Abstract
A cloned gene, designated nfo, caused overproduction of an EDTA-resistant endonuclease specific for apurinic-apyrimidinic sites in DNA. The sedimentation coefficient of the enzyme was similar to that of endonuclease IV. An insertion mutation was constructed in vitro and transferred from a plasmid to the Escherichia coli chromosome. nfo mutants had an increased sensitivity to the alkylating agents methyl methanesulfonate and mitomycin C and to the oxidants tert-butyl hydroperoxide and bleomycin. The nfo mutation enhanced the killing of xth (exonuclease III) mutants by methyl methanesulfonate, H2O2, tert-butyl hydroperoxide, and gamma rays, and it enhanced their mutability by methyl methanesulfonate. It also increased the temperature sensitivity of an xth dut (dUTPase) mutant that is defective in the repair of uracil-containing DNA. These results are consistent with earlier findings that endonuclease IV and exonuclease III both cleave DNA 5' to an apurinic-apyrimidinic site and that exonuclease III is more active. However, nfo mutants were more sensitive to tert-butyl hydroperoxide and to bleomycin than were xth mutants, suggesting that endonuclease IV might recognize some lesions that exonuclease III does not. The mutants displayed no marked increase in sensitivity to 254-nm UV radiation, and the addition of an nth (endonuclease III) mutation to nfo or nfo xth mutants did not significantly increase their sensitivity to any of the agents tested.
Full text
PDF





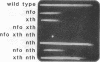
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bachmann B. J. Linkage map of Escherichia coli K-12, edition 7. Microbiol Rev. 1983 Jun;47(2):180–230. doi: 10.1128/mr.47.2.180-230.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Breimer L. H., Lindahl T. DNA glycosylase activities for thymine residues damaged by ring saturation, fragmentation, or ring contraction are functions of endonuclease III in Escherichia coli. J Biol Chem. 1984 May 10;259(9):5543–5548. [PubMed] [Google Scholar]
- Bryson V., Szybalski W. Microbial Selection. Science. 1952 Jul 18;116(3003):45–51. doi: 10.1126/science.116.3003.45. [DOI] [PubMed] [Google Scholar]
- Clarke L., Carbon J. A colony bank containing synthetic Col El hybrid plasmids representative of the entire E. coli genome. Cell. 1976 Sep;9(1):91–99. doi: 10.1016/0092-8674(76)90055-6. [DOI] [PubMed] [Google Scholar]
- Clements J. E., Rogers S. G., Weiss B. A DNase for apurinic/apyrimidinic sites associated with exonuclease III of Hemophilus influenzae. J Biol Chem. 1978 May 10;253(9):2990–2999. [PubMed] [Google Scholar]
- Cunningham R. P., Weiss B. Endonuclease III (nth) mutants of Escherichia coli. Proc Natl Acad Sci U S A. 1985 Jan;82(2):474–478. doi: 10.1073/pnas.82.2.474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demple B., Halbrook J., Linn S. Escherichia coli xth mutants are hypersensitive to hydrogen peroxide. J Bacteriol. 1983 Feb;153(2):1079–1082. doi: 10.1128/jb.153.2.1079-1082.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duncan B. K., Weiss B. Specific mutator effects of ung (uracil-DNA glycosylase) mutations in Escherichia coli. J Bacteriol. 1982 Aug;151(2):750–755. doi: 10.1128/jb.151.2.750-755.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giloni L., Takeshita M., Johnson F., Iden C., Grollman A. P. Bleomycin-induced strand-scission of DNA. Mechanism of deoxyribose cleavage. J Biol Chem. 1981 Aug 25;256(16):8608–8615. [PubMed] [Google Scholar]
- Goffin C., Verly W. G. T4 DNA ligase can seal a nick in double-stranded DNA limited by a 5'-phosphorylated base-free deoxyribose residue. Nucleic Acids Res. 1983 Nov 25;11(22):8103–8109. doi: 10.1093/nar/11.22.8103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gossard F., Verly W. G. Properties of the main endonuclease specific for apurinic sites of Escherichia coli (endonuclease VI). Mechanism of apurinic site excision from DNA. Eur J Biochem. 1978 Jan 16;82(2):321–332. doi: 10.1111/j.1432-1033.1978.tb12026.x. [DOI] [PubMed] [Google Scholar]
- Henner W. D., Grunberg S. M., Haseltine W. A. Enzyme action at 3' termini of ionizing radiation-induced DNA strand breaks. J Biol Chem. 1983 Dec 25;258(24):15198–15205. [PubMed] [Google Scholar]
- Henner W. D., Rodriguez L. O., Hecht S. M., Haseltine W. A. gamma Ray induced deoxyribonucleic acid strand breaks. 3' Glycolate termini. J Biol Chem. 1983 Jan 25;258(2):711–713. [PubMed] [Google Scholar]
- Hochhauser S. J., Weiss B. Escherichia coli mutants deficient in deoxyuridine triphosphatase. J Bacteriol. 1978 Apr;134(1):157–166. doi: 10.1128/jb.134.1.157-166.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katcher H. L., Wallace S. S. Characterization of the Escherichia coli X-ray endonuclease, endonuclease III. Biochemistry. 1983 Aug 16;22(17):4071–4081. doi: 10.1021/bi00286a013. [DOI] [PubMed] [Google Scholar]
