Abstract
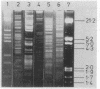
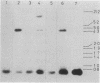
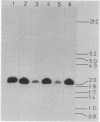
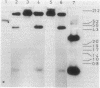
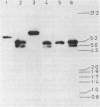
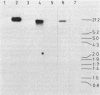
Structural genes for catechol 2,3-oxygenase (C23O) were cloned from the TOL plasmids pWW5, pWW14, pWW74, pWW84, and pWW88 isolated from Pseudomonas strains of diverse geographical origins. Each pKT230-based C23O+ recombinant plasmid carried a 2.05-kilobase XhoI insert which showed strong homology in Southern hybridizations with the xylE gene from the archetype TOL plasmid pWW0. Fragments were mapped for restriction endonuclease sites and were classified into two closely related groups on the basis of restriction maps. C23O structural genes were located on cloned fragments by a combination of subcloning and site-specific mutagenesis. All five TOL plasmids examined yielded clones whose maps differed from that of xylE of pWW0 by only a single XbaI site, but in addition plasmids pWW5, pWW74, and pWW88 carried a second, homologous C23O gene with seven further restriction site differences. The remaining plasmids, pWW14 and pWW84, carried a second nonhomologous C23O gene related to the second C23O gene (C23OII) of TOL plasmid pWW15 described previously (H. Keil, M. R. Lebens, and P. A. Williams, J. Bacteriol. 163:248-255, 1985). Thus, each naturally occurring TOL plasmid in this study appears to carry genes for two meta cleavage dioxygenases.
Full text
PDF
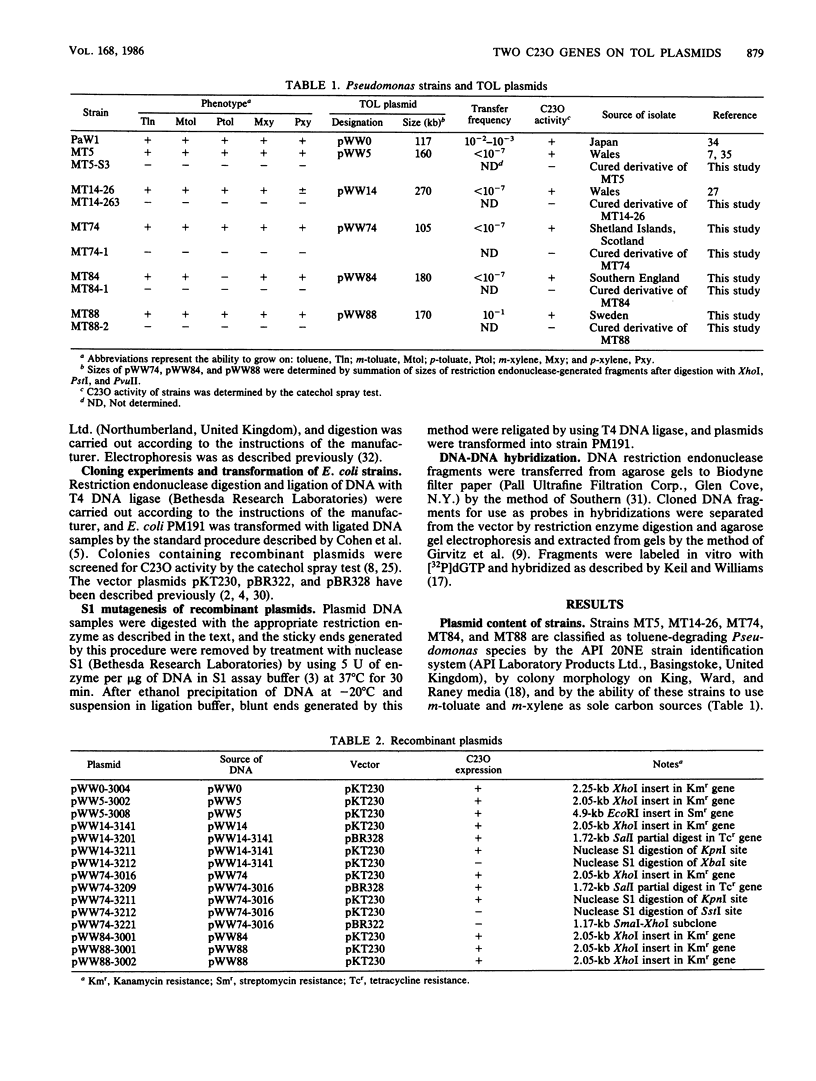


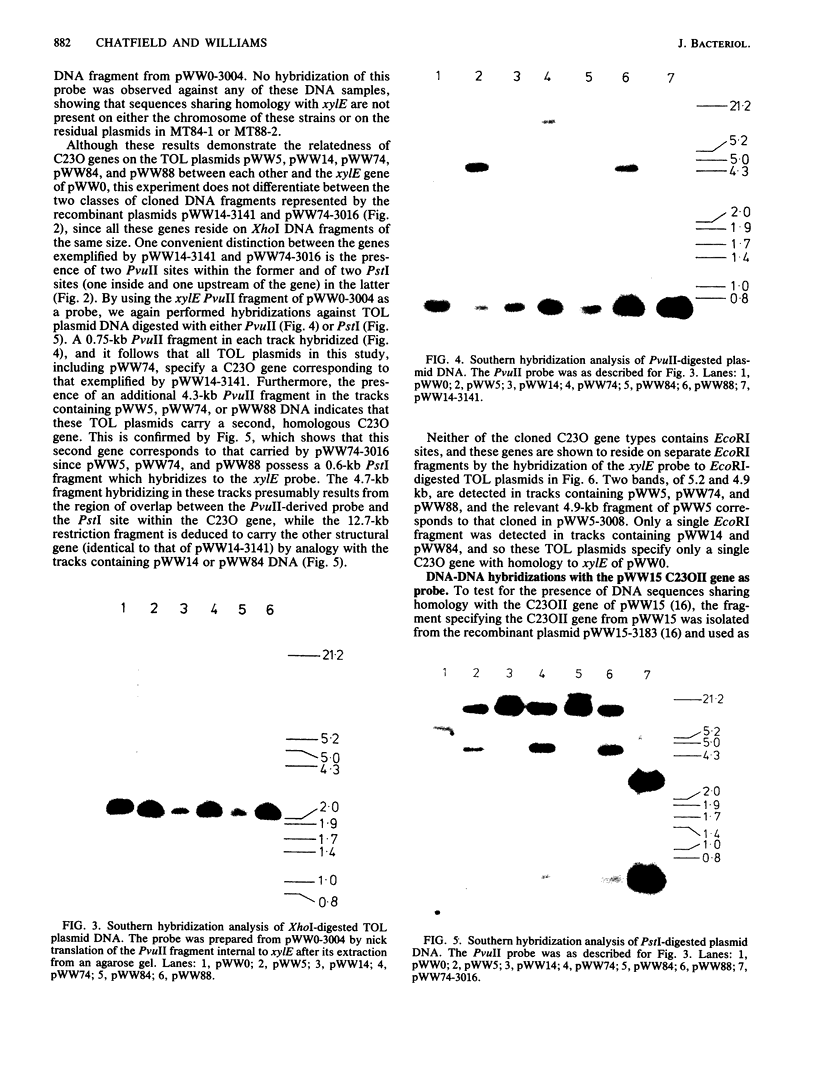



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bachmann B. J. Pedigrees of some mutant strains of Escherichia coli K-12. Bacteriol Rev. 1972 Dec;36(4):525–557. doi: 10.1128/br.36.4.525-557.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bagdasarian M., Lurz R., Rückert B., Franklin F. C., Bagdasarian M. M., Frey J., Timmis K. N. Specific-purpose plasmid cloning vectors. II. Broad host range, high copy number, RSF1010-derived vectors, and a host-vector system for gene cloning in Pseudomonas. Gene. 1981 Dec;16(1-3):237–247. doi: 10.1016/0378-1119(81)90080-9. [DOI] [PubMed] [Google Scholar]
