus. Dev Biol. 1978 Jun;64(2):284–296. doi: 10.1016/0012-1606(78)90079-9. [DOI] [PubMed] [Google Scholar]
- Hagen T. J., Shimkets L. J. Nucleotide sequence and transcriptional products of the csg locus of Myxococcus xanthus. J Bacteriol. 1990 Jan;172(1):15–23. doi: 10.1128/jb.172.1.15-23.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hodgkin J., Kaiser D. Cell-to-cell stimulation of movement in nonmotile mutants of Myxococcus. Proc Natl Acad Sci U S A. 1977 Jul;74(7):2938–2942. doi: 10.1073/pnas.74.7.2938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunkapiller M. W., Lujan E., Ostrander F., Hood L. E. Isolation of microgram quantities of proteins from polyacrylamide gels for amino acid sequence analysis. Methods Enzymol. 1983;91:227–236. doi: 10.1016/s0076-6879(83)91019-4. [DOI] [PubMed] [Google Scholar]
- Janssen G. R., Dworkin M. Cell-cell interactions in developmental lysis of Myxococcus xanthus. Dev Biol. 1985 Nov;112(1):194–202. doi: 10.1016/0012-1606(85)90133-2. [DOI] [PubMed] [Google Scholar]
- Kim S. K., Kaiser D. C-factor: a cell-cell signaling protein required for fruiting body morphogenesis of M. xanthus. Cell. 1990 Apr 6;61(1):19–26. doi: 10.1016/0092-8674(90)90211-v. [DOI] [PubMed] [Google Scholar]
- Kim S. K., Kaiser D. Purification and properties of Myxococcus xanthus C-factor, an intercellular signaling protein. Proc Natl Acad Sci U S A. 1990 May;87(10):3635–3639. doi: 10.1073/pnas.87.10.3635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kroos L., Kaiser D. Expression of many developmentally regulated genes in Myxococcus depends on a sequence of cell interactions. Genes Dev. 1987 Oct;1(8):840–854. doi: 10.1101/gad.1.8.840. [DOI] [PubMed] [Google Scholar]
- Kroos L., Kuspa A., Kaiser D. A global analysis of developmentally regulated genes in Myxococcus xanthus. Dev Biol. 1986 Sep;117(1):252–266. doi: 10.1016/0012-1606(86)90368-4. [DOI] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Marston F. A. The purification of eukaryotic polypeptides synthesized in Escherichia coli. Biochem J. 1986 Nov 15;240(1):1–12. doi: 10.1042/bj2400001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Rhie H. G., Shimkets L. J. Developmental bypass suppression of Myxococcus xanthus csgA mutations. J Bacteriol. 1989 Jun;171(6):3268–3276. doi: 10.1128/jb.171.6.3268-3276.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sancar A., Hack A. M., Rupp W. D. Simple method for identification of plasmid-coded proteins. J Bacteriol. 1979 Jan;137(1):692–693. doi: 10.1128/jb.137.1.692-693.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimkets L. J., Asher S. J. Use of recombination techniques to examine the structure of the csg locus of Myxococcus xanthus. Mol Gen Genet. 1988 Jan;211(1):63–71. doi: 10.1007/BF00338394. [DOI] [PubMed] [Google Scholar]
- Shimkets L. J. Control of morphogenesis in myxobacteria. Crit Rev Microbiol. 1987;14(3):195–227. doi: 10.3109/10408418709104439. [DOI] [PubMed] [Google Scholar]
- Shimkets L. J., Gill R. E., Kaiser D. Developmental cell interactions in Myxococcus xanthus and the spoC locus. Proc Natl Acad Sci U S A. 1983 Mar;80(5):1406–1410. doi: 10.1073/pnas.80.5.1406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimkets L. J., Kaiser D. Induction of coordinated movement of Myxococcus xanthus cells. J Bacteriol. 1982 Oct;152(1):451–461. doi: 10.1128/jb.152.1.451-461.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimkets L. J., Kaiser D. Murein components rescue developmental sporulation of Myxococcus xanthus. J Bacteriol. 1982 Oct;152(1):462–470. doi: 10.1128/jb.152.1.462-470.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimkets L. J. The role of the cell surface in social and adventurous behaviour of myxobacteria. Mol Microbiol. 1989 Sep;3(9):1295–1299. doi: 10.1111/j.1365-2958.1989.tb00280.x. [DOI] [PubMed] [Google Scholar]
- Silverton E. W., Navia M. A., Davies D. R. Three-dimensional structure of an intact human immunoglobulin. Proc Natl Acad Sci U S A. 1977 Nov;74(11):5140–5144. doi: 10.1073/pnas.74.11.5140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. A bacteriophage T7 RNA polymerase/promoter system for controlled exclusive expression of specific genes. Proc Natl Acad Sci U S A. 1985 Feb;82(4):1074–1078. doi: 10.1073/pnas.82.4.1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Talian J. C., Olmsted J. B., Goldman R. D. A rapid procedure for preparing fluorescein-labeled specific antibodies from whole antiserum: its use in analyzing cytoskeletal architecture. J Cell Biol. 1983 Oct;97(4):1277–1282. doi: 10.1083/jcb.97.4.1277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Towbin H., Staehelin T., Gordon J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci U S A. 1979 Sep;76(9):4350–4354. doi: 10.1073/pnas.76.9.4350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]