- Kow Y. W., Wallace S. S. Exonuclease III recognizes urea residues in oxidized DNA. Proc Natl Acad Sci U S A. 1985 Dec;82(24):8354–8358. doi: 10.1073/pnas.82.24.8354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuhnlein U., Penhoet E. E., Linn S. An altered apurinic DNA endonuclease activity in group A and group D xeroderma pigmentosum fibroblasts. Proc Natl Acad Sci U S A. 1976 Apr;73(4):1169–1173. doi: 10.1073/pnas.73.4.1169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindahl T., Andersson A. Rate of chain breakage at apurinic sites in double-stranded deoxyribonucleic acid. Biochemistry. 1972 Sep 12;11(19):3618–3623. doi: 10.1021/bi00769a019. [DOI] [PubMed] [Google Scholar]
- Ljungquist S. A new endonuclease from Escherichia coli acting at apurinic sites in DNA. J Biol Chem. 1977 May 10;252(9):2808–2814. [PubMed] [Google Scholar]
- Ljungquist S., Lindahl T., Howard-Flanders P. Methyl methane sulfonate-sensitive mutant of Escherichia coli deficient in an endonuclease specific for apurinic sites in deoxyribonucleic acid. J Bacteriol. 1976 May;126(2):646–653. doi: 10.1128/jb.126.2.646-653.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milcarek C., Weiss B. Mutants of Escherichia coli with altered deoxyribonucleases. I. Isolation and characterization of mutants for exonuclease 3. J Mol Biol. 1972 Jul 21;68(2):303–318. doi: 10.1016/0022-2836(72)90215-x. [DOI] [PubMed] [Google Scholar]
- Murugesan N., Xu C., Ehrenfeld G. M., Sugiyama H., Kilkuskie R. E., Rodriguez L. O., Chang L. H., Hecht S. M. Analysis of products formed during bleomycin-mediated DNA degradation. Biochemistry. 1985 Oct 8;24(21):5735–5744. doi: 10.1021/bi00342a008. [DOI] [PubMed] [Google Scholar]
- Rao R. N., Rogers S. G. Plasmid pKC7: a vector containing ten restriction endonuclease sites suitable for cloning DNA segments. Gene. 1979 Sep;7(1):79–82. doi: 10.1016/0378-1119(79)90044-1. [DOI] [PubMed] [Google Scholar]
- Smith P. K., Krohn R. I., Hermanson G. T., Mallia A. K., Gartner F. H., Provenzano M. D., Fujimoto E. K., Goeke N. M., Olson B. J., Klenk D. C. Measurement of protein using bicinchoninic acid. Anal Biochem. 1985 Oct;150(1):76–85. doi: 10.1016/0003-2697(85)90442-7. [DOI] [PubMed] [Google Scholar]
- Spitzer E. D., Weiss B. dfp Gene of Escherichia coli K-12, a locus affecting DNA synthesis, codes for a flavoprotein. J Bacteriol. 1985 Dec;164(3):994–1003. doi: 10.1128/jb.164.3.994-1003.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Talpaert-Borlé M., Liuzzi M. Reaction of apurinic/apyrimidinic sites with [14C]methoxyamine. A method for the quantitative assay of AP sites in DNA. Biochim Biophys Acta. 1983 Sep 9;740(4):410–416. doi: 10.1016/0167-4781(83)90089-1. [DOI] [PubMed] [Google Scholar]
- Taylor A. F., Weiss B. Role of exonuclease III in the base excision repair of uracil-containing DNA. J Bacteriol. 1982 Jul;151(1):351–357. doi: 10.1128/jb.151.1.351-357.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tye B. K., Nyman P. O., Lehman I. R., Hochhauser S., Weiss B. Transient accumulation of Okazaki fragments as a result of uracil incorporation into nascent DNA. Proc Natl Acad Sci U S A. 1977 Jan;74(1):154–157. doi: 10.1073/pnas.74.1.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Walker G. C. Plasmid (pKM101)-mediated enhancement of repair and mutagenesis: dependence on chromosomal genes in Escherichia coli K-12. Mol Gen Genet. 1977 Mar 28;152(1):93–103. doi: 10.1007/BF00264945. [DOI] [PubMed] [Google Scholar]
- Warner H. R., Demple B. F., Deutsch W. A., Kane C. M., Linn S. Apurinic/apyrimidinic endonucleases in repair of pyrimidine dimers and other lesions in DNA. Proc Natl Acad Sci U S A. 1980 Aug;77(8):4602–4606. doi: 10.1073/pnas.77.8.4602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss B., Cunningham R. P. Genetic mapping of nth, a gene affecting endonuclease III (thymine glycol-DNA glycosylase) in Escherichia coli K-12. J Bacteriol. 1985 May;162(2):607–610. doi: 10.1128/jb.162.2.607-610.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss B. Endonuclease II of Escherichia coli is exonuclease III. J Biol Chem. 1976 Apr 10;251(7):1896–1901. [PubMed] [Google Scholar]
- White B. J., Hochhauser S. J., Cintron N. M., Weiss B. Genetic mapping of xthA, the structural gene for exonuclease III in Escherichia coli K-12. J Bacteriol. 1976 Jun;126(3):1082–1088. doi: 10.1128/jb.126.3.1082-1088.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yajko D. M., Weiss B. Mutations simultaneously affecting endonuclease II and exonuclease III in Escherichia coli. Proc Natl Acad Sci U S A. 1975 Feb;72(2):688–692. doi: 10.1073/pnas.72.2.688. [DOI] [PMC free article] [PubMed] [Google Scholar]