- Barth P. T., Grinter N. J. Assay of deoxyribonucleic acid homology using a single-strand-specific nuclease at 75 C. J Bacteriol. 1975 Feb;121(2):434–441. doi: 10.1128/jb.121.2.434-441.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Cohen S. N., Chang A. C., Hsu L. Nonchromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-factor DNA. Proc Natl Acad Sci U S A. 1972 Aug;69(8):2110–2114. doi: 10.1073/pnas.69.8.2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DAGLEY S., GIBSON D. T. THE BACTERIAL DEGRADATION OF CATECHOL. Biochem J. 1965 May;95:466–474. doi: 10.1042/bj0950466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duggleby C. J., Bayley S. A., Worsey M. J., Williams P. A., Broda P. Molecular sizes and relationships of TOL plasmids in Pseudomonas. J Bacteriol. 1977 Jun;130(3):1274–1280. doi: 10.1128/jb.130.3.1274-1280.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franklin F. C., Bagdasarian M., Bagdasarian M. M., Timmis K. N. Molecular and functional analysis of the TOL plasmid pWWO from Pseudomonas putida and cloning of genes for the entire regulated aromatic ring meta cleavage pathway. Proc Natl Acad Sci U S A. 1981 Dec;78(12):7458–7462. doi: 10.1073/pnas.78.12.7458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Girvitz S. C., Bacchetti S., Rainbow A. J., Graham F. L. A rapid and efficient procedure for the purification of DNA from agarose gels. Anal Biochem. 1980 Aug;106(2):492–496. doi: 10.1016/0003-2697(80)90553-9. [DOI] [PubMed] [Google Scholar]
- Guerry P., LeBlanc D. J., Falkow S. General method for the isolation of plasmid deoxyribonucleic acid. J Bacteriol. 1973 Nov;116(2):1064–1066. doi: 10.1128/jb.116.2.1064-1066.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harayama S., Lehrbach P. R., Timmis K. N. Transposon mutagenesis analysis of meta-cleavage pathway operon genes of the TOL plasmid of Pseudomonas putida mt-2. J Bacteriol. 1984 Oct;160(1):251–255. doi: 10.1128/jb.160.1.251-255.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Inouye S., Nakazawa A., Nakazawa T. Nucleotide sequence of the promoter region of the xylDEGF operon on TOL plasmid of Pseudomonas putida. Gene. 1984 Sep;29(3):323–330. doi: 10.1016/0378-1119(84)90061-1. [DOI] [PubMed] [Google Scholar]
- Jeenes D. J., Williams P. A. Excision and integration of degradative pathway genes from TOL plasmid pWW0. J Bacteriol. 1982 Apr;150(1):188–194. doi: 10.1128/jb.150.1.188-194.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- KING E. O., WARD M. K., RANEY D. E. Two simple media for the demonstration of pyocyanin and fluorescin. J Lab Clin Med. 1954 Aug;44(2):301–307. [PubMed] [Google Scholar]
- Keil H., Keil S., Pickup R. W., Williams P. A. Evolutionary conservation of genes coding for meta pathway enzymes within TOL plasmids pWW0 and pWW53. J Bacteriol. 1985 Nov;164(2):887–895. doi: 10.1128/jb.164.2.887-895.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keil H., Lebens M. R., Williams P. A. TOL plasmid pWW15 contains two nonhomologous, independently regulated catechol 2,3-oxygenase genes. J Bacteriol. 1985 Jul;163(1):248–255. doi: 10.1128/jb.163.1.248-255.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunz D. A., Chapman P. J. Isolation and characterization of spontaneously occurring TOL plasmid mutants of Pseudomonas putida HS1. J Bacteriol. 1981 Jun;146(3):952–964. doi: 10.1128/jb.146.3.952-964.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meulien P., Broda P. Identification of chromosomally integrated TOL DNA in cured derivatives of Pseudomonas putida PAW1. J Bacteriol. 1982 Nov;152(2):911–914. doi: 10.1128/jb.152.2.911-914.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakai C., Kagamiyama H., Nozaki M., Nakazawa T., Inouye S., Ebina Y., Nakazawa A. Complete nucleotide sequence of the metapyrocatechase gene on the TOI plasmid of Pseudomonas putida mt-2. J Biol Chem. 1983 Mar 10;258(5):2923–2928. [PubMed] [Google Scholar]
- Nakazawa T., Hayashi E., Yokota T., Ebina Y., Nakazawa A. Isolation of TOL and RP4 recombinants by integrative suppression. J Bacteriol. 1978 Apr;134(1):270–277. doi: 10.1128/jb.134.1.270-277.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Negoro S., Taniguchi T., Kanaoka M., Kimura H., Okada H. Plasmid-determined enzymatic degradation of nylon oligomers. J Bacteriol. 1983 Jul;155(1):22–31. doi: 10.1128/jb.155.1.22-31.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okada H., Negoro S., Kimura H., Nakamura S. Evolutionary adaptation of plasmid-encoded enzymes for degrading nylon oligomers. Nature. 1983 Nov 10;306(5939):203–206. doi: 10.1038/306203a0. [DOI] [PubMed] [Google Scholar]
- Pankhurst E. S. A spot test for catechol 2:3-oxygenase in bacteria. J Appl Bacteriol. 1965 Aug;28(2):309–315. doi: 10.1111/j.1365-2672.1965.tb02158.x. [DOI] [PubMed] [Google Scholar]
- Pickup R. W., Williams P. A. Spontaneous deletions in the TOL plasmid pWW20 which give rise to the B3 regulatory mutants of Pseudomonas putida MT20. J Gen Microbiol. 1982 Jul;128(7):1385–1390. doi: 10.1099/00221287-128-7-1385. [DOI] [PubMed] [Google Scholar]
- Reanney D. Extrachromosomal elements as possible agents of adaptation and development. Bacteriol Rev. 1976 Sep;40(3):552–590. doi: 10.1128/br.40.3.552-590.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roussel A. F., Chabbert Y. A. Taxonomy and epidemiology of gram-negative bacterial plasmids studied by DNA-DNA filter hybridization in formamide. J Gen Microbiol. 1978 Feb;104(2):269–276. doi: 10.1099/00221287-104-2-269. [DOI] [PubMed] [Google Scholar]
- Soberon X., Covarrubias L., Bolivar F. Construction and characterization of new cloning vehicles. IV. Deletion derivatives of pBR322 and pBR325. Gene. 1980 May;9(3-4):287–305. doi: 10.1016/0378-1119(90)90328-o. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Wheatcroft R., Williams P. A. Rapid methods for the study of both stable and unstable plasmids in Pseudomonas. J Gen Microbiol. 1981 Jun;124(2):433–437. doi: 10.1099/00221287-124-2-433. [DOI] [PubMed] [Google Scholar]
- White G. P., Dunn N. W. Apparent fusion of the TOL plasmid with the R91 drug resistance plasmid in Pseudomonas aeruginosa. Aust J Biol Sci. 1977 Aug;30(4):345–355. doi: 10.1071/bi9770345. [DOI] [PubMed] [Google Scholar]
- Williams P. A., Murray K. Metabolism of benzoate and the methylbenzoates by Pseudomonas putida (arvilla) mt-2: evidence for the existence of a TOL plasmid. J Bacteriol. 1974 Oct;120(1):416–423. doi: 10.1128/jb.120.1.416-423.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams P. A., Worsey M. J. Plasmids and catabolism. Biochem Soc Trans. 1976;4(3):466–468. doi: 10.1042/bst0040466. [DOI] [PubMed] [Google Scholar]
- Williams P. A., Worsey M. J. Ubiquity of plasmids in coding for toluene and xylene metabolism in soil bacteria: evidence for the existence of new TOL plasmids. J Bacteriol. 1976 Mar;125(3):818–828. doi: 10.1128/jb.125.3.818-828.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Worsey M. J., Williams P. A. Characterization of a spontaneously occurring mutant of the TOL20 plasmid in Pseudomonas putida MT20: possible regulatory implications. J Bacteriol. 1977 Jun;130(3):1149–1158. doi: 10.1128/jb.130.3.1149-1158.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Worsey M. J., Williams P. A. Metabolism of toluene and xylenes by Pseudomonas (putida (arvilla) mt-2: evidence for a new function of the TOL plasmid. J Bacteriol. 1975 Oct;124(1):7–13. doi: 10.1128/jb.124.1.7-13.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yano K., Nishi T. pKJ1, a naturally occurring conjugative plasmid coding for toluene degradation and resistance to streptomycin and sulfonamides. J Bacteriol. 1980 Aug;143(2):552–560. doi: 10.1128/jb.143.2.552-560.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zukowski M. M., Gaffney D. F., Speck D., Kauffmann M., Findeli A., Wisecup A., Lecocq J. P. Chromogenic identification of genetic regulatory signals in Bacillus subtilis based on expression of a cloned Pseudomonas gene. Proc Natl Acad Sci U S A. 1983 Feb;80(4):1101–1105. doi: 10.1073/pnas.80.4.1101. [DOI] [PMC free article] [PubMed] [Google Scholar]